1. Introduction
Cotton (
Gossypium hirsutum L.) is an important commercial crop that is grown worldwide as a source of fiber and edible oil [
1,
2,
3,
4]. Cotton fiber serves as the important raw material in the global textile industry and is a better alternative to synthetic fiber, annually consuming billions of barrels of fossil oil worldwide [
5,
6,
7,
8,
9]. Cotton seed contains 28-40% oil, which could be used as edible oil, and raw material for chemical production as well as a feedstock for biodiesel production [
5,
10]. Refined as edible oil, cotton seed oil is typically composed of about 26 % palmitic acid (C16: 0), 15 % oleic acid (C18: 1), and 58 % linoleic acid (C18: 2) [
11], with less total serum cholesterol but more effectively usage compared with corn (
Zea mays) oil [
12], and provides several kinds of benefits and essential fatty acids. So cotton seed oil is a valued raw material in the food industry because it contains a high quantity of saturated palmitic acid and lacks unstable linolenic acids imparting good stability and flavor properties [
13].
Fatty acid synthase (FAS) catalyzes the synthesis of straight-chain fatty acids with 16 or 18 carbon atoms [
14]. The chains are extended further by an endoplasmic reticulum-localized extension system. Two types of FAS are known, of which Type II FAS is found in plant plastids [
15]. Plant plastid FASII consists of six kinds of enzymes: acetyl-coenzyme A (CoA)-acyl carrier protein (ACP) transferase (ACAT), malonic acid single acyl-CoA-ACP transferase (MCAT), β-ketone fatty acyl-ACP synthase (KAS), β-ketone fatty acyl-ACP reductase (KAR), β-hydroxy fatty acyl-ACP dehydratase (HAD), and enoyl-ACP reductase (ENR) [
16].
After carbon-chain condensation by KAS, carbon-chain reduction, dehydration, and reduction reactions are catalyzed by KAR, HAD, and ENR, respectively [
17]. The four-step reaction occurs in the ACP and completes carbon-chain extension. The few studies of KAR, HAD, and ENR have predominantly used transcriptome sequencing. Transcriptome sequencing of
Eustigmatos cf.
polyphem (an oleaginous microalga) and analysis of the anabolic mode of fatty acids and triglyceride metabolism reveals that
KAR,
HAD, and
ENR are key genes in fatty acid synthesis and metabolism [
18]. Transcriptome sequencing indicates that the mesocarp of oil and date palms show considerable differences in oil content. Analysis at five stages of mesocarp development reveals that the total transcript levels of
KAR,
HAD, and
ENR at each stage are significantly higher in the oil palm by 44, 34, and 17-fold, respectively, compared with those of the date palm [
19]. Troncoso–Ponce studied changes in transcription level in four oilseed plants (seed rape, castor, nasturtium, and
Euonymus) at four stages of seed development and observed that
KAR,
HAD, and
ENR showed consistent trends, which suggests that these genes may show similar expression patterns in different plant species [
20]. In peanut, the open reading frame of the genes
AhKAR,
AhHAD, and
AhENR has been cloned. Analysis of the transcript levels of these genes at different stages of ovule development, and in different tissues and organs, shows that the expression pattern of
AhHAD differs from those of
AhKAR and
AhENR [
21]. In cotton, the full-length cDNAs of
GhKAR,
GhHAD, and
GhENR have been cloned and each encodes a 283, 221, and 394 amino acid protein, respectively. Bioinformatic analysis of
GhKAR,
GhHAD, and
GhENR (see phylogenetic tree in the attached table) indicates that the three genes serve important functions in oil accumulation and are involved in the response to physiological stress [
22].
Recently some major genes that involved in fatty acid biosynthesis in oilseeds have been cloned, which were dehydrogenase in
Brassica napus L [
23],
Oryza sativa [
24] and
Hansenula polymorpha [
25], acyltransferase in
Brassicaceae [
26,
27,
28] and the transcription factor genes in
Arabidopsis thaliana [
29]
, Brassica napus [
30] and
Nicotiana tabacum [
31]. In this study we examined the expression patterns of
GhKAR,
GhHAD, and
GhENR in cotton during fiber and seed development. The results provide a foundation for understanding the mechanism of the seed oil accumulation and fiber quality in cotton.
2. Results
2.1. Identification of overexpressing GhKAR, GhHAD and GhENR genes plants
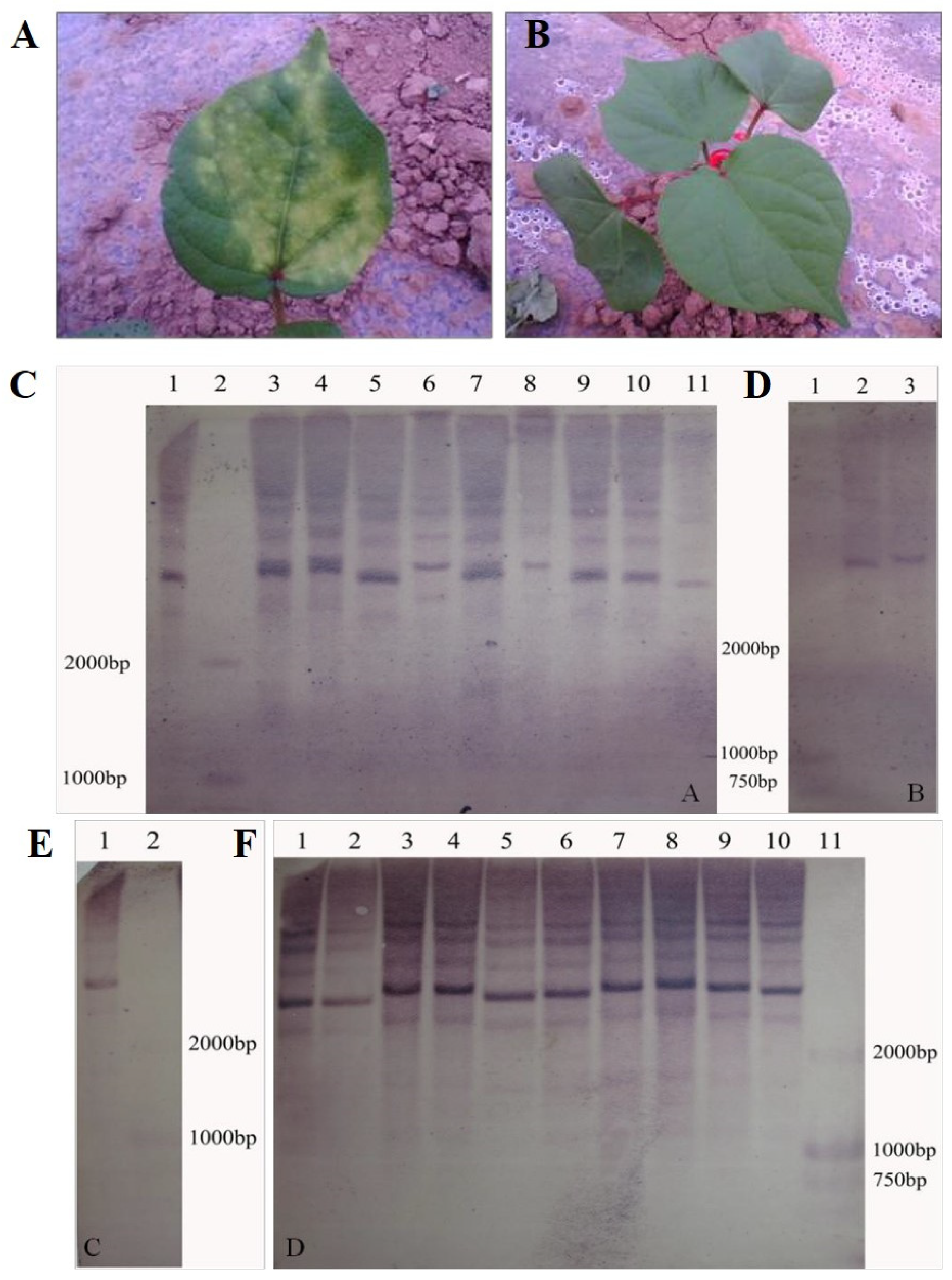
We primarily used 5g/L kanamycin solution to identify the transgenic cotton plants. After 3-5 days of application on leaves, those with yellow spots on the leaves were transgenic negative plants (
Figure 1A), while those without yellow spots were transgenic positive candidate plants (
Figure 1B). Then, the transgenic positive plants were extracted from leaves and the genomic DNA was digested by BamHI-HF endonuclease. The labeled probe was the fragment of kanamycin resistance encoding gene Kan. The copy number of transgenic positive plants was identified by Roche Southern blot kit, and it was found that the copy number of the labeled probe in transgenic plants was between 1-4, as shown in the
Figure 1C–F. The results showed that
GhKAR and
GhENR overexpressed lines contained 3 and 4 copy transgene insertions (
Figure 1C,F), while the copy number of overexpressed
GhHAD plants was single (
Figure 1D,E).
2.2. Phenotypic traits of the plant materials
Based on the seed oil content and gene copy number, four wild-type cotton cultivars (2074B, 10H1007, 10H1014, and 10H1041), and six T
3 transgenic lines that overexpression
GhKAR (13MW012, 13MW124, and 13MW125) or
GhENR (13MW005, 13MW039, and 13MW070), each with single gene copy and high seed oil content were selected to study the functions of
GhKAR, and
GhENR in cotton oil metabolism (
Table 1). The results showed that two wild-type cotton cultivars (10H1014 and 10H1041) contained high oil contents up to 30.73% and 35.97%, respectively, however, another two wild-types (10H1007 and 2074B) contained oil contents low to 26.09% and 27.32%, respectively. In this study, 2074B was the background material for transgene receptor. Compared to 2074B, the oil contents of three
GhKAR overexpression lines increased 1.05~1.07 folds, while
GhENR overexpression lines increased 1.06~1.08 folds, respectively.
The other important agronomic and economic traits of four wild-type cotton cultivars were also recorded. Among the four cultivars, 10H1041 was distinct with the plant height 86.8±4.73 cm and the height of first branch 18.3±2.21 cm, which were significantly lower than the others (
Table 2). Meanwhile, plants produce numerous branches with bolls on each fruiting branch. As for the six transgenic lines, the plant height, height of first branch, branch number, and lint percentage were significantly higher, whereas the seed indices were significantly lower than those of the control line 2074B (
Table 3). Among them, 13MW125 was a favorable line in which the plant height, seed oil content, and lint percentage were significantly higher compared with those of 2074B; however, its fiber quality was not significantly different (
Table 3).
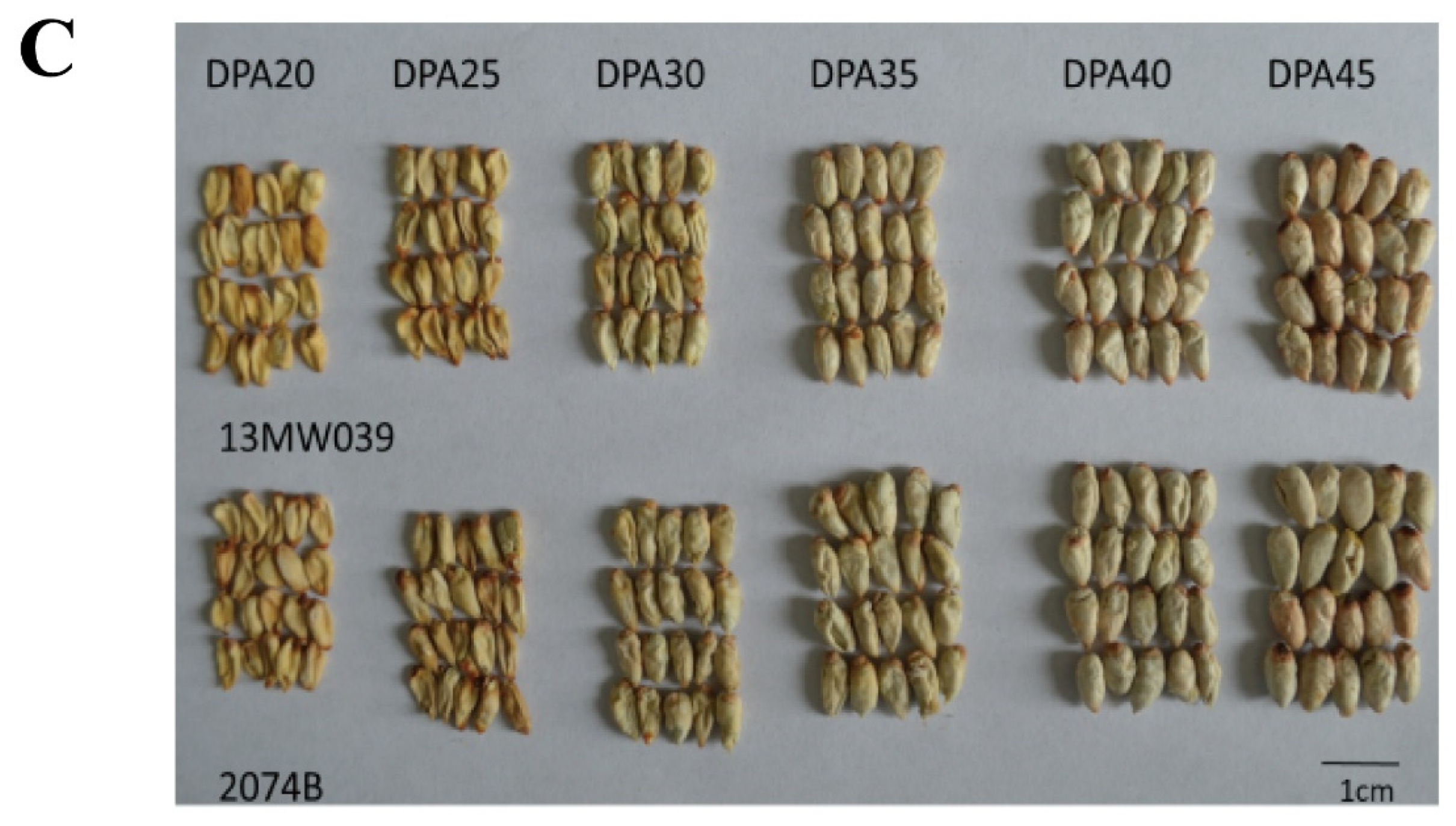
2.3. Morphological changes of immature ovules
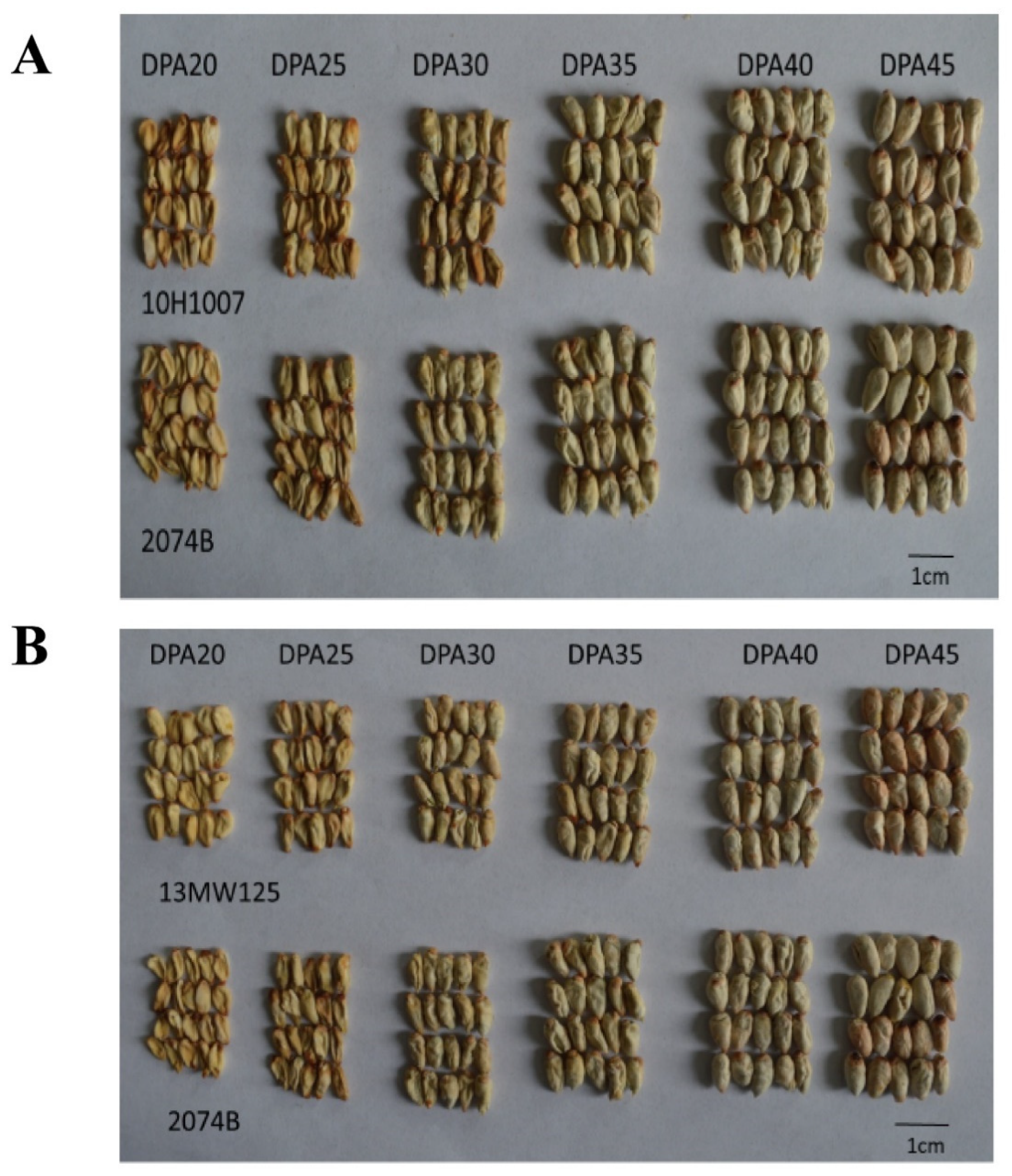
Similar morphologically immature ovules at different stages were indistinguishable as seed maturity differed among bolls collected from different parts of the same plant. After the ovules were shelled and dried, the morphology of the ovules at different seed developmental stages differed notably. The sizes of dried ovules at different developmental stages between 10H1007 and 2074B were consistent (
Figure 2A). Similarly, the sizes of dried ovules at different developmental stages also showed no significant difference between
GhKAR-overexpression line (13MW125),
GhENR-overexpression line (13MW039) and 2074B (
Figure 2B,C).
2.4. Moisture content, grain weight and oil content of four wild-type cotton cultivars and six T3 transgenic lines during seed development
Seed samples from the four wild-type cotton cultivars were collected at 20, 25, 30, 35, 40, and 45 DPA and mature stage. The seed moisture content at each developmental stage was determined. The changes in seed moisture content at the different seed developmental stages were similar among the four wild type cotton cultivars (
Table 4). The moisture content of ovules ranged from 81 % to 87 % at 20 and 25 DPA, 71-76 % at 30 DPA, 61-65 % at 35 DPA, about 55-59 % at 40 DPA and 46-50% at 45 DPA. However, at maturity the moisture content of ovules had decreased to 5–6 %. When the cotton bolls developed after 45 DPA, the seeds were already mature. However, between 45 DPA and maturity stage, the moisture content decreased by almost 50 %, suggesting that seed ripening occurred.
Moreover, the grain weight in four wild-type cotton cultivars increased by about 1 g every 5 d but there existed no significant correlations between the grain weight and oil content in each line (
Table 4). As also shown in
Table 4, there had similar oil content accumulation trends among the four wild type cultivars with different accumulation rates. Oil accumulation rapidly increased in ovules during the period of 20–30 DPA and peaked at 40 and 45 DPA, then decreased slightly at mature stage. Among the four cultivars, 10H1041 contained the highest oil content with oil content declined less at maturity stage than that in the other three lines.
At the 20 DPA and 25 DPA, the grain weights of the six T
3 transgenic lines were significantly higher than that of 2074B (
Table 5). However, at the late developmental stages (30-45 DPA), most grain weights of the six T
3 transgenic lines were significantly lower than that of in 2074B, which were mainly caused by the decreased moisture content at the late stages (
Table 5).
The oil content at the different stages of ovular developmental of the transgenic lines was measured using the Soxhlet extraction method. The oil contents of
GhKAR, GhHAD and
GhENR overexpression transgenic cotton lines presented similar patterns (
Table 5). The oil content of 13MW125 was the highest among the six transgenic lines, and could be used as an important germplasm resource material. The net growth rate of oil content in each
GhKAR and
GhENR overexpression transgenic lines was higher than their receptor material (2074B) at 30–35 DPA (
Table 6), indicating that
GhKAR and
GhENR may have important functions in oil accumulation at 30–35 DPA.
2.5. Expression patterns of GhKAR, GhHAD, and GhENR in 2074B
The expression patterns of
GhKAR,
GhHAD, and
GhENR genes in different tissues were analyzed using quantitative real-time RT-PCR (qRT-PCR). To investigate the expression patterns of these three genes, which are associated with fatty acid synthesis and metabolism, in 2074B (
Figure 3), total RNAs isolated from the roots, stems, leaves, fibers (5, 10, 15, and 20 DPA), and seeds (10, 15, 20, 25, 30, 35, 40, and 45 DPA), were used as templates, respectively. The housekeeping gene
GhUBQ7 was used as an internal comparison gene.
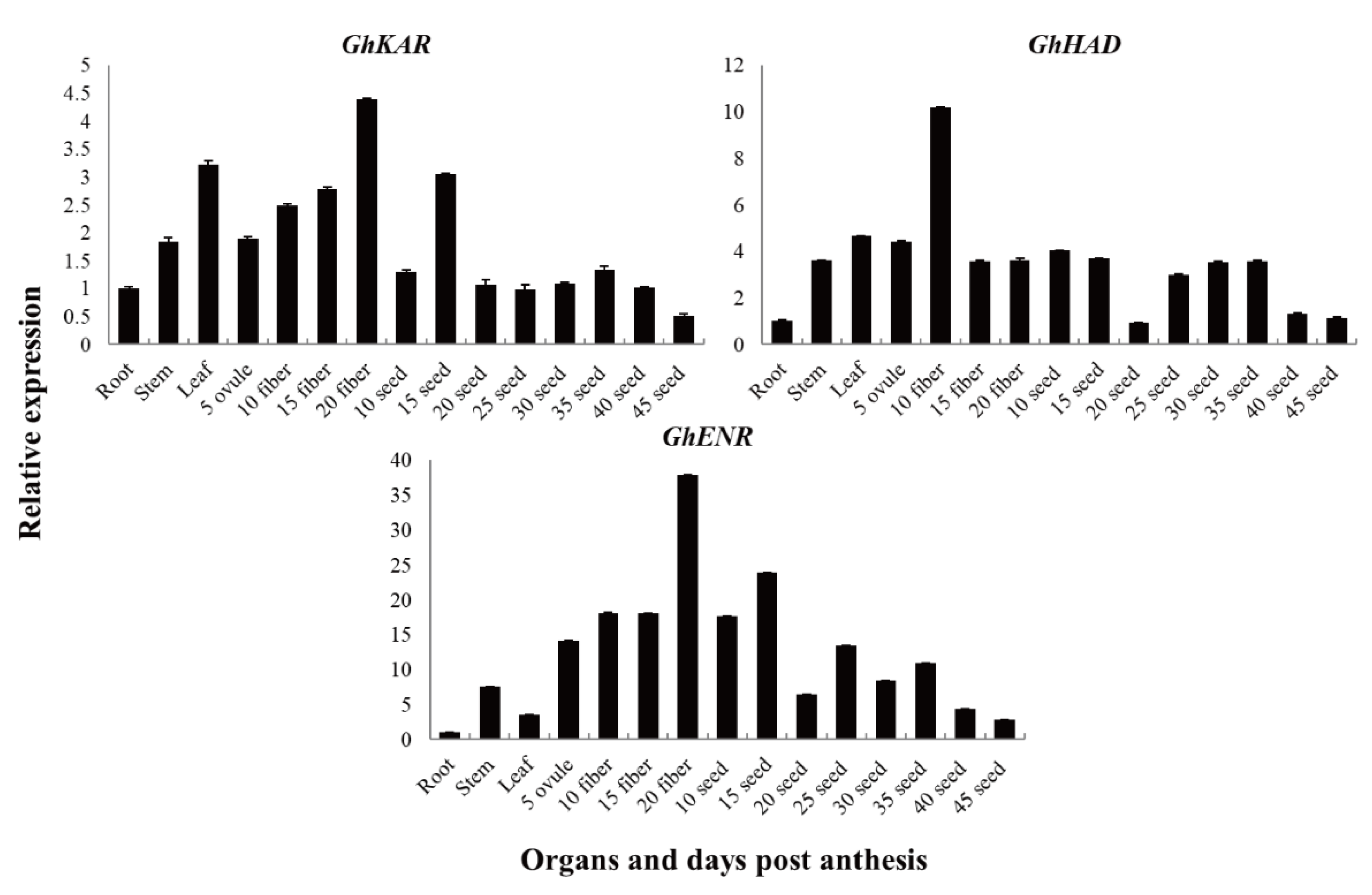
The genes
GhKAR,
GhHAD, and
GhENR were constitutively expressed. The transcript levels of
GhKAR and
GhHAD were higher in the leaves than in the stems and roots, whereas the transcript levels of
GhENR were significantly higher in the stems than in the leaves and roots (
Figure 3). The transcript levels of
GhKAR and
GhENR were high in the fiber (especially in 20 DPA fiber tissue) and 15 DPA seeds. We therefore speculated that expression of
GhKAR and
GhENR has important functions in seed oil synthesis and fiber elongation. The transcript level of
GhHAD was high in fibers and seeds, and especially high in 10 DPA fibers.
GhHAD may have a similar function as
GhKAR and
GhENR, but showed a different expression pattern.
2.6. Expression patterns of GhKAR, GhHAD, and GhENR in four cotton cultivars
2.6.1. Expression patterns in fibers
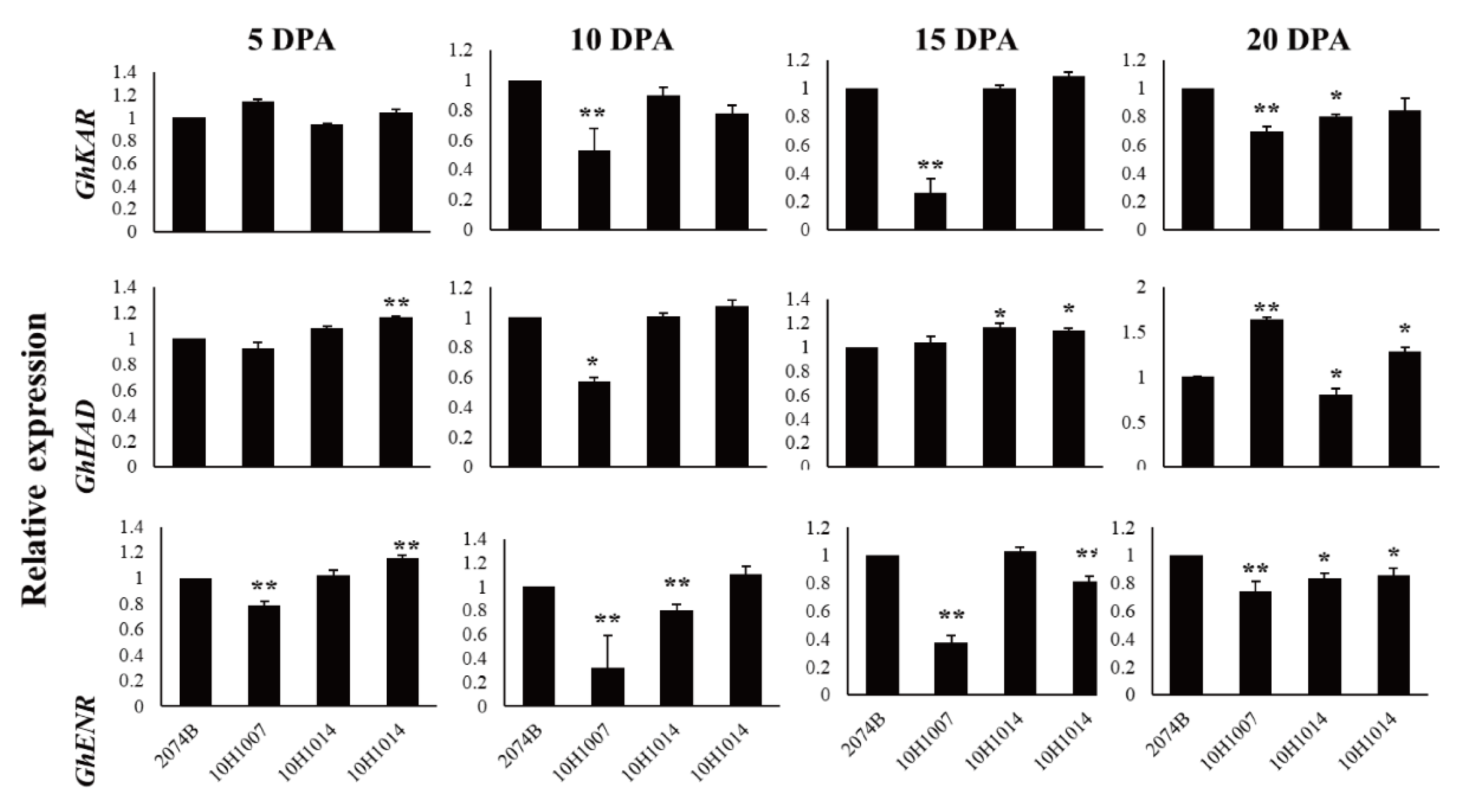
The expression patterns of
GhKAR,
GhHAD, and
GhENR in fibers (5, 10, 15, and 20 DPA) were analyzed using qRT-PCR with the housekeeping gene
GhUBQ7 as the internal comparison gene. Among the four materials, the expression patterns of the three genes varied during fiber development. The expression pattern of
GhKAR and
GhENR both showed an increasing trend that peaked at 20 DPA. However, the expression pattern of
GhHAD differed with peaking at 10 DPA (
Figure 4).
2.6.2. Expression patterns in ovules
To investigate the gene expression patterns in relation to ovule development, the expression patterns of GhKAR, GhHAD, and GhENR genes in ovules (5, 10, 15, 20, 25, 30, 35, 40, and 45 DPA) were also analyzed using qRT-PCR with the housekeeping gene GhUBQ7 as an internal comparison gene.
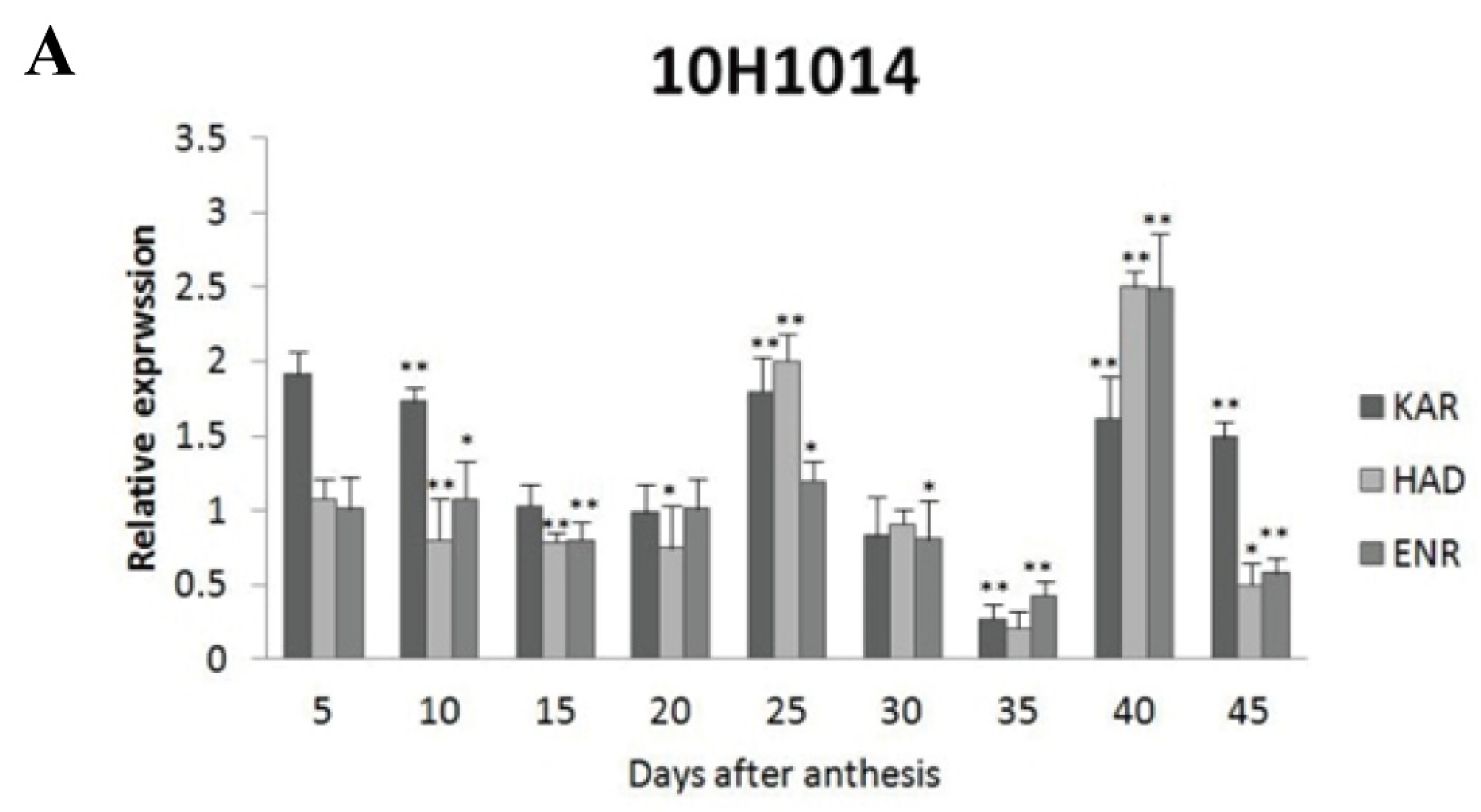
Among the four wild type cultivars, seed oil contents were 26.41% in 2074B, 28.09% in 10H1007, 30.88% in 10H1014, and 35.29% in 10H1041. Meanwhile, the expression patterns of
GhKAR,
GhHAD and
GhENR genes were analyzed in developing ovules of the three wild type cultivars compared to 2074B (
Figure 5). Compared to the 2074B,
GhKAR showed high expression levels at 5, 10, 25, 40 and 45 DPA, while the transcript level of
GhHAD and
GhENR were higher at 25 and 40 DPA in 10H1014 (
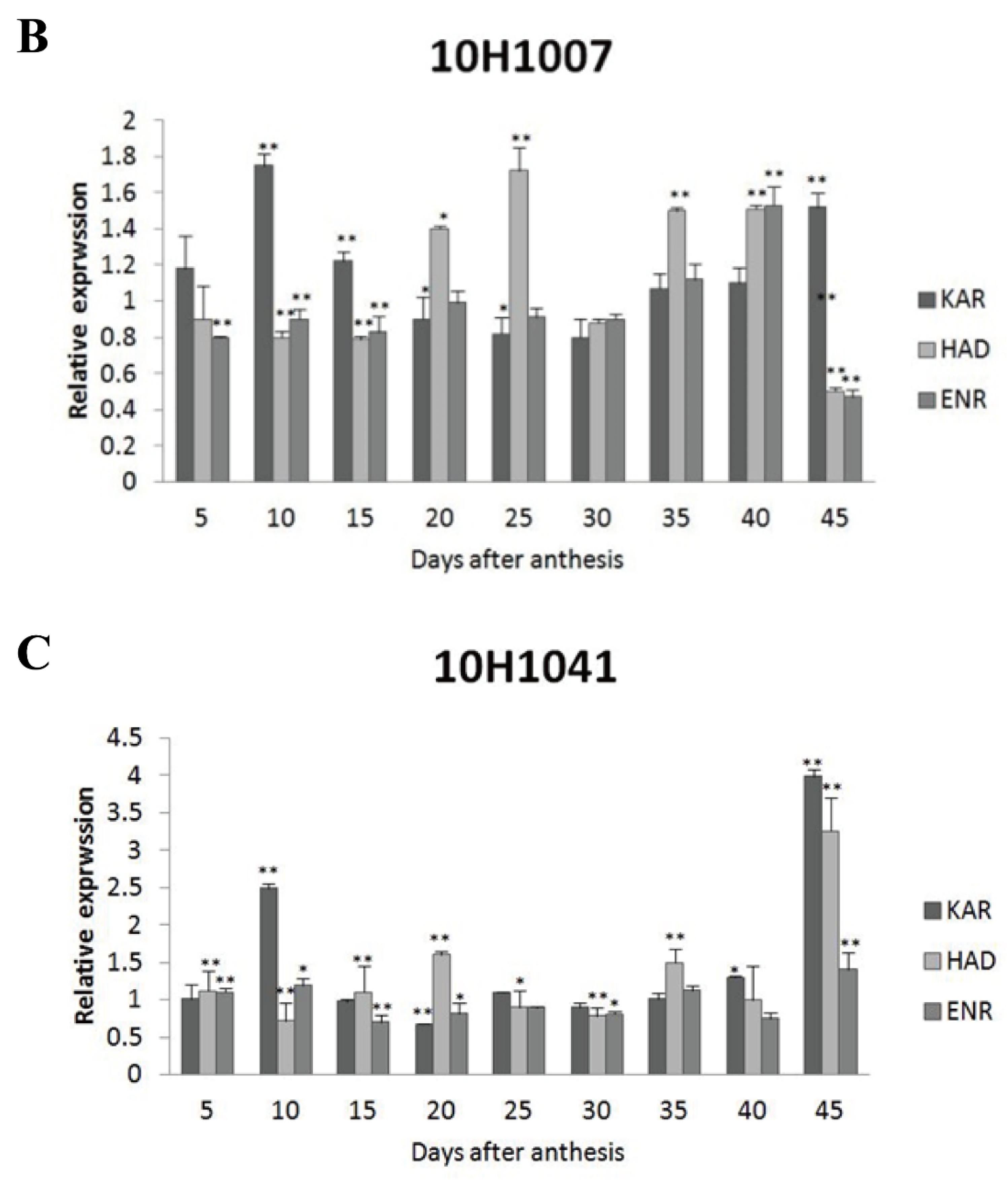
Figure 5A). As to 10H1007, the transcript level of
GhKAR were almost two-fold at 10, 15 and 45 DPA than 2074B,
GhHAD showed high expression levels at 20, 25, 35 and 40 DPA, while
GhENR only increased its transcript level at 40 DPA (
Figure 5B). For 10H1041,
GhKAR were expressed highly at 10, 40 and 45 DPA,
GhHAD elevated its expressions at 20, 35 and 45 DPA and
GhENR increased only at 45 DPA compared with 2074B (
Figure 5C). The above results showed that
GhKAR,
GhHAD and
GhENR genes almost presented higher expression levels at 10H1007, 10H1014 and 10H1041 cultivars with higher seed oil contents than 2074B. However, at this stage, the cultivars that showed high gene transcript levels were not necessarily consistent with the cultivars that showed low oil contents. Interestingly, at the late stage of oil accumulation (40–45 DPA), cultivars with high oil contents exhibited high gene transcript levels. We therefore hypothesized that the high transcript levels of
GhKAR,
GhHAD, and
GhENR at the late oil accumulation stage may affect oil accumulation.
3. Discussion
3.1. The formation of vegetable oils.
Vegetable oil is mainly synthesized during the maturation phase of seed development and is the major energy reserve for later growth and development prior to full establishment of the photosynthetic capacity of the seedling [
30,
32,
33]. The storage oil in seeds mainly consists of TAG synthesized from glycerol-3-phosphate and fatty acids [
34,
35,
36]. In higher plants, biosynthesis of fatty acids and lipids are well characterized in biochemical and molecular studies [
37,
38]. Fatty acid is
de novo synthesized mainly from acetyl-CoA catalyzed in plastids by a series of enzymes, of which ACCase and the FAS multienzyme complex are the most critical enzymes. ACCase, a rate-limiting enzyme of fatty acid synthesis, catalyzes the formation of malonyl-CoA from acetyl-CoA. Subsequently, FAS catalyzes the transfer of the malonyl moiety of malonyl-CoA to ACP by adding two carbons to the growing chain, eventually resulting in the formation of C16:0 and C18:0 acyl-ACP, which are then released from the FAS complex and transferred to the cytoplasm [
39,
40]. In the cytoplasm, fatty acid dehydrogenase (FAD) catalyzes the formation of unsaturated fatty acid, and fatty acid elongase (FAE) sequentially adds two-carbon units to the growing acyl chain to form long-chain fatty acids [
41]. Finally, TAG synthesis is initiated in the endoplasmic reticulum by glycerol-3-phosphate acyltransferase, and the reaction is sequentially completed by lysophosphatidic acid acyltransferase and diacylglycerol acyltransferase [
34,
35,
36].
Triacylglycerol (TAG), the main component of plant oil, represents a compact molecule for energy and carbon storage in organisms because of its highly reduced state [
42]. TAG is typically accumulated in seeds. Besides serving as a source of dietary calories for humans, TAG from plants is increasingly used as a basis for renewable biomaterials and fuels [
43]. Thus, improvement in TAG production would be valuable, and much previous research has focused on this aim within the biotechnology field [
44,
45,
46].
De novo fatty acid synthesis shows no tissue and organ specificity, and occurs in all cells, mainly in the plastids. Acetyl-CoA carboxylase (ACCase) is catalyzed from acetyl-CoA to form malonyl CoA (Mal-CoA). Overexpression the subunits of heteromeric
GhACCase increased the oil content in Upland cotton [
47]. The carbon chain elongation processed condensation, reduction, dehydration, and reduction through the FAS enzymatic system. Finally, acyl is released from ACP through thioesterase activity. Three types of KAS exist, namely KASI, KASII, and KASIII. These three enzymes have different functions in the carbon-chain-extension condensation reaction [
48]. KASIII catalyzes Mal-CoA and acetyl-CoA to form 4:0-ACP, and the starting substrates of the condensation reaction, acetyl-CoA and Mal-CoA, are formed by ACAT and MCAT, respectively. KASI catalyzes 4:0-ACP to form 16:0-ACP, and the fatty acid carbon chain-extending carbon units of the direct donor is Mal-CoA. KASII catalyzes the condensation of 16:0-ACP to 18:0-ACP, and this reaction determines the ratio of 16:0-ACP and 18:0-ACP [
49]. Overexpression of
Spinacia oleracea KASIII (
SoKASIII) in tobacco leaves under control of the 35S CaMV promoter, and of
Jatropha Curcas KASIII (
ChKASIII) in
A. thaliana and oilseed rape seeds under control of the napin promoter, indicates that the 16:0 fatty acid contents of both transgenic plants is significantly increased [
50]. KASI has multifunctional effects on plant growth and development, and also affects oil synthesis, chloroplast division and development. Plants of an
A. thaliana T-DNA insertion deletion mutant,
kas1, show multiple morphological defects, including chlorotic and curly leaves, reduced fertility, and semi-dwarfism [
51]. KASII affects not only the 16:0-ACP content, but also seed development. In the
A. thaliana heterozygous mutant
fab1-2, the seeds are aborted and
fab1-2 homozygous mutants are inviable, which indicates that complete absence of
fab1-2 is lethal for
A. thaliana [
52].
3.2. The application of fatty acid synthase in improving seed oil content.
In recent years, efforts have been made to manipulate key fatty acid synthetic genes in various species using transgenic technology [
26,
53,
54,
55,
56]. Transformations using transgenes encoding key enzymes or enzyme subunits have resulted in alteration of lipid levels to varying degrees [
57,
58], and in some cases the oil content was reduced [
49]. Four key enzymes are involved in the fatty acid carbon-chain extension process, namely KAS, KAR, HAD, and ENR. KAS can be divided into three categories (KASI, KASII, and KASIII) based on its catalytic substrates. KASIII serves an important function in the synthesis of C16:0 fatty acids [
49]. KASII affects the C16:0 fatty acid content [
52]. Mutation or deletion of KAS not only affects fatty acid synthesis, but also seed development. The mutation or deletion of KAS will not only affect the fatty acid synthesis, but also the seed development [
51,
52]. Studies on KAR, HAD, and ENR are relatively limited. Transcriptional research works have shown that KAR, HAD, and ENR are involved in seed oil and fatty acid syntheses [
19]. The ENR deletion mutant of
A. thaliana suffers seed abortion and decreased fatty acid content. The deletion mutant of ENR in
A. thaliana will cause seed abortion and fatty acid content decrement [
22]. The mechanisms by which KAR, HAD, and ENR affect seed oil synthesis remain unclear, therefore further studies are needed.
3.3. GhKAR, GhHAD, and GhENR are key enzymes in the synthesis of fatty acids.
Oil accumulation is part of the seed maturation process, a highly controlled developmental program that sets in ovule tissues once morphogenesis has been achieved. The maturation process is characterized by accumulation of storage compounds, acquisition of desiccation tolerance, and entry into a dormancy period of variable length [
59]. Gene expression programs associated with these processes are activated during the maturation phase and are switched off during vegetative phases of plant development. Studies of developing seeds and/or embryos have established that the biosynthetic pathways for fatty acids and TAGs are regulated at the transcription level [
60,
61,
62]. In the present study, four cotton cultivars with different seed oil contents showed consistent oil accumulation trends during seed development. The overall trend of oil accumulation was that at 20–30 DPA the oil content rapidly increased and thereafter showed slow accumulation. At 40 DPA, the oil content peaked and then slightly decreased. The decrease in oil content at maturity may be attributable to β-oxidation and several other physiological and biochemical processes, suggesting that the decline in oil accumulation during seed maturation might be overcome using molecular breeding techniques to achieve the goal of developing lines with high seed oil contents.
GhKAR,
GhHAD, and
GhENR are crucial genes involved in fatty acid carbon-chain extension that have been cloned from immature ovules of upland cotton [
19]. The present analysis of transcript levels of these genes in different tissues and organs of 2074B indicated that the genes are highly expressed during fiber development, and show higher transcription levels than in most developing seeds. Therefore, these genes are not only involved in plant seed oil synthesis, but also in the cotton fiber elongation process. The transcript levels of
GhKAR and
GhENR were particularly high at 20 DPA during fiber development and at 15 DPA during seed development. However, the transcript level of
GhHAD was high only at 10 DPA during fiber development, which indicated that
GhHAD, together with
GhKAR and
GhENR, showed different expression patterns during seed and fiber development. Analysis of the fatty acid compositions of the homozygous transgenic T
3 and T
4 generations of
A. thaliana seed (data not shown) showed that overexpression of
GhKAR,
GhHAD, or
GhENR could increase the total content of fatty acids in the seed. The total fatty acid content of transgenic
GhKAR,
GhHAD, and
GhENR lines were 6.59 %, 7.76 %, and 3.86 % higher than that of wild-type
A. thaliana, respectively. Thus, overexpression of
GhKAR,
GhHAD, or
GhENR may improve the total fatty acid content by increasing the concentration of every fatty acid because
GhKAR,
GhHAD, and
GhENR are genes involved in fatty acid synthesis. Further studies are needed to verify this hypothesis and the molecular functions of the three genes.
4. Materials and methods
4.1. Plant materials and growth conditions
Four cotton cultivars (2074B, 10H1004, 10H1007, and 10H1041), and T3 transgenic cotton lines that overexpression GhKAR (13MW012, 13MW124, and 13MW125) and GhENR (13MW005, 13MW039, and 13MW070), were used in our experiments.
Plants were grown on soil in a greenhouse (16 h light/8 h dark) at 28°C. After three weeks, the roots, stems, and leaves of seedlings at the three-leaf stage were harvested, frozen in liquid nitrogen, and stored at −80°C for DNA and total RNA extraction. Seeds were grown in the field in Hejian, from which samples of fibers (5, 10, 15, and 20 DPA), and ovules (5, 10, 15, 20, 25, 30, 35, 40, and 45 DPA) of the different materials were collected.
4.2. Field experiments and agronomic trait investigation
For field experiments, non- transgenic and the transgenic plants were sown directly on April 27th at Hejian, Hebei Province (38°43′N, 116°09′E). The field planting followed a randomized complete block design with three replications. Two-row plots with 80 cm and 50 cm row spacing were used. The length of each plot was 4m. Field management followed conventional standard field practices. Data were collected from at least 10 plants of in each line. Self-pollinated bells were harvested from each primary transgenic (T
3) plant and analyzed for quality character of fibers and seed oil content. The cotton seed oil content was determined using the Soxhlet extraction method and near-infrared spectroscopy [
22]. All results were statistical analyzed by three times repeats. LSD test and difference significance test of statistics method were adopted in finally result analysis.
4.3. RNA isolation and cDNA synthesis
Total RNA was extracted using a modified CTAB-SDS method [
63]. RNA samples were treated with DNase I (Ambion, Austin, TX, USA) in accordance with the manufacturer’s instructions to remove genomic DNA contaminants. Total RNA samples (1 µg per reaction) were reverse transcribed into cDNA by avian myeloblastosis virus (AMV) reverse transcriptase. The cDNAs were used as the template in subsequent qPCR reactions.
4.4. Quantitative real-time RT-PCR
Total RNA was extracted from young leaves, roots, stems, and developing fibers (5, 10, 15, and 20 DPA) and ovules (5, 10, 15, 20, 25, 30, 35, 40, and 45 DPA) as indicated above. Gene-specific primers were designed to amplify PCR products of ~200 bp in length (see
Supplementary Table S1). The relative level of gene expression was estimated using the 2
-△△CT method [
64]. The analyses were performed with three biological replicates using samples from different plants. The SYBR
® Premix Ex Taq™ II (Tli RNaseH Plus) (TaKaRa, Biotechnology (Dalian) Co., Ltd.) was used for RT-PCR.
The expression patterns of GhKAR, GhHAD, and GhENR in different tissues were studied using qRT-PCR. To investigate the expression patterns of the three genes in relation to fatty acid synthesis and metabolism in 2074B, total RNAs isolated from the root, stem, young leaves, and ovule, as well as from fibers (5, 10, 15, and 20 DPA), and ovules (5, 10, 15, 20, 25, 30, 35, 40, and 45 DPA), of 2074B were used as templates. In addition, G. hirsutum UBQ7 gene used to normalize served as an endogenous reference. Data are presented as the means (±SD) of three independent experiments.
4.5. Southern blotting analysis
Procedure for Southern blot was described briefly as follows.Kan fragment was amplified by PCR (500bp) and the concentration of recovered DNA was diluted to 60-70ng/µL. Add 16µL Kan fragment in a 200µL centrifuge tube, bathe in boiling water for 10 min, quickly place on ice containing sodium chloride for 10 min, and centrifuge instantaneously. Adding 4µL DIG-High-Prime, instantaneous centrifugation, and 37℃ water bath for 20h; then, 2µL 0.2M EDTA (pH8.0) (or 10min in 65℃ water bath) was added to terminate the reaction and stored at -20℃. Probe denaturation: Add 5µL labeled probe into 200µL centrifuge tube, denaturate in water bath at 68℃ for 10 min after sealing with sealing film, and cool rapidly in ice water for 5 min (denaturation before use). The subsequent enzymatic digestion of cotton genomic DNA, treatment of digested products, electrophoresis, transmembrane and crosslinking, prehybridization and hybridization, rigor elution, detection reaction and color reaction were finished according to the protocol of Roche Southern blot kit.
4.6. Statistical analysis
The SPSS 13.0 statistical package (IBM Corporation, New York, USA) was used for the analysis of variance and Student’s t test. The significance was tested using the least significant difference (LSD) at the 5% level. Each sample included in the analysis was based on three biological replicates.
5. Conclusions
In this study, we observed that GhKAR and GhENR showed similar expression trends during fiber and seed development in three cultivars with different oil contents. The transcript levels of GhKAR and GhENR gradually increased and peaked at 20 DPA during fiber development. Considering that 10-20 DPA is a period of rapid fiber elongation, GhKAR and GhENR might have important functions at the late stage of rapid fiber elongation. However, the expression level of GhHAD peaked at 10 DPA, which suggested that GhHAD plays an important role in the early stage of rapid fiber elongation. Our results provided new insights into the fatty acid biosynthesis in cotton.
Supplementary Materials
The following supporting information can be downloaded at the website of this paper posted on Preprints.org. Figures S1. Phylogram of GhKAR protein sequences from different organisms; Figures S2. Phylogram of Gh HAD protein sequences from different organisms; Figures S3. Phylogram of GhENR protein sequences from different organisms; Table S1 Primers used in the experiment.
Author Contributions
Conceptualization, Li Liu, Jinping Hua, Yu Yu and Zhiwen Chen; Data curation, Li Liu, Dan Wang, Fuxiang Zhao and Wenhao Liu; Formal analysis, Dan Wang, Xuwen Wang, Juan Wang, Aijun Si and Zhiwen Chen; Funding acquisition, Jinping Hua and Yu Yu; Investigation, Li Liu, Dan Wang, Xianhui Kong, Xuwen Wang, Aijun Si and Fuxiang Zhao; Methodology, Li Liu, Dan Wang, Xianhui Kong, Xuwen Wang, Juan Wang, Aijun Si and Fuxiang Zhao; Project administration, Jinping Hua and Yu Yu; Resources, Li Liu, Dan Wang and Wenhao Liu; Software, Li Liu, Dan Wang, Xianhui Kong, Xuwen Wang, Aijun Si and Zhiwen Chen; Supervision, Jinping Hua, Yu Yu and Zhiwen Chen; Validation, Li Liu, Dan Wang, Xianhui Kong, Xuwen Wang, Juan Wang, Fuxiang Zhao, Wenhao Liu and Zhiwen Chen; Visualization, Jinping Hua and Zhiwen Chen; Writing – original draft, Li Liu and Zhiwen Chen; Writing – review & editing, Jinping Hua, Yu Yu and Zhiwen Chen.
Funding
This work was supported in part by the National Natural Science Foundation of China (NO. 31360349), and the National Key Research and Development Program of China (No. 2022YFD1200300). The funding bodies did not participate in the design of the study; the collection, analysis, or interpretation of the data; or in the writing of the manuscript.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Acknowledgments
We are deeply indebted to Professor Lida Zhang for helpful suggestions and comments on bioinformatic analyses, and we thank Professor Yi Huang for valuable comments on previous versions of the manuscript. We are also grateful to two anonymous reviewers for their helpful suggestions and comments.
Conflicts of Interest
The authors declare no conflict of interest.
Abbreviations
TAG: Triacylglycerol; FAS, Fatty acid synthase; β-ketone fatty acyl-ACP reductase (KAR); β-hydroxy fatty acyl-ACP dehydratase (HAD), and enoyl-ACP reductase (ENR); β-ketone fatty acyl-ACP synthase (KAS); Acetyl-CoA carboxylase (ACCase); acetyl-coenzyme A (CoA)-acyl carrier protein (ACP) transferase (ACAT); malonic acid single acyl-CoA-ACP transferase (MCAT); DPA, days post-anthesis; G. hirsutum, Gossypium hirsutum; A. thaliana, Arabidopsis thaliana; qRT-PCR, quantitative real-time polymerase chain reaction.
References
- Zhang, F.; Li, S.; Yang, S.; Wang, L.; Guo, W., Overexpression of a cotton annexin gene, GhAnn1, enhances drought and salt stress tolerance in transgenic cotton. Plant Mol Biol 2015, 87, 47-67. [CrossRef]
- Gao, L.; Chen, W.; Xu, X.; Zhang, J.; Singh, T. K.; Liu, S.; Zhang, D.; Tian, L.; White, A.; Shrestha, P.; Zhou, X. R.; Llewellyn, D.; Green, A.; Singh, S. P.; Liu, Q., Engineering trienoic fatty acids into cottonseed oil improves low-temperature seed germination, plant photosynthesis and cotton fiber quality. Plant Cell Physiol 2020, 61, 1335-1347. [CrossRef]
- Khan, A.; Tan, D. K. Y.; Munsif, F.; Afridi, M. Z.; Shah, F.; Wei, F.; Fahad, S.; Zhou, R., Nitrogen nutrition in cotton and control strategies for greenhouse gas emissions: A review. Environ Sci Pollut Res Int 2017, 24, 23471-23487. [CrossRef]
- Mubeen, M.; Ahmad, A.; Hammad, H. M.; Awais, M.; Farid, H. U.; Saleem, M.; Din, M. S. u.; Amin, A.; Ali, A.; Fahad, S.; Nasim, W., Evaluating the climate change impact on water use efficiency of cotton-wheat in semi-arid conditions using DSSAT model. Journal of Water and Climate Change 2019, 11, 1661-1675. [CrossRef]
- Lee, M. K.; Zhang, Y.; Zhang, M.; Goebel, M.; Kim, H. J.; Triplett, B. A.; Stelly, D. M.; Zhang, H. B., Construction of a plant-transformation-competent BIBAC library and genome sequence analysis of polyploid Upland cotton (Gossypium hirsutum L.). BMC Genomics 2013, 14, 208. [CrossRef]
- Wendel, J. F.; Grover, C. E., Taxonomy and evolution of the cotton genus. In Cotton, Agronomy, Fang, D.; Percy, R., Eds. Monograph 24, ASA-CSSA-SSSA: Madison, WI, 2015.
- Wang, K.; Wendel, J. F.; Hua, J., Designations for individual genomes and chromosomes in Gossypium. Journal of Cotton Research 2018, 1, 3. [CrossRef]
- Chen, Z.; Grover, C. E.; Li, P.; Wang, Y.; Nie, H.; Zhao, Y.; Wang, M.; Liu, F.; Zhou, Z.; Wang, X.; Cai, X.; Wang, K.; Wendel, J. F.; Hua, J., Molecular evolution of the plastid genome during diversification of the cotton genus. Mol Phylogenet Evol 2017, 112, 268-276. [CrossRef]
- Huang, G.; Huang, J. Q.; Chen, X. Y.; Zhu, Y. X., Recent advances and future perspectives in cotton research. Annu Rev Plant Biol 2021, 72, 437-462. [CrossRef]
- Khan, A.; Tan, D. K. Y.; Afridi, M. Z.; Luo, H.; Tung, S. A.; Ajab, M.; Fahad, S., Nitrogen fertility and abiotic stresses management in cotton crop: A review. Environ Sci Pollut Res Int 2017, 24, 14551-14566. [CrossRef]
- Liu, Q.; Singh, S. P.; Green, A. G., High-stearic and High-oleic cottonseed oils produced by hairpin RNA-mediated post-transcriptional gene silencing. Plant Physiol 2002, 129, 1732-43. [CrossRef]
- Radcliffe, J. D.; King, C. C.; Czajka-Narins, D. M.; Imrhan, V., Serum and liver lipids in rats fed diets containing corn oil, cottonseed oil, or a mixture of corn and cottonseed oils. Plant Foods Hum Nutr 2001, 56, 51-60. [CrossRef]
- Royon, D.; Daz, M.; Ellenrieder, G.; Locatelli, S., Enzymatic production of biodiesel from cotton seed oil using t-butanol as a solvent. Bioresour Technol 2007, 98, 648-53. [CrossRef]
- Brown, A. P.; Slabas, A. R.; Rafferty, J. B., Fatty acid biosynthesis in plants — metabolic pathways, structure and organization. In Lipids in Photosynthesis: Essential and Regulatory Functions, Wada, H.; Murata, N., Eds. Springer Netherlands: Dordrecht, 2009; pp 11-34. [CrossRef]
- Ryall, K.; Harper, J. T.; Keeling, P. J., Plastid-derived Type II fatty acid biosynthetic enzymes in chromists. Gene 2003, 313, 139-148. [CrossRef]
- Rogalski, M.; Carrer, H., Engineering plastid fatty acid biosynthesis to improve food quality and biofuel production in higher plants. Plant Biotechnol J 2011, 9, 554-64. [CrossRef]
- Cui, Y.; Zhao, Y.; Wang, Y.; Liu, Z.; Ijaz, B.; Huang, Y.; Hua, J., Genome-wide identification and expression analysis of the biotin carboxyl carrier subunits of heteromeric acetyl-CoA carboxylase in Gossypium. Front Plant Sci 2017, 8, 624. [CrossRef]
- Wan, L.; Han, J.; Sang, M.; Li, A.; Wu, H.; Yin, S.; Zhang, C., De novo transcriptomic analysis of an oleaginous microalga: Pathway description and gene discovery for production of next-generation biofuels. PLoS ONE 2012, 7, e35142. [CrossRef]
- Bourgis, F.; Kilaru, A.; Cao, X.; Ngando-Ebongue, G. F.; Drira, N.; Ohlrogge, J. B.; Arondel, V., Comparative transcriptome and metabolite analysis of oil palm and date palm mesocarp that differ dramatically in carbon partitioning. Proc Natl Acad Sci U S A 2011, 108, 12527-32. [CrossRef]
- Troncoso-Ponce, M. A.; Kilaru, A.; Cao, X.; Durrett, T. P.; Fan, J.; Jensen, J. K.; Thrower, N. A.; Pauly, M.; Wilkerson, C.; Ohlrogge, J. B., Comparative deep transcriptional profiling of four developing oilseeds. Plant J 2011, 68, 1014-27. [CrossRef]
- Zhang, H.; Ji, Y.; Wang, Y.; Zhang, X.; Yu, Y., Cloning and characterization of a novel β-ketoacyl-ACP reductase from Comamonas testosteroni. Chem Biol Interact 2015, 234, 213-20. [CrossRef]
- LIU Li, Z. P., WANG Dan, LIU Zheng-Jie, WANG Yu-Mei, HUA Jin-Ping, Molecular cloning and expression analysis of three chain extension genes related to fatty acid synthesis in Upland cotton (Gossypium hirsutum L.). Scientia Agricultura Sinica 2013, 46, 3523-3533. [CrossRef]
- Vigeolas, H.; Waldeck, P.; Zank, T.; Geigenberger, P., Increasing seed oil content in oil-seed rape (Brassica napus L.) by over-expression of a yeast glycerol-3-phosphate dehydrogenase under the control of a seed-specific promoter. Plant Biotechnol J 2007, 5, 431-41. [CrossRef]
- Shi, J.; Cao, Y.; Fan, X.; Li, M.; Wang, Y.; Ming, F., A rice microsomal delta-12 fatty acid desaturase can enhance resistance to cold stress in yeast and Oryza sativa. Mol Breeding 2012, 29, 743-757. [CrossRef]
- Sangwallek, J.; Kaneko, Y.; Tsukamoto, T.; Marui, M.; Sugiyama, M.; Ono, H.; Bamba, T.; Fukusaki, E.; Harashima, S., Cloning and functional analysis of HpFAD2 and HpFAD3 genes encoding Delta12- and Delta15-fatty acid desaturases in Hansenula polymorpha. Gene 2014, 533, 110-8. [CrossRef]
- Zou, J.; Katavic, V.; Giblin, E. M.; Barton, D. L.; MacKenzie, S. L.; Keller, W. A.; Hu, X.; Taylor, D. C., Modification of seed oil content and acyl composition in the brassicaceae by expression of a yeast sn-2 acyltransferase gene. Plant Cell 1997, 9, 909-23. [CrossRef]
- Jain, R. K.; Coffey, M.; Lai, K.; Kumar, A.; MacKenzie, S. L., Enhancement of seed oil content by expression of glycerol-3-phosphate acyltransferase genes. Biochem Soc Trans 2000, 28, 958-61.
- Maisonneuve, S.; Bessoule, J. J.; Lessire, R.; Delseny, M.; Roscoe, T. J., Expression of rapeseed microsomal lysophosphatidic acid acyltransferase isozymes enhances seed oil content in Arabidopsis. Plant Physiol 2010, 152, 670-84. [CrossRef]
- Wang, H. W.; Zhang, B.; Hao, Y. J.; Huang, J.; Tian, A. G.; Liao, Y.; Zhang, J. S.; Chen, S. Y., The soybean Dof-type transcription factor genes, GmDof4 and GmDof11, enhance lipid content in the seeds of transgenic Arabidopsis plants. Plant J 2007, 52, 716-29. [CrossRef]
- Tan, H.; Yang, X.; Zhang, F.; Zheng, X.; Qu, C.; Mu, J.; Fu, F.; Li, J.; Guan, R.; Zhang, H.; Wang, G.; Zuo, J., Enhanced seed oil production in canola by conditional expression of Brassica napus LEAFY COTYLEDON1 and LEC1-LIKE in developing seeds. Plant Physiol 2011, 156, 1577-88. [CrossRef]
- Fan, Z.; Li, J.; Lu, M.; Li, X.; Yin, H., Overexpression of phosphoenolpyruvate carboxylase from Jatropha curcas increases fatty acid accumulation in Nicotiana tabacum. Acta Physiol Plant 2013, 35, 2269-2279. [CrossRef]
- Rahman, M. H. u.; Ahmad, A.; Wajid, A.; Hussain, M.; Rasul, F.; Ishaque, W.; Islam, M. A.; Shelia, V.; Awais, M.; Ullah, A.; Wahid, A.; Sultana, S. R.; Saud, S.; Khan, S.; Fahad, S.; Hussain, M.; Hussain, S.; Nasim, W., Application of CSM-CROPGRO-Cotton model for cultivars and optimum planting dates: Evaluation in changing semi-arid climate. Field Crop Res 2019, 238, 139-152. [CrossRef]
- Nazim, M.; Ali, M.; Shahzad, K.; Ahmad, F.; Nawaz, F.; Amin, M.; Anjum, S.; Nasif, O.; Ali Alharbi, S.; Fahad, S.; Danish, S.; Datta, R., Kaolin and Jasmonic acid improved cotton productivity under water stress conditions. Saudi Journal of Biological Sciences 2021, 28, 6606-6614. [CrossRef]
- Slabas, A. R.; Fawcett, T., The biochemistry and molecular biology of plant lipid biosynthesis. Plant Mol Biol 1992, 19, 169-191. [CrossRef]
- Ohlrogge, J.; Browse, J., Lipid biosynthesis. The Plant Cell 1995, 7, 957-970. [CrossRef]
- Voelker, T.; Kinney, A. J., Variations in the biosynthesis of seed-storage lipids. Annu Rev Plant Physiol Plant Mol Biol 2001, 52, 335-361. [CrossRef]
- Harwood, J. L., Recent advances in the biosynthesis of plant fatty acids. Biochim Biophys Acta 1996, 1301, (1-2), 7-56. [CrossRef]
- Beisson, F. d. r.; Koo, A. J. K.; Ruuska, S.; Schwender, J. r.; Pollard, M.; Thelen, J. J.; Paddock, T.; Salas, J. n. J.; Savage, L.; Milcamps, A.; Mhaske, V. B.; Cho, Y.; Ohlrogge, J. B., Arabidopsis genes involved in acyl lipid metabolism. A 2003 census of the candidates, a study of the distribution of expressed sequence tags in organs, and a web-based database. Plant Physiol 2003, 132, 681-697. [CrossRef]
- Ohlrogge, J. B.; Jaworski, J. G., Regulation of fatty acid synthesis. Annu Rev Plant Physiol Plant Mol Biol 1997, 48, 109-136.
- Sasaki, Y.; Nagano, Y., Plant acetyl-CoA carboxylase: Structure, biosynthesis, regulation, and gene manipulation for plant breeding. Biosci Biotechnol Biochem 2004, 68, 1175-84. [CrossRef]
- Zhao, Y.; Wang, Y.; Huang, Y.; Cui, Y.; Hua, J., Gene network of oil accumulation reveals expression profiles in developing embryos and fatty acid composition in Upland cotton. J Plant Physiol 2018, 228, 101-112. [CrossRef]
- Li-Beisson, Y.; Shorrosh, B.; Beisson, F.; Andersson, M. X.; Arondel, V.; Bates, P. D.; Baud, S.; Bird, D.; Debono, A.; Durrett, T. P.; Franke, R. B.; Graham, I. A.; Katayama, K.; Kelly, A. A.; Larson, T.; Markham, J. E.; Miquel, M.; Molina, I.; Nishida, I.; Rowland, O.; Samuels, L.; Schmid, K. M.; Wada, H.; Welti, R.; Xu, C.; Zallot, R.; Ohlrogge, J., Acyl-lipid metabolism. Arabidopsis Book 2013, 11, e0161. [CrossRef]
- Chapman, K. D.; Ohlrogge, J. B., Compartmentation of triacylglycerol accumulation in plants. J Biol Chem 2012, 287, 2288-94. [CrossRef]
- Andrianov, V.; Borisjuk, N.; Pogrebnyak, N.; Brinker, A.; Dixon, J.; Spitsin, S.; Flynn, J.; Matyszczuk, P.; Andryszak, K.; Laurelli, M.; Golovkin, M.; Koprowski, H., Tobacco as a production platform for biofuel: Overexpression of Arabidopsis DGAT and LEC2 genes increases accumulation and shifts the composition of lipids in green biomass. Plant Biotechnol J 2010, 8, 277-87. [CrossRef]
- Taylor, D. C.; Falk, K. C.; Palmer, C. D.; Hammerlindl, J.; Babic, V.; Mietkiewska, E.; Jadhav, A.; Marillia, E.-F.; Francis, T.; Hoffman, T.; Giblin, E. M.; Katavic, V.; Keller, W. A., Brassica carinata – a new molecular farming platform for delivering bio-industrial oil feedstocks: Case studies of genetic modifications to improve very long-chain fatty acid and oil content in seeds. Biofuels, Bioproducts and Biorefining 2010, 4, 538-561. [CrossRef]
- Guiheneuf, F.; Leu, S.; Zarka, A.; Khozin-Goldberg, I.; Khalilov, I.; Boussiba, S., Cloning and molecular characterization of a novel acyl-CoA:diacylglycerol acyltransferase 1-like gene (PtDGAT1) from the diatom Phaeodactylum tricornutum. FEBS J 2011, 278, 3651-66. [CrossRef]
- Cui, Y.; Liu, Z.; Zhao, Y.; Wang, Y.; Huang, Y.; Li, L.; Wu, H.; Xu, S.; Hua, J., Overexpression of heteromeric GhACCase subunits enhanced oil accumulation in Upland cotton. Plant Mol Biol Rep 2017, 35, 287-297. [CrossRef]
- Gonzalez-Mellado, D.; von Wettstein-Knowles, P.; Garces, R.; Martinez-Force, E., The role of beta-ketoacyl-acyl carrier protein synthase III in the condensation steps of fatty acid biosynthesis in sunflower. Planta 2010, 231, 1277-89. [CrossRef]
- Dehesh, K.; Tai, H.; Edwards, P.; Byrne, J.; Jaworski, J. G., Overexpression of 3-ketoacyl-acyl-carrier protein synthase IIIs in plants reduces the rate of lipid synthesis. Plant Physiol 2001, 125, 1103-14. [CrossRef]
- Yu, N.; Xiao, W. F.; Zhu, J.; Chen, X. Y.; Peng, C. C., The Jatropha curcas KASIII gene alters fatty acid composition of seeds in Arabidopsis thaliana. Biol Plantarum 2015, 59, 773-782. [CrossRef]
- Wu, G. Z.; Xue, H. W., Arabidopsis β-ketoacyl-[acyl carrier protein] synthase i is crucial for fatty acid synthesis and plays a role in chloroplast division and embryo development. Plant Cell 2010, 22, 3726-44. [CrossRef]
- Pidkowich, M. S.; Nguyen, H. T.; Heilmann, I.; Ischebeck, T.; Shanklin, J., Modulating seed β-ketoacyl-acyl carrier protein synthase II level converts the composition of a temperate seed oil to that of a palm-like tropical oil. Proceedings of the National Academy of Sciences 2007, 104, 4742-4747. [CrossRef]
- Arias-Gaguancela, O.; Aziz, M.; Chapman, K. D., Fatty acid amide hydrolase and 9-lipoxygenase modulate cotton seedling growth by ethanolamide oxylipin levels. Plant Physiol 2022. [CrossRef]
- Shi, Z.; Chen, X.; Xue, H.; Jia, T.; Meng, F.; Liu, Y.; Luo, X.; Xiao, G.; Zhu, S., GhBZR3 suppresses cotton fiber elongation by inhibiting very-long-chain fatty acid biosynthesis. Plant J 2022, 111, 785-799. [CrossRef]
- Shorrosh, B. S.; Roesler, K. R.; Shintani, D.; van de Loo, F. J.; Ohlrogge, J. B., Structural analysis, plastid localization, and expression of the biotin carboxylase subunit of acetyl-coenzyme A carboxylase from tobacco. Plant Physiol 1995, 108, 805-12. [CrossRef]
- Jako, C.; Kumar, A.; Wei, Y.; Zou, J.; Barton, D. L.; Giblin, E. M.; Covello, P. S.; Taylor, D. C., Seed-specific over-expression of an Arabidopsis cDNA encoding a diacylglycerol acyltransferase enhances seed oil content and seed weight. Plant Physiol 2001, 126, 861-74. [CrossRef]
- Zhao, Y.; Huang, Y.; Wang, Y.; Cui, Y.; Liu, Z.; Hua, J., RNA interference of GhPEPC2 enhanced seed oil accumulation and salt tolerance in Upland cotton. Plant Sci 2018, 271, 52-61. [CrossRef]
- Zhu, Y.; Hu, X.; Wang, P.; Gao, L.; Pei, Y.; Ge, Z.; Ge, X.; Li, F.; Hou, Y., GhPLP2 positively regulates cotton resistance to verticillium wilt by modulating fatty acid accumulation and jasmonic acid signaling pathway. Front Plant Sci 2021, 12, 749630. [CrossRef]
- Baud, S.; Lepiniec, L., Physiological and developmental regulation of seed oil production. Prog Lipid Res 2010, 49, 235-49. [CrossRef]
- Baud, S.; Graham, I. A., A spatiotemporal analysis of enzymatic activities associated with carbon metabolism in wild-type and mutant embryos of Arabidopsis using in situ histochemistry. Plant J 2006, 46, 155-69. [CrossRef]
- Fawcett, T.; Simon, W. J.; Swinhoe, R.; Shanklin, J.; Nishida, I.; Christie, W. W.; Slabas, A. R., Expression of mRNA and steady-state levels of protein isoforms of enoyl-ACP reductase from Brassica napus. Plant Mol Biol 1994, 26, 155-63. [CrossRef]
- O’Hara, P.; Slabas, A. R.; Fawcett, T., Fatty acid and lipid biosynthetic genes are expressed at constant molar ratios but different absolute levels during embryogenesis. Plant Physiol 2002, 129, 310-320. [CrossRef]
- Vennapusa, A. R.; Somayanda, I. M.; Doherty, C. J.; Jagadish, S. V. K., A universal method for high-quality RNA extraction from plant tissues rich in starch, proteins and fiber. Sci Rep-Uk 2020, 10, 16887. [CrossRef]
- Livak, K. J.; Schmittgen, T. D., Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402-8. [CrossRef]
Figure 1.
Identification of overexpressing GhKAR, GhHAD and GhENR genes plants. A: transgenic negative plants identified by kanamycin. B: transgenic positive plants identified by kanamycin. C: Southern blotting detection of transgenic GhENR plants. 1, 3-11 were the results of GhENR and 2 was Marker D2000; D: Southern blotting detection of transgenic GhHAD plants. 1: Marker D2000; 2 and 3: Southern blotting results of GhHAD plants. E: Southern blotting detection of transgenic GhHAD plants. 1: Southern blotting results of GhHAD plants; 2: Marker D2000.F: Southern blotting detection of transgenic GhKAR plants. 1-10: Southern blotting results; 11: Marker D2000.
Figure 1.
Identification of overexpressing GhKAR, GhHAD and GhENR genes plants. A: transgenic negative plants identified by kanamycin. B: transgenic positive plants identified by kanamycin. C: Southern blotting detection of transgenic GhENR plants. 1, 3-11 were the results of GhENR and 2 was Marker D2000; D: Southern blotting detection of transgenic GhHAD plants. 1: Marker D2000; 2 and 3: Southern blotting results of GhHAD plants. E: Southern blotting detection of transgenic GhHAD plants. 1: Southern blotting results of GhHAD plants; 2: Marker D2000.F: Southern blotting detection of transgenic GhKAR plants. 1-10: Southern blotting results; 11: Marker D2000.
Figure 2.
Morphological changes in immature ovules of cotton at different developmental stages. A: 10H1007 and 2074 (non-transgenic lines); B: 13MW125 (GhKAR-overexpression line) and 2074B; C: 13MW039 (GhENR-overexpression line) and 2074B. DPA, days post-anthesis.
Figure 2.
Morphological changes in immature ovules of cotton at different developmental stages. A: 10H1007 and 2074 (non-transgenic lines); B: 13MW125 (GhKAR-overexpression line) and 2074B; C: 13MW039 (GhENR-overexpression line) and 2074B. DPA, days post-anthesis.
Figure 3.
Transcription patterns of GhKAR, GhHAD, and GhENR in vegetative organs, and during fiber and seed development in upland cotton 2074B. Numerals in fiber and ovule development stages indicate the number of days post-anthesis; the values and error bars the mean +/- SE of three biological replicates.
Figure 3.
Transcription patterns of GhKAR, GhHAD, and GhENR in vegetative organs, and during fiber and seed development in upland cotton 2074B. Numerals in fiber and ovule development stages indicate the number of days post-anthesis; the values and error bars the mean +/- SE of three biological replicates.
Figure 4.
Expression levels of GhKAR, GhHAD, and GhENR at different stages of fiber development in four non-transgenic cotton lines showing different seed oil contents. Values presented are the mean ± SE of three biological replicates. * and ** indicate significant differences in test of statistics (p < 0.05 and p < 0.01, respectively) compared with the 2074B value set at 1. Oil contents of the four lines were: 2074B, 26.41 %; 10H1007, 28.09 %; 10H1014, 30.88 %; 10H1041, 35.29 %. DPA, days post-anthesis.
Figure 4.
Expression levels of GhKAR, GhHAD, and GhENR at different stages of fiber development in four non-transgenic cotton lines showing different seed oil contents. Values presented are the mean ± SE of three biological replicates. * and ** indicate significant differences in test of statistics (p < 0.05 and p < 0.01, respectively) compared with the 2074B value set at 1. Oil contents of the four lines were: 2074B, 26.41 %; 10H1007, 28.09 %; 10H1014, 30.88 %; 10H1041, 35.29 %. DPA, days post-anthesis.
Figure 5.
Expression levels of GhKAR, GhHAD, and GhENR at different stages of ovular development in three cotton cultivars showing different seed oil contents. A: 10H1014; B: 10H1007; C: 10H1041. Values presented are the mean ± SE of three biological replicates. * and ** indicate significant differences in test of statistics (p < 0.05 and p < 0.01, respectively) compared with the 2074B value set at 1. Seed oil contents of the four lines were: 2074B, 26.41 %; 10H1007, 28.09 %; 10H1014, 30.88 %; and 10H1041, 35.29 %. DPA, days post-anthesis.
Figure 5.
Expression levels of GhKAR, GhHAD, and GhENR at different stages of ovular development in three cotton cultivars showing different seed oil contents. A: 10H1014; B: 10H1007; C: 10H1041. Values presented are the mean ± SE of three biological replicates. * and ** indicate significant differences in test of statistics (p < 0.05 and p < 0.01, respectively) compared with the 2074B value set at 1. Seed oil contents of the four lines were: 2074B, 26.41 %; 10H1007, 28.09 %; 10H1014, 30.88 %; and 10H1041, 35.29 %. DPA, days post-anthesis.
Table 1.
Oil content of the four wild-type and six T3 transgenic cotton lines used in the study.
Table 1.
Oil content of the four wild-type and six T3 transgenic cotton lines used in the study.
| Materials ID |
Genotypes |
Oil contents /% |
| 10H1041 |
wild-type |
35.97 |
| 10H1014 |
wild-type |
30.73 |
| 10H1007 |
wild-type |
26.09 |
| 2074B |
wild-type |
27.32 |
| 13MW012 |
KAR OE line |
28.56 |
| 13MW124 |
KAR OE line |
29.12 |
| 13MW125 |
KAR OE line |
28.87 |
| 13MW005 |
ENR OE line |
29.36 |
| 13MW039 |
ENR OE line |
29.61 |
| 13MW070 |
ENR OE line |
28.93 |
Table 2.
Phenotypic traits of the four non-transgenic cotton lines.
Table 2.
Phenotypic traits of the four non-transgenic cotton lines.
| Traits |
10H1041 |
10H1014 |
10H1007 |
2074B |
| Plant height (cm) |
86.8±4.73c
|
103.1±16.88ab
|
109.9±11.33a
|
92.9±13.74bc
|
| Height of first branch (cm) |
18.3±2.21b
|
30.5±6.65a
|
29.7±5.17a
|
28.9±3.00a
|
| Branch number |
15.2±1.14a
|
14.5±1.72ab
|
15.9±1.97a
|
12.9±1.79b
|
| lint percent (%) |
36.04±1.55b
|
36.72±0.82b
|
43.13±1.51a
|
38.95±1.22ab
|
| Seed index (g) |
11.70±0.07ab
|
11.88±0.02a
|
10.89±0.61b
|
11.23±0.34ab
|
| Fiber length (mm) |
29.63 ± 0.47a
|
29.76 ± 0.18a
|
29.40 ± 1.69a
|
30.63 ± 0.29a
|
| Fiber uniformity ratio (%) |
85.80 ± 0.28a |
85.10 ± 1.98a
|
86.45 ± 1.06a
|
86.85 ± 0.21a
|
| Fiber strength (cN/tex) |
29.70 ± 0.57b
|
29.70 ± 0.14b
|
30.05 ± 1.06ab
|
31.45 ± 0.07a
|
| Fiber elongation (%) |
6.25 ± 0.07a
|
6.35 ± 0.07a
|
6.40 ± 0.14a
|
6.30 ± 0.00a
|
| Micronaire |
4.83 ± 0.31a
|
4.27 ± 0.15a
|
4.60 ± 0.28a
|
4.37 ± 0.19a
|
Table 3.
Phenotypic traits of six transgenic cotton lines and non-transgenic cotton 2074B.
Table 3.
Phenotypic traits of six transgenic cotton lines and non-transgenic cotton 2074B.
| Traits |
2074B |
13MW012 |
13MW124 |
13MW125 |
13MW005 |
13MW039 |
13MW070 |
| Plant height (cm) |
92.9±13.74 |
95.1±8.20 |
108.9±9.48**
|
109.2±10.03**
|
99.6±9.41 |
112.3±4.81**
|
103.4±9.38 |
| Height of first branch (cm) |
28.9±3.00 |
33.0±6.88 |
36.3±4.97**
|
34.54±9.14 |
31.2±4.92 |
26.1±3.28 |
27.7±3.97 |
| Branch number |
12.9±1.79 |
13.8±2.35 |
13.8±2.94 |
12.8±4.34 |
13.4±1.96 |
17.1±0.88**
|
15.1±1.20**
|
| Lint percent (%) |
38.95±1.22 |
36.92±2.57 |
39.78±1.59 |
43.85±2.06**
|
39.18±2.04 |
44.14±1.71**
|
37.29±1.83 |
| Seed index (g) |
11.23±0.34 |
10.73±1.07 |
10.16±0.80 |
10.01±0.90 |
9.34±0.80**
|
9.64±1.54 |
9.75±0.73* |
| Fiber length(mm) |
30.63 ± 0.29 |
30.06 ± 0.48 |
29.63 ± 0.75 |
30.43 ± 0.80 |
30.97 ± 0.83 |
27.69 ± 0.93*
|
30.81 ± 0.88 |
| Fiber uniformity ratio (%) |
86.85 ± 0.21 |
85.63 ± 0.86 |
85.57 ± 0.76 |
87.54 ± 0.83 |
85.85 ± 0.07*
|
86.23 ± 0.46 |
84.94 ± 1.25 |
| Fiber strength (cN/tex) |
31.45 ± 0.07 |
33.27 ± 1.63 |
32.53 ± 0.67 |
31.14 ± 1.47 |
30.6- ± 0.71 |
29.13 ± 1.66 |
32.96 ± 0.68*
|
| Fiber elongation (%) |
6.30 ± 0.00 |
6.13 ± 0.12 |
6.20 ± 0.17 |
6.26 ± 0.23 |
6.20 ± 0.00**
|
6.48 ± 0.19 |
6.00 ± 0.07**
|
| Micronaire |
4.37 ± 0.19 |
4.35 ± 0.62 |
4.34 ± 0.28 |
4.71 ± 0.28 |
4.03 ± 0.77 |
4.70 ± 0.41 |
3.96 ± 0.31 |
Table 4.
Phenotypic traits for the ovule of four wild type cotton cultivars.
Table 4.
Phenotypic traits for the ovule of four wild type cotton cultivars.
| Plant lines and traits |
20 DPA |
25 DPA |
30 DPA |
35 DPA |
40 DPA |
45 DPA |
Mature |
| 10H1041 |
| Moisture content (%) |
86.15 |
85.05 |
76.90 |
65.01 |
55.05 |
46.96 |
5.12 |
| Grain weight (g) |
1.52±0.05c |
1.95±0.03a |
2.91±0.03a |
4.37±0.04a |
5.66±0.01a |
7.06±0.03a |
|
| Oil content (%) |
1.37d
|
14.57ab
|
24.12c
|
32.53a
|
36.27a
|
35.29a
|
35.29a
|
| 10H1014 |
| Moisture content (%) |
83.12 |
84.80 |
71.17 |
65.42 |
56.76 |
50.74 |
4.57 |
| Grain weight (g) |
1.74±0.01a |
1.81±0.01b |
2.90±0.01a |
4.15±0.01b |
4.29±0.09d |
5.70±0.18d |
|
| Oil content (%) |
6.25a
|
11.67c
|
29.31a
|
29.49b
|
33.55b
|
30.88b
|
30.88b
|
| 10H1007 |
| Moisture content (%) |
84.66 |
82.91 |
73.84 |
61.62 |
59.08 |
50.55 |
5.90 |
| Grain weight (g) |
1.61±0.01b |
1.91±0.01a |
2.98±0.02a |
3.97±0.02c |
4.53±0.04c |
6.31±0.01c |
|
| Oil content (%) |
4.29 b
|
14.03 b
|
24.05 c
|
28.31 c
|
29.69 c
|
24.72 c
|
28.09 c
|
| 2074B |
| Moisture content (%) |
85.73 |
84.00 |
73.40 |
62.44 |
56.23 |
48.71 |
4.24 |
| Grain weight (g) |
1.42±0.02d |
1.82±0.02b |
2.94±0.11a |
3.96±0.05c |
4.70±0.05b |
6.70±0.05b |
|
| Oil content (%) |
2.68c
|
15.06a
|
26.90b
|
29.52b
|
27.54d
|
29.86b
|
26.41d
|
Table 5.
Phenotypic traits of six T3 transgenic lines and 2074B.
Table 5.
Phenotypic traits of six T3 transgenic lines and 2074B.
| Plant lines and traits |
20 DPA/% |
25 DPA/% |
30 DPA /% |
35 DPA/% |
40 DPA/% |
45 DPA/% |
Mature/% |
| 2074B |
| Moisture content (%) |
85.73 |
84.00 |
73.40 |
62.44 |
56.23 |
48.71 |
4.24 |
| Grain weight (g) |
1.42±0.02d |
1.82±0.02b |
2.94±0.11a |
3.96±0.05c |
4.70±0.05b |
6.70±0.05b |
|
| Oil content (%) |
2.68c
|
15.06a
|
26.90b
|
29.52b
|
27.54d
|
29.86b
|
26.41d
|
|
13MW012 (GhKAR OE line) |
| Moisture content (%) |
86.12 |
84.89 |
76.01 |
67.98 |
54.18 |
54.69 |
6.30 |
| Grain weight (g) |
1.76±0.01** |
1.94±0.01** |
3.06±0.01 |
3.98±0.08 |
4.34±0.01** |
6.44±0.09* |
|
| Oil content (%) |
1.41 **
|
11.88 **
|
24.86 *
|
29.66 |
30.47 **
|
26.84 **
|
25.85 |
|
13MW124 (GhKAR OE line) |
| Moisture content (%) |
87.02 |
85.44 |
75.10 |
68.67 |
55.83 |
52.92 |
5.97 |
| Grain weight (g) |
1.42±0.01 |
1.62±0.01** |
2.90±0.01 |
3.60±0.05** |
4.90±0.07 * |
5.63±0.03** |
|
| Oil content (%) |
1.42 **
|
12.69 *
|
27.50 |
28.94 |
30.44 *
|
26.37 *
|
27.18 |
|
13MW125 (GhKAR OE line) |
| Moisture content (%) |
84.14 |
83.29 |
75.93 |
64.35 |
55.25 |
49.18 |
5.93 |
| Grain weight (g) |
1.71±0.01** |
2.04±0.02** |
2.74±0.01 * |
3.85±0.03 * |
5.06±0.01** |
5.54±0.01** |
|
| Oil content (%) |
1.25 **
|
14.01 **
|
27.34 |
30.77 *
|
30.17 *
|
28.80 |
31.38 **
|
|
13MW005 (GhENR OE line) |
| Moisture content (%) |
86.05 |
81.89 |
70.92 |
59.25 |
52.69 |
52.77 |
6.06 |
| grain weight (g) |
1.51±0.04 * |
1.97±0.04** |
2.98±0.01 |
3.70±0.03** |
4.64±0.03 |
5.05±0.02** |
|
| Oil content (%) |
3.73 **
|
16.47 **
|
25.43 *
|
30.45 *
|
29.41 *
|
27.20 **
|
27.77 *
|
|
13MW039 (GhENR OE line) |
| Moisture content (%) |
86.25 |
83.74 |
76.67 |
64.52 |
52.03 |
48.82 |
4.48 |
| Grain weight |
1.50±0.03 * |
1.71±0.03** |
2.43±0.02** |
3.60±0.01** |
4.63±0.01 |
5.05±0.05** |
|
| Oil content (%) |
2.04 **
|
14.91 |
26.51 |
31.07 *
|
31.01 *
|
28.63 **
|
27.16 |
|
13MW070 (GhENR OE line) |
| Moisture content (%) |
85.66 |
83.56 |
76.67 |
61.63 |
56.15 |
49.76 |
5.49 |
| Grain weight (g) |
1.53±0.03** |
1.86±0.01 |
2.75±0.03* |
3.82±0.050* |
3.85±0.02** |
5.31±0.01** |
|
| Oil content (%) |
3.12 *
|
14.50 |
24.07 *
|
29.59 |
30.19 *
|
27.76 *
|
25.88 |
Table 6.
Net increment in oil content of ovules of T3 transgenic cotton lines at three stages of ovular development.
Table 6.
Net increment in oil content of ovules of T3 transgenic cotton lines at three stages of ovular development.
| Material |
Gene |
Net increment of 20DPA-25DPA (%) |
Net increment of 25DPA-30DPA (%) |
Net increment of 30DPA-35DPA (%) |
| 2074B |
Contrast |
12.38 |
11.84 |
2.63 |
| 13MW012 |
OEGhKAR |
10.47**
|
12.99 |
4.79*
|
| 13MW124 |
11.27 |
14.81*
|
1.43*
|
| 13MW125 |
12.75*
|
13.33 |
3.43*
|
| 13MW005 |
OEGhENR |
12.74 |
8.96*
|
5.02*
|
| 13MW039 |
12.87 |
11.60 |
4.57*
|
| 13MW070 |
11.38 |
9.57 |
5.53*
|
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).