1. Introduction
A key aim in translational research for cancer treatment is to focus on targeting mechanisms that allow malignant cells to resist cytotoxic chemotherapy. One general aspect of cancer cell resistance mechanisms to chemotherapy is the development of clones with increased capacity to respond to cellular stress. Chemotherapy is administered to kill cancer cells, and proves especially effective in killing cancer cells that operate error-prone systems of biomolecular synthesis and processing. However, exposure of cancer cells to chemotherapy or any other cytotoxic conditions tends to select for clones that operate efficient cell stress adaptation mechanisms, which often act by accelerating the removal of mediators of cell death, or by preventing accumulation of cytotoxic metabolites. The latter is particularly important for leukemia cells exposed to chemotherapy [
1].
The clearest manifestation of cancer cell resistance mechanisms to chemotherapy should be found in patients who develop recurrent neoplastic disease. An example of a recurrent hematologic cancer is acute myeloid leukemia (AML). In AML, it was previously hypothesized that aldehyde dehydrogenase (ALDH) enzymatic activity marks a positive outcome, because a drop in activity was found in cancer patients when compared to healthy study volunteers; yet soon after, in addition to identifying non-malignant stem cells within some AML samples, high ALDH activity was also a marker of CD34
+/ CD38
- leukemic stem cells in some patients [
2,
3,
4,
5,
6]. However, it was difficult to reach a definitive conclusion on the role of ALDH in leukemia development, as ALDH activity assays could not distinguish between different members of this protein family. However, patients with leukemia cells lacking ALDH1A1 expression were later found to have a positive prognosis, and their leukemia cells could be killed by chemotherapy [
7]. ALDH1A1 is a vital enzyme for AML cell detoxification from toxic aldehydes that arise after chemotherapy, although it has a similar role in normal hematopoietic cells [
8]. Additionally, we found previously that favorable prognosis patients were generally expressing lower RNA levels from the gene that encodes ALDH1A1; furthermore, high expression of ALDH1A1 RNA had a significant negative association with survival for AML patients [
9].
Nevertheless, a redundancy between the activities of subgroup members of the ALDH family has hampered research efforts to find conclusive evidence for addressing the role of specific ALDH genes in cancer. Herein, we analyzed publicly available data from leading AML studies with the objective of characterizing the relationship between ALDH gene RNA expression with patient risk and survival, in an effort to focus on therapeutically relevant findings and specifically on pharmaceutically actionable target genes.
2. Materials and Methods
We searched the literature for AML datasets containing RNA expression and risk or survival information. Datasets were obtained from the Gene Expression Omnibus (GEO) [
10], the Genomic Data Commons [
11] and cBioPortal [
12], and were processed as described previously [
9].
RNA-seq data was processed using TMM (
edgeR package) [
13]. Counts were converted to log counts per million (log CPM) and genes with < 15 read counts across all samples were removed. Processed microarray datasets were downloaded from GEO. One dataset (BEAT AML) was retrieved using the
beatAML package [
14]. Beat AML samples were separated into patients with bone marrow aspirate (BMA) samples and patients with peripheral blood (PB) samples. Patients with both BMA and PB samples were excluded, and each sample type was analyzed separately. In the TARGET cohort, when comparing expression between primary and recurrent tumors, patients with paired primary and recurrent tumor samples were analyzed separately from additional independent primary and recurrent tumor samples. Only primary tumors were included for all other TARGET analyses. For genes with multiple probes, the probe with the highest mean expression was used. Patients with risk assignments of ‘favorable or ‘low’, were considered ‘low risk’, while patients with ‘adverse’, ‘high’, or ‘poor’ risk were considered ‘high risk’. Samples labeled ‘intermediate’, ‘normal’, and ‘standard’ were excluded from the risk analysis. Association with risk is measured by the area under the receiver operating characteristics curve (AUC), using the ROCR package [
15]. P-values comparing groups are calculated using a paired or independent two-sample t-test. For survival analysis, log rank P-values were used to assess statistical significance of survival curves. Confidence intervals for hazard ratios were calculated using the
confint package in R. The
forestplot package was used to generate the forest plots.
3. Results
Our analysis is based on gene expression and clinical data from 9 independent patient cohorts, with 8 cohorts containing risk information (low- or high-risk, N = 860) and 7 cohorts containing overall survival information (N = 1170). These datasets are summarized in
Table 1. In section 3.1, we first compare
ALDH1A1 expression between primary and recurrent tumors; in section 3.2, we evaluate the association of all
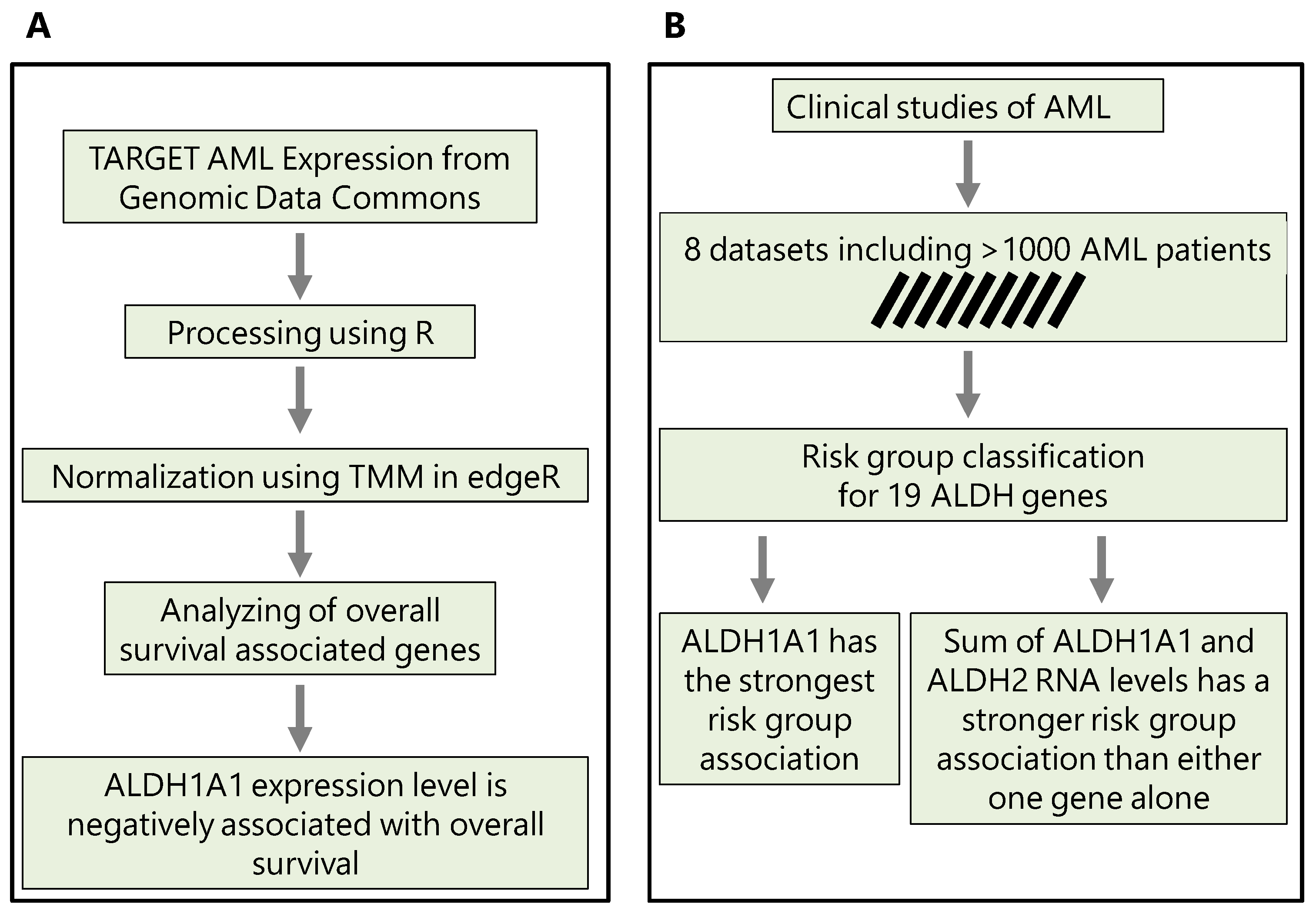
ALDH genes with risk; in section 3.3, we consider the top two genes, and evaluate whether their combined expression is a better marker of risk and survival than either gene alone. The workflows that we use and key findings are summarized in
Figure 1.
3.1. Implication of RNA expression from the ALDH1A1 gene in AML resistance to chemotherapy
We first focused on
ALDH1A1 expression, since we previously demonstrated that in pediatric AML,
ALDH1A1 gene RNA had a stronger association with risk group classification than the established biomarker CALCRL [
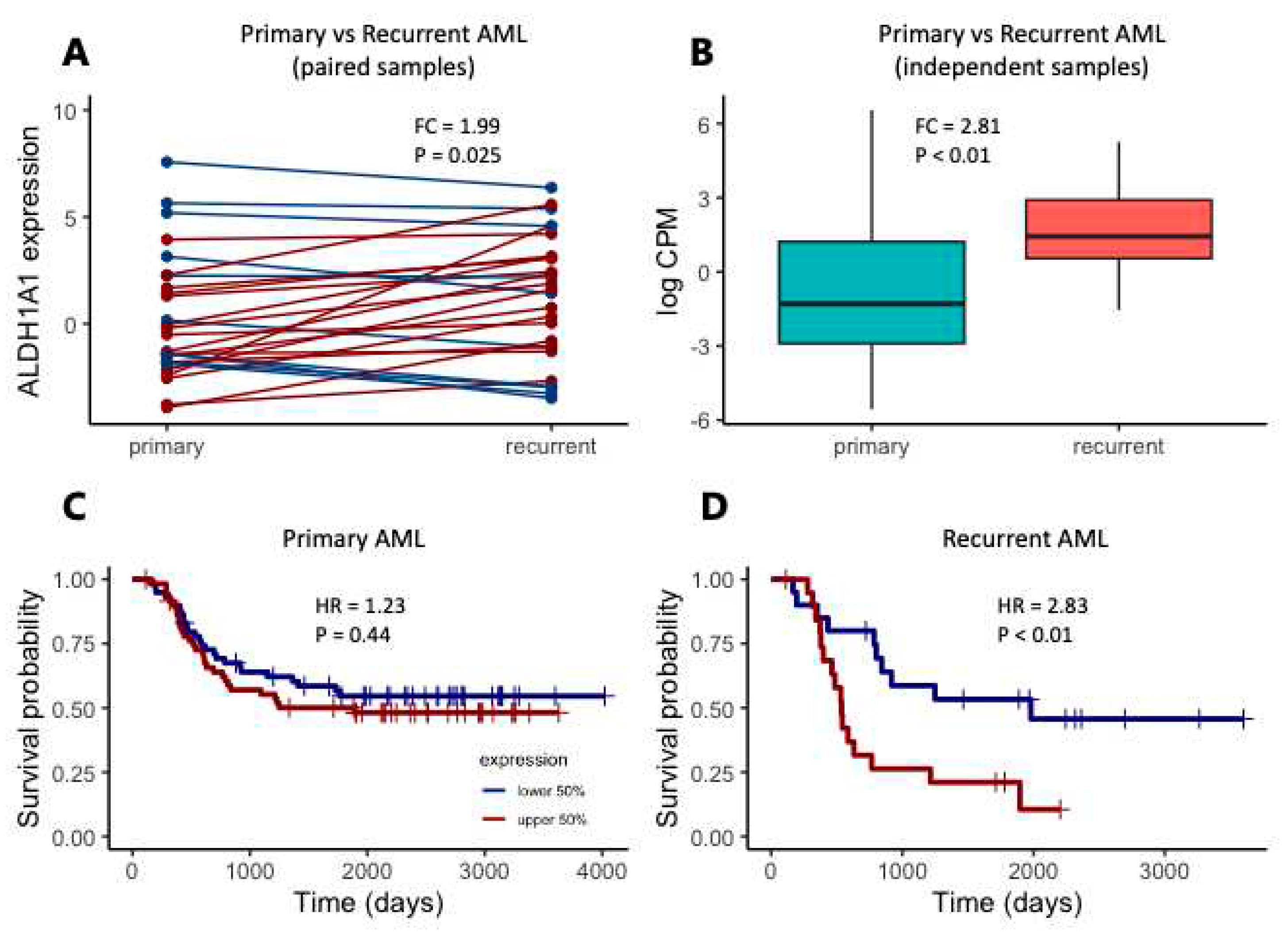
8]. We also focus initially on the TARGET cohort, considering the 27 primary-recurrent paired samples, and 105 additional independent patients with either primary (N = 92) or recurrent (N = 13) tumors. In the paired sample analysis, expression is higher in recurrent AML samples (
Figure 2A, FC = 1.99, P = 0.025). Similarly, when comparing ALDH1A1 expression between additional primary and recurrent tumors, we also find that
ALDH1A1 expression is higher in recurrent tumors than in primary tumors (
Figure 2B, FC = 2.81, P < 0.01). We then evaluated the association of
ALDH1A1 expression with overall survival and found that the hazard ratio is nearly twice as high in patients with recurrent tumors (HR = 2.38, P < 0.01) compared to patients with primary tumors (HR = 1.23, P = 0.44) (
Figure 2C-D). Our findings are consistent with
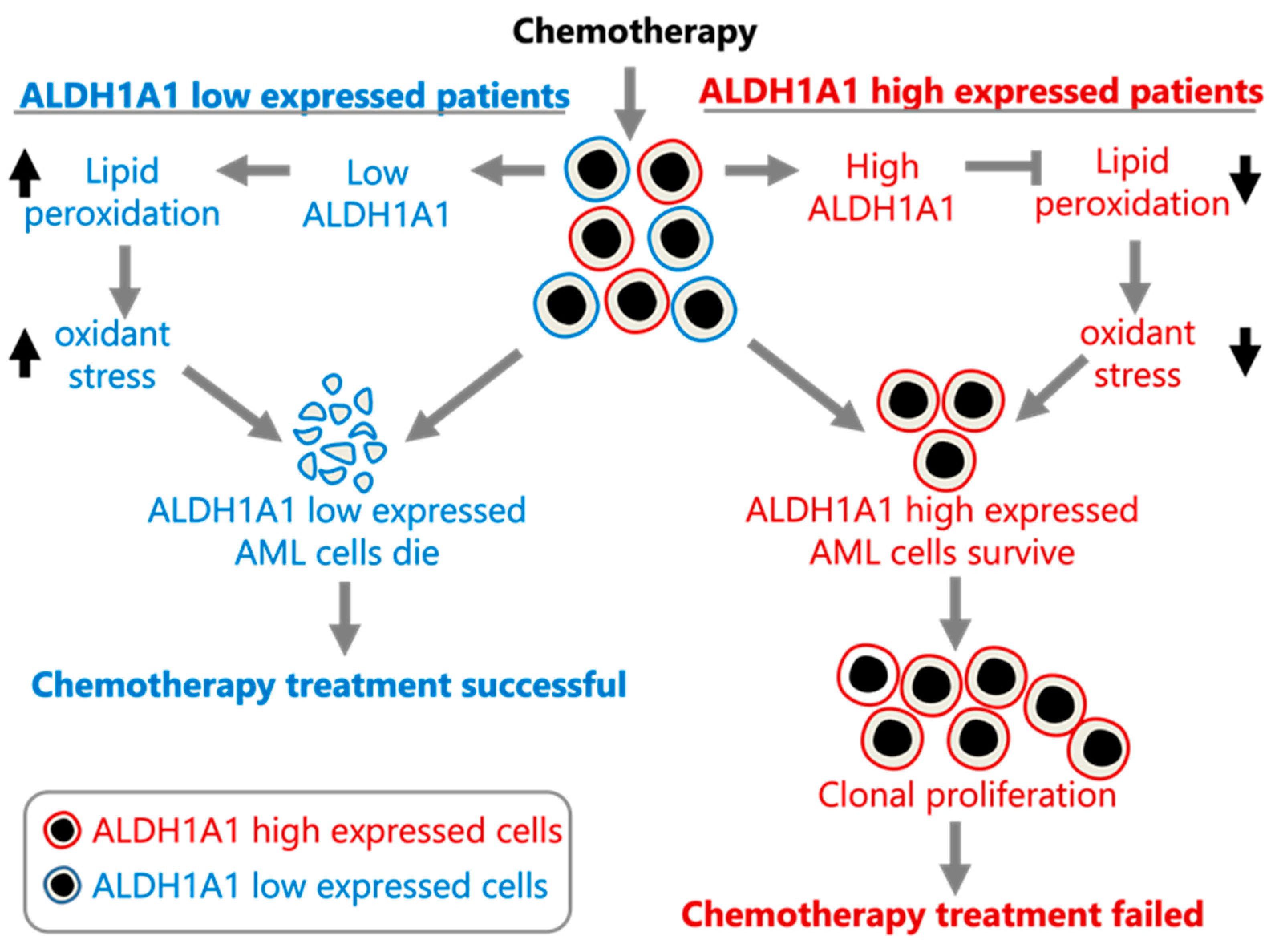
ALDH1A1 gene RNA expression being involved in the development of AML cell clones that are resistant to chemotherapy, which allows them to establish recurrent AML (
Figure 3).
3.2. ALDH1A1 is the ALDH gene with the strongest risk group association
To gain an understanding of the association of
ALDH1A1 gene expression with AML risk group classification and patient survival, we take into consideration 9 independent datasets that are derived from 6 clinical studies of AML, which enrolled a total of over 1000 patients. When examining the relationship of
ALDH1A1 RNA expression level and risk group, we have previously found that expression consistently differs across risk groups (P < 0.01) in the 8 patient cohorts with risk information. In all cases,
ALDH1A1 gene expression is the lowest in the “favorable” or “low” risk group [
9]. When we now compare all nineteen
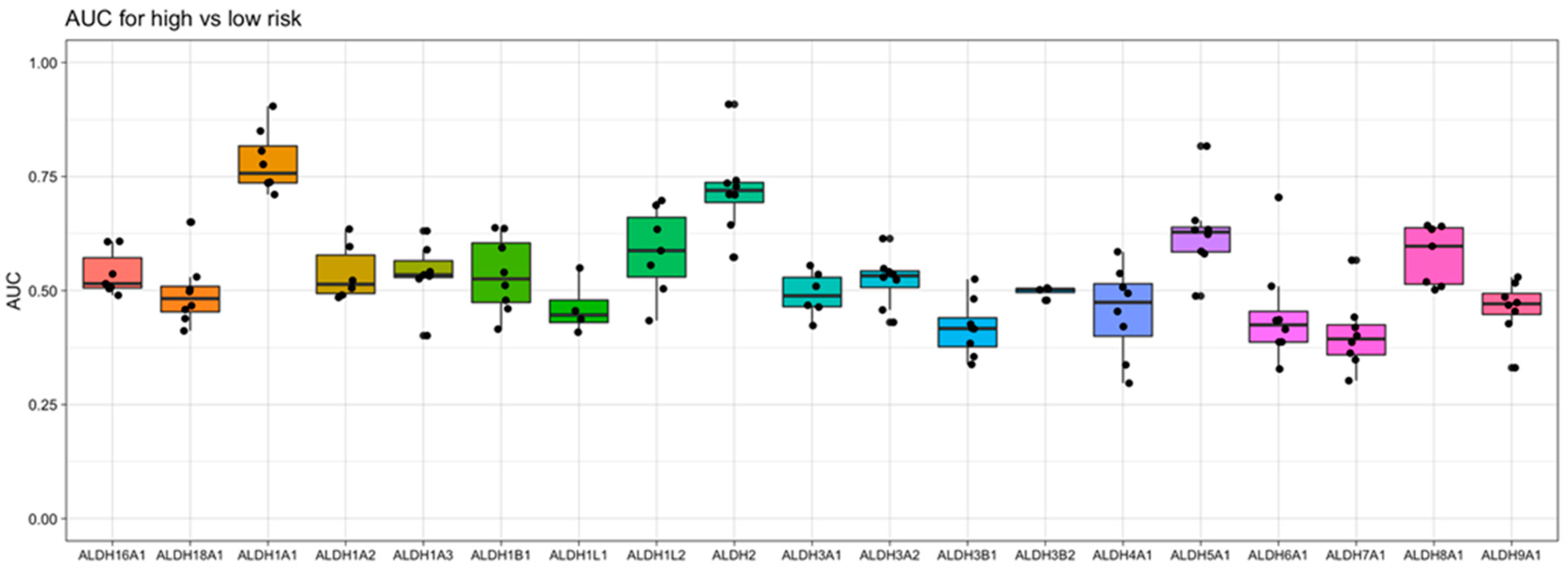
ALDH genes for their association with risk group classification, we observe that
ALDH1A1 gives the strongest diagnostic separation between patients with a favorable prognosis and patients with an adverse prognosis (median AUC = 0.76), while
ALDH2 gives the second strongest separation (median AUC = 0.72) (
Figure 4). In fact,
ALDH1A1 had the 7th highest AUC value of all 20,330 genes profiled (in at least 4 datasets) in AML (
Supporting Table S1).
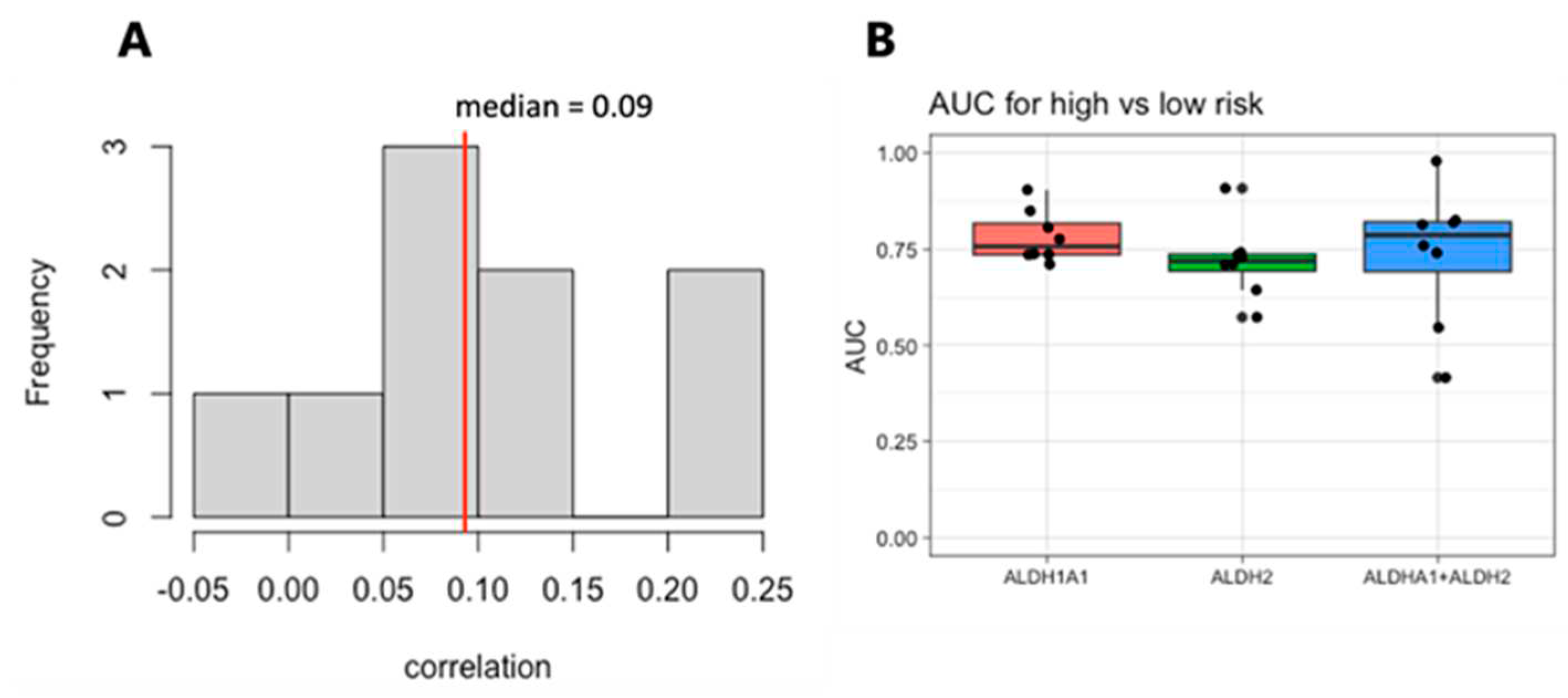
3.3. Combined RNA expression levels from the genes ALDH1A1 and ALDH2 have a stronger risk group and survival association than either gene alone
We next focused on the top two genes,
ALDH1A1 and
ALDH2. The expression of these genes is weakly correlated within each dataset, with a median correlation of 0.09 (
Figure 5A), suggesting that they may be independent markers of risk. We therefore evaluated the combined expression of
ALDH1A1 and
ALDH2 for its association with risk and survival. The sum of the expression values for the RNA from the genes
ALDH1A1 and
ALDH2 shows a stronger risk group association than either one gene alone (
Figure 5B).
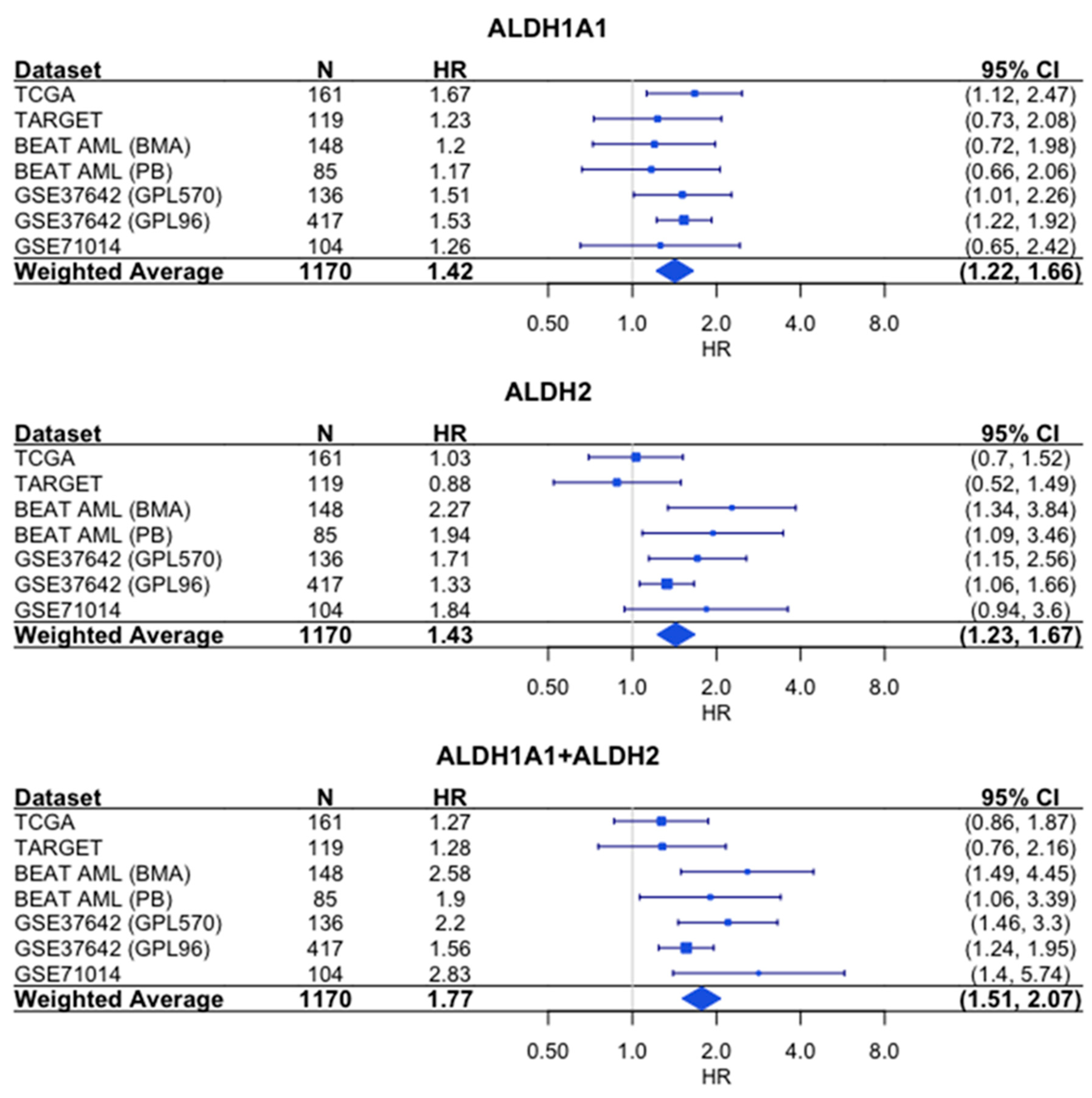
Similarly, the sum of the expression values shows a stronger association with survival than either gene alone, based on the hazard ratios (HR = 1.77 for combined expression, compared to HR = 1.42 for
ALDH1A1 and HR = 1.43 for
ALDH2) (
Figure 6).
4. Discussion
Our analysis of the association of
ALDH1A1 RNA expression levels with recurrent AML leads to the suggestion that ALDH1A1 is involved in AML cell resistance to chemotherapy, potentially through reduction of oxidative stress [
16]. A model consistent with our findings is provided in
Figure 3. If true, then targeting ALDH1A1 in AML may sensitize tumor cells to chemotherapy. In fact, ALDH1A1 inhibition does sensitize colon, breast, pancreas, and ovarian cancer cells to chemotherapy [
17,
18,
19,
20] but further evidence is needed to conclusively determine whether this is also the case in AML. Our analysis of recurrent AML is also based on a single dataset with both primary and recurrent tumors, and it remains to be seen whether this result can be validated in additional prospective studies.
However, the association of ALDH1A1 and ALDH2 RNA expression levels with risk group classification and survival in patients with primary AML draws conclusions from a large sample size (>1000 patients) over 9 independent cohorts (8 cohorts with risk information and 8 cohorts with survival information), and therefore cannot be ignored. With up to 57% of AML patients having refractory AML, or experiencing relapse or death within 12 months of diagnosis [
18], what is important is the fact that both ALDH1A1 and ALDH2 enzymes can be targeted by molecules such as disulfiram that inhibit both activities. Additionally, based on our survival and risk analyses, targeting these genes would likely provide the greatest benefit in high-risk, poor outcome patients whose RNA expression of these genes is up-regulated.
Treatment for refractory and relapsed AML is a challenge, but efforts are underway to target genetic mutations such as FLT3-ITD and IDH1/IDH2 [
19]. ALDH1A1 and ALDH2 can also be targeted. One molecule that targets ALDH1A1 is disulfiram, a substance approved for maintenance of abstinence from alcohol. Disulfiram has been shown to target AML stem cells in cell lines and in primary AML samples [
20], and evaluation of disulfiram in combination with chemotherapy has shown promise in other cancers. In a phase II clinical trial for patients with metastatic non-small cell lung cancer, patients receiving disulfiram in addition to cisplatin and vinorelbine chemotherapy had a modest improvement in survival (10 months vs 7.1 months) compared to patients receiving chemotherapy alone [
21]. Furthermore, in a phase II clinical trial for patients with recurrent temozolomide (TMZ)-resistant glioblastoma, 14% of patients receiving disulfiram in addition to TMZ had clinical benefit, though the objective response rate was 0 [
22]. Importantly, neither study selected patients on the basis of biomarkers that might predict disulfiram efficacy.
While disulfiram mainly inhibits ALDH1A1, in the organism it is readily metabolized to substances that inhibit mainly ALDH2 [
23]. Therefore, patients with high ALDH1A1 or ALDH2 activity would be expected to benefit from disulfiram treatment. The challenge would be to optimize delivery of disulfiram to eradicate leukemia cell clones that escaped the cytotoxic effects of chemotherapy.
Additionally, other molecules exist that target more than one member of the ALDH family. Examples are diethylaminobenzaldehyde (DEAB), which can induce the expansion of normal human hematopoietic stem cells [
24], and dimethyl ampal thiolester (DIMATE), which can eradicate leukemia stem cells while sparing normal progenitors, both
in vitro as well as in mouse xenografts of human AML cells [
25]. Additional ALDH2 inhibitors include daidzin, an isoflavone found in the kudzu plant [
26], and CVT-10216, which is derived from daidzin [
27]. However currently only disulfiram is approved for clinical use, even though it needs to be repurposed for AML.
5. Conclusions
Chemotherapy is one of the major weapons in the fight against cancer. However, in a significant proportion of patients, some sub-populations of cancer cells develop various molecular and/or cellular resistance mechanisms to these agents and thereby treatment fails. Consequently, the sensitive cells die depending on the chemotherapy treatment, whereas the resistant cell sub-populations survive and proliferate, and the disease recurs.
Our study suggests that ALDH family member ALDH1A1 is one of the likely causes of recurrent AML, and also that ALDH1A1 is the ALDH gene with the highest association with patient risk group classification during primary AML. Drugs that inhibit both the ALDH1A1 and ALDH2 enzymes are potential preclinical development candidates for AML, and our results suggest that simultaneous targeting of both ALDH1A1 and ALDH2 will be more efficacious than targeting either enzyme alone.
Supplementary Materials
The following supporting information can be downloaded at the website of this paper posted on Preprints.org. Table S1: All genes evaluated as markers of risk in AML.
Author Contributions
Conceptualization, G.M.D., V.T., L.V., and S.V.; methodology, G.M.D.; formal analysis, G.M.D; data curation, G.M.D. and S.V.; writing—original draft preparation, G.M.D., V.T., L.V., and S.V; writing—review and editing, G.M.D., V.T., L.V., and S.V. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Data Availability Statement
Acknowledgments
The authors wish to thank the AMLCG study group for help in this project.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Dancik, G.M.; Varisli, L.; Vlahopoulos, S.A. The Molecular Context of Oxidant Stress Response in Cancer Establishes ALDH1A1 as a Critical Target: What This Means for Acute Myeloid Leukemia. Int. J. Mol. Sci. 2023, 24, 9372. [Google Scholar] [CrossRef]
- Schuurhuis, G.J.; Meel, M.H.; Wouters, F.; Min, L.A.; Terwijn, M.; de Jonge, N.A.; Kelder, A.; Snel, A.N.; Zweegman, S.; Ossenkoppele, G.J.; et al. Normal Hematopoietic Stem Cells within the AML Bone Marrow Have a Distinct and Higher ALDH Activity Level than Co-Existing Leukemic Stem Cells. PloS One 2013, 8, e78897. [Google Scholar] [CrossRef]
- Smith, C.; Gasparetto, M.; Humphries, K.; Pollyea, D.A.; Vasiliou, V.; Jordan, C.T. Aldehyde Dehydrogenases in Acute Myeloid Leukemia. Ann. N. Y. Acad. Sci. 2014, 1310, 58–68. [Google Scholar] [CrossRef]
- Hoang, V.T.; Hoffmann, I.; Borowski, K.; Zepeda-Moreno, A.; Ran, D.; Buss, E.C.; Wuchter, P.; Eckstein, V.; Ho, A.D. Identification and Separation of Normal Hematopoietic Stem Cells and Leukemia Stem Cells from Patients with Acute Myeloid Leukemia. Methods Mol. Biol. Clifton NJ 2013, 1035, 217–230. [Google Scholar] [CrossRef]
- Yang, X.; Yao, R.; Wang, H. Update of ALDH as a Potential Biomarker and Therapeutic Target for AML. BioMed Res. Int. 2018, 2018, 9192104. [Google Scholar] [CrossRef] [PubMed]
- Pearce, D.J.; Taussig, D.; Simpson, C.; Allen, K.; Rohatiner, A.Z.; Lister, T.A.; Bonnet, D. Characterization of Cells with a High Aldehyde Dehydrogenase Activity from Cord Blood and Acute Myeloid Leukemia Samples. Stem Cells Dayt. Ohio 2005, 23, 752–760. [Google Scholar] [CrossRef] [PubMed]
- Gasparetto, M.; Pei, S.; Minhajuddin, M.; Khan, N.; Pollyea, D.A.; Myers, J.R.; Ashton, J.M.; Becker, M.W.; Vasiliou, V.; Humphries, K.R.; et al. Targeted Therapy for a Subset of Acute Myeloid Leukemias That Lack Expression of Aldehyde Dehydrogenase 1A1. Haematologica 2017, 102, 1054–1065. [Google Scholar] [CrossRef] [PubMed]
- Dancik, G.M.; Voutsas, I.F.; Vlahopoulos, S. Aldehyde Dehydrogenase Enzyme Functions in Acute Leukemia Stem Cells. Front. Biosci. Sch. Ed. 2022, 14, 8. [Google Scholar] [CrossRef]
- Dancik, G.M.; Voutsas, I.F.; Vlahopoulos, S. Lower RNA Expression of ALDH1A1 Distinguishes the Favorable Risk Group in Acute Myeloid Leukemia. Mol. Biol. Rep. 2022, 49, 3321–3331. [Google Scholar] [CrossRef]
- Barrett, T.; Wilhite, S.E.; Ledoux, P.; Evangelista, C.; Kim, I.F.; Tomashevsky, M.; Marshall, K.A.; Phillippy, K.H.; Sherman, P.M.; Holko, M.; et al. NCBI GEO: Archive for Functional Genomics Data Sets--Update. Nucleic Acids Res. 2013, 41, D991–995. [Google Scholar] [CrossRef]
- Jensen, M.A.; Ferretti, V.; Grossman, R.L.; Staudt, L.M. The NCI Genomic Data Commons as an Engine for Precision Medicine. Blood 2017, 130, 453–459. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Aksoy, B.A.; Dogrusoz, U.; Dresdner, G.; Gross, B.; Sumer, S.O.; Sun, Y.; Jacobsen, A.; Sinha, R.; Larsson, E.; et al. Integrative Analysis of Complex Cancer Genomics and Clinical Profiles Using the CBioPortal. Sci. Signal. 2013, 6, pl1. [Google Scholar] [CrossRef]
- Robinson, M.D.; Oshlack, A. A Scaling Normalization Method for Differential Expression Analysis of RNA-Seq Data. Genome Biol. 2010, 11, R25. [Google Scholar] [CrossRef]
- Radivoyevitch, T. AMLbeatR 2019.
- Sing, T.; Sander, O.; Beerenwinkel, N.; Lengauer, T. ROCR: Visualizing Classifier Performance in R. Bioinformatics 2005, 21, 3940–3941. [Google Scholar] [CrossRef] [PubMed]
- Calleja, L.F.; Yoval-Sánchez, B.; Hernández-Esquivel, L.; Gallardo-Pérez, J.C.; Sosa-Garrocho, M.; Marín-Hernández, Á.; Jasso-Chávez, R.; Macías-Silva, M.; Salud Rodríguez-Zavala, J. Activation of ALDH1A1 by Omeprazole Reduces Cell Oxidative Stress Damage. FEBS J. 2021, 288, 4064–4080. [Google Scholar] [CrossRef]
- Kozovska, Z.; Patsalias, A.; Bajzik, V.; Durinikova, E.; Demkova, L.; Jargasova, S.; Smolkova, B.; Plava, J.; Kucerova, L.; Matuskova, M. ALDH1A Inhibition Sensitizes Colon Cancer Cells to Chemotherapy. BMC Cancer 2018, 18, 656. [Google Scholar] [CrossRef] [PubMed]
- Croker, A.K.; Rodriguez-Torres, M.; Xia, Y.; Pardhan, S.; Leong, H.S.; Lewis, J.D.; Allan, A.L. Differential Functional Roles of ALDH1A1 and ALDH1A3 in Mediating Metastatic Behavior and Therapy Resistance of Human Breast Cancer Cells. Int. J. Mol. Sci. 2017, 18, 2039. [Google Scholar] [CrossRef] [PubMed]
- Duong, H.-Q.; Yi, Y.W.; Kang, H.J.; Bae, I.; Jang, Y.-J.; Kwak, S.-J.; Seong, Y.-S. Combination of Dasatinib and Gemcitabine Reduces the ALDH1A1 Expression and the Proliferation of Gemcitabine-Resistant Pancreatic Cancer MIA PaCa-2 Cells. Int. J. Oncol. 2014, 44, 2132–2138. [Google Scholar] [CrossRef]
- Landen, C.N.; Goodman, B.; Katre, A.A.; Steg, A.D.; Nick, A.M.; Stone, R.L.; Miller, L.D.; Mejia, P.V.; Jennings, N.B.; Gershenson, D.M.; et al. Targeting Aldehyde Dehydrogenase Cancer Stem Cells in Ovarian Cancer. Mol. Cancer Ther. 2010, 9, 3186–3199. [Google Scholar] [CrossRef]
- Walter, R.B.; Othus, M.; Burnett, A.K.; Löwenberg, B.; Kantarjian, H.M.; Ossenkoppele, G.J.; Hills, R.K.; Ravandi, F.; Pabst, T.; Evans, A.; et al. Resistance Prediction in AML: Analysis of 4601 Patients from MRC/NCRI, HOVON/SAKK, SWOG and MD Anderson Cancer Center. Leukemia 2015, 29, 312–320. [Google Scholar] [CrossRef]
- Ma, J.; Ge, Z. Recent Advances of Targeted Therapy in Relapsed/Refractory Acute Myeloid Leukemia. Bosn. J. Basic Med. Sci. 2021, 21, 409–421. [Google Scholar] [CrossRef] [PubMed]
- Xu, B.; Wang, S.; Li, R.; Chen, K.; He, L.; Deng, M.; Kannappan, V.; Zha, J.; Dong, H.; Wang, W. Disulfiram/Copper Selectively Eradicates AML Leukemia Stem Cells in Vitro and in Vivo by Simultaneous Induction of ROS-JNK and Inhibition of NF-ΚB and Nrf2. Cell Death Dis. 2017, 8, e2797. [Google Scholar] [CrossRef] [PubMed]
- Nechushtan, H.; Hamamreh, Y.; Nidal, S.; Gotfried, M.; Baron, A.; Shalev, Y.I.; Nisman, B.; Peretz, T.; Peylan-Ramu, N. A Phase IIb Trial Assessing the Addition of Disulfiram to Chemotherapy for the Treatment of Metastatic Non-Small Cell Lung Cancer. The Oncologist 2015, 20, 366–367. [Google Scholar] [CrossRef]
- Huang, J.; Chaudhary, R.; Cohen, A.L.; Fink, K.; Goldlust, S.; Boockvar, J.; Chinnaiyan, P.; Wan, L.; Marcus, S.; Campian, J.L. A Multicenter Phase II Study of Temozolomide plus Disulfiram and Copper for Recurrent Temozolomide-Resistant Glioblastoma. J. Neurooncol. 2019, 142, 537–544. [Google Scholar] [CrossRef] [PubMed]
- Lam, J.P.; Mays, D.C.; Lipsky, J.J. Inhibition of Recombinant Human Mitochondrial and Cytosolic Aldehyde Dehydrogenases by Two Candidates for the Active Metabolites of Disulfiram. Biochemistry 1997, 36, 13748–13754. [Google Scholar] [CrossRef]
- Chute, J.P.; Muramoto, G.G.; Whitesides, J.; Colvin, M.; Safi, R.; Chao, N.J.; McDonnell, D.P. Inhibition of Aldehyde Dehydrogenase and Retinoid Signaling Induces the Expansion of Human Hematopoietic Stem Cells. Proc. Natl. Acad. Sci. U. S. A. 2006, 103, 11707–11712. [Google Scholar] [CrossRef]
- Venton, G.; Pérez-Alea, M.; Baier, C.; Fournet, G.; Quash, G.; Labiad, Y.; Martin, G.; Sanderson, F.; Poullin, P.; Suchon, P.; et al. Aldehyde Dehydrogenases Inhibition Eradicates Leukemia Stem Cells While Sparing Normal Progenitors. Blood Cancer J. 2016, 6, e469. [Google Scholar] [CrossRef]
- Lowe, E.D.; Gao, G.-Y.; Johnson, L.N.; Keung, W.M. Structure of Daidzin, a Naturally Occurring Anti-Alcohol-Addiction Agent, in Complex with Human Mitochondrial Aldehyde Dehydrogenase. J. Med. Chem. 2008, 51, 4482–4487. [Google Scholar] [CrossRef]
- Arolfo, M.P.; Overstreet, D.H.; Yao, L.; Fan, P.; Lawrence, A.J.; Tao, G.; Keung, W.-M.; Vallee, B.L.; Olive, M.F.; Gass, J.T.; et al. Suppression of Heavy Drinking and Alcohol Seeking by a Selective ALDH-2 Inhibitor. Alcohol. Clin. Exp. Res. 2009, 33, 1935–1944. [Google Scholar] [CrossRef]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).