Submitted:
28 April 2026
Posted:
30 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Identification and Retrieval Information of Hsp20 Genomic Sequences
2.2. Peptide Physicochemical Features and Subcellular Localization Prediction
2.3. Gene Structure, Protein Motif Prediction and Prediction of Transcription Factor Binding Sites
2.4. Phylogenetic Analysis
2.5. Chromosomal Organization, Gene Duplications and Evolutionary Analysis
2.6. Identification of Hsp20 Under Different Stress Conditions and Quantification Pattern Analysis
3. Results
3.1. Identification and Descriptions of CqHsp20 Genomic Sequences
3.2. Physicochemical Properties and Peptide Subcellular Localization
3.3. Gene Structure, Protein Motif Prediction and Prediction of Transcription Factor Binding Sites
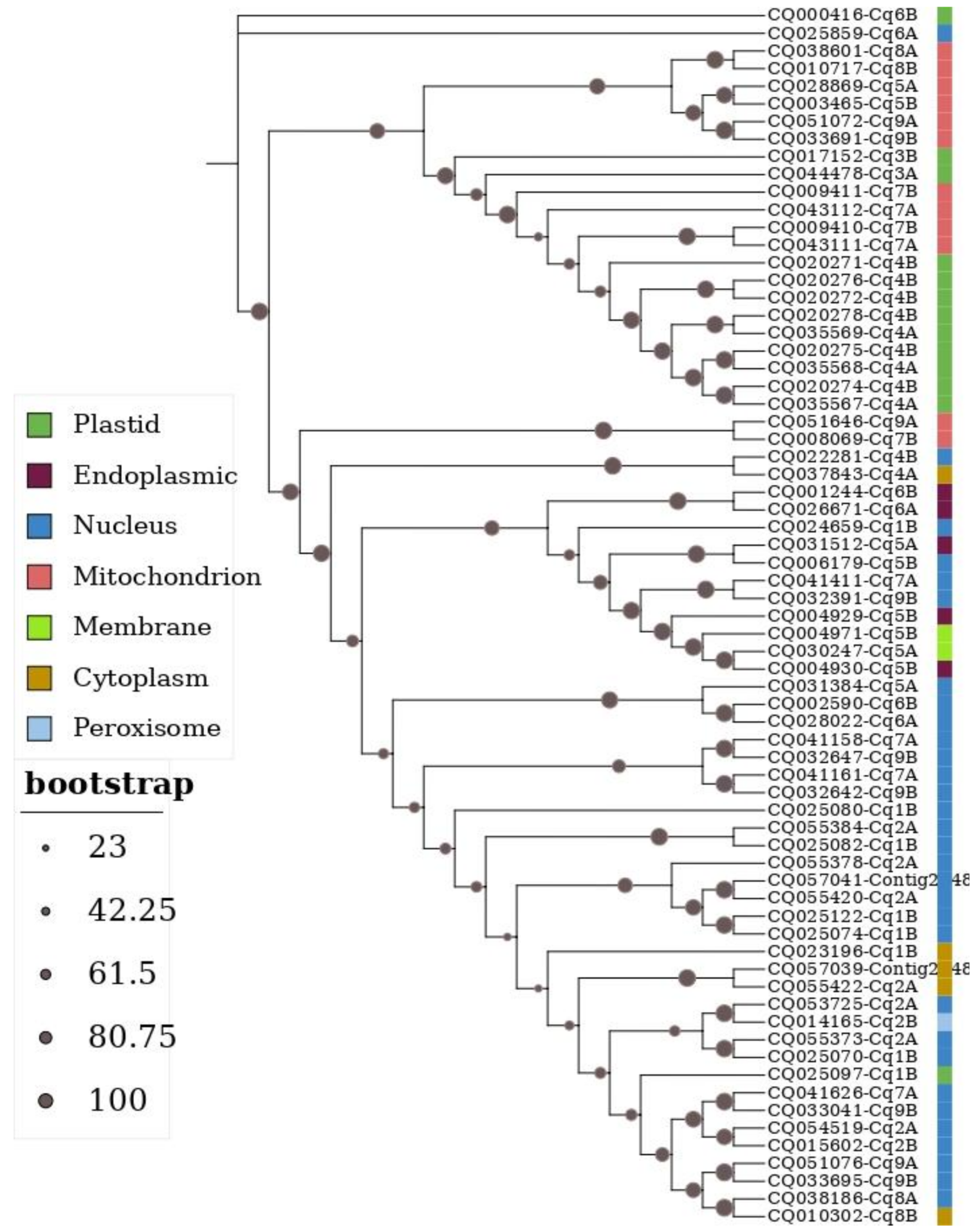
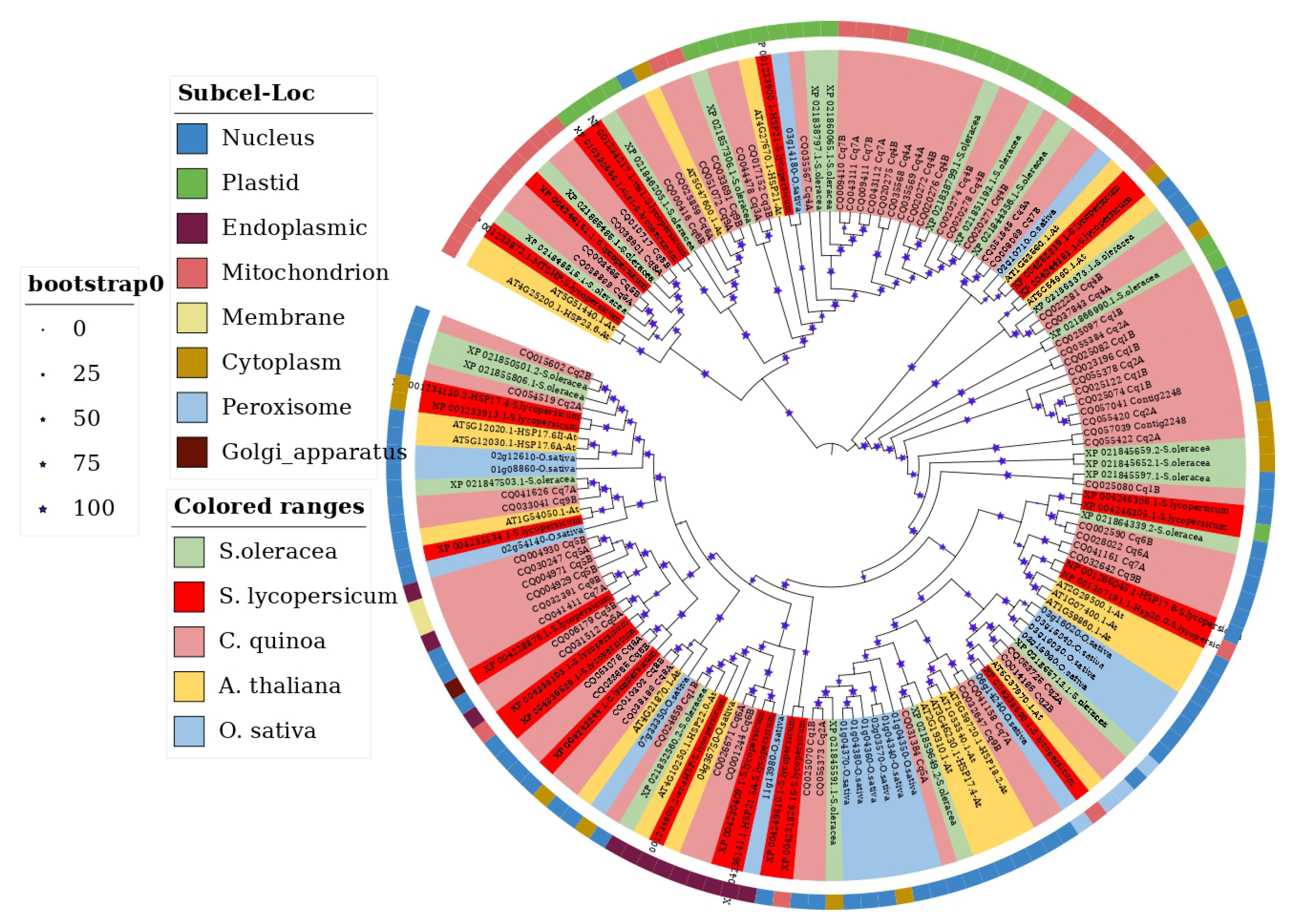
3.4. Phylogenetic Analysis
3.5. Chromosomal Organization, Gene Duplications and Evolutionary Analysis
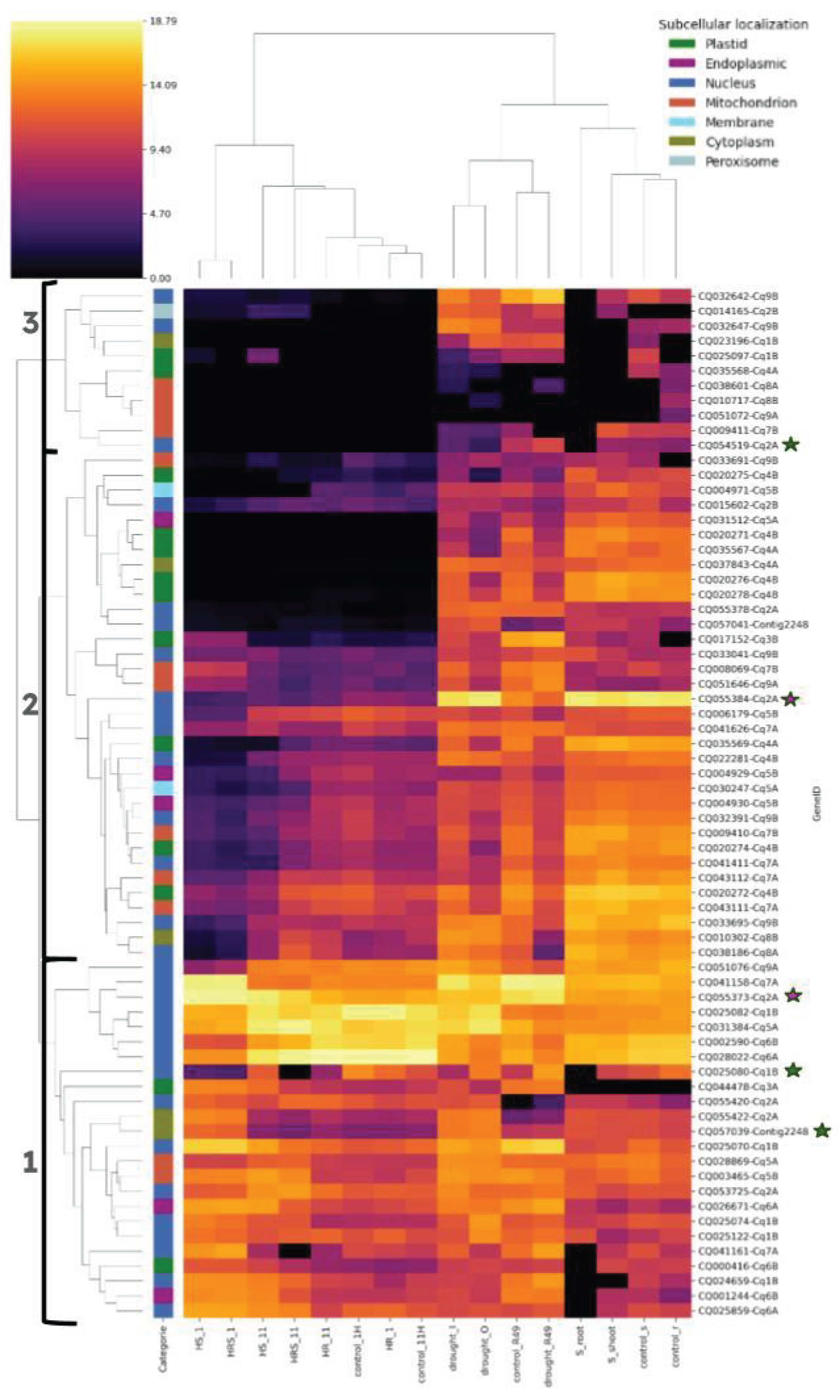
3.6. Expression Profiling of the CqHsp20 Genes in Different Abiotic Stresses
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lichtenthaler, H.K. The Stress Concept in Plants: An Introduction. Ann. NY. Acad. Sci. 1998, 851, 187–198. [Google Scholar] [CrossRef]
- Shao, H.B.; Guo, Q.J.; Chu, L.Y.; Zhao, X.N.; Su, Z.L.; Hu, Y.C.; Cheng, J.F. Understanding molecular mechanism of higher plant plasticity under abiotic stress. Colloids Surf. B Biointerfaces 2007, 54, 37–45. [Google Scholar] [CrossRef]
- Waters, E.R. The evolution, function, structure, and expression of the plant sHSPs. J. Exp. Bot. 2013, 64, 391–403. [Google Scholar] [CrossRef]
- Carra, S.; Alberti, S.; Benesch, J.L.; Boelens, W.; Buchner, J.; Carver, J.A.; Cecconi, C.; Ecroyd, H.; Gusev, N.; Hightower, L.E.; et al. Small heat shock proteins: multifaceted proteins with important implications for life. Cell. Stress Chaperones 2019, 24, 295–308. [Google Scholar] [CrossRef] [PubMed]
- Rizzo, A.J.; Palacios, M.B.; Vale, E.M.; Zelada, A.M.; Silveira, V.; Burrieza, H.P. Snapshot of four mature quinoa (Chenopodium quinoa) seeds: a shotgun proteomics analysis with emphasis on seed maturation, reserves and early germination. Physiol. Mol. Biol. Plan. 2023, 29, 319–334. [Google Scholar] [CrossRef]
- Arce, D.P.; De Las Rivas, J.; Pratta, G.R. Interactomic analysis of the sHSP family during tomato fruit ripening. Plant Gene 2020, 21. [Google Scholar] [CrossRef]
- Cacchiarelli, P.; Arce, D.P.; Tapia, E.; Pratta, G. Structural and functional analysis of two sHSP subfamilies in tomato ripening. Plant Gene 2021, 27, 100297. [Google Scholar] [CrossRef]
- Goytia Bertero, V.; Cacchiarelli, P.; Pratta, G.R.; Arce, D.P. Comparative and integrative omic analysis focused on chaperones and interactors in a cultivated and an exotic tomato at different fruit ripening stages. Plant Gene 2024, 37, 100448. [Google Scholar] [CrossRef]
- Sun, W.; Van Montagu, M.; Verbruggen, N. Small heat shock proteins and stress tolerance in plants. Biochim. Et. Biophys. Acta (BBA) -Gene Struct. Expr. 2002, 1577, 1–9. [Google Scholar] [CrossRef]
- Al-Whaibi, M.H. Plant heat-shock proteins: A mini review. J. King Saud. Univ.-Sci. 2011, 23, 139–150. [Google Scholar] [CrossRef]
- Scharf, K.D.; Siddique, M.; Vierling, E. The expanding family of Arabidopsis thaliana small heat stress proteins and a new family of proteins containing alfa-crystalin domains (Acd proteins). Cell. Stress Soc. Int. 2001, 6, 225–237. [Google Scholar] [CrossRef]
- Waters, E.R. The molecular evolution of the small heat-shock proteins in plants. Genetics 1995, 141, 785–795. [Google Scholar] [CrossRef] [PubMed]
- Waters, E.R.; Aevermann, B.D.; Sanders-Reed, Z. Comparative analysis of the small heat shock proteins in three angiosperm genomes identifies new subfamilies and reveals diverse evolutionary patterns. Cell. Stress Chaperones 2008, 13, 127–142. [Google Scholar] [CrossRef]
- Haslbeck, M.; Vierling, E. A First Line of Stress Defense: Small Heat Shock Proteins and Their Function in Protein Homeostasis. J. Mol. Biol. 2015, 427, 1537–1548. [Google Scholar] [CrossRef] [PubMed]
- Lallemand, T.; Leduc, M.; Landès, C.; Rizzon, C.; Lerat, E. An Overview of Duplicated Gene Detection Methods: Why the Duplication Mechanism Has to Be Accounted for in Their Choice. Genes 2020, 11, 1046. [Google Scholar] [CrossRef] [PubMed]
- Panchy, N.; Lehti-Shiu, M.; Shiu, S.H. Evolution of Gene Duplication in Plants. Plant Physiol. 2016, 171, 2294–2316. [Google Scholar] [CrossRef]
- Krsticevic, F.J.; Arce, D.P.; Ezpeleta, J.; Tapia, E. Tandem Duplication Events in the Expansion of the Small Heat Shock Protein Gene Family in Solanum lycopersicum (cv. Heinz 1706). G3 Genes|Genomes|Genetics 2016, 6, 3027–3034. [Google Scholar] [CrossRef]
- Liu, J.; Wang, R.; Liu, W.; Zhang, H.; Guo, Y.; Wen, R. Genome-Wide Characterization of Heat-Shock Protein 70s from Chenopodium quinoa and Expression Analyses of Cqhsp70s in Response to Drought Stress. In Genes; 2018; Volume 9, ISBN 8613935230446. [Google Scholar] [CrossRef]
- De Souza Resende, J.S.; Dos Santos, T.B.; Souza, S.G.H.D. Small heat shock protein (Hsp20) gene family in Phaseolus vulgaris L.: Genome-wide identification, evolutionary and expression analysis. Plant Gene 2022, 31, 100370. [Google Scholar] [CrossRef]
- Ramakrishna, G.; Kaur, P.; Singh, A.; Yadav, S.S.; Sharma, S.; Singh, N.K.; Gaikwad, K. Comparative transcriptome analyses revealed different heat stress responses in pigeonpea (Cajanus cajan) and its crop wild relatives. Plant Cell. Rep. 2021, 40, 881–898. [Google Scholar] [CrossRef]
- Do, J.M.; Kim, H.J.; Shin, S.Y.; Park, S.I.; Kim, J.J.; Yoon, H.S. OsHSP 17.9, a Small Heat Shock Protein, Confers Improved Productivity and Tolerance to High Temperature and Salinity in a Natural Paddy Field in Transgenic Rice Plants. Agriculture 2023, 13, 931. [Google Scholar] [CrossRef]
- Guo, M.; Liu, J.H.; Lu, J.P.; Zhai, Y.F.; Wang, H.; Gong, Z.H.; Wang, S.B.; Lu, M.H. Genome-wide analysis of the CaHsp20 gene family in pepper: comprehensive sequence and expression profile analysis under heat stress. Front. Plant Sci. 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- Biondi, E.; Ruiz, K.B.; Martinez, E.A.; Zurita-Silva, A.; Orsini, F.; Antognoni, F.; Dinelli, G.; Marotti, I.; Giaquinto, G.; Maldonado, S.; et al. Tolerance to saline conditions. In Estado del arte de la quinua en el mundo en 2013; FAO and CIRAD: Santiago de Chile, Chile; Montpellier, Francia, 2014. [Google Scholar]
- Ruiz, K.B.; Biondi, S.; Oses, R.; Acuña-Rodríguez, I.S.; Antognoni, F.; Martinez-Mosqueira, E.A.; Coulibaly, A.; Canahua-Murillo, A.; Pinto, M.; Zurita-Silva, A.; et al. Quinoa biodiversity and sustainability for food security under climate change. A review. Agron. Sustain. Dev. 2014, 34, 349–359. [Google Scholar] [CrossRef]
- Pickersgill, B. Domestication of plants in the Americas: Insights from Mendelian and molecular genetics. Ann. Bot. 2007, 100, 925–940. [Google Scholar] [CrossRef]
- Mu, H.; Xue, S.; Sun, Q.; Shi, J.; Zhang, D.; Wang, D.; Wei, J. Research Progress of Quinoa Seeds (Chenopodium quinoa Wild.): Nutritional Components, Technological Treatment, and Application. Foods 2023, 12, 2087. [Google Scholar] [CrossRef]
- Pereira, E.; Encina-Zelada, C.; Barros, L.; Gonzales-Barron, U.; Cadavez, V.; C.F.R. Ferreira, I. Chemical and nutritional characterization of Chenopodium quinoa Willd (quinoa) grains: A good alternative to nutritious food. Food Chem. 2019, 280, 110–114. [Google Scholar] [CrossRef]
- Jellen, E.N.; Jarvis, D.E.; Benet-Pierce, N.; Maughan, P.J. Botanical Context for Domestication in North America; Springer International Publishing, 2021. [Google Scholar] [CrossRef]
- Wilson, H.; Manhart, J. Crop/weed gene flow:Chenopodium quinoa Willd. andC. berlandieri Moq. Theor. Appl. Genet. 1993, 86, 642–648. [Google Scholar] [CrossRef]
- Jarvis, D.E.; Ho, Y.S.; Lightfoot, D.J.; Schmöckel, S.M.; Li, B.; Borm, T.J.A.; Ohyanagi, H.; Mineta, K.; Michell, C.T.; Saber, N.; et al. The genome of Chenopodium quinoa. Nature 2017, 542, 307–312. [Google Scholar] [CrossRef]
- Maughan, P.J.; Chaney, L.; Lightfoot, D.J.; Cox, B.J.; Tester, M.; Jellen, E.N.; Jarvis, D.E. Mitochondrial and chloroplast genomes provide insights into the evolutionary origins of quinoa (Chenopodium quinoa Willd.). Sci. Rep. 2019, 9, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Rey, E.; Maughan, P.J.; Maumus, F.; Lewis, D.; Wilson, L.; Fuller, J.; Schmöckel, S.M.; Jellen, E.N.; Tester, M.; Jarvis, D.E. A chromosome-scale assembly of the quinoa genome provides insights into the structure and dynamics of its subgenomes. Commun. Biol. 2023, 6, 1263. [Google Scholar] [CrossRef]
- Young, L.A.; Maughan, P.J.; Jarvis, D.E.; Hunt, S.P.; Warner, H.C.; Durrant, K.K.; Kohlert, T.; Curti, R.N.; Bertero, D.; Filippi, G.A.; et al. A chromosome-scale reference of Chenopodium watsonii helps elucidate relationships within the North American A-genome Chenopodium species and with quinoa. Plant Genome 2023, 16, e20349. [Google Scholar] [CrossRef]
- Alandia, G.; Rodriguez, J.P.; Jacobsen, S.E.E.; Bazile, D.; Condori, B. Global expansion of quinoa and challenges for the Andean region. Glob. Food Secur. 2020, 26, 100429. [Google Scholar] [CrossRef]
- Bazile, D.; Jacobsen, S.E.; Verniau, A. The Global Expansion of Quinoa: Trends and Limits. Front. Plant Sci. 2016, 7. [Google Scholar] [CrossRef]
- Hinojosa, L.; González, J.A.; Barrios-Masias, F.H.; Fuentes, F.; Murphy, K.M. Quinoa abiotic stress responses: A review. Plants 2018, 7. [Google Scholar] [CrossRef] [PubMed]
- The Quinoa Genome. In Compendium of Plant Genomes; Schmöckel, S.M., Ed.; Springer International Publishing: Cham, 2021. [Google Scholar] [CrossRef]
- García-Parra, M.; Zurita-silva, A.; Stechauner-rohringer, R.; Roa-acosta, D.; Jacobsen, S.E.E.; García-Parra, M.; Zurita-silva, A.; Stechauner-rohringer, R.; Roa-acosta, D.; Jacobsen, S.E.E. Quinoa ( Chenopodium quinoa Willd.) and its relationship with agroclimatic characteristics: A Colombian perspective. Chil. J. Agric. Res. 2020, 80, 290–302. [Google Scholar] [CrossRef]
- Maughan, P.J.; Turner, T.B.; Coleman, C.E.; Elzinga, D.B.; Jellen, E.N.; Morales, J.A.; Udall, J.A.; Fairbanks, D.J.; Bonifacio, A. Characterization of Salt Overly Sensitive 1 (SOS1) gene homoeologs in quinoa (Chenopodium quinoa Willd.). Genome 2009, 52, 647–657. [Google Scholar] [CrossRef]
- Ruiz-Carrasco, K.; Antognoni, F.; Coulibaly, A.K.; Lizardi, S.; Covarrubias, A.; Martínez, E.A.; Molina-Montenegro, M.A.; Biondi, S.; Zurita-Silva, A. Variation in salinity tolerance of four lowland genotypes of quinoa (Chenopodium quinoa Willd.) as assessed by growth, physiological traits, and sodium transporter gene expression. Plant Physiol. Biochem. 2011, 49, 1333–1341. [Google Scholar] [CrossRef] [PubMed]
- Morales, A.; Zurita-Silva, A.; Maldonado, J.; Silva, H. Transcriptional Responses of Chilean Quinoa (Chenopodium quinoa Willd.) Under Water Deficit Conditions Uncovers ABA-Independent Expression Patterns. Front. Plant Sci. 2017, 8, 1–13. [Google Scholar] [CrossRef]
- Tovar, J.C.; Quillatupa, C.; Callen, S.T.; Castillo, S.E.; Pearson, P.; Shamin, A.; Schuhl, H.; Fahlgren, N.; Gehan, M.A. Heating quinoa shoots results in yield loss by inhibiting fruit production and delaying maturity. Plant J. 2020, 102, 1058–1073. [Google Scholar] [CrossRef]
- Schmöckel, S.M.; Lightfoot, D.J.; Razali, R.; Tester, M.; Jarvis, D.E. Identification of Putative Transmembrane Proteins Involved in Salinity Tolerance in Chenopodium quinoa by Integrating Physiological Data, RNAseq, and SNP Analyses. Front. Plant Sci. 2017, 8, 1023. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Guo, X.; Liu, J.; Zhou, F.; Liu, W.; Wu, J.; Zhang, H.; Cao, H.; Su, H.; Wen, R. Genome-Wide Identification, Characterization, and Expression Analysis of the NAC Transcription Factor in Chenopodium quinoa. Genes 2019, 10, 500. [Google Scholar] [CrossRef]
- Li, F.; Liu, J.; Guo, X.; Yin, L.; Zhang, H.; Wen, R. Genome-wide survey, characterization, and expression analysis of bZIP transcription factors in Chenopodium quinoa. BMC Plant Biol. 2020, 20, 405. [Google Scholar] [CrossRef]
- Tashi, G.; Zhan, H.; Xing, G.; Chang, X.; Zhang, H.; Nie, X.; Ji, W. Genome-Wide Identification and Expression Analysis of Heat Shock Transcription Factor Family in Chenopodium quinoa Willd. Agronomy 2018, 8, 103. [Google Scholar] [CrossRef]
- Wu, Q.; Bai, X.; Zhao, W.; Shi, X.; Xiang, D.; Wan, Y.; Wu, X.; Sun, Y.; Zhao, J.; Peng, L.; et al. Investigation into the underlying regulatory mechanisms shaping inflorescence architecture in Chenopodium quinoa. BMC Genom. 2019, 20, 1–20. [Google Scholar] [CrossRef]
- Yue, H.; Chang, X.; Zhi, Y.; Wang, L.; Xing, G.; Song, W.; Nie, X. Evolution and identification of the WRKY gene family in Quinoa (Chenopodium quinoa). Genes 2019, 10. [Google Scholar] [CrossRef] [PubMed]
- Golicz, A.A.; Steinfort, U.; Arya, H.; Singh, M.B.; Bhalla, P.L. Analysis of the quinoa genome reveals conservation and divergence of the flowering pathways. Funct. Integr. Genom. 2020, 20, 245–258. [Google Scholar] [CrossRef]
- Patiranage, D.S.; Asare, E.; Maldonado-Taipe, N.; Rey, E.; Emrani, N.; Tester, M.; Jung, C. Haplotype variations of major flowering time genes in quinoa unveil their role in the adaptation to different environmental conditions. Plant Cell. Environ. 2021, 44, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Patiranage, D.S.R.; Rey, E.; Emrani, N.; Wellman, G.; Schmid, K.; Schmöckel, S.M.; Tester, M.; Jung, C. Genome-wide association study in the pseudocereal quinoa reveals selection pattern typical for crops with a short breeding history. eLife 2022, 1–22. [Google Scholar] [CrossRef]
- Paysan-Lafosse, T.; Andreeva, A.; Blum, M.; Chuguransky, S.; Grego, T.; Pinto, B.; Salazar, G.; Bileschi, M.; Llinares-López, F.; Meng-Papaxanthos, L.; et al. The Pfam protein families database: embracing AI/ML. Nucleic Acids Res. 2025, 53, D523–D534. Available online: https://academic.oup.com/nar/article-pdf/53/D1/D523/60667756/gkae997.pdf. [CrossRef]
- Bateman, A.; Martin, M.J.; Orchard, S.; Magrane, M.; Ahmad, S.; Alpi, E.; Bowler-Barnett, E.H.; Britto, R.; Bye-A-Jee, H.; et al.; The UniProt Consortium UniProt: the Universal Protein Knowledgebase in 2023. Nucleic Acids Res. 2023, 51, D523–D531. [Google Scholar] [CrossRef]
- Gasteiger, E.; Hoogland, C.; Gattiker, A.; Duvaud, S.; Wilkins, M.R.; Appel, R.D.; Bairoch, A. Protein Identification and Analysis Tools on the ExPASy Server. In The Proteomics Protocols Handbook; Walker, J.M., Ed.; Humana Press: Totowa, NJ, 2005; pp. 571–607. [Google Scholar] [CrossRef]
- Jiang, Y.; Wang, D.; Yao, Y.; Eubel, H.; Künzler, P.; Møller, I.M.; Xu, D. MULocDeep: A deep-learning framework for protein subcellular and suborganellar localization prediction with residue-level interpretation. Comput. Struct. Biotechnol. J. 2021, 19, 4825–4839. [Google Scholar] [CrossRef]
- Jiang, Y.; Jiang, L.; Akhil, C.S.; Wang, D.; Zhang, Z.; Zhang, W.; Xu, D. MULocDeep web service for protein localization prediction and visualization at subcellular and suborganellar levels. Nucleic Acids Res. 2023, 51, W343–W349. [Google Scholar] [CrossRef] [PubMed]
- Hu, B.; Jin, J.; Guo, A.Y.; Zhang, H.; Luo, J.; Gao, G. GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar] [CrossRef]
- Bailey, T.L.; Johnson, J.; Grant, C.E.; Noble, W.S. The MEME Suite. Nucleic Acids Res. 2015, 43, W39–W49. [Google Scholar] [CrossRef]
- O’Malley, R.; Huang, S.s.; Song, L.; Lewsey, M.; Bartlett, A.; Nery, J.; Galli, M.; Gallavotti, A.; Ecker, J. Cistrome and Epicistrome Features Shape the Regulatory DNA Landscape. Cell 2016, 165, 1280–1292. [Google Scholar] [CrossRef]
- Xue, G.; Fan, Y.; Zheng, C.; Yang, H.; Feng, L.; Chen, X.; Yang, Y.; Yao, X.; Weng, W.; Kong, L.; et al. bHLH transcription factor family identification, phylogeny, and its response to abiotic stress in Chenopodium quinoa. Front Plant Sci. 2023, 14, 1171518. [Google Scholar] [CrossRef]
- Edgar, R.C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef]
- Minh, B.Q.; Schmidt, H.A.; Chernomor, O.; Schrempf, D.; Woodhams, M.D.; von Haeseler, A.; Lanfear, R. IQ-TREE 2: New Models and Efficient Methods for Phylogenetic Inference in the Genomic Era. Mol. Biol. Evol. 2020, 37, 1530–1534. [Google Scholar] [CrossRef] [PubMed]
- Kalyaanamoorthy, S.; Minh, B.Q.; Wong, T.K.F.; von Haeseler, A.; Jermiin, L.S. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat. Methods 2017, 14, 587–589. [Google Scholar] [CrossRef]
- Hoang, D.T.; Chernomor, O.; von Haeseler, A.; Minh, B.Q.; Vinh, L.S. UFBoot2: Improving the Ultrafast Bootstrap Approximation. Mol. Biol. Evol. 2018, 35, 518–522. [Google Scholar] [CrossRef] [PubMed]
- Rice, P.; Longden, I.; Bleasby, A. EMBOSS: the European Molecular Biology Open Software Suite. Trends Genet 2000, 16, 276–277. [Google Scholar] [CrossRef]
- Wang, D.; Zhang, Y.; Zhang, Z.; Zhu, J.; Yu, J. KaKs_Calculator 2.0: a toolkit incorporating gamma-series methods and sliding window strategies. Genom. Proteom. Bioinform. 2010, 8, 77–80. [Google Scholar] [CrossRef]
- Suyama, M.; Torrents, D.; Bork, P. PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res. 2006, 34, W609–612. [Google Scholar] [CrossRef]
- Li, W.H.; Wu, C.I.; Luo, C.C. Nonrandomness of point mutation as reflected in nucleotide substitutions in pseudogenes and its evolutionary implications. J. Mol. Evol. 1984, 21, 58–71. [Google Scholar] [CrossRef] [PubMed]
- Raney, J.A.; Reynolds, D.J.; Elzinga, D.B.; Page, J.; A. Udall, J.; Jellen, E.N.; Bonfacio, A.; Fairbanks, D.J.; Maughan, P.J. Transcriptome Analysis of Drought Induced Stress in Chenopodium quinoa. AJPS 2014, 05, 338–357. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Paggi, J.M.; Park, C.; Bennett, C.; Salzberg, S.L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 2019, 37, 907–915. [Google Scholar] [CrossRef] [PubMed]
- Danecek, P.; Bonfield, J.K.; Liddle, J.; Marshall, J.; Ohan, V.; Pollard, M.O.; Whitwham, A.; Keane, T.; McCarthy, S.A.; Davies, R.M.; et al. Twelve years of SAMtools and BCFtools. Gigascience 2021, 10. [Google Scholar] [CrossRef]
- Liao, Y.; Smyth, G.K.; Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef]
- Liao, Y.; Smyth, G.K.; Shi, W. The R package Rsubread is easier, faster, cheaper and better for alignment and quantification of RNA sequencing reads. Nucleic Acids Res. 2019, 47, e47. [Google Scholar] [CrossRef]
- Conesa, A.; Madrigal, P.; Tarazona, S.; Gomez-Cabrero, D.; Cervera, A.; McPherson, A.; Szcześniak, M.W.; Gaffney, D.J.; Elo, L.L.; Zhang, X.; et al. A survey of best practices for RNA-seq data analysis. Genome Biol. 2016, 17, 13. [Google Scholar] [CrossRef]
- Zhao, Y.; Li, M.C.; Konaté, M.M.; Chen, L.; Das, B.; Karlovich, C.; Williams, P.M.; Evrard, Y.A.; Doroshow, J.H.; McShane, L.M. TPM, FPKM, or Normalized Counts? A Comparative Study of Quantification Measures for the Analysis of RNA-seq Data from the NCI Patient-Derived Models Repository. J. Transl. Med. 2021, 19, 269. [Google Scholar] [CrossRef]
- Kumar, P.; Paul, D.; Jhajhriya, S.; Kumar, R.; Dutta, S.; Siwach, P.; Das, S. Understanding heat-shock proteins’ abundance and pivotal function under multiple abiotic stresses. J. Plant Biochem. Biotechnol. 2024, 33, 492–513. [Google Scholar] [CrossRef]
- Mukhopadhyay, R.; Boro, P.; Karmakar, K.; Pradhan, P.; Saha Chowdhury, R.; Das, B.; Mandal, R.; Kumar, D. Advances in the understanding of heat shock proteins and their functions in reducing abiotic stress in plants. J. Plant Biochem. Biotechnol. 2024, 33, 474–491. [Google Scholar] [CrossRef]
- Waters, E.R.; Vierling, E. Plant small heat shock proteins - evolutionary and functional diversity. New Phytol. 2020, 227, 24–37. [Google Scholar] [CrossRef]
- Glover, N.M.; Redestig, H.; Dessimoz, C. Homoeologs: What Are They and How Do We Infer Them? Trends Plant Sci. 2016, 21, 609–621. [Google Scholar] [CrossRef]
- Yu, J.; Cheng, Y.; Feng, K.; Ruan, M.; Ye, Q.; Wang, R.; Li, Z.; Zhou, G.; Yao, Z.; Yang, Y.; et al. Genome-Wide Identification and Expression Profiling of Tomato Hsp20 Gene Family in Response to Biotic and Abiotic Stresses. Front. Plant Sci. 2016, 7. [Google Scholar] [CrossRef]
- Huang, J.; Hai, Z.; Wang, R.; Yu, Y.; Chen, X.; Liang, W.; Wang, H. Genome-wide analysis of HSP20 gene family and expression patterns under heat stress in cucumber (Cucumis sativus L.). Front. Plant Sci. 2022, 13, 968418. [Google Scholar] [CrossRef]
- Jaggi, K.E.; Krak, K.; Štorchová, H.; Mandák, B.; Marcheschi, A.; Belyayev, A.; Jellen, E.N.; Sproul, J.; Jarvis, D.; Maughan, P.J. A pangenome reveals LTR repeat dynamics as a major driver of genome evolution in Chenopodium. Plant Genome 2025, 18, e70010. [Google Scholar] [CrossRef] [PubMed]
- Maughan, P.J.; Jarvis, D.E.; De La Cruz-Torres, E.; Jaggi, K.E.; Warner, H.C.; Marcheschi, A.K.; Bertero, H.D.; Gomez-Pando, L.; Fuentes, F.; Mayta-Anco, M.E.; et al. North American pitseed goosefoot (Chenopodium berlandieri) is a genetic resource to improve Andean quinoa (C. quinoa). Sci. Rep. 2024, 14, 12345. [Google Scholar] [CrossRef]
- Rey, E.; Abrouk, M.; Dufau, I.; Rodde, N.; Saber, N.; Cizkova, J.; Fiene, G.; Stanschewski, C.; Jarvis, D.E.; Jellen, E.N.; et al. Genome assembly of a diversity panel of Chenopodium quinoa. Sci. Data 2024, 11, 1366. [Google Scholar] [CrossRef]
- Stephensen, K.B.; Costa-Tártara, S.M.; Roser, R.L.; Jarvis, D.E.; Maughan, P.J.; Jellen, E.N. Germplasm Pools for Quinoa Improvement. Crops 2025, 6, 4. [Google Scholar] [CrossRef]
- Matías, J.; Cruz, V.; Reguera, M. Heat Stress Impact on Yield and Composition of Quinoa Straw under Mediterranean Field Conditions. In Plants (Basel); 2021. [Google Scholar] [CrossRef]
- Grenfell-Shaw, L.; Tester, M. Abiotic Stress Tolerance in Quinoa. In The Quinoa Genome; Schmöckel, S.M., Ed.; Springer International Publishing: Cham, 2021; pp. 139–167. [Google Scholar] [CrossRef]
- Ohama, N.; Sato, H.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Transcriptional Regulatory Network of Plant Heat Stress Response. Trends Plant Sci. 2017, 22, 53–65. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Farooq, H.U.; Hashim, M.; Rey, E.; Curti, R.; Morris, A.; Maughan, P.J.; Jellen, E.N.; Jarvis, D.E.; Bertero, D.; et al. Wild relatives to improve heat tolerance of cultivated quinoa (Chenopodium quinoa): pollen viability and grain number. J. Exp. Bot. 2025, 76, 5117–5128. [Google Scholar] [CrossRef] [PubMed]
| Cqhsp20 | ID Gene | Chr | Start | End | strand | Lenght (bp) | Note | Description | Subcellular localization | Suborganellar localization |
|---|---|---|---|---|---|---|---|---|---|---|
| CqHsp20_1 | CQ000416 | Cq6B | 4760615 | 4762599 | + | 1984 | HSP21 | Small heat shock protein%2C chloroplastic (Pisum sativum OX%3D3888) | Plastid | chloroplast stroma, plastid chloroplast thylakoid membrane |
| CqHsp20_2 | CQ001244 | Cq6B | 29417175 | 29417741 | - | 566 | HSP22.7 | 22.7 kDa class IV heat shock protein (Pisum sativum OX%3D3888) | Endoplasmic | endoplasmic reticulum lumen |
| CqHsp20_3 | CQ002590 | Cq6B | 78911925 | 78912416 | - | 491 | HSP18.2 | 18.2 kDa class I heat shock protein (Medicago sativa OX%3D3879) | Nucleus | nucleolus |
| CqHsp20_4 | CQ003465 | Cq5B | 31785 | 32884 | + | 1099 | HSP23 | Small heat shock protein%2C chloroplastic (Oxybasis rubra OX%3D3560) | Mitochondrion | mitocondrion matrix |
| CqHsp20_5 | CQ004929 | Cq5B | 19870323 | 19872162 | + | 1839 | HSP17.5-E | 17.5 kDa class I heat shock protein (Glycine max OX%3D3847) | Endoplasmic | endoplasmnic reticulum membrane |
| CqHsp20_6 | CQ004930 | Cq5B | 19873649 | 19875601 | + | 1952 | Putative Hsp20 | Protein of unknown function | Endoplasmic | endoplasmnic reticulum membrane |
| CqHsp20_7 | CQ004971 | Cq5B | 21350497 | 21352202 | + | 1705 | HSP22.0 | 22.0 kDa heat shock protein (Arabidopsis thaliana OX%3D3702) | Membrane | cell membrane |
| CqHsp20_8 | CQ006179 | Cq5B | 70325544 | 70331841 | + | 6297 | Putative Hsp20 | Protein of unknown function | Nucleus | nucleolus, chromosome |
| CqHsp20_9 | CQ008069 | Cq7B | 9790716 | 9792735 | - | 2019 | HSP26.5 | 26.5 kDa heat shock protein%2C mitochondrial (Arabidopsis thaliana OX%3D3702) | Mitochondrion | inner membrane, mitocondrion matrix |
| CqHsp20_10 | CQ009410 | Cq7B | 61723203 | 61724706 | + | 1503 | HSP22 | Small heat shock protein%2C chloroplastic (Fragment) (Glycine max OX%3D3847) | Mitochondrion | mitocondrion matrix |
| CqHsp20_11 | CQ009411 | Cq7B | 61736730 | 61738298 | + | 1568 | HSP22 | Small heat shock protein%2C chloroplastic (Fragment) (Glycine max OX%3D3847) | Mitochondrion | mitocondrion matrix |
| CqHsp20_12 | CQ010302 | Cq8B | 2128538 | 2130966 | + | 2428 | HSP15.4 | 15.4 kDa class V heat shock protein (Arabidopsis thaliana OX%3D3702) | Cytoplasm | cytosol |
| CqHsp20_13 | CQ010717 | Cq8B | 6525197 | 6526685 | + | 1488 | HSP26.2 | 26.2 kDa heat shock protein%2C mitochondrial (Oryza sativa subsp. japonica OX%3D39947) | Mitochondrion | inner membrane, mitocondrion matrix |
| CqHsp20_14 | CQ014165 | Cq2B | 20867278 | 20869174 | - | 1896 | HSP15.7 | 15.7 kDa heat shock protein%2C peroxisomal (Arabidopsis thaliana OX%3D3702) | Peroxisome | peroxisome membrane |
| CqHsp20_15 | CQ015602 | Cq2B | 65554231 | 65556160 | - | 1929 | HSP17.9 | 17.9 kDa class II heat shock protein (Helianthus annuus OX%3D4232) | Nucleus | nucleolus, nucleus speckle |
| CqHsp20_16 | CQ017152 | Cq3B | 6543782 | 6544985 | + | 1203 | HSP22 | Small heat shock protein%2C chloroplastic (Petunia hybrida OX%3D4102) | Plastid | chloroplast stroma, plastid chloroplast thylakoid membrane, chloroplast thylakoid lumen |
| CqHsp20_17 | CQ020271 | Cq4B | 5841595 | 5843390 | - | 1795 | HSP22 | Small heat shock protein%2C chloroplastic (Fragment) (Glycine max OX%3D3847) | Plastid | chloroplast stroma, plastid chloroplast thylakoid membrane |
| CqHsp20_18 | CQ020272 | Cq4B | 5845203 | 5846496 | - | 1293 | HSP22 | Small heat shock protein%2C chloroplastic (Fragment) (Glycine max OX%3D3847) | Plastid | chloroplast stroma, plastid chloroplast thylakoid membrane |
| CqHsp20_19 | CQ020274 | Cq4B | 5849685 | 5851242 | - | 1557 | HSP22 | Small heat shock protein%2C chloroplastic (Fragment) (Glycine max OX%3D3847) | Plastid | chloroplast stroma, plastid chloroplast thylakoid membrane, chloroplast thylakoid lumen |
| CqHsp20_20 | CQ020275 | Cq4B | 5853034 | 5855059 | - | 2025 | HSP22 | Small heat shock protein%2C chloroplastic (Fragment) (Glycine max OX%3D3847) | Plastid | chloroplast stroma, plastid chloroplast thylakoid membrane, chloroplast thylakoid lumen |
| CqHsp20_21 | CQ020276 | Cq4B | 5855447 | 5858290 | - | 2843 | HSP22 | Small heat shock protein%2C chloroplastic (Fragment) (Glycine max OX%3D3847) | Plastid | chloroplast stroma |
| CqHsp20_22 | CQ020278 | Cq4B | 5859801 | 5861206 | - | 1405 | HSP22 | Small heat shock protein%2C chloroplastic (Fragment) (Glycine max OX%3D3847) | Plastid | chloroplast stroma, plastid chloroplast thylakoid membrane, chloroplast thylakoid lumen |
| CqHsp20_23 | CQ022281 | Cq4B | 66507791 | 66510378 | + | 2587 | HSP21.7 | 21.7 kDa class VI heat shock protein (Arabidopsis thaliana OX%3D3702) | Nucleus | nucleolus |
| CqHsp20_24 | CQ023196 | Cq1B | 9076468 | 9077151 | - | 683 | HSP18.2 | 18.2 kDa class I heat shock protein (Medicago sativa OX%3D3879) | Cytoplasm | cytosol |
| CqHsp20_25 | CQ024659 | Cq1B | 63625415 | 63625801 | + | 386 | HSP22.0 | 22.0 kDa heat shock protein (Arabidopsis thaliana OX%3D3702) | Nucleus | nucleolus |
| CqHsp20_26 | CQ025070 | Cq1B | 67633753 | 67634238 | + | 485 | HSP18 | 18.3 kDa class I heat shock protein (Oxybasis rubra OX%3D3560) | Nucleus | nucleolus |
| CqHsp20_27 | CQ025074 | Cq1B | 67663581 | 67664066 | + | 485 | HSP18.2 | 18.2 kDa class I heat shock protein (Medicago sativa OX%3D3879) | Nucleus | nucleolus |
| CqHsp20_28* | CQ025080* | Cq1B | 67789198 | 67789677 | + | 479 | HSP17.6C | 17.6 kDa class I heat shock protein 3 (Arabidopsis thaliana OX%3D3702) | Nucleus | nucleolus |
| CqHsp20_29 | CQ025082 | Cq1B | 67796702 | 67797133 | - | 431 | HSP17.6C | 17.6 kDa class I heat shock protein 3 (Arabidopsis thaliana OX%3D3702) | Nucleus | nucleolus |
| CqHsp20_30 | CQ025097 | Cq1B | 67976470 | 67977066 | + | 596 | HSP17.6C | 17.6 kDa class I heat shock protein 3 (Arabidopsis thaliana OX%3D3702) | Plastid | chloroplast thylakoid lumen, plastid chloroplast thylakoid membrane |
| CqHsp20_31* | CQ025122* | Cq1B | 68262816 | 68263304 | + | 488 | HSP18.2 | 18.2 kDa class I heat shock protein (Medicago sativa OX%3D3879) | Nucleus | nucleolus |
| CqHsp20_32 | CQ025859 | Cq6A | 5459407 | 5460351 | + | 944 | HSP21 | Small heat shock protein%2C chloroplastic (Pisum sativum OX%3D3888) | Nucleus | nucleolus |
| CqHsp20_33 | CQ026671 | Cq6A | 24294981 | 24295547 | + | 566 | HSP22.0 | 22.0 kDa heat shock protein (Arabidopsis thaliana OX%3D3702) | Endoplasmic | endoplasmnic reticulum lumen |
| CqHsp20_34 | CQ028022 | Cq6A | 58683254 | 58683745 | - | 491 | HSP18.2 | 18.2 kDa class I heat shock protein (Medicago sativa OX%3D3879) | Nucleus | nucleolus |
| CqHsp20_35 | CQ028869 | Cq5A | 240412 | 241889 | + | 1477 | HSP23 | Small heat shock protein%2C chloroplastic (Oxybasis rubra OX%3D3560) | Mitochondrion | mitocondrion matrix |
| CqHsp20_36 | CQ030247 | Cq5A | 18530915 | 18532643 | - | 1728 | HSP22.0 | 22.0 kDa heat shock protein (Arabidopsis thaliana OX%3D3702) | Membrane | cell membrane |
| CqHsp20_37 | CQ031384 | Cq5A | 55145998 | 55146510 | + | 512 | HSP18.5-C | 18.5 kDa class I heat shock protein (Glycine max OX%3D3847) | Nucleus | nucleolus |
| CqHsp20_38 | CQ031512 | Cq5A | 56496464 | 56498160 | + | 1696 | Putative Hsp20 | Protein of unknown function-SHSP domain-containing protein | Endoplasmic | endoplasmnic reticulum membrane |
| CqHsp20_39 | CQ032391 | Cq9B | 1045640 | 1047217 | + | 1577 | Putative Hsp20 | SHSP domain-containing protein | Nucleus | nucleolus |
| CqHsp20_40 | CQ032642 | Cq9B | 3213204 | 3215697 | + | 2493 | HSP18.2 | 18.2 kDa class I heat shock protein (Medicago sativa OX%3D3879) | Nucleus | nucleolus |
| CqHsp20_41 | CQ032647 | Cq9B | 3248233 | 3248712 | + | 479 | HSP17.6C | 17.6 kDa class I heat shock protein 3 (Arabidopsis thaliana OX%3D3702) | Nucleus | nucleolus |
| CqHsp20_42 | CQ033041 | Cq9B | 6925014 | 6927875 | + | 2861 | HSP17.4B | 17.4 kDa class III heat shock protein (Arabidopsis thaliana OX%3D3702) | Nucleus | nucleolus |
| CqHsp20_43 | CQ033691 | Cq9B | 16252267 | 16254671 | + | 2404 | HSP14.7 | 14.7 kDa heat shock protein (Arabidopsis thaliana OX%3D3702) | Mitochondrion | inner membrane, mitocondrion matrix |
| CqHsp20_44 | CQ033695 | Cq9B | 16261504 | 16265597 | - | 4093 | hspC2 | Small heat shock protein C2 (Rickettsia felis (strain ATCC VR-1525 / URRWXCal2) OX%3D315456) | Nucleus | chromosome |
| CqHsp20_45 | CQ035567 | Cq4A | 5774532 | 5791524 | - | 16992 | HSP22 | Small heat shock protein%2C chloroplastic (Fragment) (Glycine max OX%3D3847) | Plastid | chloroplast stroma, chloroplast thylakoid lumen |
| CqHsp20_46 | CQ035568 | Cq4A | 5797559 | 5798553 | - | 994 | HSP22 | Small heat shock protein%2C chloroplastic (Fragment) (Glycine max OX%3D3847) | Plastid | chloroplast stroma, plastid chloroplast thylakoid membrane, chloroplast thylakoid lumen |
| CqHsp20_47 | CQ035569 | Cq4A | 5799657 | 5804216 | - | 4559 | HSP22 | Small heat shock protein%2C chloroplastic (Fragment) (Glycine max OX%3D3847) | Plastid | chloroplast stroma, plastid chloroplast thylakoid membrane, chloroplast thylakoid lumen |
| CqHsp20_48 | CQ037843 | Cq4A | 57885839 | 57888383 | - | 2544 | HSP21.7 | 21.7 kDa class VI heat shock protein (Arabidopsis thaliana OX%3D3702) | Cytoplasm | cytosol |
| CqHsp20_49 | CQ038186 | Cq8A | 2335050 | 2336484 | + | 1434 | HSP15.4 | 15.4 kDa class V heat shock protein (Arabidopsis thaliana OX%3D3702) | Nucleus | nucleolus |
| CqHsp20_50 | CQ038601 | Cq8A | 6861985 | 6863544 | + | 1559 | HSP26.2 | 26.2 kDa heat shock protein%2C mitochondrial (Oryza sativa subsp. japonica OX%3D39947) | Mitochondrion | inner membrane, mitocondrion matrix |
| CqHsp20_51 | CQ041158 | Cq7A | 2563866 | 2564345 | - | 479 | HSP17.6C | 17.6 kDa class I heat shock protein 3 (Arabidopsis thaliana OX%3D3702) | Nucleus | nucleolus |
| CqHsp20_52 | CQ041161 | Cq7A | 2584994 | 2585461 | - | 467 | HSP17.6C | 17.6 kDa class I heat shock protein 3 (Arabidopsis thaliana OX%3D3702) | Nucleus | nucleolus |
| CqHsp20_53 | CQ041411 | Cq7A | 4638026 | 4639642 | - | 1616 | Putative Hsp20 | SHSP domain-containing protein | Nucleus | nucleolus |
| CqHsp20_54 | CQ041626 | Cq7A | 6818599 | 6821690 | + | 3091 | HSP17.4B | 17.4 kDa class III heat shock protein (Arabidopsis thaliana OX%3D3702) | Nucleus | nucleolus |
| CqHsp20_55 | CQ043111 | Cq7A | 46482205 | 46483574 | + | 1369 | HSP22 | Small heat shock protein%2C chloroplastic (Fragment) (Glycine max OX%3D3847) | Mitochondrion | mitocondrion matrix |
| CqHsp20_56 | CQ043112 | Cq7A | 46507093 | 46508453 | + | 1360 | HSP22 | Small heat shock protein%2C chloroplastic (Fragment) (Glycine max OX%3D3847) | Mitochondrion | mitocondrion matrix |
| CqHsp20_57 | CQ044478 | Cq3A | 6321631 | 6323853 | + | 2222 | HSP22 | Small heat shock protein%2C chloroplastic (Petunia hybrida OX%3D4102) | Plastid | chloroplast stroma, plastid chloroplast thylakoid membrane, chloroplast thylakoid lumen |
| CqHsp20_58 | CQ051072 | Cq9A | 36959210 | 36962360 | + | 3150 | HSP14.7 | 14.7 kDa heat shock protein (Arabidopsis thaliana OX%3D3702) | Mitochondrion | inner membrane, mitocondrion matrix |
| CqHsp20_59 | CQ051076 | Cq9A | 36970639 | 36975236 | - | 4597 | hspC2 | Small heat shock protein C2 (Rickettsia felis (strain ATCC VR-1525 / URRWXCal2) OX%3D315456) | Nucleus | chromosome |
| CqHsp20_60 | CQ051646 | Cq9A | 44858084 | 44859935 | + | 1851 | HSP26.5 | 26.5 kDa heat shock protein%2C mitochondrial (Arabidopsis thaliana OX%3D3702) | Mitochondrion | inner membrane, mitocondrion matrix |
| CqHsp20_61 | CQ053725 | Cq2A | 15091148 | 15093074 | - | 1926 | HSP15.7 | 15.7 kDa heat shock protein%2C peroxisomal (Arabidopsis thaliana OX%3D3702) | Nucleus | nucleolus |
| CqHsp20_62 | CQ054519 | Cq2A | 40101462 | 40104471 | + | 3009 | HSP17.9-D | 17.9 kDa class II heat shock protein (Glycine max OX%3D3847) | Nucleus | nucleolus |
| CqHsp20_63 | CQ055373 | Cq2A | 49930925 | 49931410 | - | 485 | HSP18 | 18.3 kDa class I heat shock protein (Oxybasis rubra OX%3D3560) | Nucleus | nucleolus |
| CqHsp20_64 | CQ055378 | Cq2A | 49989213 | 49989705 | + | 492 | HSP17.6C | 17.6 kDa class I heat shock protein 3 (Arabidopsis thaliana OX%3D3702) | Nucleus | nucleolus |
| CqHsp20_65 | CQ055384 | Cq2A | 50041297 | 50041788 | - | 491 | HSP17.6C | 17.6 kDa class I heat shock protein 3 (Arabidopsis thaliana OX%3D3702) | Nucleus | nucleolus |
| CqHsp20_66 | CQ055420 | Cq2A | 50431690 | 50432130 | + | 440 | HSP18.2 | 18.2 kDa class I heat shock protein (Medicago sativa OX%3D3879) | Nucleus | nucleolus |
| CqHsp20_67 | CQ055422 | Cq2A | 50435461 | 50435913 | - | 452 | HSP18.2 | 18.2 kDa class I heat shock protein (Medicago sativa OX%3D3879) | Cytoplasm | cytosol |
| CqHsp20_68 | CQ057039 | Contig2248 | 9641 | 10093 | + | 452 | HSP18.2 | 18.2 kDa class I heat shock protein (Medicago sativa OX%3D3879) | Cytoplasm | cytoskeleton |
| CqHsp20_69 | CQ057041 | Contig2248 | 13424 | 13864 | - | 440 | HSP18.2 | 18.2 kDa class I heat shock protein (Medicago sativa OX%3D3879) | Nucleus | nucleolus |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




