Submitted:
27 April 2026
Posted:
30 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Sampling, DNA Extraction Sequencing and Annotation
2.2. Calculation of Codon Nucleotide Composition and Preference Parameters
2.3. Analysis of Relative Synonymous Codon Usage
2.4. Neutrality Plot Analysis
2.5. ENC-Plot Analysis
2.6. PR2-Plot Bias Analysis
2.7. Determination of Optimal Codons
3. Results
3.1. Characteristics of the Chloroplast Genome of S. japonicum f. oligophyllum
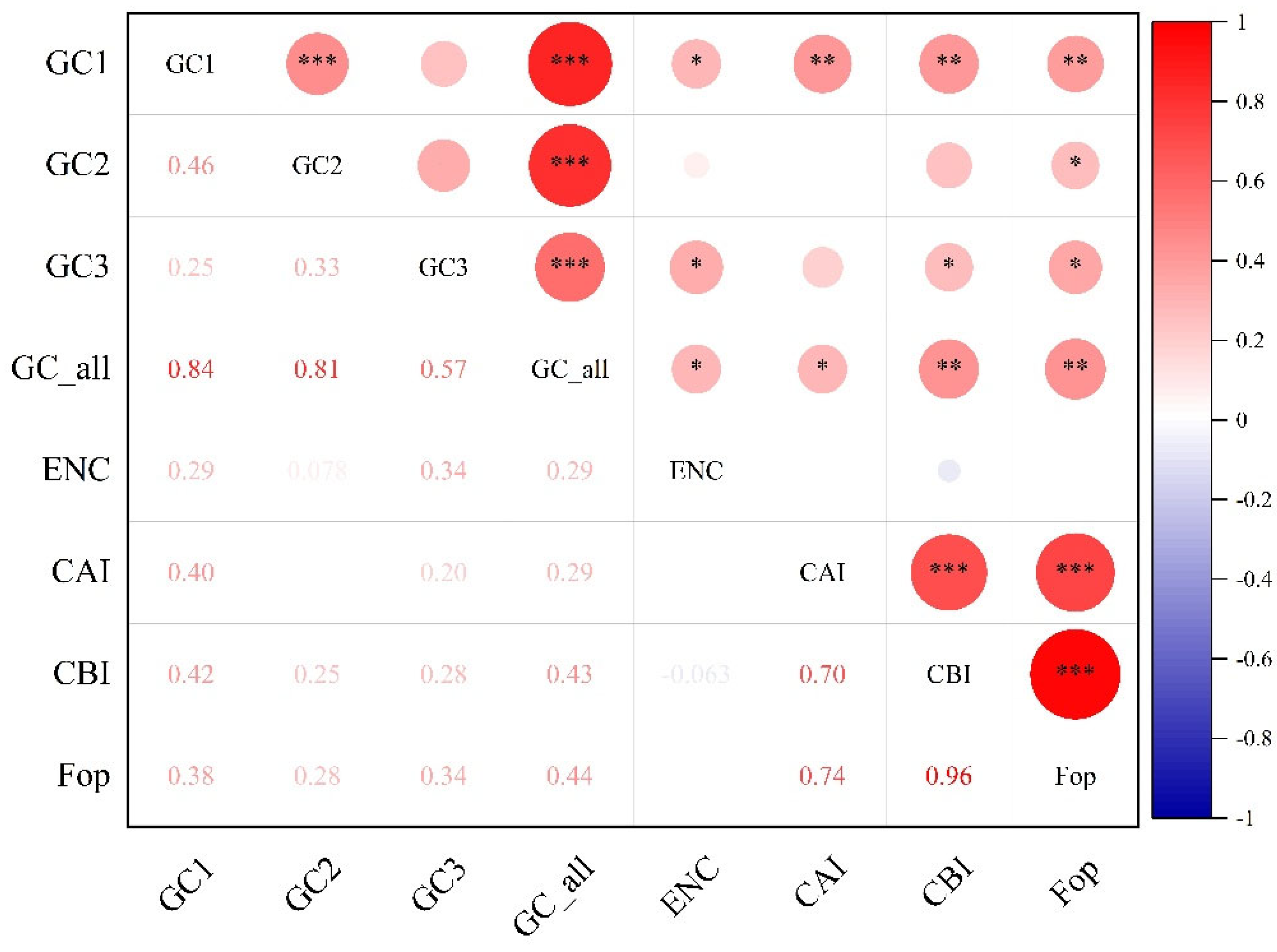
3.2. Codon Composition and Preference Parameter Analysis
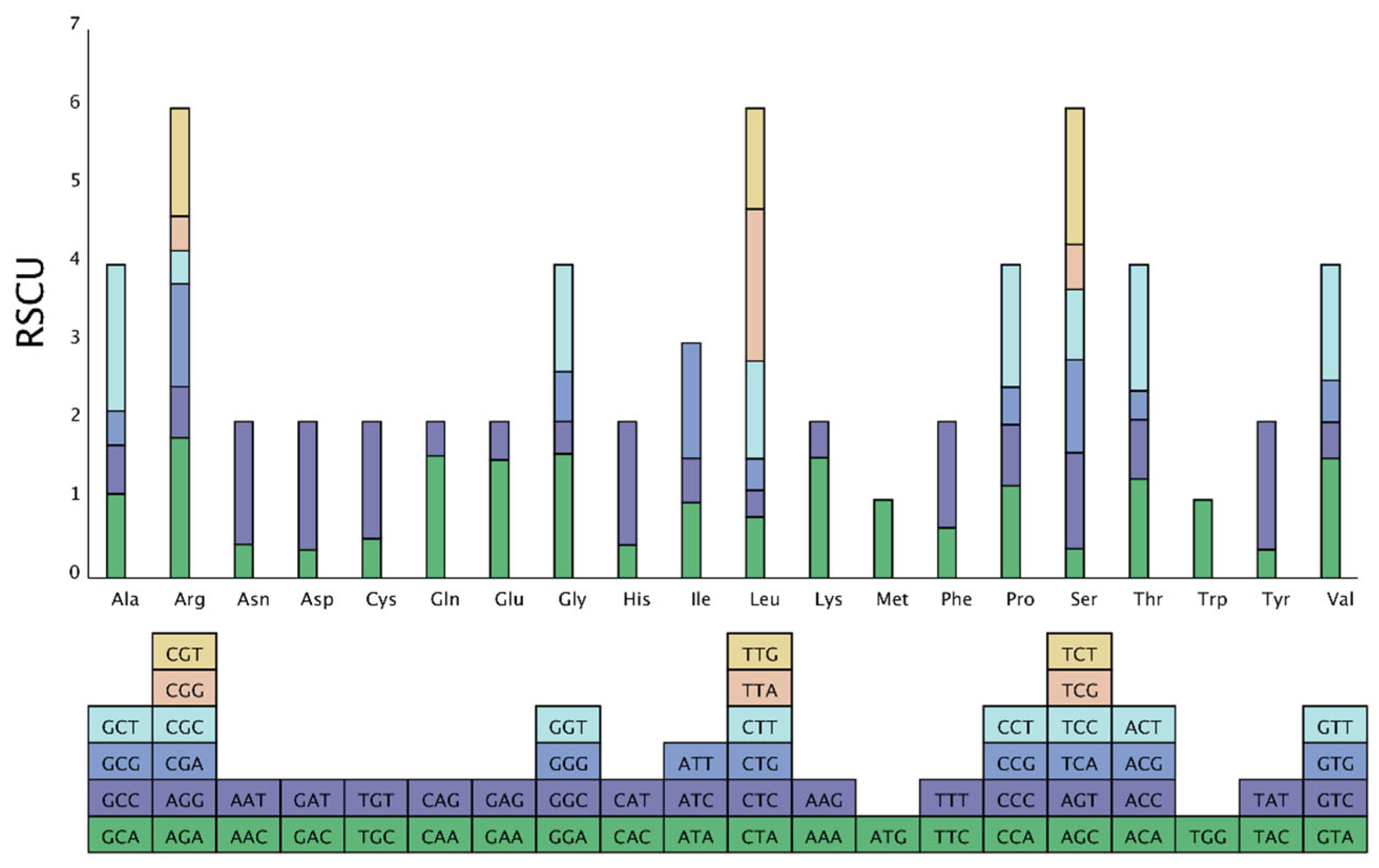
3.3. Analysis of Relative Synonymous Codon Usage
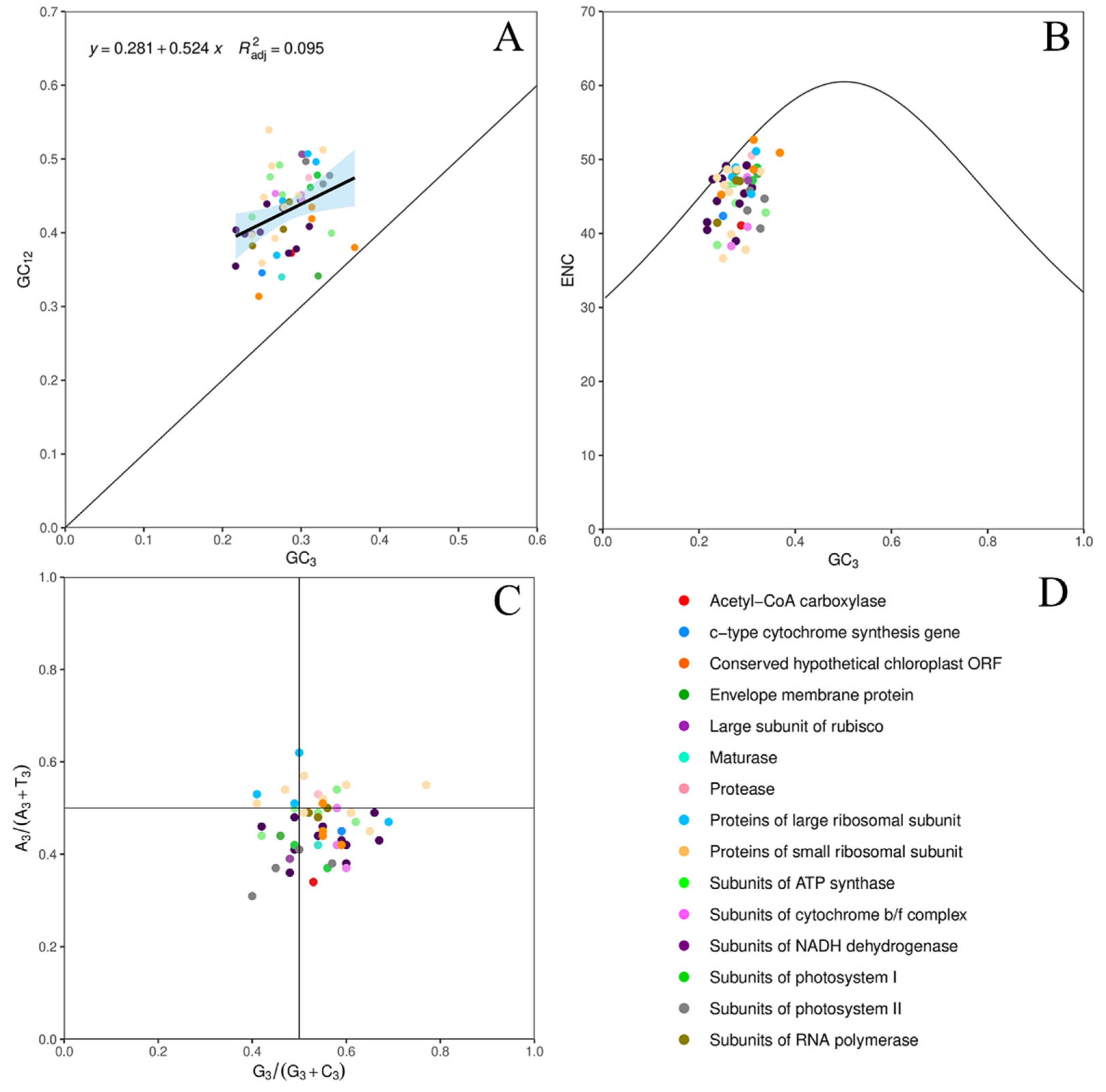
3.4. Neutrality Plot Analysis
3.5. ENC-Plot Analysis
3.6. PR2-Plot Bias Analysis
3.7. Determination of Optimal Codons
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Arimura, S.; Nakazato, I. Genome Editing of Plant Mitochondrial and Chloroplast Genomes. Plant Cell. Physiol. 2024, 65, 477–483. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.; Dong, W.; Li, W.; Lu, Y.; Xie, X.; Jin, X.; Shi, J.; He, K.; Suo, Z. Comparative Analysis of Six Lagerstroemia Complete Chloroplast Genomes. Front. Plant Sci. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Daniell, H.; Lin, C.-S.; Yu, M.; Chang, W.-J. Chloroplast Genomes: Diversity, Evolution, and Applications in Genetic Engineering. Genome Biol. 2016, 17, 134. [Google Scholar] [CrossRef]
- Raubeson, L.A.; Jansen, R.K. Chloroplast genomes of plants. In Plant Diversity and Evolution. Genotypic and Phenotypic Variation in Higher Plants; Henry, R.J., Ed.; CABI Publishing: Wallingford, UK, 2005; Volume 42, pp. 45–68. [Google Scholar]
- Daniell, H.; Kumar, S.; Dufourmantel, N. Breakthrough in Chloroplast Genetic Engineering of Agronomically Important Crops. Trends Biotechnol. 2005, 23, 238–245. [Google Scholar] [CrossRef]
- Bock, R. Genetic Engineering of the Chloroplast: Novel Tools and New Applications. Curr. Opin. Biotechnol. 2014, 26, 7–13. [Google Scholar] [CrossRef]
- Bulle, M.; Rahman, Md.M.; Kota, S.; Islam, Md.R.; Keya, S.S.; Abbagani, S.; Kirti, P.B. Advancing Chloroplast Bioengineering: Innovations, Regulatory Challenges, and Translational Pathways for Sustainable Agriculture. Int. J. Biol. Macromol. 2026, 350, 150873. [Google Scholar] [CrossRef]
- Ingvarsson, P.K. Molecular Evolution of Synonymous Codon Usage in Populus. BMC Evol. Biol. 2008, 8, 307. [Google Scholar] [CrossRef]
- Wang, J.; Qian, J.; Jiang, Y.; Chen, X.; Zheng, B.; Chen, S.; Yang, F.; Xu, Z.; Duan, B. Comparative Analysis of Chloroplast Genome and New Insights Into Phylogenetic Relationships of Polygonatum and Tribe Polygonateae. Front. Plant Sci. 2022, 13. [Google Scholar] [CrossRef]
- Li, C.-L.; Zhang, P.; Cai, C.-L.; Qin, S.-F. Genomic Characteristics and Codon Preference Analysis of Schefflera Octophylla (Lour.) Harms Chloroplasts. J. Agric. Sci. Technol. 2024, 26, 63–76. [Google Scholar]
- Geng, X.; Huang, N.; Zhu, Y.; Qin, L.; Hui, L. Codon Usage Bias Analysis of the Chloroplast Genome of Cassava. South Afr. J. Bot. 2022, 151, 970–975. [Google Scholar] [CrossRef]
- Yao, S.; Zhang, Q.; Su, M. Chloroplast Genome Capture History and Genetie Diversity of Camelia Sinensis Var. Sinensis ’Liupao. Guihaia 2025, 45, 527–541. [Google Scholar]
- Mazumdar, P.; Binti Othman, R.; Mebus, K.; Ramakrishnan, N.; Ann Harikrishna, J. Codon Usage and Codon Pair Patterns in Non-Grass Monocot Genomes. Ann. Bot. 2017, 120, 893–909. [Google Scholar] [CrossRef]
- Hershberg, R.; Petrov, D.A. Selection on Codon Bias. Annu Rev. Genet 2008, 42, 287–299. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Wang, B.; Xiao, S.; Chen, L.; Yin, F.; Li, J.; Jiang, C.; Zhang, D.; Zhong, Q.; Zhang, Y.; et al. Characterization of the Complete Chloroplast Genome and Comparative Analysis of the Phylogeny and Codon Usage Bias of Three Yunnan Wild Rice Species. Front. Plant Sci. 2025, 16. [Google Scholar] [CrossRef] [PubMed]
- Qu, Y.-Y.; Xin, J.; Feng, F.-F. Codon Usage Bais in Chloroplast Genome of Eriobotrya Fragrans Champ. Ex Benth. J. Northwest For. Univ. 2021, 36, 138–144. [Google Scholar]
- Liang, Y.; Kong, Y.-G.; Wang, Y.-H.; Yan, Y.-P. Characteristics and Codon Usage Bias in Chloroplast Genome of Robinia Neo-Mexicana Var. Luxurians. J. Cent. South Univ. For. Technol. 2026, 46, 199–210. [Google Scholar]
- Zhao, Y. Study on Taxonomy and Reproduction Characteristics of Sophora Japonica L.Abstract; Shandong Normal University, 2007. [Google Scholar]
- Mu, Z.-Q. Genetic Diversity of Ancient Styphnolobium Japonicum Trees inHenan; Henan Agricultural University, 2023. [Google Scholar]
- Mu, Z.; Zhang, Y.; Zhang, B.; Cheng, Y.; Shang, F.; Wang, H. Intraspecific Chloroplast Genome Variation and Domestication Origins of Major Cultivars of Styphnolobium Japonicum. Genes 2023, 14. [Google Scholar] [CrossRef]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A New Genome Assembly Algorithm and Its Applications to Single-Cell Sequencing. J. Comput Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An Integrated and Extendable Desktop Software Platform for the Organization and Analysis of Sequence Data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef]
- Cai, Y.-B.; Yang, X.-Y. Codon Usage Bias and Its Influencing Factors in the Chloroplast Genome of Macadamia Integrifolia Maiden & Betche. Plant Sci. J. 2022, 40, 229–239. [Google Scholar] [CrossRef]
- Li, R.-X.; Wang, B.; Xiao, S.-Q. Assembly, Codon Usage Bias, and Phylogenetic Analysis of Chloroplast Genome of Oryza Meyeriana. Chin. J. Rice Sci. 2025, 39, 801–812. [Google Scholar] [CrossRef]
- Chen, W.-Q.; Liu, Z.-H.; Wang, M.-J. Analysis of Codon Usage Bias in Chloroplast Genome of Halenia Elliptica. Mol. Plant Breed. 2024. [Google Scholar]
- Ma, Y.-D.; Yang, Y.-R.; Guo, C.-C. Analysis of Codon Bias in Chloroplast Genome of Cercis Gigantea. J. Agric. Sci. Technol. 2026. [Google Scholar] [CrossRef]
- Dai, J.-F.; Dang, L.; Zhao, X. Characteristics and Codon Usage Bias in Laria Principis-Rupprechtii Chloroplast Genome. J. Northwest For. Univ. 2026, 41, 81–92. [Google Scholar] [CrossRef]
- Lu, Y.; Li, W.-Q.; Xie, X.-M. The Complete Chloroplast Genome Sequence of Sophora Japonica Var. Violacea: Gene Organization and Genomic Resources. Conserv. Genet. Resour. 2018, 10, 1–4. [Google Scholar] [CrossRef]
- Shi, Y.; Liu, B. Complete Chloroplast Genome Sequence of Sophora Japonica ‘JinhuaiJ2’ (Papilionaceae), an Important Traditional Chinese Herb. Mitochondrial DNA B Resour. 5 319–320. [CrossRef]
- Li, P.-Y.; Hong, T.; Chen, X.-L.; He, J.-Y. Characteristics and Phylogenetic Analysis of the Complete Chloroplast Genome of Quercus Lamellosa. South China Agric. 2023, 17, 1–7. [Google Scholar] [CrossRef]
- Chen, C.-H.; Yu, X.-L.; Fu, C. Genomic Sequence Characteristics and Phylogeny of Chloroplast in Cinnamomum Mollifolium. Mol. Plant Breed. 2023, 21, 8066–8074. [Google Scholar] [CrossRef]
- Bhattacharyya, D.; Uddin, A.; Das, S.; Chakraborty, S. Mutation Pressure and Natural Selection on Codon Usage in Chloroplast Genes of Two Species in Pisum L. (Fabaceae: Faboideae). Mitochondrial DNA A DNA Mapp. Seq. Anal. 2019, 30, 664–673. [Google Scholar] [CrossRef]
- Mu, J.-J.; Zhang, J.-S. Codon Usage Bias Analysis in the Chloroplast Genome of Actinostemma Tenerum (Cucurbitaceae). Curr. Issues Mol. Biol. 2025, 47, 833. [Google Scholar] [CrossRef]
- Liang, X.-L.; Guo, S. Codon Usage Bias in the Chloroplast Genome of Sphaerophysa Salsula. J. Northwest For. Univ. 2022, 37, 121–126. [Google Scholar]
- Xiao, M.-K.; Nie, K.-H.; Shen, Z.-B. Analysis of Codon Usage Bias in the Chloroplast Genome of Koelreuteria Bipinnata. J. Southwest For. Univ. (Natural Sciences) 2023, 43, 56–63. [Google Scholar]
- Ding, S.-J.; Wei, J.-S.; Lu, Y.-L. Characteristics and Codon Usage Bias of Picea Brachytyla Var. Complanata chloroplast Genome. J. Cent. South Univ. For. Technol. 43 156–163+190. [CrossRef]
- Liu, T.; Yin, D.-P.; jIN, J.-F. Analysis of Codon Usage Bias in the Chloroplast Genome of Kadsura. J. Northwest For. Univ. 2023, 38, 102–109. [Google Scholar] [CrossRef]
- Li, J.-F.; Yi, X.-L.; Li, X.-Y. Analysis on Codon Usage Bias of Chloroplast Genome in Cladrastis Yun-Chunii X. W. Li & G. S. Fan. Mol. Plant Breed. 2023, 21, 2583–2590. [Google Scholar] [CrossRef]
- Zeng, Y.; Shen, L.; Chen, S.; Qu, S.; Hou, N. Codon Usage Profiling of Chloroplast Genome in Juglandaceae. Forests 2023, 14. [Google Scholar] [CrossRef]
- Long, T.; Dong, W.-P.; Chao, M. Codon Usage Bias Analysis in the Acer Amplum Subsp.Catalpifolium Genome. J. Northwest For. Univ. 2023, 38, 61–66+80. [Google Scholar] [CrossRef]
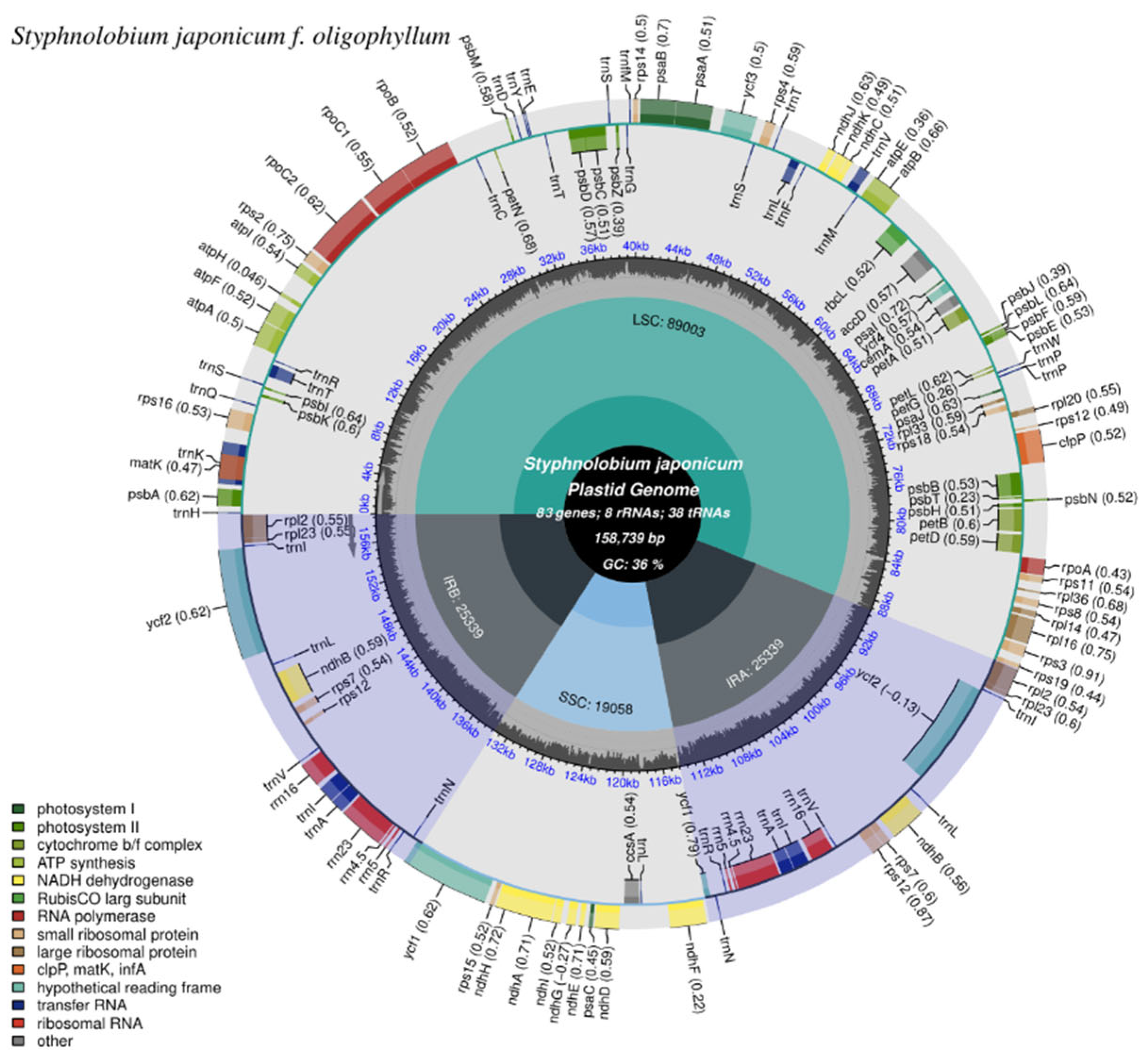
| Category | Gene group | Gene group |
| Photosynthesis | Subunits of photosystem I | psaA,psaB,psaC,psaI,psaJ |
| Subunits of photosystem II | psbA,psbB,psbC,psbD,psbE,psbF,psbH,psbI,psbJ,psbK,psbL,psbM,psbN,psbT,psbZ | |
| Subunits of NADH dehydrogenase | ndhA*,ndhB*(2),ndhC,ndhD,ndhE,ndhF,ndhG,ndhH,ndhI,ndhJ, ndhK | |
| Subunits of cytochrome b/f complex | petA,petB*,petD*,petG,petL,petN | |
| Subunits of ATP synthase | atpA,atpB,atpE,atpF*,atpH,atpI | |
| Large subunit of rubisco | rbcL | |
| Self-replication | Proteins of large ribosomal subunit | rpl14,rpl16*,rpl2*(2),rpl20,rpl23(2),rpl33,rpl36 |
| Proteins of small ribosomal subunit | rps11,rps12*(2),rps14,rps15,rps16*,rps18,rps19,rps2,rps3,rps4, rps7(2),rps8 | |
| Subunits of RNA polymerase | rpoA,rpoB,rpoC1*,rpoC2 | |
| Ribosomal RNAs | rrn16S(2),rrn23S(2),rrn4.5S(2),rrn5S(2) | |
| Transfer RNAs | trnA-UGC*(2),trnC-GCA,trnD-GUC,trnE-UUC,trnF-GAA,trnG-GCC,trnH-GUG,trnI-CAU(2),trnI-GAU*(2),trnK-UUU*,trnL-CAA(2),trnL-UAA*,trnL-UAG,trnM-CAU,trnN-GUU(2),trnP-GGG,trnP-UGG,trnQ-UUG,trnR-ACG(2),trnR-UCU,trnS-GCU,trnS-GGA,trnS-UGA,trnT-CGU*,trnT-GGU,trnT-UGU,trnV-GAC(2),trnV-UAC*,trnW-CCA,trnY-GUA,trnfM-CAU | |
| Other genes | Maturase | matK |
| Protease | clpP** | |
| Envelope membrane protein | cemA | |
| Acetyl-CoA carboxylase | accD | |
| c-type cytochrome synthesis gene | ccsA | |
| Genes of unknown function | Conserved hypothetical chloroplast ORF | ycf1(2),ycf2(2),ycf3**,ycf4 |
| Class range | Class mid value | number | Frequency |
| -0.05~0.05 | 0 | 13 | 0.25 |
| 0.05~0.15 | 0.1 | 27 | 0.52 |
| 0.15~0.25 | 0.2 | 11 | 0.21 |
| 0.25~0.35 | 0.3 | 1 | 0.02 |
| Total | 52 | 0.25 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




