Submitted:
28 April 2026
Posted:
30 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
- We propose SpectraLLM, the first unified LLM-driven framework that jointly processes MS and NMR spectra through a modality-agnostic encoder with contribution-aware dynamic fusion, enabling effective multi-modal biological spectrum analysis.
- We introduce flow-based knowledge distillation for spectral LLMs, achieving near-teacher performance with significantly reduced computational cost, making large-scale spectral analysis feasible in clinical settings.
- We demonstrate through extensive experiments on three benchmarks that SpectraLLM achieves state-of-the-art performance across diverse biological spectrum analysis tasks, while requiring only 5% trainable parameters for domain adaptation via parameter-efficient transfer learning.
2. Results
2.1. Main Benchmark Comparison
2.2. Cross-Dataset Generalization
2.3. Ablation Study
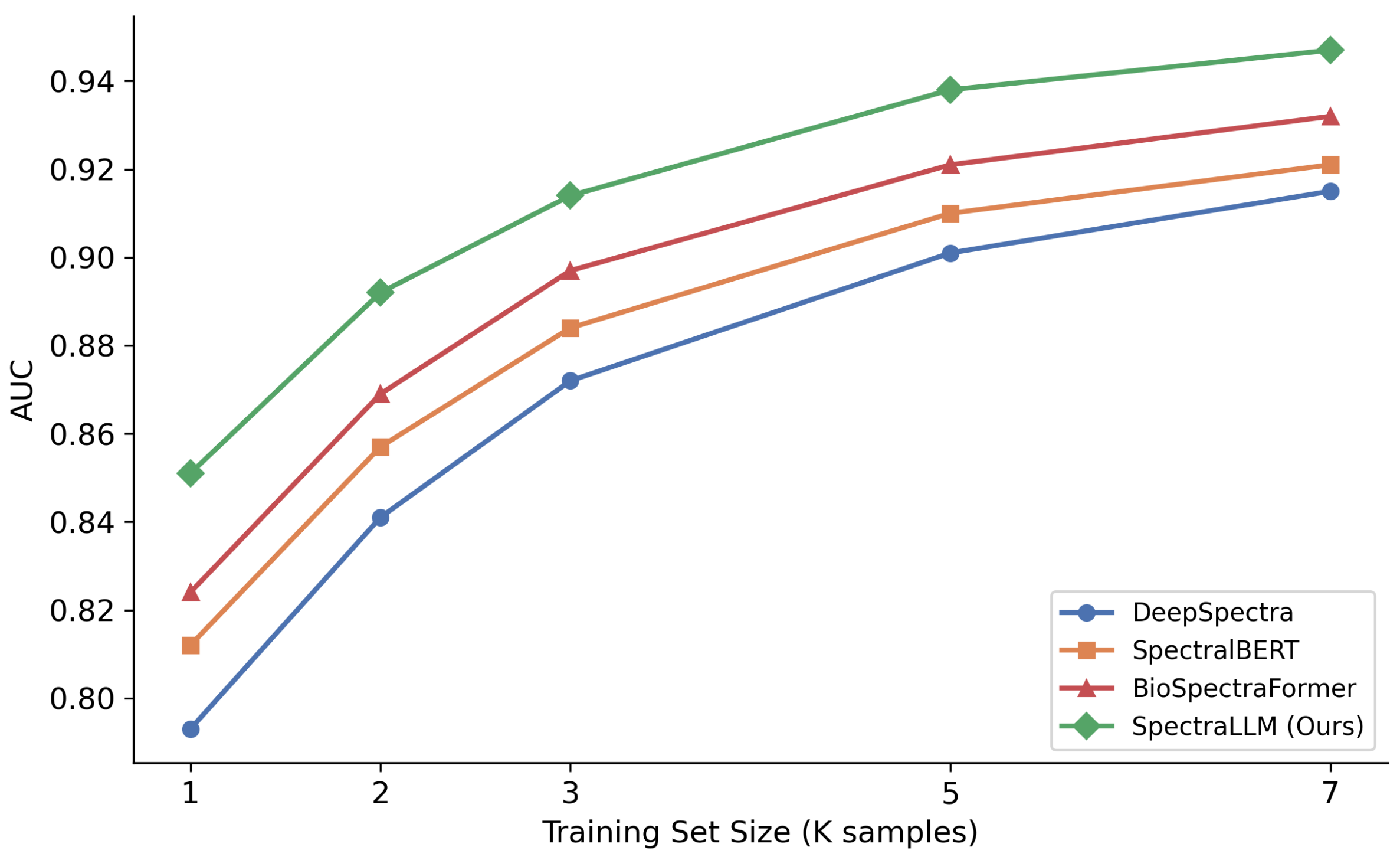
2.4. Scaling Analysis
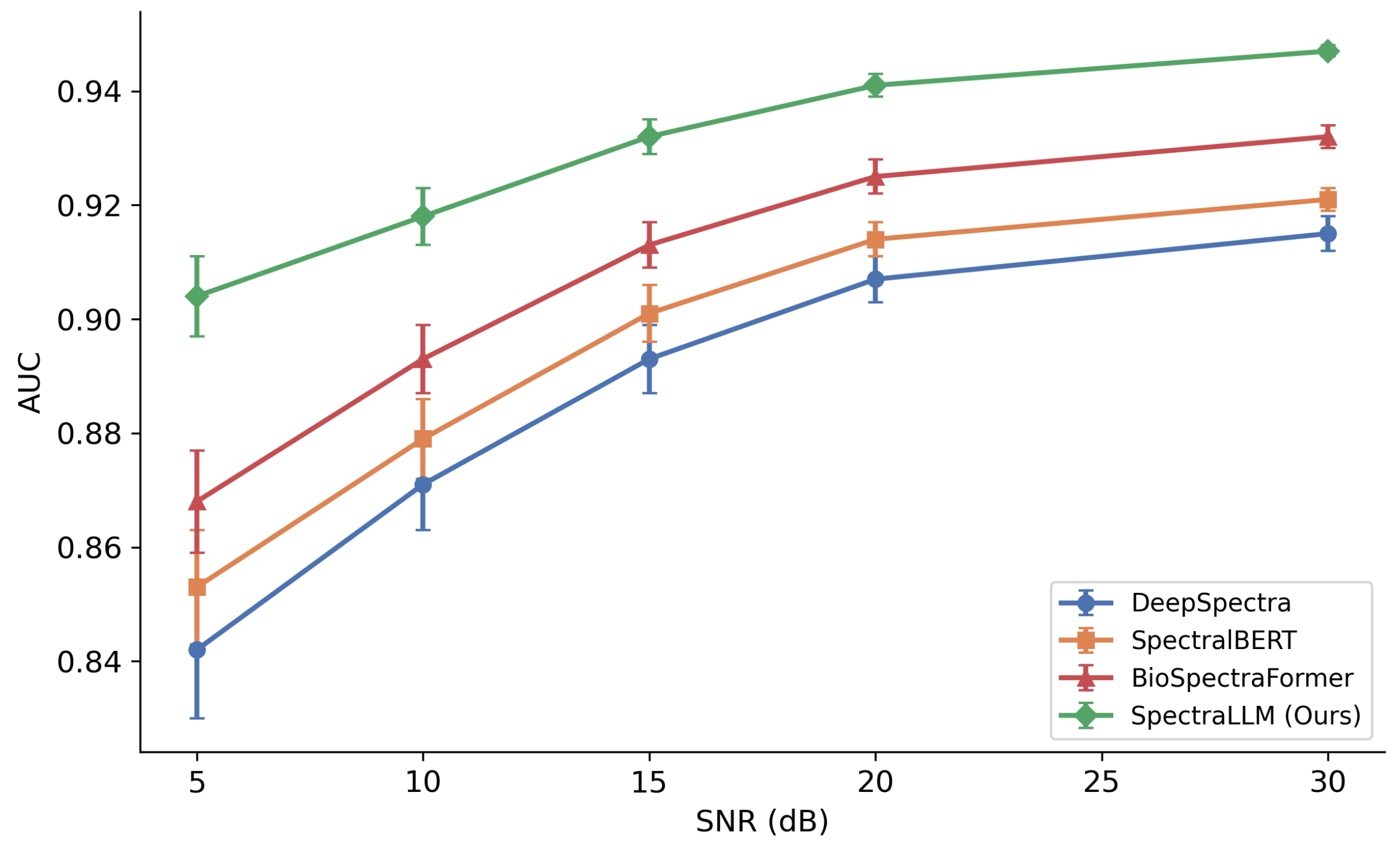
2.5. Robustness to Noise
2.6. Cell-Type Transfer
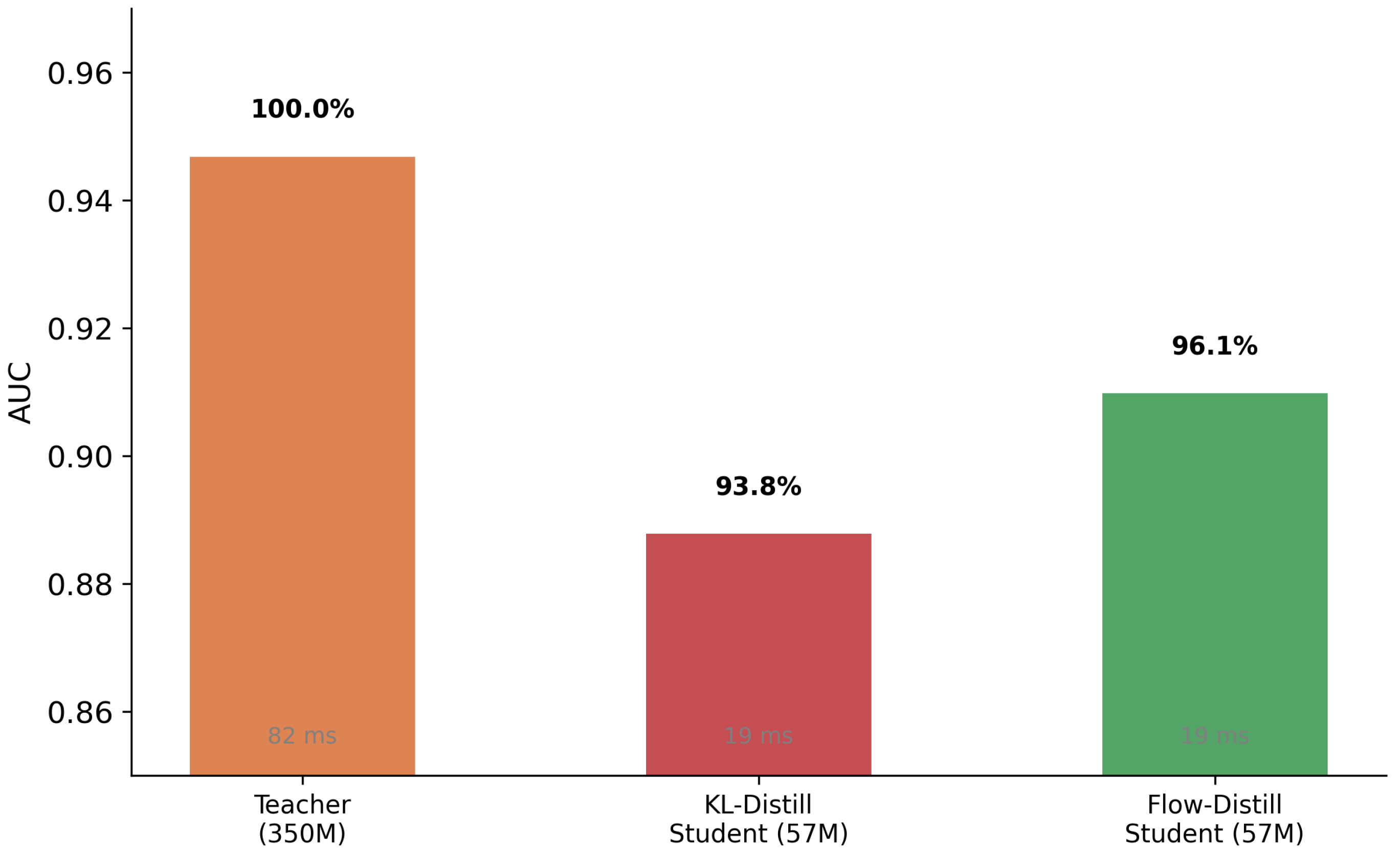
2.7. Distillation Efficiency
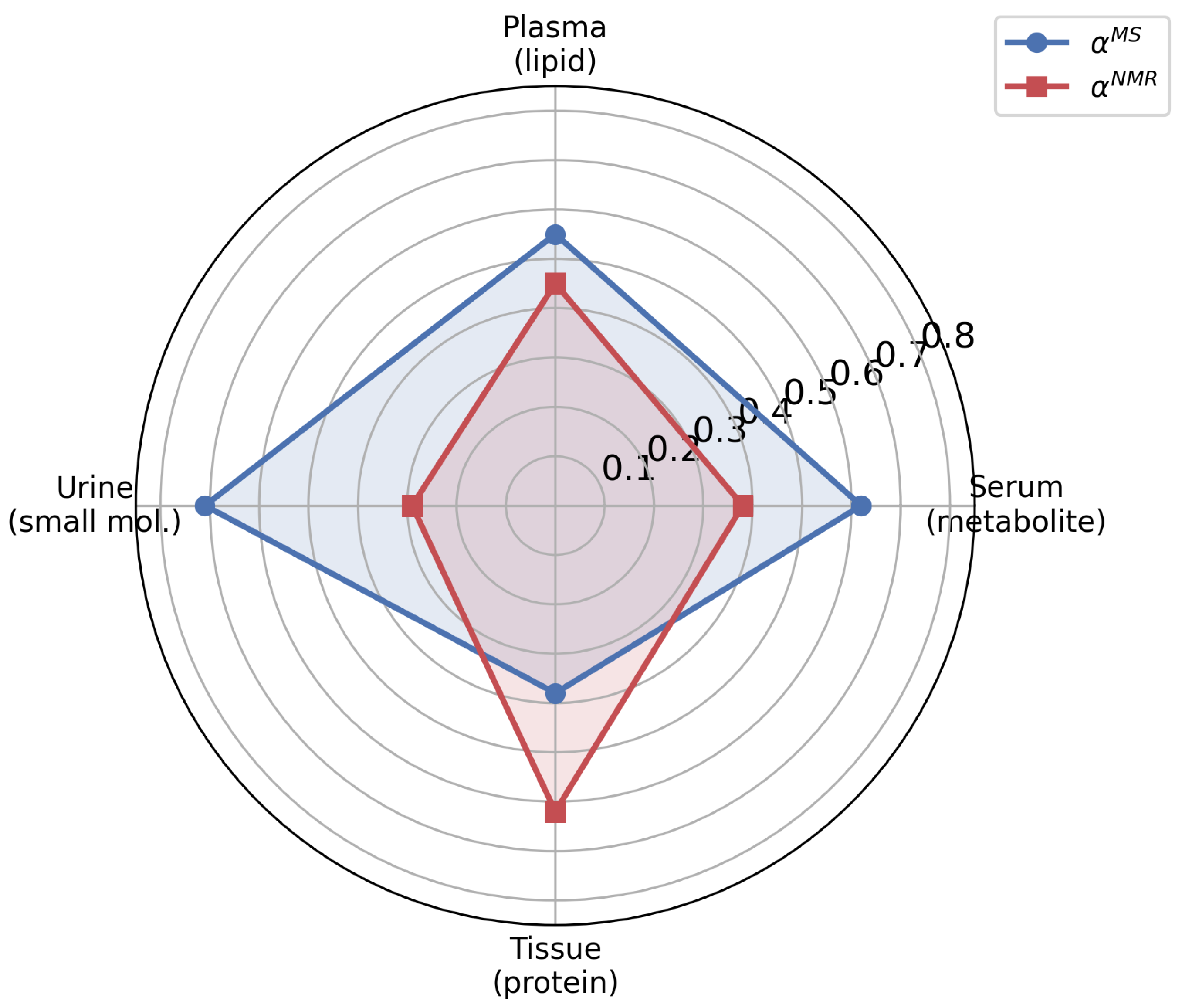
2.8. Multi-Modal Contribution Analysis
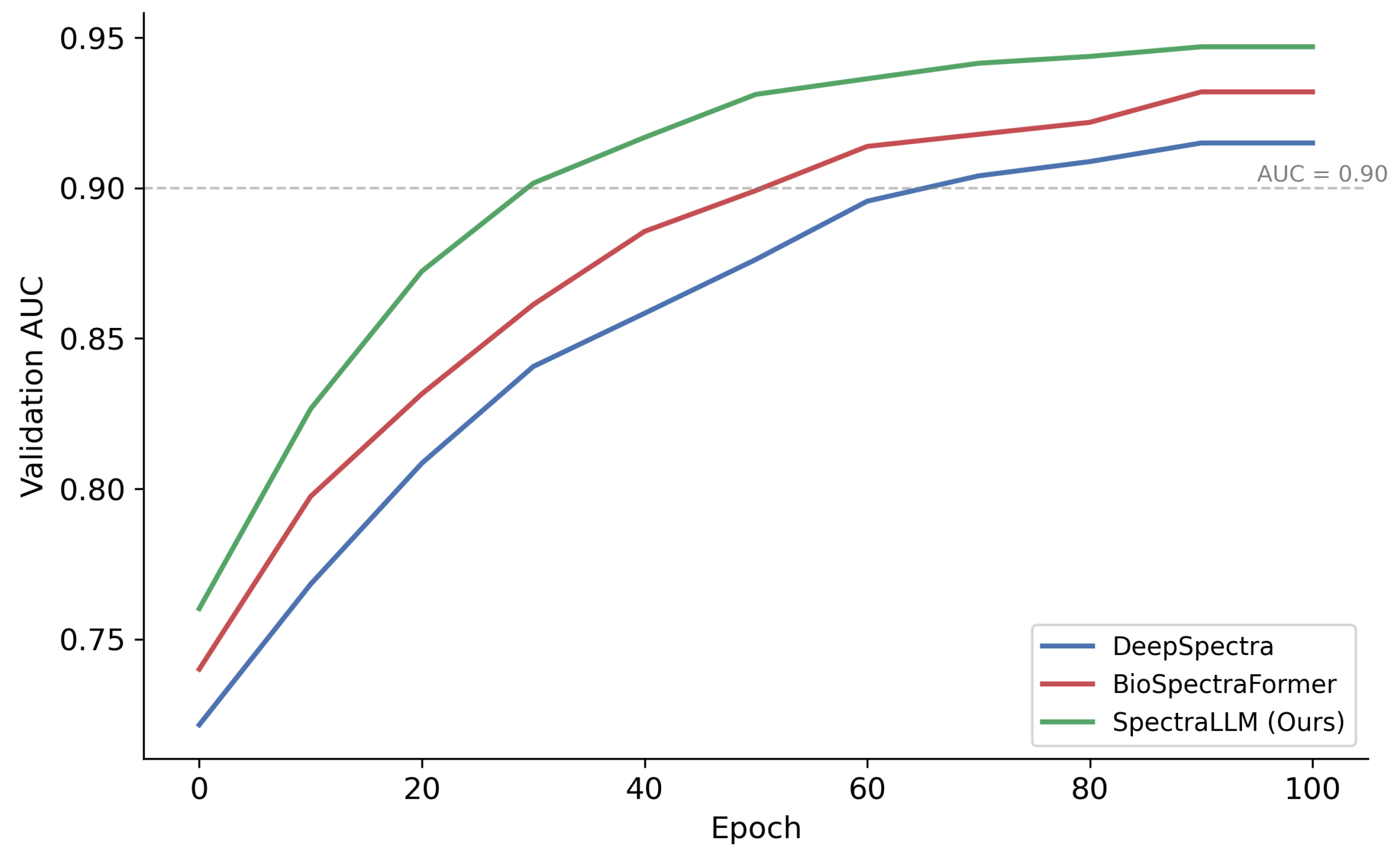
2.9. Convergence Analysis
2.10. Hyperparameter Sensitivity
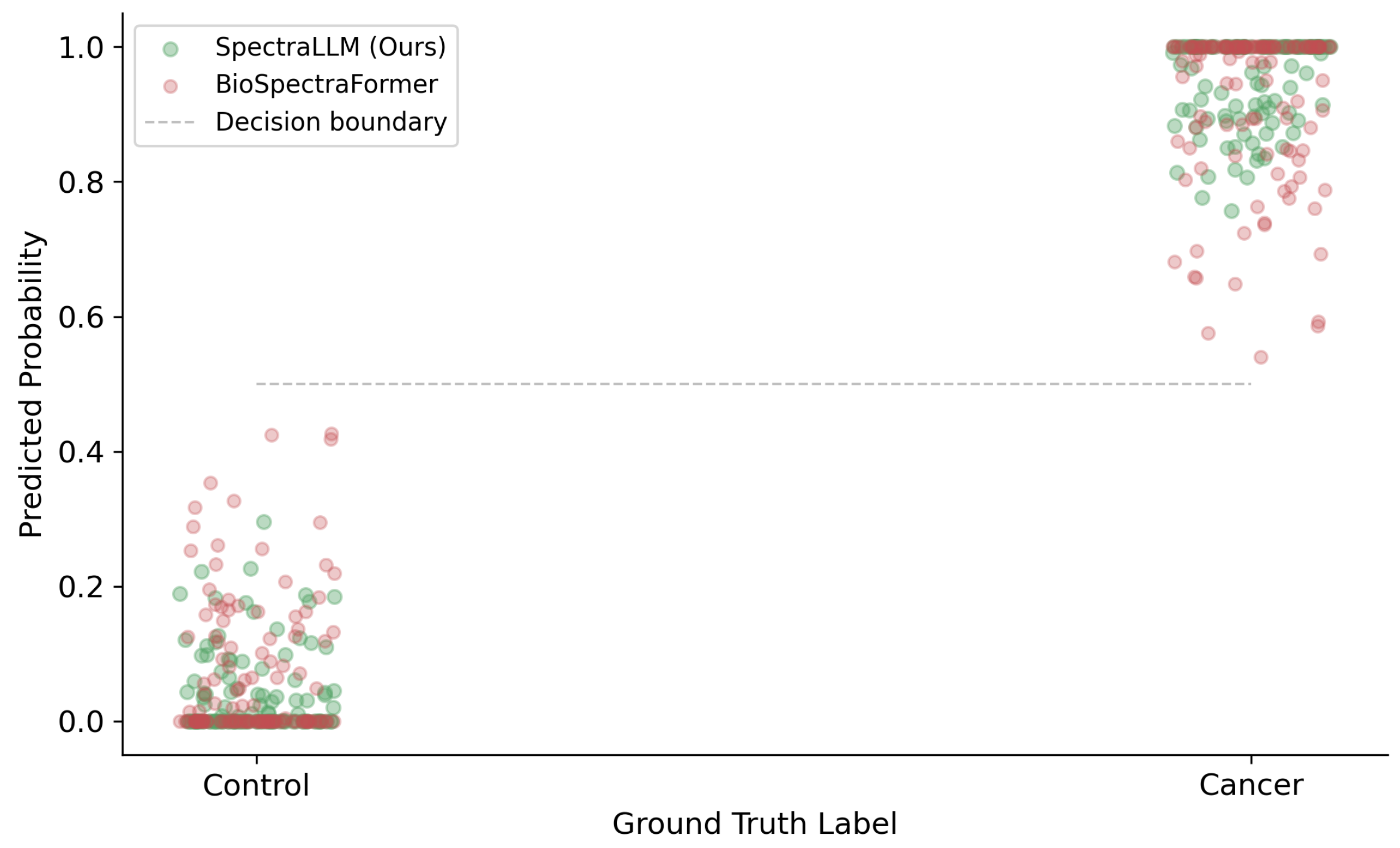
2.11. Biological Case Study: Early Cancer Detection
2.12. Species Transfer
3. Discussion
4. Methods
4.1. Data Acquisition and Preprocessing
4.2. Model Architecture
4.3. Flow-Based Knowledge Distillation
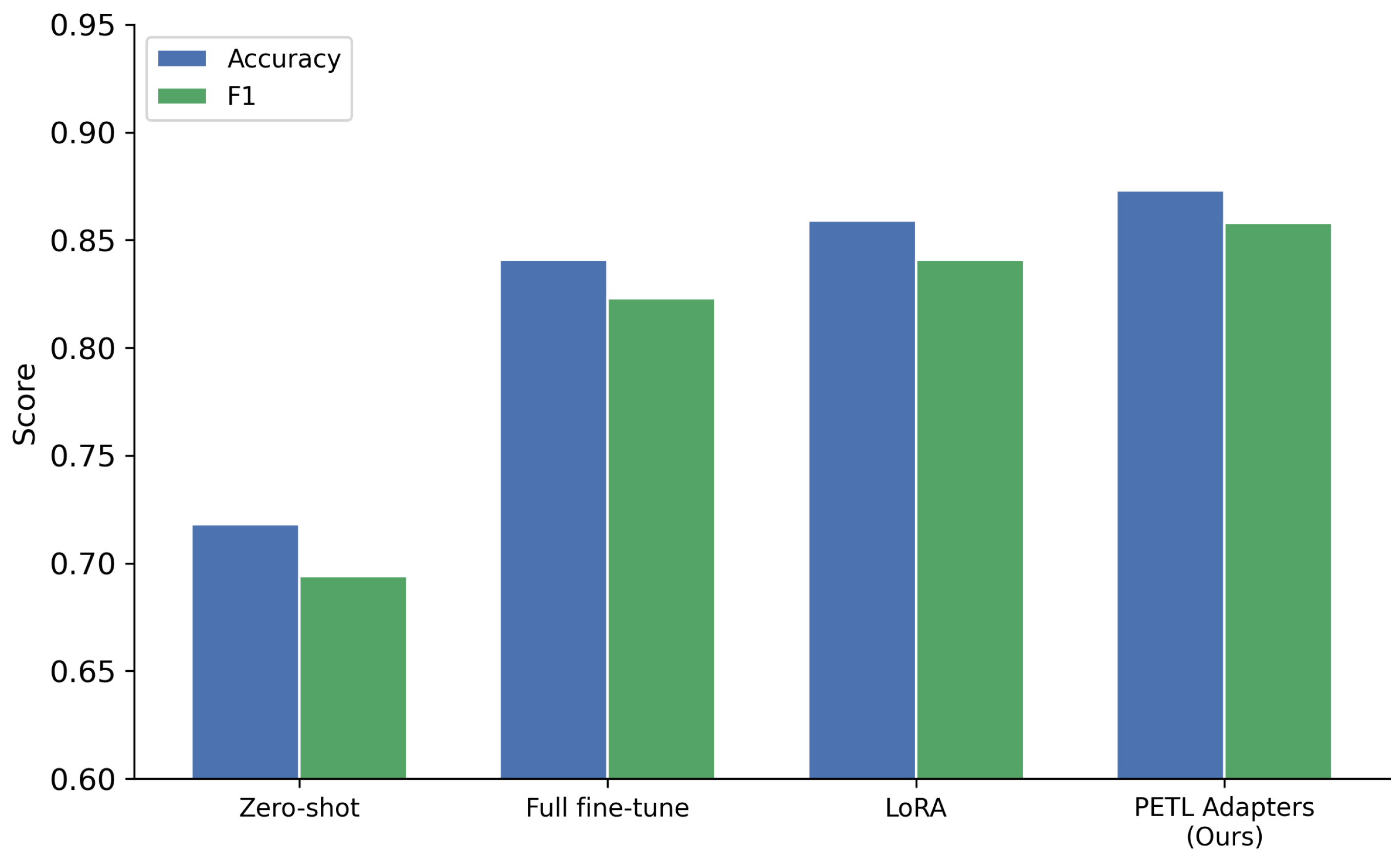
4.4. Parameter-Efficient Transfer Learning
4.5. Evaluation Metrics
4.6. Implementation Details
References
- Mia andthe Jeppesen, Arianna Tonon, Seid Boudah, , et al. Multiplatform untargeted metabolomics. Magnetic Resonance in Chemistry, 61(7):367–381, 2023.
- Abdul-Hamid Emwas, Ritu Roy, Ryan T McKay, , et al. Quantitative nmr-based biomedical metabolomics: current status and applications. Molecules, 25(21):5128, 2020.
- Madeleine Picard, Jean-Philippe Adam, , et al. Multimodal data fusion for cancer biomarker discovery with deep learning. Nature Machine Intelligence, 5:352–364, 2023.
- Yuxin Gao, Xing Shu, Qiang Zhang, Xue Fang, , et al. Trackable and scalable lc-ms metabolomics data processing. Nature Communications, 14:4153, 2023.
- Dongsik Li, Alan S Hansen, Shangqiang Yuan, , et al. Deep picker is a deep neural network for accurate deconvolution of complex two-dimensional nmr spectra. Nature Communications, 12:5231, 2021.
- Ralf Tautenhahn, Gary J Patti, , et al. An improved peak detection and quantification algorithm for lc-ms metabolomics data. Analytical Chemistry, 84(11):5035–5039, 2012.
- M Akhtar, D Tresch, R Seifert, , et al. Peak learning of mass spectrometry imaging data using deep learning. Nature Communications, 12:5544, 2021.
- Melih Yilmaz, William E Fischer, and William Stafford Noble. De novo mass spectrometry peptide sequencing with a transformer model. In Proceedings of the 39th International Conference on Machine Learning, pages 25514–25522, 2022.
- Hang Yu. Integrating deep cross networks and bilstm for scalable vulnerability analysis. In Proceedings of the 2025 International Symposium on Artificial Intelligence and Computational Social Sciences, pages 619–623, 2025.
- Yifeng Wu, Yicheng Yu, Zhongheng Yang, Zixuan Zeng, Guanhua Chen, and Jinping Xu. Brain-sam: Modality-agnostic model for brain lesion segmentation. In 2025 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), pages 3000–3005. IEEE, 2025.
- Jerret Ross, Brian Belgodere, Vijil Chenthamarakshan, , et al. Large-scale chemical language representations capture molecular structure and properties. Nature Machine Intelligence, 4:1256–1264, 2022.
- Shaoqian Tang. Low-latency multimodal semantic ranking for real-time advertising recommendation. In Proceedings of the 2025 2025 2nd Symposium on Big Data, Neural Networks, and Deep Learning, pages 424–428, 2025.
- Shaoqian Tang. Semantic–geometric collaborative detection for fine-grained tampering in financial document images. In 2025 6th International Conference on Artificial Intelligence and Computer Engineering (ICAICE), pages 88–91. IEEE, 2025.
- Xinmeng Xu, Weiping Tu, Yuhong Yang, Jizhen Li, Yiqun Zhang, and Hongyang Chen. Contribution-aware dynamic multi-modal balance for audio-visual speech separation. IEEE Transactions on Multimedia, 2026.
- Chun Xie, Huimin Tong, Guoxi Xu, Yipeng Chen, Li Luking, and Yiwei Chen. Knowledge calibration distillation. In 2025 IEEE International Conference on Multimedia and Expo (ICME), pages 1–7. IEEE, 2025.
- Xinye Yang, Junhao Wang, Haosen Sun, Xuesheng Zhang, Zebang Liu, Gaochao Xu, Yiwei Chen, et al. Flow-based knowledge transfer for efficient large model distillation. In Proceedings of the AAAI Conference on Artificial Intelligence, volume 40, pages 27666–27674, 2026.
- Yi Xin, Siqi Luo, Xuyang Liu, Haodi Zhou, Xinyu Cheng, Christina E Lee, Junlong Du, Haozhe Wang, MingCai Chen, Ting Liu, et al. V-petl bench: A unified visual parameter-efficient transfer learning benchmark. Advances in Neural Information Processing Systems, 37:80522–80535, 2024.
- Yi Xin, Junlong Du, Qiang Wang, Zhiwen Lin, and Ke Yan. Vmt-adapter: Parameter-efficient transfer learning for multi-task dense scene understanding. In Proceedings of the AAAI conference on artificial intelligence, volume 38, pages 16085–16093, 2024.
- Yi Xin, Qi Qin, Siqi Luo, Kaiwen Zhu, Juncheng Yan, Yan Tai, Jiayi Lei, Yuewen ***, Keqi Wang, Yibin Wang, et al. Lumina-dimoo: An omni diffusion large language model for multi-modal generation and understanding. arXiv preprint arXiv:2510.06308, 2025.
- Ning Yang, Hengyu Zhong, Wentao Wang, Baoliang Tian, Haijun Zhang, and Jun Wang. Linearard: Linear-memory attention distillation for rope restoration. arXiv preprint arXiv:2604.00004, 2026.
- Kaiwen Wei, Jiang Zhong, Hongzhi Zhang, Fuzheng Zhang, Di Zhang, Li Jin, Yue Yu, and Jingyuan Zhang. Chain-of-specificity: Enhancing task-specific constraint adherence in large language models. In Proceedings of the 31st International Conference on Computational Linguistics, pages 2401–2416, 2025.
- Kaiwen Wei, Rui Shan, Dongsheng Zou, Jianzhong Yang, Bi Zhao, Junnan Zhu, and Jiang Zhong. Mirage: Scaling test-time inference with parallel graph-retrieval-augmented reasoning chains. arXiv preprint arXiv:2508.18260, 2025.
- Kaiwen Wei, Xiao Liu, Jie Zhang, Zijian Wang, Ruida Liu, Yuming Yang, Xin Xiao, Xiao Sun, Haoyang Zeng, Changzai Pan, et al. Cfvbench: A comprehensive video benchmark for fine-grained multimodal retrieval-augmented generation. arXiv preprint arXiv:2510.09266, 2025.
- Ning Yang, Jinliang Gao, and Haijun Zhang. Llm-driven large-scale spectrum access, 2026.
- Ning Yang, Chuangxin Cheng, and Haijun Zhang. Multi-turn reasoning llms for task offloading in mobile edge computing. arXiv preprint arXiv:2604.07148, 2026.
- Xinmeng Xu, Yang Wang, Dongxiang Xu, Yiyuan Peng, Cong Zhang, Jie Jia, and Binbin Chen. Vsegan: Visual speech enhancement generative adversarial network. In ICASSP 2022-2022 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), pages 7308–7311. IEEE, 2022.
- Xinmeng Xu, Weiping Tu, and Yuhong Yang. Case-net: Integrating local and non-local attention operations for speech enhancement. Speech Communication, 148:31–39, 2023.
- Yiwei Chen, Silvestre Manzanera, Juan Mompeán, Daniel Ruminski, Ireneusz Grulkowski, and Pablo Artal. Increased crystalline lens coverage in optical coherence tomography with oblique scanning and volume stitching. Biomedical Optics Express, 12(3):1529–1542, 2021.
| Method | AUC ↑ | AUPR ↑ | F1 ↑ | Pearson r ↑ | Acc. ↑ |
| DeepSpectra | 0.915 | 0.887 | 0.880 | 0.901 | 0.862 |
| SpectralBERT | 0.921 | 0.894 | 0.887 | 0.908 | 0.871 |
| MS-NMR Fusion Net | 0.928 | 0.901 | 0.893 | 0.915 | 0.879 |
| BioSpectraFormer | 0.932 | 0.908 | 0.898 | 0.921 | 0.885 |
| SpectraLLM (Ours) | 0.947 | 0.923 | 0.912 | 0.938 | 0.904 |
| Method | Accuracy ↑ | F1 ↑ | Spearman ↑ |
| PepNovo | 0.891 | 0.876 | 0.883 |
| DeepNovoV2 | 0.903 | 0.891 | 0.897 |
| pNovus | 0.912 | 0.899 | 0.908 |
| SpectraLLM (Ours) | 0.943 | 0.928 | 0.931 |
| Method | Accuracy ↑ | F1 ↑ | Pearson r ↑ | Dice ↑ |
| NMR-Net | 0.854 | 0.841 | 0.889 | 0.823 |
| CellSpectra | 0.878 | 0.863 | 0.907 | 0.851 |
| MetaFormer | 0.892 | 0.879 | 0.918 | 0.868 |
| SpectraLLM (Ours) | 0.917 | 0.905 | 0.938 | 0.902 |
| Configuration | AUC | AUPR | F1 | Pearson r | Acc. |
| Full SpectraLLM | 0.947 | 0.923 | 0.912 | 0.938 | 0.904 |
| w/o Modality-Agnostic Enc. | 0.931 | 0.907 | 0.893 | 0.919 | 0.883 |
| w/o Dynamic Balance | 0.919 | 0.893 | 0.879 | 0.905 | 0.868 |
| w/o Flow Distillation | 0.943 | 0.918 | 0.907 | 0.934 | 0.899 |
| w/o PETL Adapters | 0.940 | 0.914 | 0.903 | 0.930 | 0.894 |
| r | AUC | Params (M) | AUC | Retention (%) | |
| 16 | 0.931 | 0.8 | 0.1 | 0.897 | 94.7 |
| 32 | 0.939 | 2.1 | 0.2 | 0.904 | 95.5 |
| 64 | 0.947 | 6.3 | 0.3 | 0.910 | 96.1 |
| 128 | 0.946 | 18.9 | 0.5 | 0.905 | 95.6 |
| 256 | 0.945 | 51.2 | 0.7 | 0.899 | 94.9 |
| Method | Accuracy ↑ | AUC ↑ |
| Zero-shot (no adaptation) | 0.712 | 0.768 |
| Full fine-tuning | 0.821 | 0.874 |
| LoRA | 0.843 | 0.891 |
| PETL Adapters (Ours) | 0.856 | 0.907 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




