Submitted:
01 May 2026
Posted:
04 May 2026
You are already at the latest version
Abstract

Keywords:
1. Introduction
2. Results
2.1. Signal Model and Sensitivity in D-FFOCT
2.2. Optical Flux Optimization: Ratio-Free D-FFOCT
2.2.1. Conventional NPBS Configuration
2.2.2. Ratio-Free Polarization Architecture
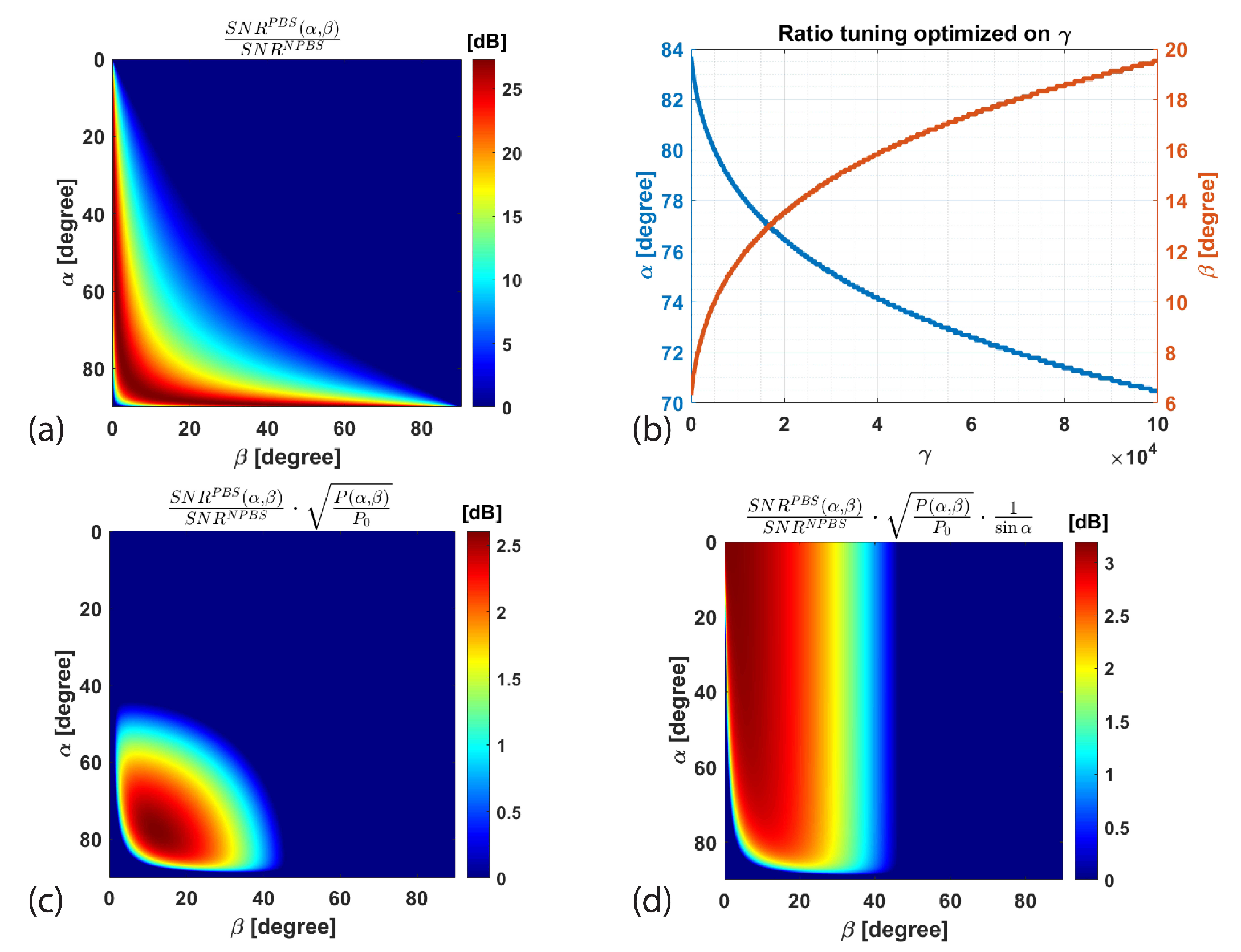
2.3. Sensitivity Gain Mechanisms
2.3.1. Accumulation Gain Under Full-Well Constraint
2.3.2. Dose-Limited Operation
2.3.3. Global Performance
2.4. Biological Validation in Retinal Organoids and Müller Glial Cells
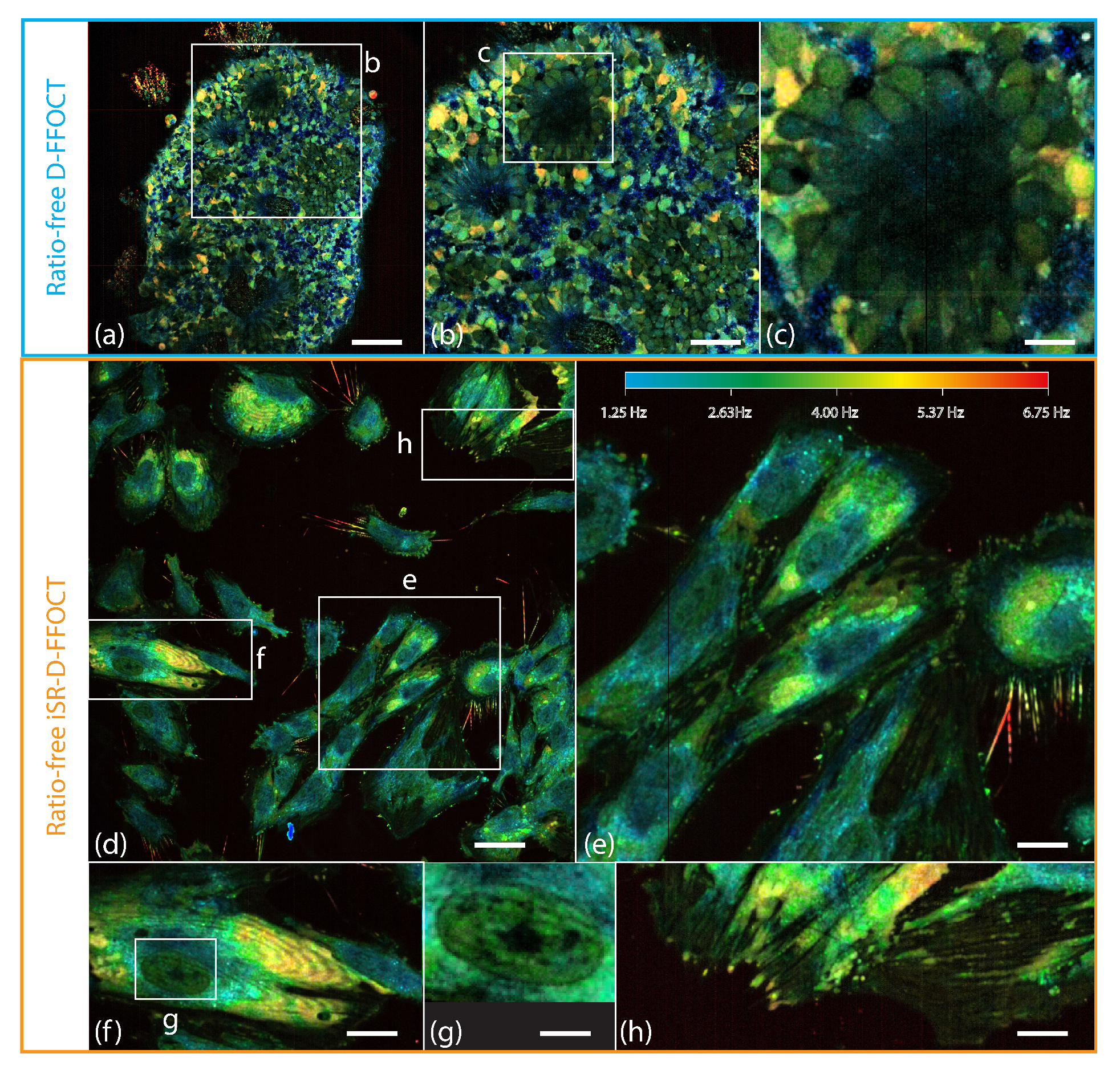
2.4.1. Ratio-Free D-FFOCT Imaging of Retinal Organoids
2.4.2. Interface Self-Referenced D-FFOCT of Müller Glial Cells
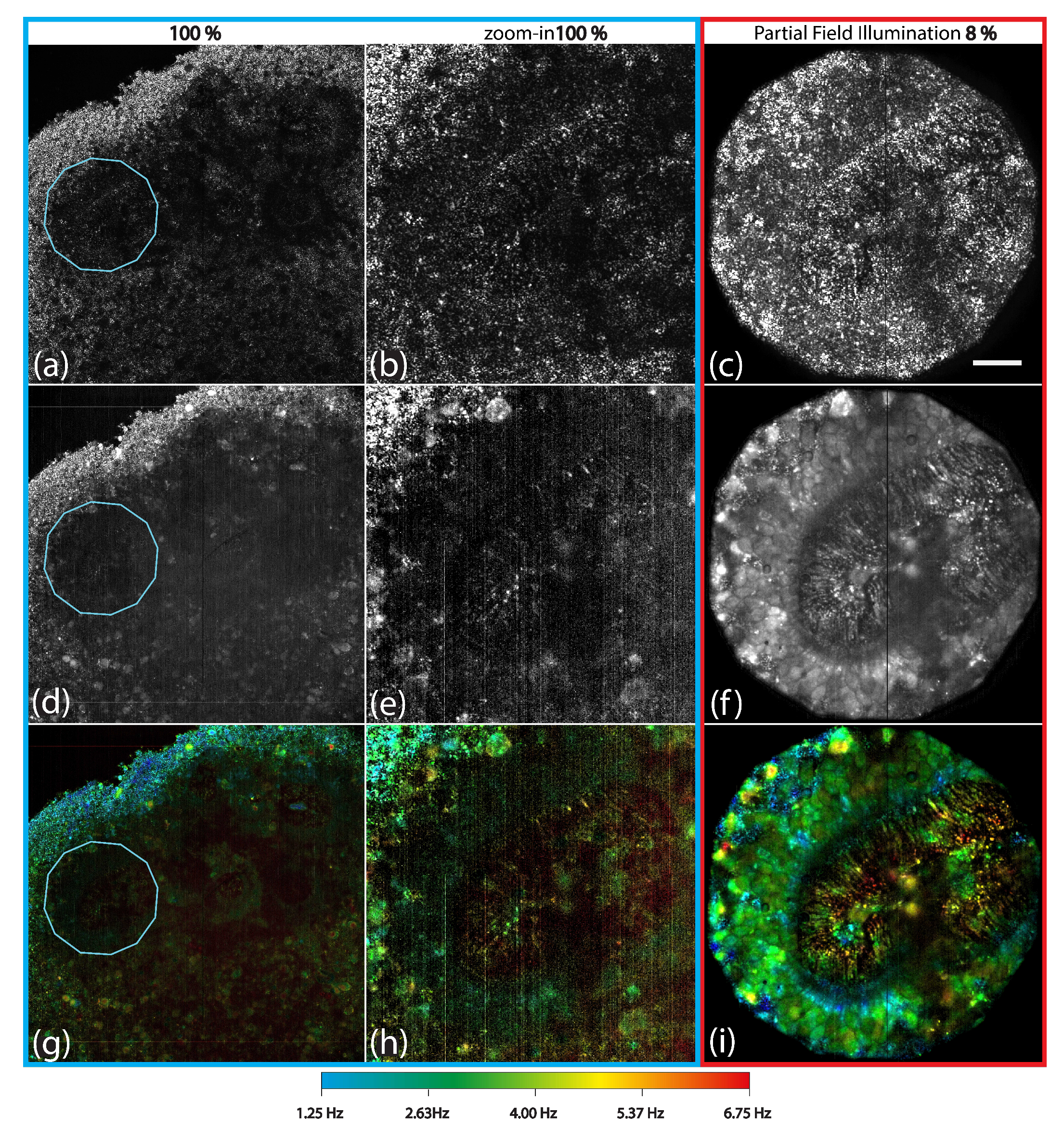
2.5. Multi-Scattering Mitigation Using Partial Field Illumination
2.5.1. Partial Field Illumination Strategy
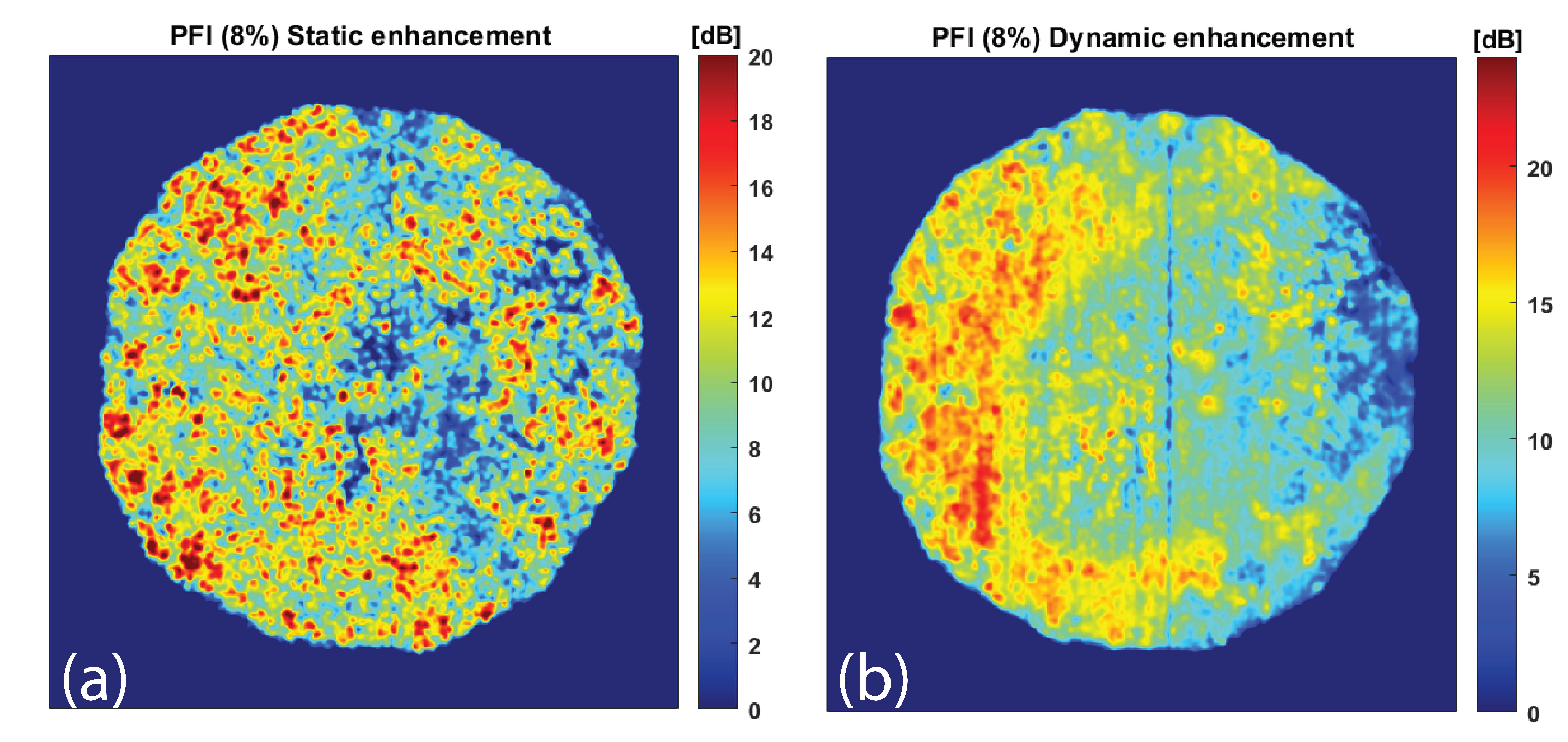
2.5.2. Static and Dynamic Signal Enhancement
2.5.3. Quantitative Sensitivity Gain
3. Discussion
4. Materials and Methods
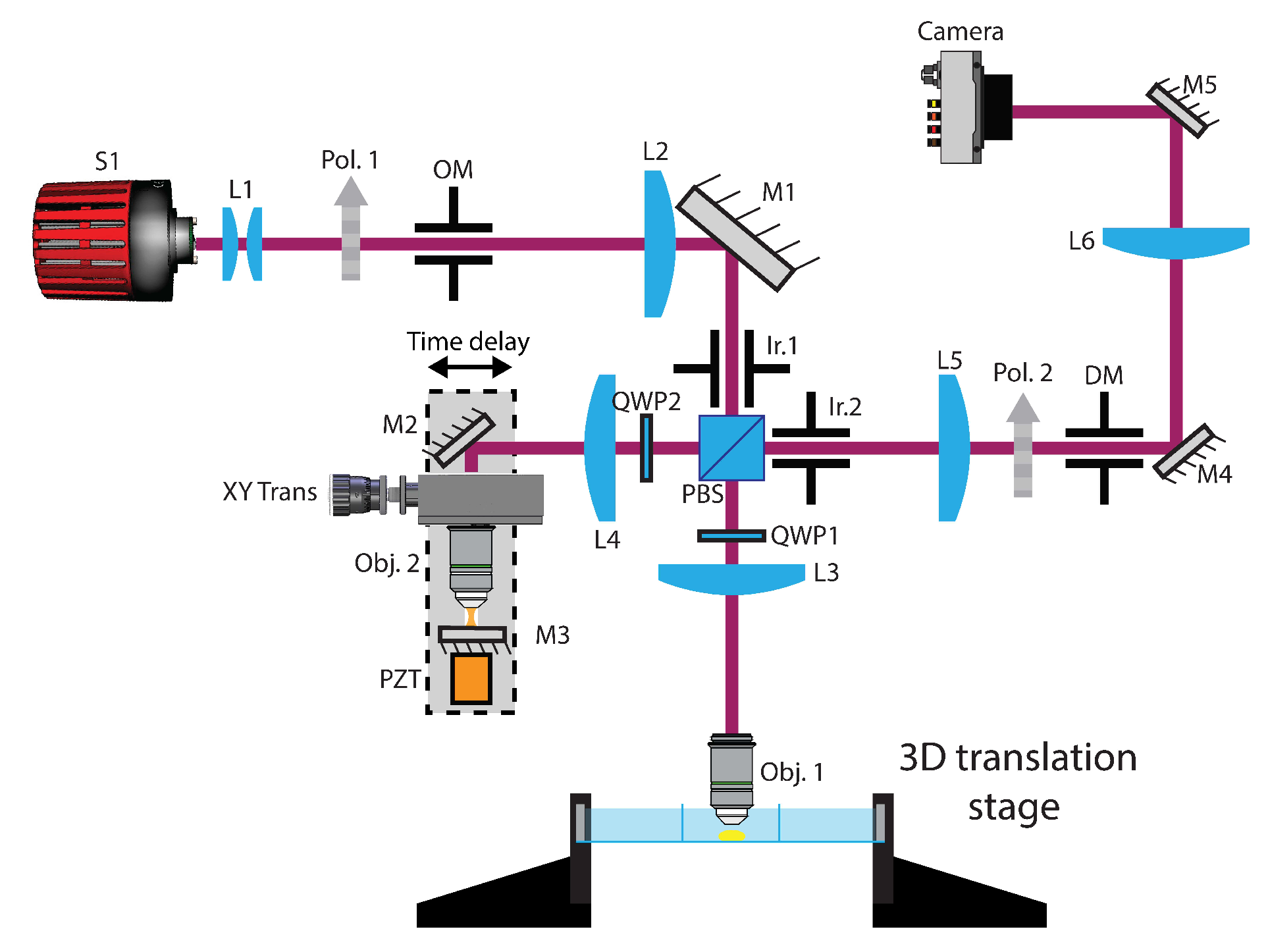
4.1. Optical Setup
4.2. Simulation
4.3. Image Acquisition
4.4. Rolling-Phase Detection and Signal Demodulation
4.5. 2-Phase Imaging (Static Imaging)
4.6. Dynamic Image Rendering
- dynamic magnitude through Phase Fluctuation Index (PhFI),
- quantitative transport speed through mean MSD,
- motility versus Brownian behavior through MSD standard deviation.
- hue: mean MSD (),
- saturation: standard deviation of MSD (),
- brightness: PhFI.
- hue: 0.1% to 99.9%,
- saturation: 5% to 99.9%,
- brightness: 5% to 99.9%.
4.7. PFI Quantification
4.8. Sample Preparation
4.8.1. Retinal Organoids
4.8.2. Müller Glial Cells
5. Code Availability
Author Contributions
Funding
Data Availability Statement
Acknowledgments
References
- Zhang, B.; Korolj, A.; Lai, B. F. L.; Radisic, M. Advances in organ-on-a-chip engineering. Nat. Rev. Mater. 2018, vol. 3(no. 8), 257–278. [Google Scholar] [CrossRef]
- Rios, A. C.; Clevers, H. Imaging organoids: a bright future ahead. Nat. Methods 2018, vol. 15(no. 1), 24–26. [Google Scholar] [CrossRef]
- Fatehullah, A.; Tan, S. H.; Barker, N. Organoids as an in vitro model of human development and disease. Nat. Cell Biol. 2016, vol. 18(no. 3), 246–254. [Google Scholar] [CrossRef]
- Wells, W. A. In vivo and ex vivo microscopy: Moving toward the integration of optical imaging technologies into pathology practice. Arch. Pathol. Lab. Med. 2018, vol. 143(no. 3), 288–298. [Google Scholar] [CrossRef]
- Krishnamurthy, S. Ex vivo microscopy: A promising next-generation digital microscopy tool for surgical pathology practice. Arch. Pathol. Lab. Med. 2019, vol. 143(no. 9), 1058–1068. [Google Scholar] [CrossRef]
- Darche, M. Three-dimensional characterization of developing and adult ocular vasculature in mice using in toto clearing. Commun. Biol. 2022, vol. 5(no. 1). [Google Scholar] [CrossRef] [PubMed]
- Monfort, T. Dynamic full-field optical coherence tomography module adapted to commercial microscopes allows longitudinal in vitro cell culture study. Commun. Biol. 2023, vol. 6(no. 1). [Google Scholar] [CrossRef]
- Sahl, S. J.; Hell, S. W.; Jakobs, S. Fluorescence nanoscopy in cell biology. Nat. Rev. Mol. Cell Biol. 2017, vol. 18(no. 11), 685–701. [Google Scholar] [CrossRef]
- Vazquez, A. The genetics of the p53 pathway, apoptosis and cancer therapy. Nat. Rev. Drug Discov. 2008, vol. 7(no. 12), 979–987. [Google Scholar] [CrossRef]
- Aknoun, S. Quantitative phase microscopy for non-invasive live cell population monitoring. Sci. Rep. 2021, vol. 11(no. 1). [Google Scholar] [CrossRef] [PubMed]
- Caldon, C. E.; Burgess, A. Label free, quantitative single-cell fate tracking of time-lapse movies. MethodsX 2019, vol. 6, 2468–2475. [Google Scholar] [CrossRef]
- Joseph, A.; et al. Label-free imaging of immune cell dynamics in the living retina using adaptive optics. eLife 2020, vol. 9. [Google Scholar] [CrossRef] [PubMed]
- Jeknic, S.; Kudo, T.; Covert, M. W. Techniques for studying decoding of single cell dynamics. Front. Immunol. 2019, vol. 10. [Google Scholar] [CrossRef] [PubMed]
- Avogaro, L. Live-cell imaging reveals the dynamics and function of single-telomere TERRA molecules in cancer cells. RNA Biol. 2018, 1–10. [Google Scholar] [CrossRef]
- Gilmore, A. C. An in vitro tumorigenesis model based on live-cell-generated oxygen and nutrient gradients. Commun. Biol. 2021, vol. 4(no. 1). [Google Scholar] [CrossRef]
- Alzeeb, G. Gastric cancer cell death analyzed by live cell imaging of spheroids. Sci. Rep. 2022, vol. 12(no. 1). [Google Scholar] [CrossRef]
- Chen, H.-C.; Reich, N. C. Live cell imaging reveals continuous STAT6 nuclear trafficking. J. Immunol. 2010, vol. 185(no. 1), 64–70. [Google Scholar] [CrossRef] [PubMed]
- Shin, H. Y.; et al. Using automated live cell imaging to reveal early changes during human motor neuron degeneration. eNeuro 2018, May 2018, vol. 5(no. 3), ENEURO.0001–18. [Google Scholar] [CrossRef]
- Mathias, J. R. Live imaging of chronic inflammation caused by mutation of zebrafish hai1. J. Cell Sci. 2007, vol. 120(no. 19), 3372–3383. [Google Scholar] [CrossRef]
- Bakota, L.; Brandt, R. Chapter 2 live-cell imaging in the study of neurodegeneration; 2009; pp. 49–103. [Google Scholar]
- Arous, J. B. Single myelin fiber imaging in living rodents without labeling by deep optical coherence microscopy. J. Biomed. Opt. 2011, vol. 16(no. 11), 116012. [Google Scholar] [CrossRef]
- Beaurepaire, E.; Boccara, A. C.; Lebec, M.; Blanchot, L.; Saint-Jalmes, H. Full-field optical coherence microscopy. Opt. Lett. 1998, vol. 23(no. 4), 244. [Google Scholar] [CrossRef]
- Dalimier, E.; Salomon, D. Full-field optical coherence tomography: A new technology for 3D high-resolution skin imaging. Dermatology 2012, vol. 224(no. 1), 84–92. [Google Scholar] [CrossRef] [PubMed]
- Heldt, N.; et al. Guide to dynamic OCT data analysis. Biomed. Opt. Express 2025, vol. 16(no. 11), 4851. [Google Scholar] [CrossRef]
- Monfort, T.; Grieve, K.; Thouvenin, O. Rolling phase modulation regime for dynamic full field OCT. Opt. Lett. 2025, vol. 50(no. 7), 2239. [Google Scholar] [CrossRef]
- Oldenburg, A. L. Motility-, autocorrelation-, and polarization-sensitive optical coherence tomography discriminates cells and gold nanorods within 3D tissue cultures. Opt. Lett. 2013, vol. 38(no. 15), 2923. [Google Scholar] [CrossRef]
- Apelian, C.; Harms, F.; Thouvenin, O.; Boccara, A. C. Dynamic full field optical coherence tomography: subcellular metabolic contrast revealed in tissues by interferometric signals temporal analysis. Biomed. Opt. Express 2016, vol. 7(no. 4), 1511–1524. [Google Scholar] [CrossRef] [PubMed]
- Scholler, J. Automatic diagnosis and classification of breast surgical samples with dynamic full-field OCT and machine learning. J. Med. Imaging 2023, vol. 10(no. 03). [Google Scholar] [CrossRef] [PubMed]
- Thouvenin, O. Cell motility as contrast agent in retinal explant imaging with full-field optical coherence tomography. Investig. Ophthalmol. Vis. Sci. 2017, vol. 58(no. 11), 4605. [Google Scholar] [CrossRef]
- Auksorius, E.; Boccara, A. C. High-throughput dark-field full-field optical coherence tomography. Opt. Lett. 2020, vol. 45(no. 2), 455. [Google Scholar] [CrossRef]
- Adomavicius, K. High-throughput spatio-temporal optical coherence tomography (STOC-T). Opt. Contin. 2025, vol. 4(no. 9), 2099. [Google Scholar] [CrossRef]
- Auksorius, E. Light-efficient beamsplitter for Fourier-domain full-field optical coherence tomography. Opt. Lett. 2020, vol. 45(no. 5), 1240. [Google Scholar] [CrossRef]
- Krafft, L.; et al. Partial-field illumination ophthalmoscope: improving the contrast of a camera-based retinal imager. Appl. Opt. 2021, vol. 60(no. 31), 9951. [Google Scholar] [CrossRef]
- Monfort, T.; Azzollini, S.; Ben Yacoub, T.; Audo, I.; Reichman, S.; Grieve, K.; Thouvenin, O. Interface self-referenced dynamic full-field optical coherence tomography. Biomed. Opt. Express 2023, vol. 14(no. 7), 3491–3505. [Google Scholar] [CrossRef]
- Lippok, N.; Villiger, M.; Lasser, T. Improving the efficiency of optical coherence tomography by use of a variable-ratio coupler and the principle of least detectable reflectivity. Opt. Express 2011, vol. 19(no. 8), 7161–7175. [Google Scholar] [CrossRef] [PubMed]
- Senée, P.; Krafft, L.; Loukili, I.; Castro Farias, D.; Thouvenin, O.; Atlan, M.; Paques, M.; Meimon, S.; Mecê, P. Revealing neurovascular coupling at a high spatial and temporal resolution in the living human retina. Sci. Adv. 2025, vol. 11(no. 26), eadx2941. [Google Scholar] [CrossRef] [PubMed]
- Couturier, A.; Blot, G.; Vignaud, L.; Nanteau, C.; Slembrouck-Brec, C.; Fradot, V.; Acar, N.; Sahel, J. A.; Tadayoni, R.; Thuret, G.; Sennlaub, F.; Roger, J. E.; Goureau, O.; Guillonneau, X.; Reichman, S. Reproducing diabetic retinopathy features using newly developed human induced-pluripotent stem cell-derived retinal Müller glial cells. Glia 2021, vol. 69(no. 7), 1679–1693. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



