Submitted:
29 April 2026
Posted:
30 April 2026
You are already at the latest version
Abstract
Rice (Oryza sativa L.) is a staple food, and its eating quality is largely determined by amylose content (AC), which is controlled by the Wx gene. Current low-AC alleles often lead to excessive reductions, limiting their utility. The promoter region, particularly the pyrimidine-rich region (PRR) from -45 to -7, is hypothesized to fine-tune Wx expression, yet its functional motifs remain unexplored. In this study, we applied prime editing (PE) to precisely edit the Wx promoter in the japonica cultivar Huaidao 5. We generated two novel alleles, Wx1T/32d (deleting PRR1 [-45 to -19] and PRR2 [-11 to -7]) and Wx5d (deleting only PRR2), which are absent in natural germplasm. These edits resulted in significant, fine-tuned reductions in AC (3.95% and 2.77%, respectively) without affecting grain transparency. Our results identify PRR1 and PRR2 as important functional motifs whose deletion synergistically downregulates AC. Furthermore, the edited lines exhibited improved taste without compromising agronomic performance. This study not only provides novel insights into the regulatory motifs of the Wx promoter but also demonstrates the precision and potential of PE for improving rice grain quality.
Keywords:
1. Introduction
2. Materials and Methods
2.1. Plant Materials
2.2. Prime Editing Workflow
2.2.1. Target Design
2.2.2. Construction of Gene Editing Vectors
2.2.3. Genetic Transformation
2.3. Analysis of Wx Gene Expression and AC in Transgenic Lines
2.3.1. AC Determination
2.3.2. qRT-PCR ANALYSIS
2.3.3. GBSSI Protein Quantification
2.4. Investigation of Agronomic Traits in Transgenic Lines
3. Results
3.1. Screening of Homozygous Gene Editing Lines
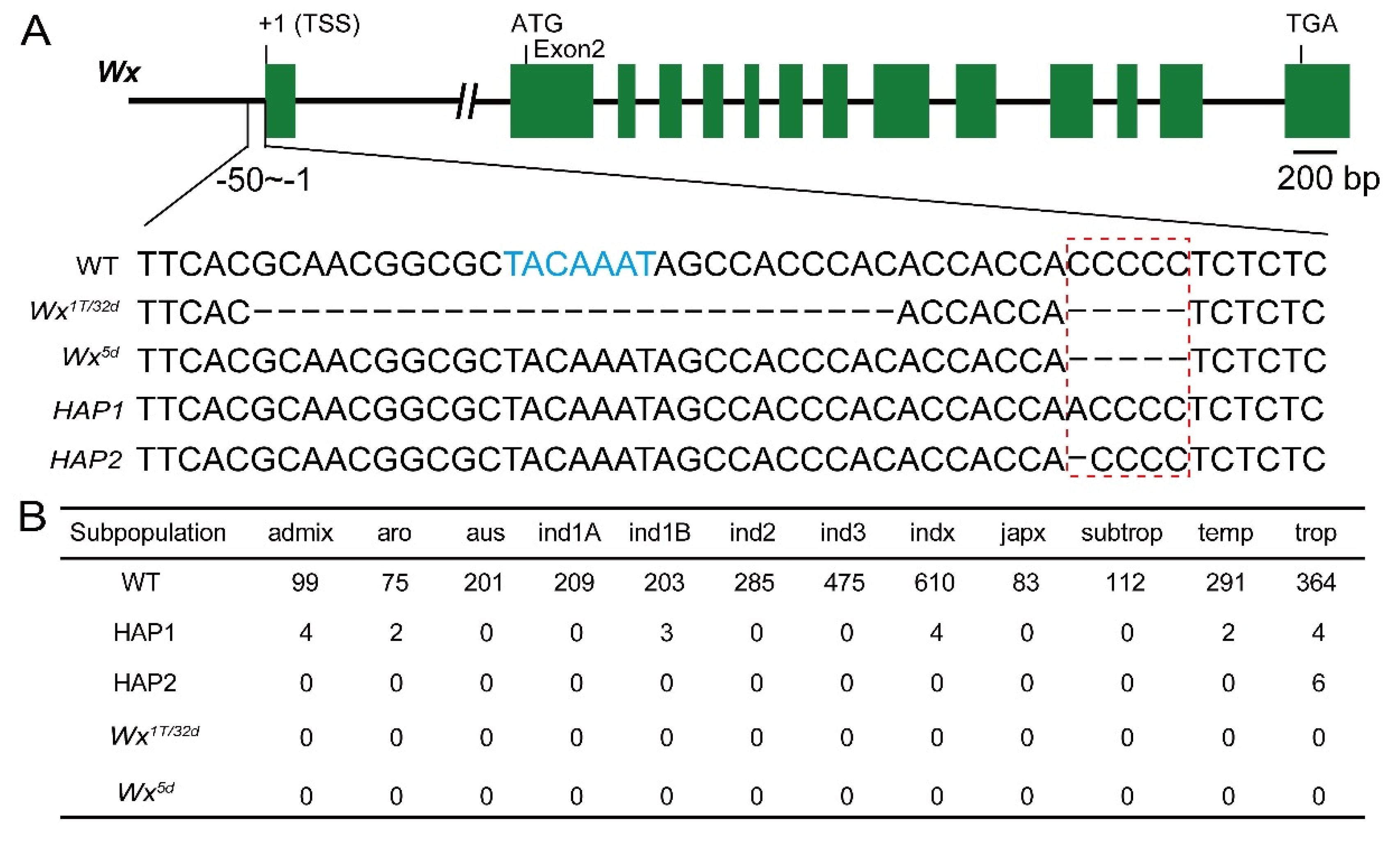
3.2. Wx novel Alleles Created by Prime Editing Are Extremely Rare in Natural Rice Populations
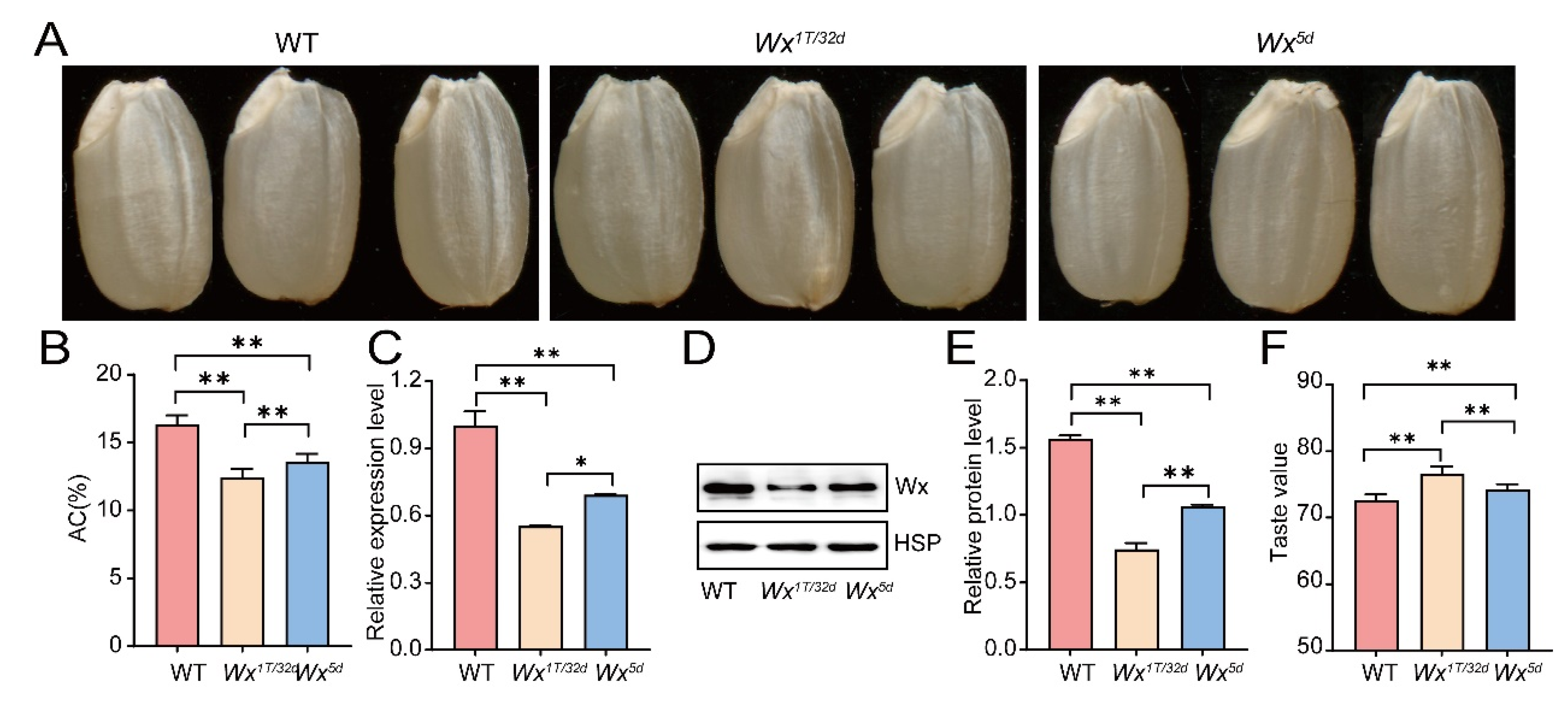
3.3. Deletion of PRR1 and PRR2 Motifs in the Wx Promoter Reduces AC and Enhances Eating Quality
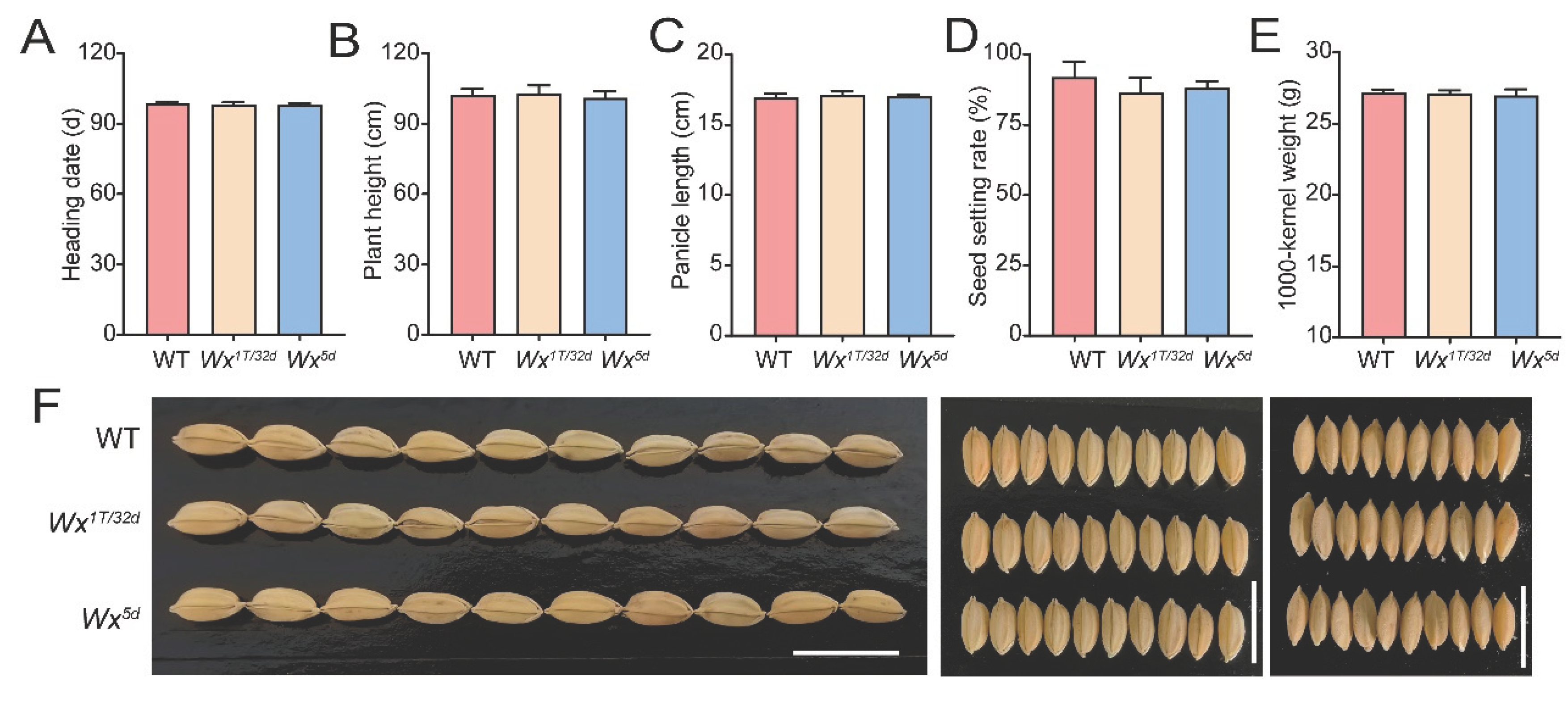
3.4. Prime Editing of the Wx Promoter Improves Grain Quality Without Compromising Agronomic Traits
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| AC | amylose content |
| PRR | pyrimidine-rich region |
| OsGBSSI | granule-bound starch synthase I |
| PE | prime editing |
| DSBs | double-strand breaks |
| pegRNAs | Prime editing guide RNAs |
| PBS | primer binding site |
| RTT | reverse transcription template |
| DAF | days after grain filling |
| qRT-PCR | quantitative real-time PCR |
| ANOVA | Analysis of variance |
References
- Fan, M. S.; Shen, J. B.; Yuan, L. X.; Jiang, R. F.; Chen, X. P.; Davies, W. J.; Zhang, F. S. Improving crop productivity and resource use efficiency to ensure food security and environmental quality in China. J. Exp. Bot. 2012, 63(1), 13–24. [Google Scholar] [CrossRef]
- He, P.; Shen, L. S.; Lu, C. F.; Chen, Y.; Zhu, L. H. Analysis of quantitative trait loci which contribute to anther culturability in rice (Oryza sativa L.). Mol. Breed. 1998, 4, 165–172. [Google Scholar] [CrossRef]
- Zhang, Q. F. Strategies for developing green super rice. Proc. Natl. Acad. Sci. U.S.A. 2007, 95, 1663–1668. [Google Scholar] [CrossRef]
- Tian, Z. X.; Qian, Q.; Liu, Q. Q.; Yan, M. X.; Liu, X. F.; Yan, C. J.; Liu, G. F.; Gao, Z. Y.; Tang, S. Z.; Zeng, D. L.; Wang, Y. H.; Wang, Y. H.; Yu, J. M.; Gu, M. H.; Li, J. Y. Allelic diversities in rice starch biosynthesis lead to a diverse array of rice eating and cooking qualities. Proc. Natl. Acad. Sci. U.S.A. 2009, 106(51), 21760–21765. [Google Scholar] [CrossRef]
- Qian, Q.; Guo, L.; Smith, S. M.; Li, J. Breeding high-yield superior quality hybrid super rice by rational design. Natl. Sci. Rev. 2016, 3, 283–294. [Google Scholar] [CrossRef]
- Liu, D.; Wang, W.; Cai, X. Modulation of amylose content by structure-based modification of OsGBSS1 activity in rice (Oryza sativa L.). Plant Biotechnol. J. 2014, 12, 1297–1307. [Google Scholar] [CrossRef] [PubMed]
- Isshiki, M.; Morino, K.; Nakajima, M.; Okagaki, R. J.; Wessler, S. R.; Izawa, T.; Shimamoto, K. A naturally occurring functional allele of the rice waxy locus has a GT to TT mutation at the 5′ splice site of the first intron. Plant J. 1998, 15(1), 133–138. [Google Scholar] [CrossRef]
- Wanchana, S.; Toojinda, T.; Tragoonrung, S.; Vanavichit, A. Duplicated coding sequence in the waxy allele of tropical glutinous rice (Oryza sativa L.). Plant Sci. 2003, 165(6), 1193–1199. [Google Scholar] [CrossRef]
- Hori, Y.; Fujimoto, R.; Sato, Y.; Nishio, T. A novel Wx mutation caused by insertion of a retrotransposon-like sequence in a glutinous cultivar of rice (Oryza sativa). Theor. Appl. Genet. 2007, 115(2), 217–224. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Yang, Y.; Chen, S.; Liu, X.; Liu, Q. A rare Waxy allele coordinately improves rice eating and cooking quality and grain transparency. J. Integr. Plant Biol. 2021, 63(5), 889–901. [Google Scholar] [CrossRef]
- Zhou, H.; Xia, D.; Zhao, D.; Li, Y.; He, Y. The origin of Wxla provides new insights into the improvement of grain quality in rice. J. Integr. Plant Biol. 2021, 63(5), 878–888. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z. Y.; Zheng, F. Q.; Shen, G. Z.; Gao, J. P.; Snustad, D. P.; Li, M. G.; Zhang, J. L.; Hong, M. M. The amylose content in rice endosperm is related to the post-transcriptional regulation of the waxy gene. Plant J. 1995, 7(4), 613–622. [Google Scholar] [CrossRef] [PubMed]
- Satoh, H.; Shimizu, Y.; Sawada, M.; Itoh, T. Molecular characterization of Wx-mq, a novel mutant gene for low-amylose content in endosperm of rice (Oryza sativa L.). Breed. Sci. 2002, 52(2), 131–135. [Google Scholar] [CrossRef]
- Yang, J.; Wang, J.; Fan, F. J.; Zhu, J. Y.; Chen, T.; Wang, C. L.; Zheng, T. Q.; Zhang, J.; Zhong, W. G.; Xu, J. L. Development of AS-PCR marker based on a key mutation confirmed by resequencing of Wx-mp in Milky Princess and its application in japonica soft rice (Oryza sativa L.) breeding. Plant Breed. 2013, 132(6), 595–603. [Google Scholar] [CrossRef]
- Liu, L. L.; Ma, X. D.; Liu, S. J.; Zhu, C. L.; Jiang, L.; Wang, Y. H.; Shen, Y.; Ren, Y. L.; Dong, H.; Chen, L. M.; Liu, X.; Zhao, Z. G.; Zhai, H. Q.; Wan, J. M. Identification and characterization of a novel Waxy allele from a Yunnan rice landrace. Plant Mol. Biol. 2009, 71(6), 609–626. [Google Scholar] [CrossRef]
- Mikami, I.; Uwatoko, N.; Ikeda, Y.; Yamaguchi, J.; Hirano, H. Y.; Suzuki, Y.; Sano, Y. Allelic diversification at the Wx locus in landraces of Asian rice. Theor. Appl. Genet. 2008, 116, 979–989. [Google Scholar] [CrossRef]
- Zhang, C.; Zhu, J.; Chen, S.; Fan, X.; Liu, Q. Wxlv, the ancestral allele of rice waxy gene. Mol. Plant. 2019, 12(8), 1157–1166. [Google Scholar] [CrossRef]
- Huang, L. C.; Li, Q. F.; Zhang, C. Q.; Chu, R.; Gu, Z. W.; Tan, H. Y.; Zhao, D. S.; Fan, X. L.; Liu, Q. Q. Creating novel Wx alleles with fine-tuned amylose levels and improved grain quality in rice by promoter editing using CRISPR/Cas9 system. Plant Biotechnol. J. 2020, 18(11), 2164–2176. [Google Scholar] [CrossRef] [PubMed]
- Hemage, K. R. D.; Sumanasinghe, V. A. In silico promoter analysis reveals rice Wx promoter could be a TATA-less promoter with a putative pyrimidine rich region. Trop. Agric. Res. 2017, 29(1), 32–44. [Google Scholar] [CrossRef]
- Fei, Y. Y.; Yang, J.; Wang, F. Q.; Fan, F. J.; Li, W. Q.; Xu, Y.; Zhu, J. Y.; Zhong, W. G. Production of two elite glutinous rice varieties by editing Wx gene. Rice Sci. 2019, 26(2), 118–124. [Google Scholar]
- Fu, Y. H.; Hua, Y. H.; Luo, T. T.; Liu, C. Y.; Zhang, B. L.; Zhang, X. Y.; Liu, Y. P.; Zhu, Z. Z.; Tao, Y.; Zhu, Z. Y.; Li, P.; Zhu, J. Generating waxy rice starch with target type of amylopectin fine structure and gelatinization temperature by waxy gene editing. Carbohydr. Polym. 2023, 306, 120595. [Google Scholar] [CrossRef] [PubMed]
- Kabaria, S. R.; Bae, Y.; Ehmann, M. E.; Lende-Dorn, B. A.; Beitz, A. M.; Peterman, E. L.; Love, K. S.; Ploessl, D. S.; Galloway, K. E. Programmable promoter editing for precise control of transgene expression. Nat. Biotechnol. 2025, 1–16. [Google Scholar] [CrossRef]
- Anzalone, A. V.; Randolph, P. B.; Davis, J. R.; Sousa, A. A.; Koblan, L. W.; Levy, J. M.; Chen, P. J.; Wilson, C.; Newby, G. A.; Raguram, A.; Liu, D. R. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature. 2019, 576(7785), 149–157. [Google Scholar] [CrossRef]
- Gao, J. P.; Li, Y. Z.; Wang, Z. Y.; Hong, M. M. Identification of Transcription Start Site of Waxy Gene in O. sativa subsp. indica 232. J. Genet. Genom. 1995, 06, 431–436. [Google Scholar]
- Klösgen, R. B.; Gierl, A.; Schwarz-Sommer, Z.; Saedler, H. Molecular analysis of the waxy locus of Zea mays. Mol. Gen. Genet. MGG. 1986, 203, 237–244. [Google Scholar] [CrossRef]
- Li, R. T.; Wang, Q.; She, K. Q.; Lu, F.; Yang, Y. CRISPR/Cas systems usher in a new era of disease treatment and diagnosis. Mol. Biomed. 2022, 3, 31. [Google Scholar] [CrossRef] [PubMed]
| Component | Volume |
|---|---|
| Forward primer (10 μM) | 2 μL |
| Reverse primer (10 μM) | 2 μL |
| 2× Taq PCR Mix | 25 μL |
| ddH2O | 20 μL |
| Template | 1 μL |
| Total | 50 μL |
| Step | Cycles |
|---|---|
| 94 °C for 5 min | 1 |
| 94 °C for 30 sec | 30 |
| 50 °C for 45 sec | 30 |
| 72 °C for 22 sec | 30 |
| 72 °C for 10 min | 1 |
| 16 °C for 30 min | 1 |
| Component | Volume |
|---|---|
| Nuclease-free Water | 8 μL |
| 10× Buffer | 2 μL |
| BsaI/Eco31I | 1 μL |
| T4 DNA ligase | 10 μL |
| pHU-PE-Lncas9(K34) (4 U/μL) | 4 μL |
| rDNAT1 | 4 μL |
| Total | 20 μL |
| Step | Cycles |
|---|---|
| 37 °C for 20 min | 1 |
| 37 °C for 10 min | 5 |
| 20 °C for 10 min | 5 |
| 37 °C for 20 min | 1 |
| 80 °C for 5 min | 1 |
| Component | Volume |
|---|---|
| Forward primer (10 μM) | 1 μL |
| Reverse primer (10 μM) | 1 μL |
| 2× Taq PCR Mix | 12.5 μL |
| ddH2O | 9.5 μL |
| Template | 1 μL |
| Total | 25 μL |
| Component | Volume |
|---|---|
| Forward primer (10 μM) | 1 μL |
| Reverse primer (10 μM) | 1 μL |
| 2× Taq PCR Mix | 10 μL |
| ddH2O | 7 μL |
| Template | 1 μL |
| Total | 20 μL |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




