Submitted:
27 April 2026
Posted:
30 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
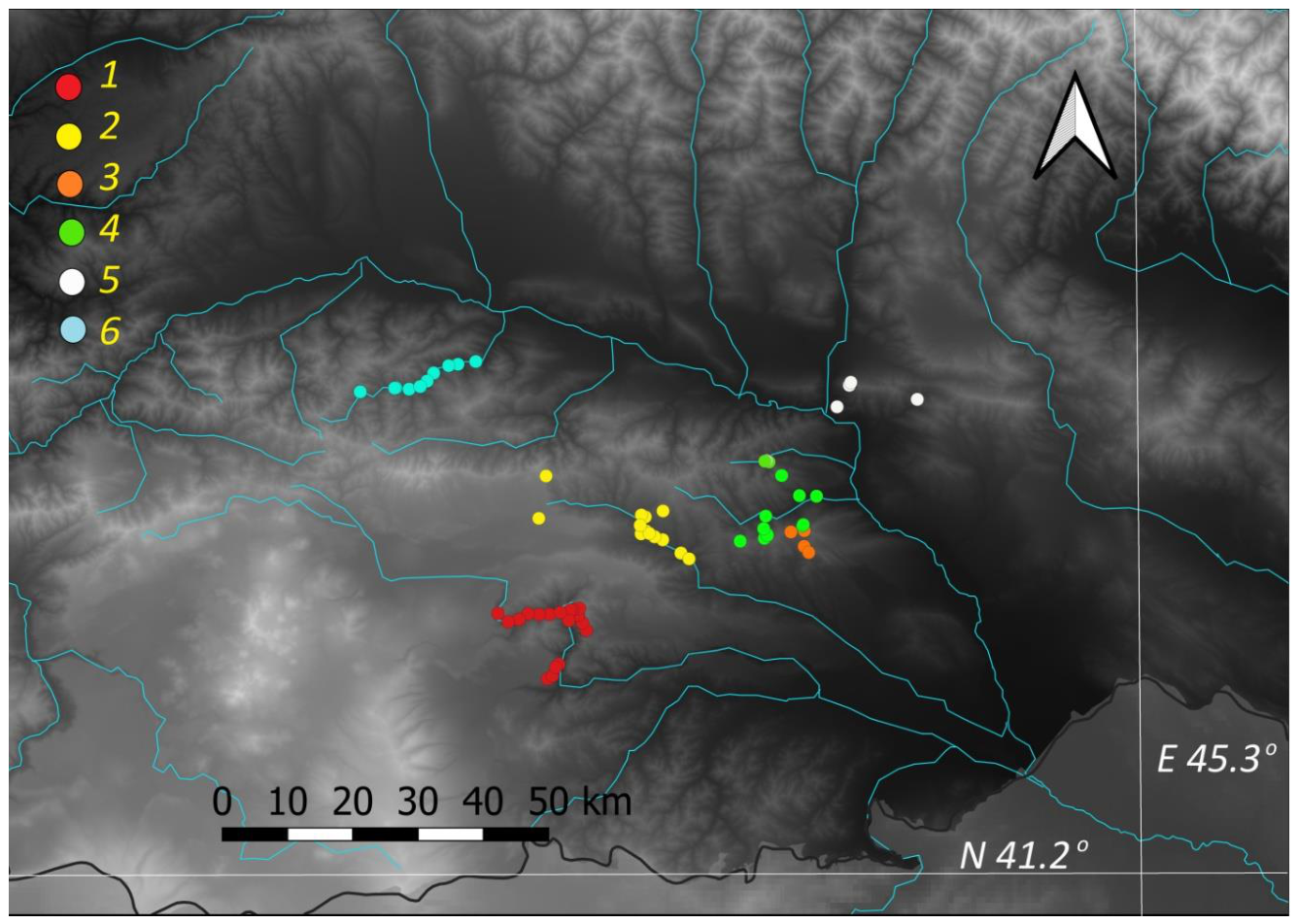
2.1. Studied System
2.2. Scoring Microsatellite Genotypes
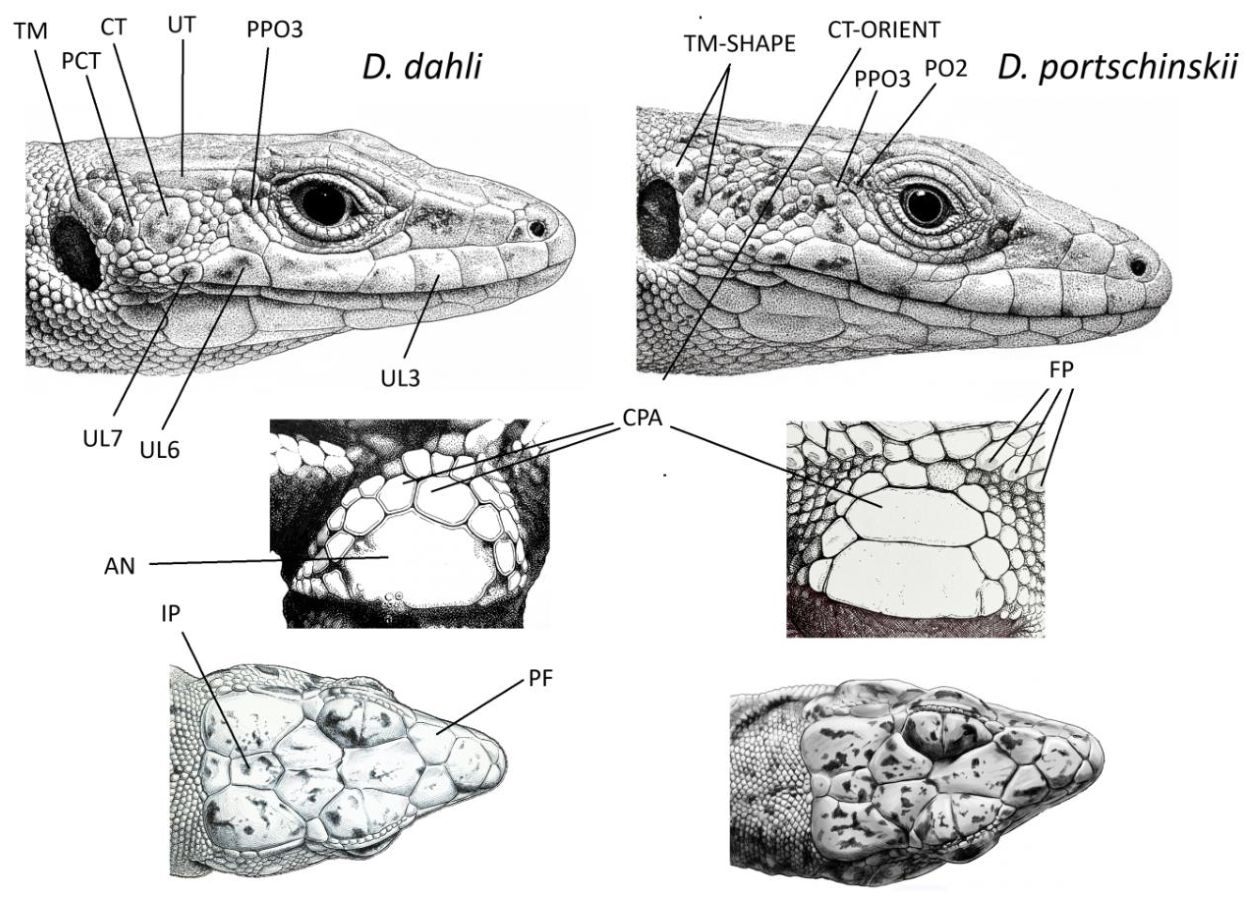
2.3. Describing Scalation
2.4. Data Analysis
2.4.1. Variability Within the Metapopulations.
2.4.2. Uniqueness of Individuals and Metapopulations.
2.4.3. Variability Among the Metapopulations.
2.4.4. Association of Dissimilarities Between Phenotypes, Genotypes and Environmental Conditions.
2.4.5. Association Analysis of Individual and Climate Differences, and Comparison of Phenotypic, Genotypic and Geographic Distances Between Metapopulations.
2.5. Software Used
3. Results
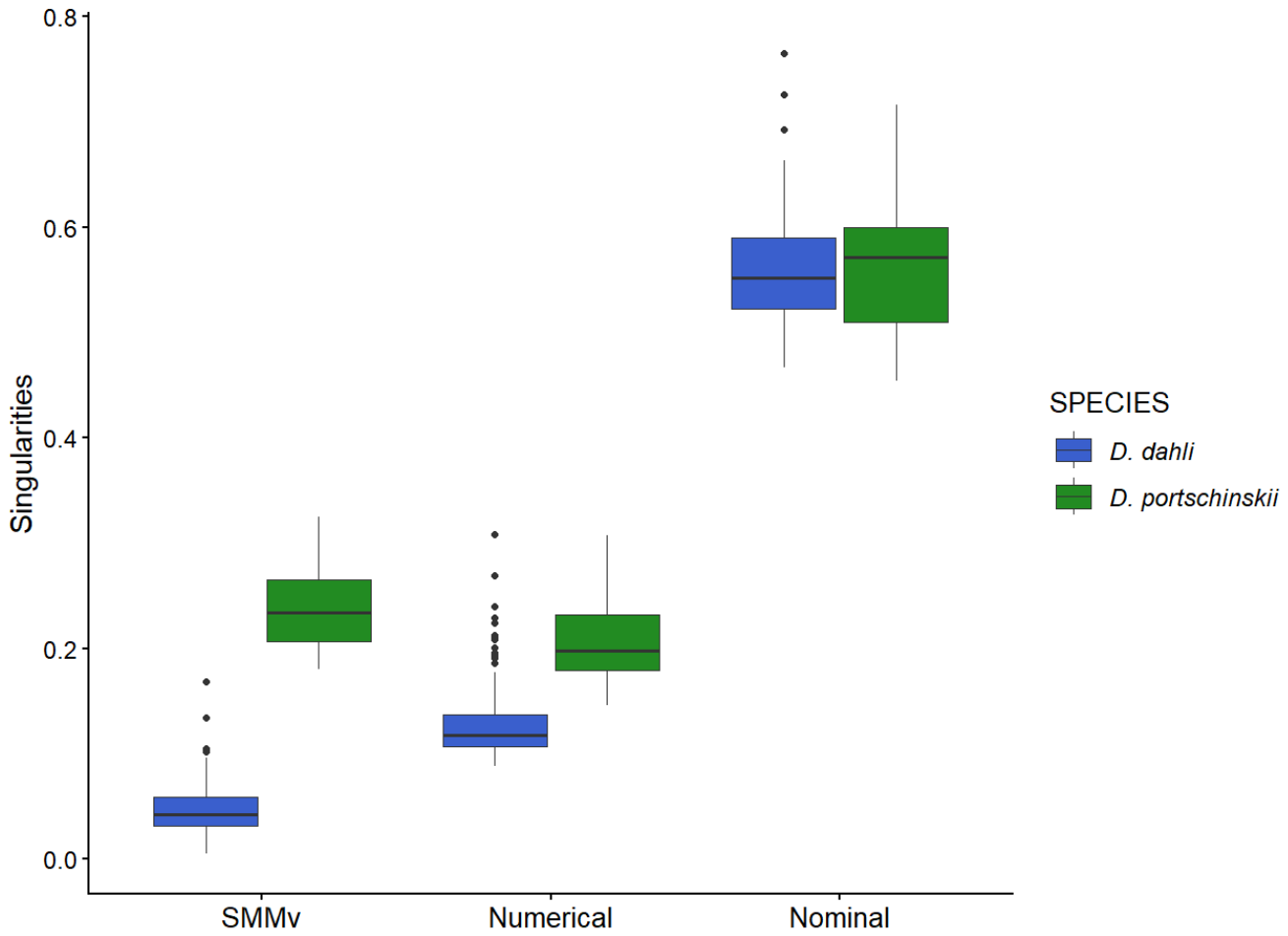
3.1. Variability Within each Species and Metapopulations
3.2. Differentiation Among Species and Metapopulations
3.3. Uniqueness of Individuals and Metapopulations
3.4. Association Between Individual Phenotypic and Genotypic Differences
3.5. Association of Genotypes and Phenotypes with Geography and Environmental Variables
4. Discussion
Supplementary Materials
Author Contributions
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Barateli, N.; Iankoshvili, G.; Tarkhnishvili, D.; Kokiashvili, L. Reproductive effort of unisexual and bisexual rock lizards (genus Darevskia). Zoologischer Anzeiger 301 2022, 196–204. [Google Scholar] [CrossRef]
- Bell, G. The Masterpiece of Nature: The Evolution and Genetics of Sexuality; University of California Press, 1982. [Google Scholar]
- Deeming, D. C. Deeming, D. C., Ed.; Post-hatching phenotypic effects of incubation in reptiles. In Reptilian incubation: Environment, evolution and behaviour; Nottingham; Nottingham University Press, 2004; pp. 229–252. [Google Scholar]
- Ferris, H.; Wilson, L. T. Concepts and principles of population dynamics. In Vistas on nematology: A commemoration of the twenty-fifth anniversary of the Society of Nematologists; 1987; pp. 372–376. [Google Scholar]
- Freitas, S.; Westram, A. M.; Schwander, T.; Arakelyan, M.; Ilgaz, Ç.; Kumlutaş, Y.; Butlin, R. K. Parthenogenesis in Darevskia lizards: A rare outcome of common hybridization, not a common outcome of rare hybridization. Evolution 2022, 76(5), 899–914. [Google Scholar] [CrossRef]
- Fujita, M. K.; McGuire, J. A.; Donnellan, S. C.; Moritz, C. Diversification and persistence at the arid–monsoonal interface: Australia-wide biogeography of the Bynoe’s gecko (Heteronotia binoei; Gekkonidae). Evolution 2010, 64(8), 2293–2314. [Google Scholar] [CrossRef]
- Falconer, D. S. Selection in different environments: effects on environmental sensitivity (reaction norm) and on mean performance. Genetics Research 1990, 56(1), 57–70. [Google Scholar] [CrossRef]
- West-Eberhard, M. J. Developmental plasticity and evolution; Oxford University Press, 2003. [Google Scholar]
- Burggren, W. W.; Mendez-Sanchez, J. F. “Bet hedging” against climate change in developing and adult animals: roles for stochastic gene expression, phenotypic plasticity, epigenetic inheritance and adaptation. Frontiers in Physiology 2023, 14, 1245875. [Google Scholar] [CrossRef] [PubMed]
- Fusco, G.; Minelli, A. Phenotypic plasticity in development and evolution: Facts and concepts. Philosophical Transactions of the Royal Society B: Biological Sciences 2010, 365(1540), 547–556. [Google Scholar] [CrossRef] [PubMed]
- Glesener, R. R.; Tilman, D. Sexuality and the components of environmental uncertainty: Clues from geographic parthenogenesis in terrestrial animals. American Naturalist 1978, 112(986), 659–673. [Google Scholar] [CrossRef] [PubMed]
- Goldschmidt, R. B. Gen und Außeneigenschaft: (Untersuchungen an Drosophila) II. Zeitschrift für induktive Abstammungs- und Vererbungslehre 69 1935, 70–131. [Google Scholar] [CrossRef]
- Goldschmidt, R. B. Phenocopies. Scientific American 1949, 181(4), 46–49. [Google Scholar] [CrossRef]
- Gultyaeva, E. I.; Shaydayuk, E. L.; Kosman, E. Regional and temporal differentiation of virulence phenotypes of Puccinia triticina Eriks. from common wheat in Russia during the period 2001–2018. Plant Pathology 69 2020, 860–871. [Google Scholar] [CrossRef]
- Heimpel, G. E.; De Boer, J. G. Sex determination in the Hymenoptera. Annual Review of Entomology 2008, 53(1), 209–230. [Google Scholar] [CrossRef]
- Jablonka, E.; Lamb, M. J. Epigenetic inheritance and evolution: The Lamarckian dimension; Oxford; Oxford University Press, 1995. [Google Scholar]
- Janick, J. Coors, J. G., Pandey, S., Eds.; Exploitation of heterosis: Uniformity and stability. In The genetics and exploitation of heterosis in crops; Madison, WI; ASA–CSSA–SSSA, 1999; pp. 319–333. [Google Scholar]
- Kearney, M.; Fujita, M. K.; Ridenour, J. Schön, I., Martens, K., van Dijk, P., Eds.; Lost sex in the reptiles: Constraints and correlations. In Lost sex: The evolutionary biology of parthenogenesis; Dordrecht; Springer, 2009; pp. 447–474. [Google Scholar]
- Kearney, M.; Shine, R. Developmental success, stability, and plasticity in closely related parthenogenetic and sexual lizards (Heteronotia, Gekkonidae). Evolution 2004, 58(7), 1560–1572. [Google Scholar] [CrossRef] [PubMed]
- Knobil, E.; Neill, J. D. (Eds.) Encyclopedia of reproduction; New York; Academic Press, 1998. [Google Scholar]
- Kondrashov, A. S. Deleterious mutations and the evolution of sexual reproduction. Nature 1988, 336(6198), 435–440. [Google Scholar] [CrossRef]
- Korchagin, V. I.; Badaeva, T. N.; Tokarskaya, O. N.; Martirosyan, I. A.; Darevsky, I. S.; Ryskov, A. P. Molecular characterization of allelic variants of (GATA)n microsatellite loci in parthenogenetic lizards Darevskia unisexualis (Lacertidae). Gene 2007, 392(1–2), 126–133. [Google Scholar] [CrossRef]
- Kosman, E. Difference and diversity of plant pathogen populations: A new approach for measuring. Phytopathology 86 1996, 1152–1155. [Google Scholar]
- Kosman, E. Measuring diversity: From individuals to populations. European Journal of Plant Pathology 138 2014, 467–486. [Google Scholar] [CrossRef]
- Kosman, E.; Ben-Yehuda, P.; Manisterski, J. Diversity of virulence phenotypes among annual populations of wheat leaf rust in Israel from 1993 to 2008. Plant Pathology 63 2014, 563–571. [Google Scholar] [CrossRef]
- Kosman, E.; Burgio, K. R.; Presley, S. J.; Willig, M. R.; Scheiner, S. M. Conservation prioritization based on trait-based metrics illustrated with global parrot distributions. Diversity and Distributions 25 2019a, 1156–1165. [Google Scholar] [CrossRef]
- Kosman, E.; Chen, X.; Dreiseitl, A.; McCallum, B.; Lebeda, A.; Ben-Yehuda, P.; et al. Functional variation of plant–pathogen interactions: New concept and methods for virulence data analyses. Phytopathology 109 2019b, 1324–1330. [Google Scholar] [CrossRef]
- Kosman, E.; Feijen, F.; Jokela, J. Effective number of different populations: A new concept and how to use it. Ecology and Evolution 14 2024, e70303. [Google Scholar] [CrossRef]
- Kosman, E.; Jokela, J. Dissimilarity of individual microsatellite profiles under different mutation models – empirical approach. Ecology and Evolution 9 2019, 4038–4054. [Google Scholar] [CrossRef]
- Kosman, E.; Leonard, K. J. Similarity coefficients for molecular markers in studies of genetic relationships between individuals for haploid, diploid, and polyploid species. Molecular Ecology 14 2005, 415–424. [Google Scholar] [CrossRef] [PubMed]
- Kosman, E.; Leonard, K. J. Conceptual analysis of methods applied to assessment of diversity within and distance between populations with asexual or mixed mode of reproduction. New Phytologist 174 2007, 683–696. [Google Scholar] [CrossRef]
- Lampert, K. P. Facultative parthenogenesis in vertebrates: Reproductive error or chance? Sexual Development 2009, 2(6), 290–301. [Google Scholar] [CrossRef] [PubMed]
- Leary, R. F.; Allendorf, F. W.; Knudsen, K. L. Developmental instability and high meristic counts in interspecific hybrids of salmonid fishes. Evolution 1985, 39(6), 1318–1326. [Google Scholar] [CrossRef] [PubMed]
- Lenz, W. Phenocopies. Journal of Medical Genetics 1973, 10(1), 34. [Google Scholar] [CrossRef]
- Lorenzon, P.; Clobert, J.; Massot, M. The contribution of phenotypic plasticity to adaptation in Lacerta vivipara. Evolution 2001, 55(2), 392–404. [Google Scholar] [CrossRef]
- MacArthur, R. H.; Wilson, E. O. The theory of island biogeography; Princeton; Princeton University Press, 2001; Vol. 1. [Google Scholar]
- Markow, T. A. Evolutionary ecology and developmental instability. Annual Review of Entomology 1995, 40(1), 105–120. [Google Scholar] [CrossRef]
- Maynard Smith, J. Evolutionary genetics; Oxford; Oxford University Press, 1998. [Google Scholar]
- Microsoft Corporation. Microsoft Excel (Version 16.0) [Computer software]. 2021. Available online: https://www.microsoft.com/.
- Miller, C. R.; Joyce, P.; Waits, L. P. Assessing allelic dropout and genotype reliability using maximum likelihood. Genetics 2002, 160(1), 357–366. [Google Scholar] [CrossRef]
- Mittwoch, U. Parthenogenesis. Journal of Medical Genetics 1978, 15(3), 165. [Google Scholar] [CrossRef]
- Moritz, C. The origin and evolution of parthenogenesis in Heteronotia binoei (Gekkonidae): Evidence for recent and localized origins of widespread clones. Genetics 1991, 129(1), 211–219. [Google Scholar] [CrossRef]
- Murphy, R. W.; Fu, J.; MacCulloch, R. D.; Darevsky, I. S.; Kupriyanova, L. A. A fine line between sex and unisexuality: The phylogenetic constraints on parthenogenesis in lacertid lizards. Zoological Journal of the Linnean Society 2000, 130(4), 527–549. [Google Scholar] [CrossRef]
- Nijhout, H. F.; Davidowitz, G. Markow, T. A., Ed.; Developmental perspectives on phenotypic variation, canalization, and fluctuating asymmetry. In Developmental instability: Causes and consequences; 2003; pp. 3–13. [Google Scholar]
- Noble, D. W.; Stenhouse, V.; Schwanz, L. E. Developmental temperatures and phenotypic plasticity in reptiles: A systematic review and meta-analysis. Biological Reviews 2018, 93(1), 72–97. [Google Scholar] [CrossRef]
- Ostberg, C. O.; Duda, J. J.; Graham, J. H.; Zhang, S.; Haywood, K. P., III; Miller, B.; Lerud, T. L. Growth, morphology, and developmental instability of rainbow trout, Yellowstone cutthroat trout, and four hybrid generations. Transactions of the American Fisheries Society 2011, 140(2), 334–344. [Google Scholar] [CrossRef]
- Pandian, T. J. Reproduction and development in Crustacea; Boca Raton; CRC Press, 2016. [Google Scholar]
- Pfennig, D. W.; Wund, M. A.; Snell-Rood, E. C.; Cruickshank, T.; Schlichting, C. D.; Moczek, A. P. Phenotypic plasticity’s impacts on diversification and speciation. Trends in Ecology & Evolution 2010, 25(8), 459–467. [Google Scholar]
- Pompanon, F.; Bonin, A.; Bellemain, E.; Taberlet, P. Genotyping errors: Causes, consequences and solutions. Nature Reviews Genetics 2005, 6(11), 847–859. [Google Scholar] [CrossRef]
- Sakich, N. B.; Tattersall, G. J. Bearded dragons (Pogona vitticeps) with reduced scalation lose water faster but do not have substantially different thermal preferences. Journal of Experimental Biology 2021, 224(12), jeb234427. [Google Scholar] [CrossRef]
- Scheiner, S. M.; Kosman, E.; Presley, S. J.; Willig, M. R. Decomposing functional diversity. Methods in Ecology and Evolution 2017, 8(7), 809–820. [Google Scholar] [CrossRef]
- Simon, J. C.; Stoeckel, S.; Tagu, D. Evolutionary and functional insights into reproductive strategies of aphids. Comptes Rendus Biologies 2010, 333(6–7), 488–496. [Google Scholar] [CrossRef]
- Slatkin, M. Inbreeding coefficients and coalescence times. Genetic Research 1991, 58(2), 167–175. [Google Scholar] [CrossRef]
- Slatkin, M. Isolation by distance in equilibrium and non-equilibrium populations. Evolution 1993, 47(1), 264–279. [Google Scholar] [CrossRef]
- Sneath, P. A.; Sokal, R. R. Numerical taxonomy; San Francisco; W. H. Freeman, 1973. [Google Scholar]
- Subramoniam, T. Mode of reproduction: Invertebrate animals. Encyclopedia of Reproduction 6 2018, 32–40. [Google Scholar]
- Sun, X.; Kosman, E.; Sharon, O.; Ezrati, S.; Sharon, A. Significant host- and environment-dependent differentiation among highly sporadic fungal endophyte communities in cereal crops-related wild grasses. Environmental Microbiology 22 2020, 3357–3374. [Google Scholar] [CrossRef] [PubMed]
- Tarkhnishvili, D.; Avaliani, A.; Gavashelishvili, A.; Murtskhvaladze, M.; Mumladze, L. Unisexual rock lizard might be outcompeting its bisexual progenitors in the Caucasus. Biological Journal of the Linnean Society 2010, 101(2), 447–460. [Google Scholar] [CrossRef]
- Tarkhnishvili, D. Evolutionary history, habitats, diversification, and speciation in Caucasian rock lizards. Advances in Zoological Research 2 2012, 79–120. [Google Scholar]
- Tarkhnishvili, D.; Murtskhvaladze, M.; Anderson, C. L. Coincidence of genotypes at two loci in two parthenogenetic rock lizards: How backcrosses might trigger adaptive speciation. Biological Journal of the Linnean Society 2017, 121(2), 365–378. [Google Scholar] [CrossRef]
- Tarkhnishvili, D.; Barateli, N.; Murtskhvaladze, M.; Iankoshvili, G. Estimating phenotypic heritability of sexual and unisexually reproducing rock lizards (genus Darevskia). Zoologischer Anzeiger 285 2020b, 105–113. [Google Scholar] [CrossRef]
- Tarkhnishvili, D.; Yanchukov, A.; Şahin, M. K.; Gabelaia, M.; Murtskhvaladze, M.; Candan, K.; et al. Genotypic similarities among the parthenogenetic Darevskia rock lizards with different hybrid origins. BMC Evolutionary Biology 20 2020a, 1–25. [Google Scholar] [CrossRef]
- Tarkhnishvili, D.; Yanchukov, A.; Böhne, A. Advantages, limitations, and evolutionary constraints of asexual reproduction: An empirical approach. Frontiers in Ecology and Evolution 11 2023, 1184306. [Google Scholar] [CrossRef]
- Thompson, K. A.; Brandvain, Y.; Coughlan, J. M.; Delmore, K. E.; Justen, H.; Linnen, C. R.; et al. The ecology of hybrid incompatibilities. Cold Spring Harbor Perspectives in Biology 2024, 16(9), a041440. [Google Scholar] [CrossRef]
- Vrijenhoek, R. C.; Parker, E. D. Schön, I., Martens, K., van Dijk, P., Eds.; Geographical parthenogenesis: General purpose genotypes and frozen niche variation. In Lost sex: The evolutionary biology of parthenogenesis; Dordrecht; Springer, 2009; pp. 99–131. [Google Scholar]
- Warren, W. C.; García-Pérez, R.; Xu, S.; Lampert, K. P.; Chalopin, D.; Stöck, M.; et al. Clonal polymorphism and high heterozygosity in the celibate genome of the Amazon molly. Nature Ecology & Evolution 2018, 2(4), 669–679. [Google Scholar] [CrossRef]
- Weir, B. S. Genetic data analysis: Methods for discrete population genetic data; New York; Springer, 1990. [Google Scholar]
- Wright, J. W.; Lowe, C. H. Weeds, polyploids, parthenogenesis, and the geographical and ecological distribution of all-female species of Cnemidophorus. Copeia 1968, 1968(1), 128–138. [Google Scholar] [CrossRef]
- Wright, S. Isolation by distance. Genetics 1943, 28(2), 114. [Google Scholar] [CrossRef]
- Wright, S. Evolution and the genetics of populations . In Genetic and biometric foundations; Chicago; University of Chicago Press, 1984; Volume 1. [Google Scholar]
- Waits, L. P.; Luikart, G.; Taberlet, P. Estimating the probability of identity among genotypes in natural populations: Cautions and guidelines. Molecular Ecology 2001, 10(1), 249–256. [Google Scholar] [CrossRef]
- Yanchukov, A.; Tarkhnishvili, D.; Erdolu, M.; Şahin, M. K.; Candan, K.; Murtskhvaladze, M.; et al. Precise paternal ancestry of hybrid unisexual ZW lizards (genus Darevskia: Lacertidae: Squamata) revealed by Z-linked genomic markers. Biological Journal of the Linnean Society 2022, 136(2), 293–305. [Google Scholar] [CrossRef]
| D. dahl | D. portschinskii a | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| KW estimate b / nENDI estimate c | KW estimate / nENDI estimate | ||||||||
| Location | N | SMMv d | Numeric e | Nominal f | N | SMMv | Numeric | Nominal | |
| 1 | 13 | 0.051 / 0.037 | 0.173 / 0.132 | 0.771 / 0.556 | 2 | 0.228 / 0.228 | 0.235 / 0.187 | 0.733 / 0.613 | |
| 2 | 8 | 0.091 / 0.088 | 0.184/ 0.132 | 0.816/ 0.570 | 4 | 0.312 / 0.307 | 0.210 / 0.202 | 0.800 / 0.650 | |
| 3 | 13 | 0.055 / 0.048 | 0.164/ 0.128 | 0.756 / 0.555 | 6 | 0.207 / 0.203 | 0.284 / 0.243 | 0.533 / 0.440 | |
| 4 | 6 | 0.125 / 0.115 | 0.163 / 0.116 | 0.787/ 0.570 | 4 | 0.220 / 0.208 | 0.237 / 0.192 | 0.775 / 0.577 | |
| 5 | 9 | 0.054 / 0.041 | 0.155/ 0.138 | 0.686 / 0.588 | 7 | 0.338 / 0.328 | 0.215 / 0.193 | 0.643/ 0.581 | |
| 6 | 5 | 0.074 / 0.067 | 0.168 / 0.142 | 0.678 / 0.557 | 4 | 0.206 / 0.201 | 0.260 / 0.220 | 0.650 / 0.518 | |
| Average | 0.075 / 0.066 | 0.168 / 0.131 | 0.749 / 0.566 | 0.252 / 0.246 | 0.240 / 0.206 | 0.689/ 0.563 | |||
| Pool | 54 | 0.083 / 0.072 | 0.192 / 0.188 | 0.834 / 0.836 | 27 | 0.349 / 0.339 | 0.297 / 0.298 | 0.830 / 0.841 | |
| D. dahli / D. portschinskii a | |||
|---|---|---|---|
| SMMv b | Numeric c | Nominal d | |
| Coefficient of differentiation | 0.020 / 0.120 | 0.019 / 0.063 | 0.059 / 0.149 |
| Significance of differentiation (p-value) | 0.001 / 0.010 | 0.001 / 0.001 | 0.001 / 0.022 |
| ENDPe | 1.22 / 1.90 | 1.45 / 1.99 | 2.98 / 3.58 |
| ED = nENDPf | 0.045 / 0.180 | 0.064 / 0.141 | 0.282 / 0.368 |
| D. dahli / D. portschinskii a | |||
|---|---|---|---|
| SMMv b | Numeric c | Nominal d | |
| Mean, M | 0.052 / 0.240 | 0.130 / 0.208 | 0.559 / 0.563 |
| Standard Deviation, SD | 0.032 / 0.040 | 0.041 / 0.041 | 0.049 / 0.064 |
| Relative range, RR e | 0.970 / 0.446 | 0.716 / 0.525 | 0.389 / 0.366 |
| Coefficient of variation, CV f | 0.615 / 0.167 | 0.315 / 0.197 | 0.088 / 0.114 |
| species | phenotypes | predictors | n_locd | Re | Pf | N_indg | R | P |
|---|---|---|---|---|---|---|---|---|
| D. dahli | numeric | geodista | 65 | 0.152 | 0.020 | 186 | 0.088 | 0.010 |
| D. dahli | numeric | BIO10b | 65 | 0.040 | 0.231 | 186 | -0.002 | 0.499 |
| D. dahli | numeric | BIO12c | 65 | 0.220 | 0.021 | 186 | 0.105 | 0.026 |
| D. dahli | nominal | geodist | 65 | 0.071 | 0.058 | 186 | 0.079 | 0.000 |
| D. dahli | nominal | BIO10 | 65 | -0.047 | 0.845 | 186 | -0.002 | 0.529 |
| D. dahli | nominal | BIO12 | 65 | 0.025 | 0.326 | 186 | 0.000 | 0.491 |
| D. portschinskii | numeric | geodist | 38 | 0.096 | 0.088 | 54 | 0.106 | 0.018 |
| D. portschinskii | numeric | BIO10 | 38 | 0.155 | 0.042 | 54 | 0.048 | 0.259 |
| D. portschinskii | numeric | BIO12 | 38 | 0.100 | 0.118 | 54 | 0.046 | 0.250 |
| D. portschinskii | nominal | geodist | 38 | 0.096 | 0.053 | 54 | 0.120 | 0.002 |
| D. portschinskii | nominal | BIO10 | 38 | 0.092 | 0.095 | 54 | 0.098 | 0.045 |
| D. portschinskii | nominal | BIO12 | 38 | 0.113 | 0.054 | 54 | 0.093 | 0.051 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




