Submitted:
03 May 2026
Posted:
05 May 2026
You are already at the latest version
Abstract
Keywords:
Introduction
Theory of Generalization
Fitness Landscapes
Discussion
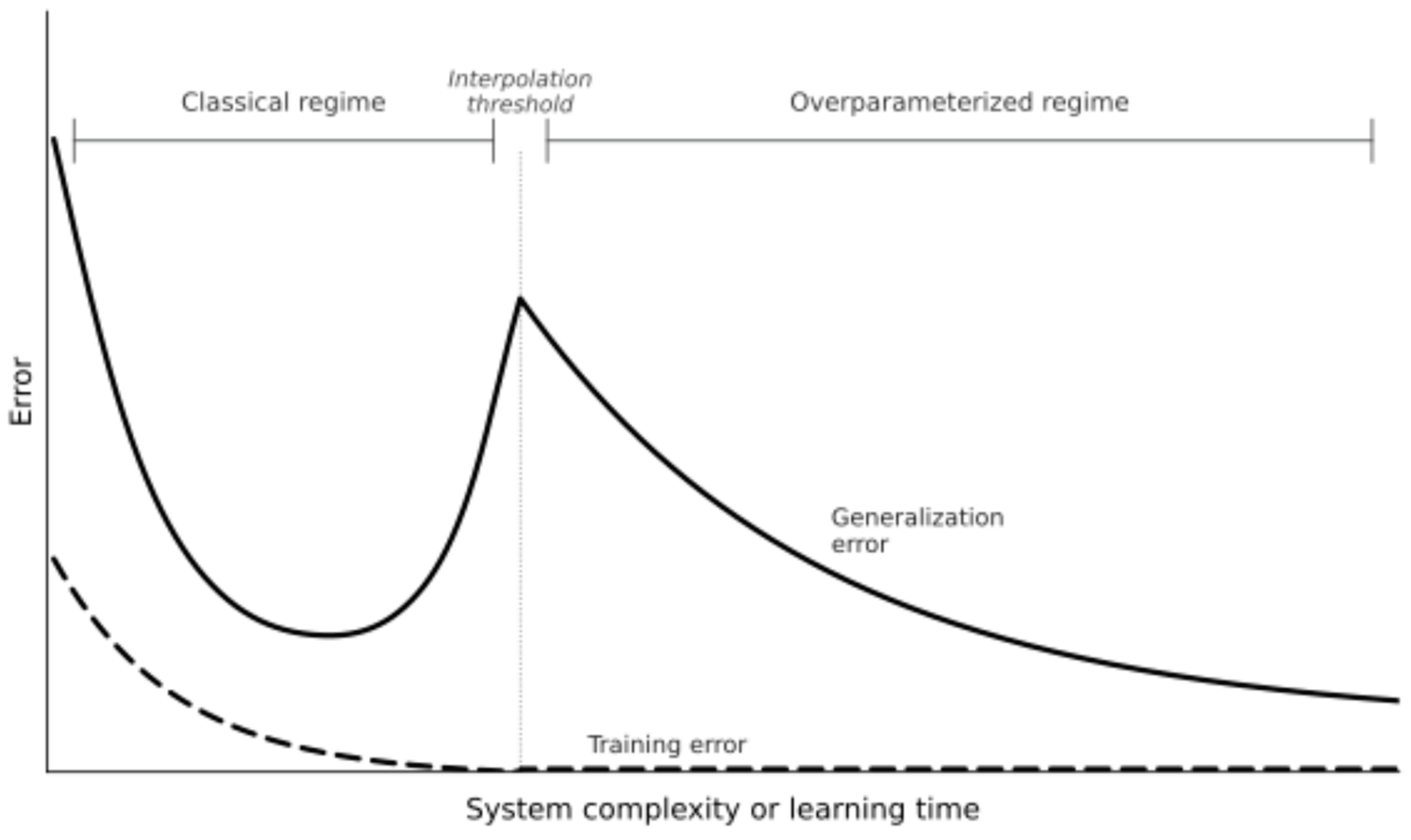
Acknowledgments
Data and code availability
References
- Valiant, L. Probably Approximately Correct: Nature’s Algorithms for Learning and Prospering in a Complex World; Basic Books, 2013. [Google Scholar]
- Watson, R.A.; Szathmáry, E. How can evolution learn? Trends Ecol. Evol. 2016, 31, 147–157. [Google Scholar] [CrossRef] [PubMed]
- Belkin, M.; et al. Reconciling modern machine-learning practice and the classical bias-variance trade-off. Proceedings of the National Academy of Sciences of the United States of America 2019, 116, 15849–15854. [Google Scholar] [CrossRef] [PubMed]
- Nakkiran, P.; et al. Deep double descent: where bigger models and more data hurt. J. Stat. Mech. Theory Exp. 2021, 2021, 124003. [Google Scholar] [CrossRef]
- Parter, M.; et al. Facilitated variation: How evolution learns from past environments to generalize to new environments. PLoS Comput. Biol. 2008, 4, e1000206. [Google Scholar] [CrossRef] [PubMed]
- Kouvaris, K.; et al. How evolution learns to generalise: Using the principles of learning theory to understand the evolution of developmental organisation. PLoS Comput Biol. 2017, 13, e1005358. [Google Scholar] [CrossRef] [PubMed]
- Wilson, A.G. Position: Deep learning is not so mysterious or different. 2025. [Google Scholar]
- Transtrum, M.K.; et al. Generalized aliasing explains double descent and informs model design. Phys. Rev. Res. 2025, 7, 043268. [Google Scholar] [CrossRef]
- Schaeffer, R.; et al. Double descent demystified: Identifying, interpreting & ablating the sources of a deep learning puzzle. arXiv 2023, arXiv:2303.14151. [Google Scholar] [CrossRef]
- Payne, J.L.; Wagner, A. The causes of evolvability and their evolution. Nat. Rev. Genet. 2019, 20, 24–38. [Google Scholar] [CrossRef] [PubMed]
- Wagner, G.P.; Altenberg, L. Complex adaptations and the evolution of evolvability. Evolution 1996, 50, 967–976. [Google Scholar] [CrossRef] [PubMed]
- Frank, S.A. Circuit design in biology and machine learning. II. Anomaly detection. Entropy 2025, 27, 896. [Google Scholar] [CrossRef] [PubMed]
- Frank, S.A. Circuit design in biology and machine learning. I. Random networks and dimensional reduction. Evolution 2025, 79, 1403–1418. [Google Scholar] [CrossRef] [PubMed]
- Xue, B.K.; et al. Environment-to-phenotype mapping and adaptation strategies in varying environments. Proceedings of the National Academy of Sciences of the United States of America 2019, 116, 13847–13855. [Google Scholar] [CrossRef] [PubMed]
- Gavrilets, S. Fitness Landscapes and the Origin of Species; Princeton University Press, 2004. [Google Scholar]
- Davidson, E.H. The regulatory genome: gene regulatory networks in development and evolution; Academic, 2006. [Google Scholar]
- Wagner, A. Robustness and Evolvability in Living Systems; Princeton University Press, 2005. [Google Scholar]
- Kirschner, M.; Gerhart, J. The Plausibility of Life: Resolving Darwin’s Dilemma; Yale University Press, 2005. [Google Scholar]
- Frank, S.A. Robustness and complexity. Cell Syst. 2023, 14, 1015–1020. [Google Scholar] [CrossRef] [PubMed]
- Elad, M.; et al. Another step toward demystifying deep neural networks. Proc. Natl. Acad. Sci. USA 2020, 117, 27070–27072. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



