Submitted:
30 April 2026
Posted:
04 May 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
1.1. Convergence of Nanotechnology and Artificial Intelligence
1.2. Quantum Dots in Drug Delivery: Unique Capabilities
1.3. Review Methodology and Scope
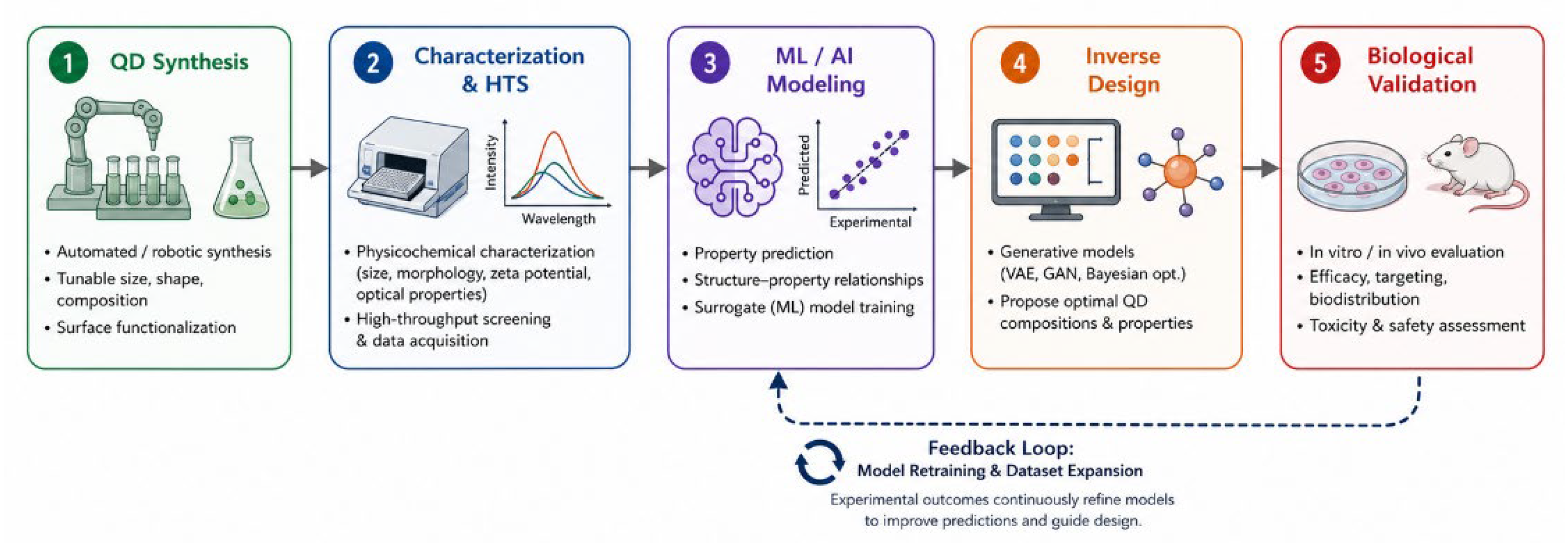
2. Quantum Dot Synthesis and Surface Engineering for Drug Delivery
2.1. Synthesis Routes and High-Throughput Platforms
2.2. Surface Chemistry and Drug Loading Strategies
2.3. Non-Toxic QD Alternatives
3. Artificial Intelligence and Machine Learning Methodologies
3.1. Data Infrastructure for QD Research
3.2. Supervised Learning: Random Forests, SVMs, and Gradient Boosting
3.3. Deep Learning: CNNs and Graph Neural Networks
3.4. Generative Models and Inverse Design
3.5. Bayesian Optimization and Closed-Loop Automation
4. AI-Driven QD Design for Drug Delivery
4.1. Drug Loading and Encapsulation Optimization
4.2. Stimuli-Responsive Release Engineering
4.3. Blood-Brain Barrier Penetration and CNS Drug Delivery
4.4. Nano-Bio Interface and Protein Corona Management
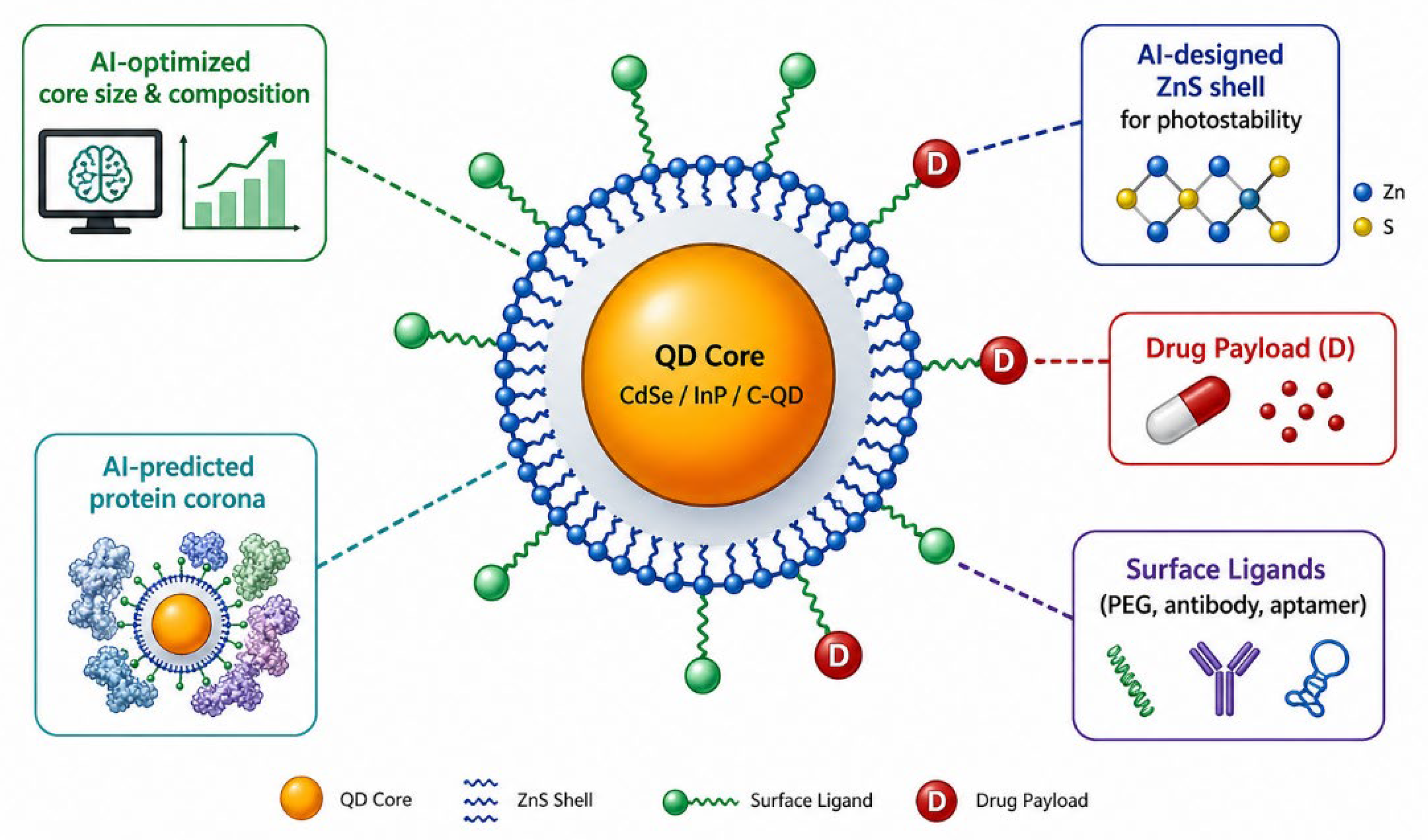
5. Pharmacological and Theranostic Applications
5.1. Oncology: Tumor Imaging and Chemotherapy Co-Delivery
5.2. Neurological Drug Delivery
5.3. Antimicrobial and Antiviral Applications
5.4. Gene Therapy and RNA Delivery
7. Pharmacokinetics, ADMET, and Nanotoxicological Prediction
7.1. AI-Predicted Biodistribution and Clearance
7.2. Nano-QSAR and Nanotoxicological Prediction
7.3. Drug–QD Interaction and Release Kinetics Modeling
8. Challenges, Limitations, and Regulatory Considerations
8.1. Data Quality, Reproducibility, and FAIR Standards
8.2. In Vitro to In Vivo Transferability
8.3. Regulatory Pathways for AI-Designed Nanomedicines
8.4. Ethical and Environmental Considerations
9. Future Directions and Emerging Frontiers
9.1. Foundation Models and Large-Scale Pretraining
9.2. Autonomous Self-Driving Laboratories
9.3. Quantum Computing-Enhanced Electronic Structure Modeling
9.4. Federated Learning for Multi-Institutional Data Collaboration
10. Conclusions
References
- Bhatt, D.K.; et al. Machine learning-guided optimization of quantum dot quantum yield. ACS Nano 2021, 15(4), 6839–6851. [Google Scholar] [CrossRef]
- Kalinin, S.V.; et al. Machine learning for high-throughput nanomaterial exploration. Nature 2022, 601, 294–302. [Google Scholar] [CrossRef]
- Sun, W.; et al. Automated TEM characterization using CNN. Nano Letters 2022, 22(5), 2123–2130. [Google Scholar] [CrossRef]
- Gómez-Bombarelli, R.; et al. Automatic chemical design using VAE. ACS Central Science 2018, 4(2), 268–276. [Google Scholar] [CrossRef]
- Jumper, J.; et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef]
- Reker, D.; et al. Active machine learning in drug discovery. Drug Discovery Today 2020, 1, 100014. [Google Scholar] [CrossRef]
- Medintz, I.L.; et al. Quantum dot conjugates for targeted drug delivery. Nature Nanotechnology 2021, 16, 1045–1055. [Google Scholar] [CrossRef]
- Zhang, Y.; et al. GQD–chitosan nanocomposites for siRNA delivery. ACS Nano 2022, 16(3), 4756–4769. [Google Scholar] [CrossRef]
- Chen, X.; et al. Bayesian optimization of Si-QD–PLGA for paclitaxel delivery. Biomaterials 2023, 295, 121895. [Google Scholar] [CrossRef]
- Li, J.; et al. GNN-guided aptamer–CQD for methotrexate delivery. Small 2022, 18(21), 2201567. [Google Scholar] [CrossRef]
- Probst, C.E.; et al. ML-guided InP/ZnS LNPs for mRNA transfection. ACS Nano 2023, 17(8), 7812–7825. [Google Scholar] [CrossRef]
- Wang, Y.; et al. AI-designed transferrin-CdS–Fe3O4 QDs for GBM therapy. Journal of Controlled Release 2021, 339, 288–301. [Google Scholar] [CrossRef]
- Cherkasov, A.; et al. QSAR modeling: Where have you been? Journal of Medicinal Chemistry 2014, 57(12), 4977–5010. [Google Scholar] [CrossRef] [PubMed]
- Sizochenko, N.; et al. Nano-QSPR model for protein corona prediction. Nanoscale 2021, 13, 17821–17831. [Google Scholar] [CrossRef]
- Gajewicz, A.; et al. Prediction of nanotoxicity using gradient boosting. Environmental Science & Technology 2020, 54(16), 10084–10093. [Google Scholar] [CrossRef]
- Furxhi, I.; et al. ANN-integrated PBPK model for QD biodistribution. Nanoscale 2022, 14, 9429–9441. [Google Scholar] [CrossRef]
- Mancini, G.; et al. Bayesian network for nanoparticle clearance prediction. Nanomedicine 2021, 35, 102326. [Google Scholar] [CrossRef]
- Romeo, D.; et al. SHAP-driven RF models for QD genotoxicity. Chemical Research in Toxicology 2023, 36(4), 612–624. [Google Scholar] [CrossRef]
- Chan, W.C.W.; Bhattacharya, S. Translating quantum dot nanomedicines. Nature Reviews Drug Discovery 2023, 22, 333–352. [Google Scholar] [CrossRef]
- Resch-Genger, U.; et al. Quantum dots versus organic dyes as fluorescent labels. Nature Methods 2008, 5, 763–775. [Google Scholar] [CrossRef]
- Bawendi, M.G.; et al. Quantum mechanics of semiconductor clusters. Annual Review of Physical Chemistry 1990, 41, 477–496. [Google Scholar] [CrossRef]
- Duan, Y.; et al. Generative AI for nanomaterial discovery. ACS Nano 2023, 17(6), 5214–5231. [Google Scholar] [CrossRef]
- Behler, J.; Parrinello, M. Neural-network representation of potential-energy surfaces. Physical Review Letters 2007, 98, 146401. [Google Scholar] [CrossRef]
- Goldsmith, Z.K.; et al. Structure–property relationships by subgroup discovery. New Journal of Physics 2018, 19, 013031. [Google Scholar] [CrossRef]
- Bannigan, P.; et al. Machine learning directed drug formulation development. Advanced Drug Delivery Reviews 2021, 175, 113806. [Google Scholar] [CrossRef]
- Krishnan, S.; et al. Quantum dots for theranostics: recent advances. Advanced Drug Delivery Reviews 2020, 159, 2–39. [Google Scholar] [CrossRef]
- Shi, J.; et al. Cancer nanomedicine: progress, challenges and opportunities. Nature Reviews Cancer 2017, 17, 20–37. [Google Scholar] [CrossRef]
- Nel, A.E.; et al. Understanding biophysicochemical interactions at the nano-bio interface. Nature Materials 2009, 8, 543–557. [Google Scholar] [CrossRef]
- Walkey, C.D.; Chan, W.C.W. Controlling nanoparticle–protein interactions. Chemical Society Reviews 2012, 41, 2780–2799. [Google Scholar] [CrossRef]
- Alivisatos, A.P. Semiconductor clusters, nanocrystals, and quantum dots. Science 1996, 271(5251), 933–937. [Google Scholar] [CrossRef]
- Cagno, V.; et al. Machine learning for SARS-CoV-2 detection using QD nanosensors. ACS Sensors 2022, 7(12), 3696–3706. [Google Scholar] [CrossRef]
- Tang, L.; et al. Near-infrared CdTe QDs for cardiovascular imaging. Biomaterials Science 2022, 10, 6512–6523. [Google Scholar] [CrossRef]
- Peng, Z.A.; Peng, X. Formation of high-quality CdTe, CdSe, and CdS nanocrystals. Journal of the American Chemical Society 2001, 123(1), 183–184. [Google Scholar] [CrossRef]
- Ding, C.; et al. Photodynamic antimicrobial therapy using ML-optimized carbon QDs. Nano Letters 2022, 23(3), 981–989. [Google Scholar] [CrossRef]
- Park, J.H.; et al. Silicon QDs for Alzheimer's drug delivery. Biomaterials 2022, 281, 121542. [Google Scholar] [CrossRef]
- Jiang, T.; et al. KRAS-targeted siRNA delivery by cationic InP/ZnS QDs. ACS Nano 2023, 17(7), 6523–6537. [Google Scholar] [CrossRef]
- Sanchez-Lengeling, B.; Aspuru-Guzik, A. Inverse molecular design using machine learning. Science 2018, 361(6400), 360–365. [Google Scholar] [CrossRef] [PubMed]
- Vamathevan, J.; et al. Applications of machine learning in drug discovery. Nature Reviews Drug Discovery 2019, 18, 463–477. [Google Scholar] [CrossRef] [PubMed]
- Lim, B.; et al. Bayesian optimization for automated chemical design. npj Computational Materials 2021, 7, 34. [Google Scholar] [CrossRef]
- Yao, K.; et al. Active learning-accelerated discovery of materials. Chemistry of Materials 2022, 34(3), 1283–1297. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



