Submitted:
27 April 2026
Posted:
28 April 2026
You are already at the latest version
Abstract
Microbial protein production from acetate represents a promising route for sustainable protein supply, yet its efficiency is constrained by limited understanding of carbon–nitrogen metabolic coordination. In this study, nitrogen availability was systematically varied to investigate its role in regulating carbon partitioning and protein biosynthesis in Yarrowia lipolytica. Nitrogen limitation markedly reduced cell growth and protein accumulation (19.56% of dry cell weight) while increasing lipid content (up to 34.16%), indicating a redistribution of carbon flux from protein to lipid synthesis. Transcriptomic analysis revealed a global downregulation of anabolic pathways under nitrogen limitation, accompanied by a shift in nitrogen assimilation from the glutamate dehydrogenase (GDH) pathway to the glutamine synthetase/glutamate synthase (GS–GOGAT) pathway, as well as significant upregulation of genes related to ammonium and amino acid transport. Guided by these findings, metabolic engineering of key nitrogen assimilation pathways was performed. Co-overexpression of GDH and GS increased protein content from 48.52% to 55.77% and improved amino acid composition, whereas GOGAT overexpression impaired growth and protein accumulation. These results demonstrate that nitrogen availability governs carbon allocation through coordinated regulation of nitrogen transport and assimilation, and that balanced enhancement of GDH and GS is an effective strategy to improve protein production from acetate, supporting the development of sustainable fermentation processes using CO₂-derived substrates.
Keywords:
1. Introduction
2. Materials and Methods
Genetic Manipulation for the Construction of Plasmids and Strains
Media and Growth Conditions
Shake Flask Culture and Determination of Cell Content
Analytical Methods
Transcriptome Sequencing and Analysis
3. Results
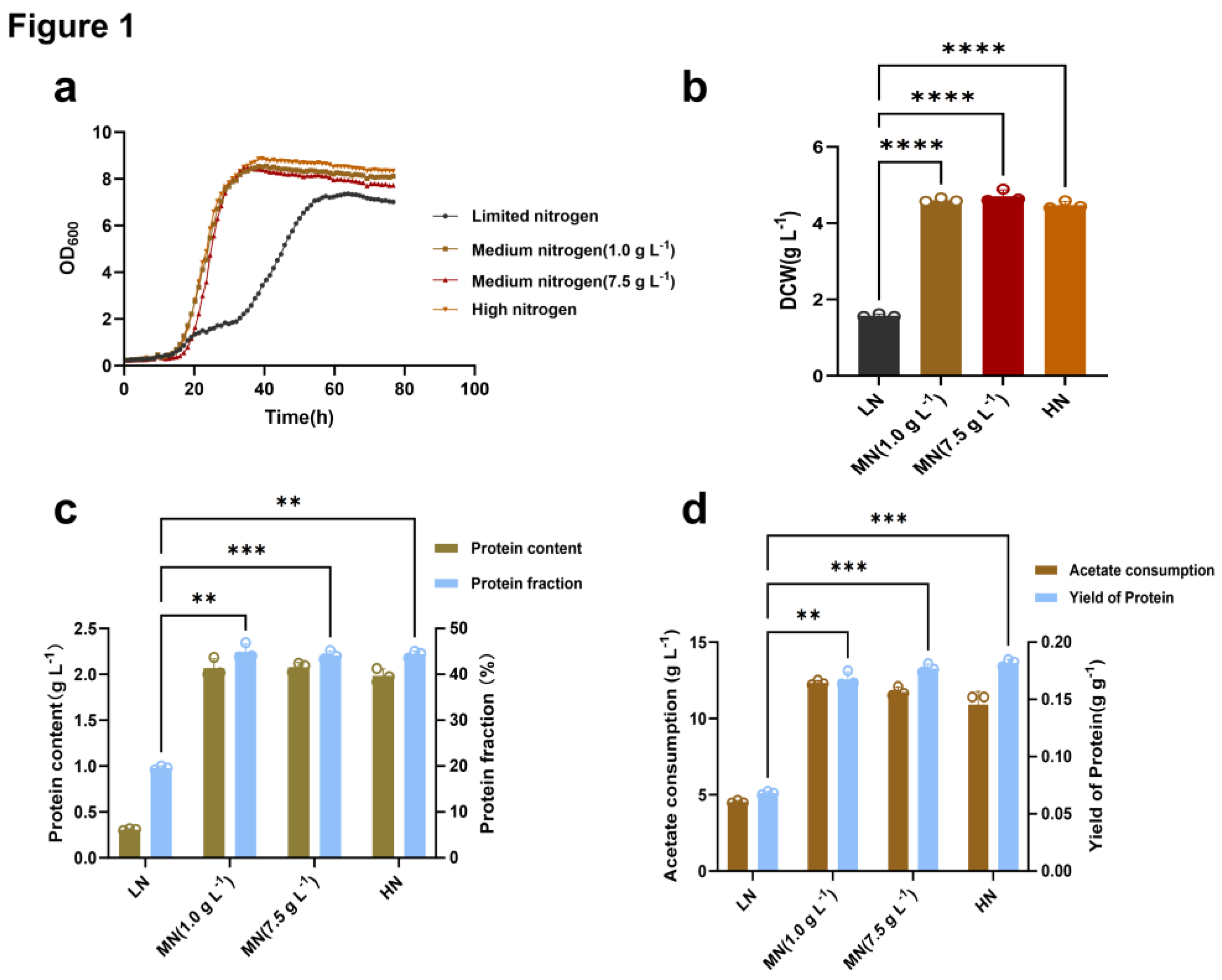
Effects of Nitrogen Availability on Growth and Protein Accumulation
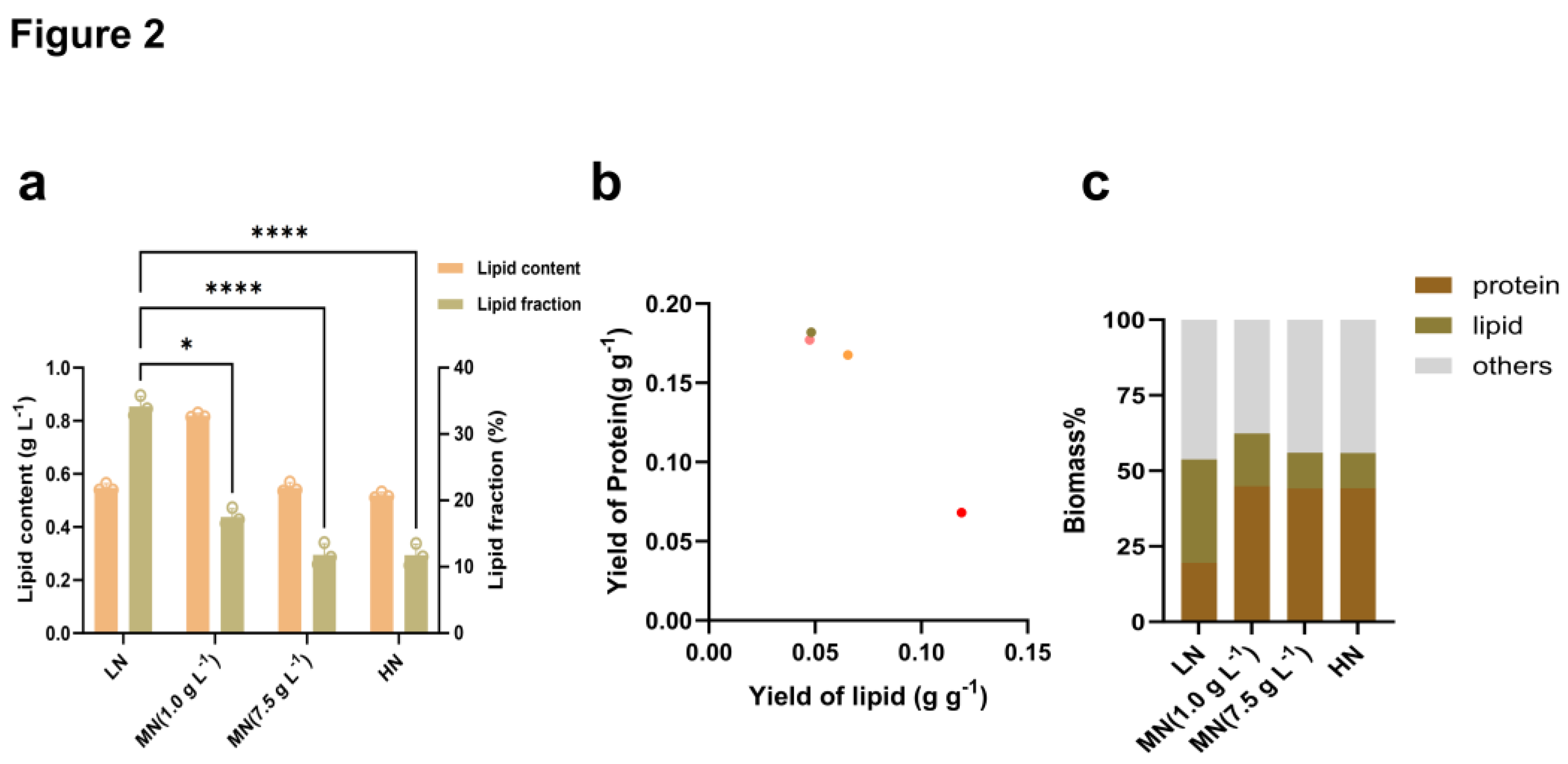
Nitrogen Availability Regulates Carbon Partitioning Between Lipid and Protein Biosynthesis
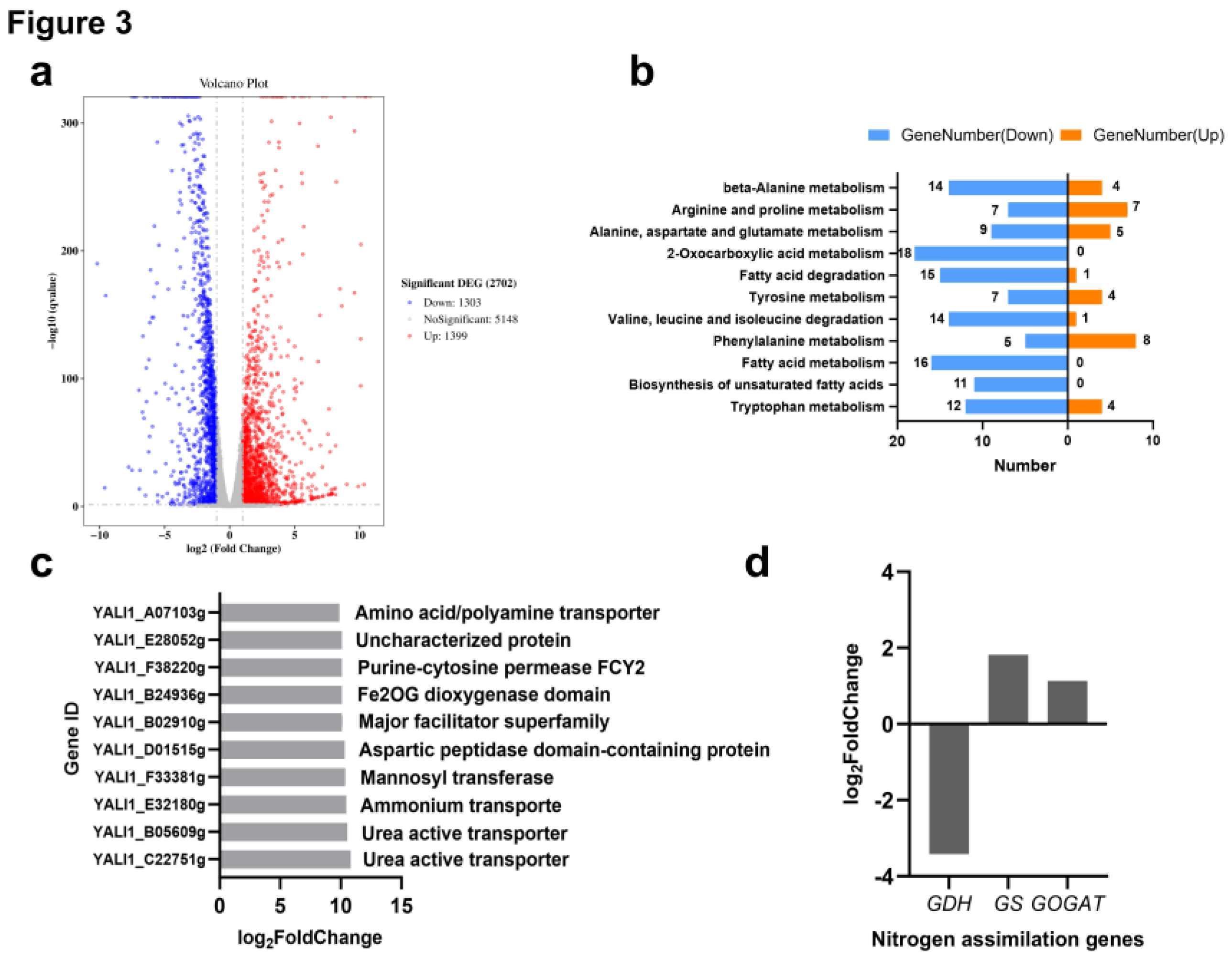
Transcriptomic Analysis Reveals Nitrogen-Dependent Regulation of Carbo–Nitrogen Metabolism
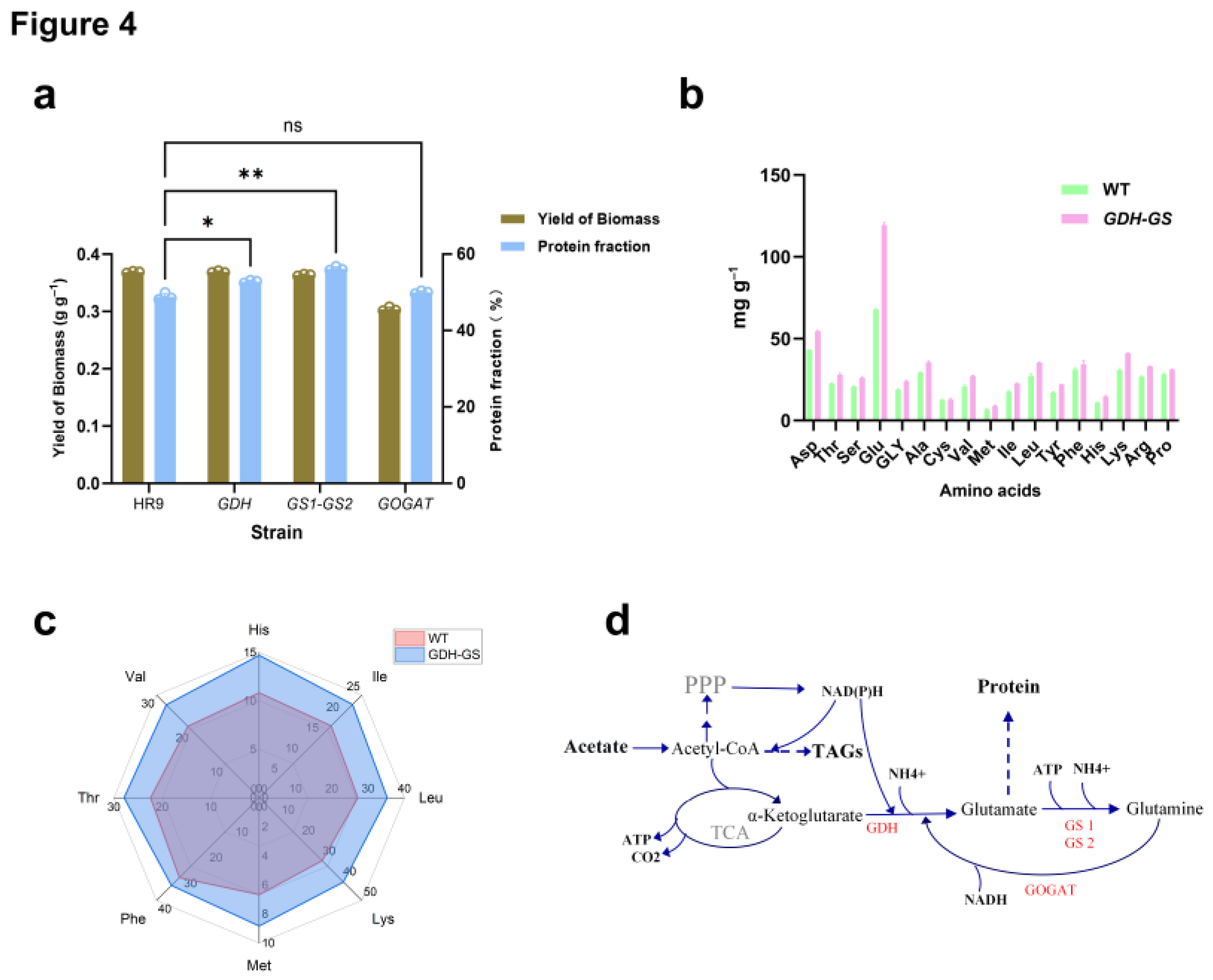
Functional Validation of Nitrogen Assimilation Pathways Through Targeted Gene Overexpression
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| GDH | glutamate dehydrogenase |
| GS | glutamine synthetase |
| GOGAT | glutamate synthase |
| DCW | Dry cell weight |
References
- Pikaar, I.; Matassa, S.; Bodirsky, B.L.; Weindl, I.; Humpenöder, F.; Rabaey, K.; Boon, N.; Bruschi, M.; Yuan, Z.; van Zanten, H.; et al. Decoupling livestock from land use through industrial feed production pathways. Environ. Sci. Technol. 2018, 52, 7351–7359. [Google Scholar] [CrossRef]
- Cedeno, F.R. P.; Olubiyo, O.J.; Ferreira, S. From microbial proteins to cultivated meat for alternative meat-like products: a review on sustainable fermentation approaches. J. Biol. Eng. 2025, 19, 44. [Google Scholar] [CrossRef] [PubMed]
- Gong, G.; Wu, B.; Liu, L.; Li, J.; Zhu, Q.; He, M.; Hu, G. Valorization of waste streams and C1 gases for sustainable food nutrients and value-added compounds production: acetate as a promising intermediate. Sci. Total Environ. 2023, 893, 164795. [Google Scholar] [CrossRef]
- Kim, Y.; Lama, S.; Agrawal, D.; Kumar, V.; Park, S. Acetate as a potential feedstock for the production of value-added chemicals: metabolism and applications. Biotechnol. Adv. 2021, 49, 107736. [Google Scholar] [CrossRef]
- Kutscha, R.; Pflügl, S. Microbial upgrading of acetate into value-added products—examining microbial diversity, bioenergetic constraints and metabolic engineering approaches. Int. J. Mol. Sci. 2020, 21, 8777. [Google Scholar] [CrossRef] [PubMed]
- Lozano Terol, G.; Gallego-Jara, J.; Sola Martínez, R.A.; Cánovas Díaz, M.; De Diego Puente, T. Engineering protein production by rationally choosing a carbon and nitrogen source using E. coli BL21 acetate metabolism knockout strains. Microb. Cell. Fact. 2019, 18, 151. [Google Scholar] [CrossRef]
- Koh, H.G.; Yook, S.; Oh, H.; Rao, C.V.; Jin, Y.S. Toward rapid and efficient utilization of nonconventional substrates by nonconventional yeast strains. Curr. Opin. Biotechnol. 2024, 85, 103059. [Google Scholar] [CrossRef]
- Mamaev, D.; Zvyagilskaya, R. Yarrowia lipolytica: a multitalented yeast species of ecological significance. FEMS Yeast Res. 2021, 21, foab008. [Google Scholar] [CrossRef]
- Kerkhoven, E.J.; Pomraning, K.R.; Baker, S.E.; Nielsen, J. Regulation of amino-acid metabolism controls flux to lipid accumulation in Yarrowia lipolytica. Npj Syst. Biol. Appl. 2016, 2, 16005. [Google Scholar] [CrossRef]
- Liu, Z.; Tian, J.; Miao, Z.; Liang, W.; Wang, G. Metabolome and transcriptome profiling reveal carbon metabolic flux changes in Yarrowia lipolytica cells to rapamycin. J. Fungi 2022, 8, 939. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Lai, Y.H.; Lu, T. Coarse-grained modeling elucidates differential metabolism of Saccharomyces cerevisiae under varied nutrient limitations. ACS Synth. Biol. 2025, 14, 1523–1532. [Google Scholar] [CrossRef] [PubMed]
- Tibocha-Bonilla, J.D.; Kumar, M.; Richelle, A.; Godoy-Silva, R.D.; Zengler, K.; Zuñiga, C. Dynamic resource allocation drives growth under nitrogen starvation in eukaryotes. Npj Syst. Biol. Appl. 2020, 6, 14. [Google Scholar] [CrossRef]
- Helling, R.B. Speed versus efficiency in microbial growth and the role of parallel pathways. J. Bacteriol. 2002, 184, 1041–1045. [Google Scholar] [CrossRef]
- Sieg, A.G.; Trotter, P.J. Differential contribution of the proline and glutamine pathways to glutamate biosynthesis and nitrogen assimilation in yeast lacking glutamate dehydrogenase. Microbiol. Res. 2014, 169, 709–716. [Google Scholar] [CrossRef] [PubMed]
- Pomraning, K.R.; Kim, Y.M.; Nicora, C.D.; Chu, R.K.; Bredeweg, E.L.; Purvine, S.O.; Hu, D.; Metz, T.O.; Baker, S.E. Multi-omics analysis reveals regulators of the response to nitrogen limitation in Yarrowia lipolytica. BMC Genom. 2016, 17, 138. [Google Scholar] [CrossRef]
- Jouny, M.; Hutchings, G.S.; Jiao, F. Carbon monoxide electroreduction as an emerging platform for carbon utilization. Nat. Catal. 2019, 2, 1062–1070. [Google Scholar] [CrossRef]
- Xia, C.; Zhu, P.; Jiang, Q.; Pan, Y.; Liang, W.; Stavitski, E.; Alshareef, H.N.; Wang, H. Continuous production of pure liquid fuel solutions via electrocatalytic CO2 reduction using solid-electrolyte devices. Nat. Energy 2019, 4, 776–785. [Google Scholar] [CrossRef]
- Hu, L.; Guo, S.; Wang, B.; Fu, R.; Fan, D.; Jiang, M.; Fei, Q.; Gonzalez, R. Bio-valorization of C1 gaseous substrates into bioalcohols: potentials and challenges in reducing carbon emissions. Biotechnol. Adv. 2022, 59, 107954. [Google Scholar] [CrossRef]
- Regis, F.; Demichelis, F.; Monteverde, A.; Fino, D. Techno-economic-environmental analysis of CO2 fermentation to acetic acid. Chem. Eng. J. 2025, 520, 166138. [Google Scholar] [CrossRef]
- Zheng, T.; Zhang, M.; Wu, L.; Guo, S.; Liu, X.; Zhao, J.; Xue, W.; Li, J.; Liu, C.; Li, X.; et al. Upcycling CO2 into energy-rich long-chain compounds via electrochemical and metabolic engineering. Nat. Catal. 2022, 5, 388–396. [Google Scholar] [CrossRef]
- Han, L.; Li, Y.; Wang, X.; Yao, Z.; Chen, Z.; Li, J.; Tan, T.; Lee, S.Y.; Lv, Y. Spatially decoupled electro-biosystem for efficient CO2-to-chemicals conversion via tandem catalysis. Adv. Energy Mater. 2025, 15, e03056. [Google Scholar] [CrossRef]
- Pan, Z.; Hu, X.; Guo, Y.; Yu, J.; Zhang, H.; Guo, K. Simultaneous production, concentration, and upgradation of acetate into single-cell protein in a hybrid microbial electrochemical system. Chem. Eng. J. 2025, 503, 158446. [Google Scholar] [CrossRef]
- Leger, D.; Matassa, S.; Noor, E.; Shepon, A.; Milo, R.; Bar-Even, A. Photovoltaic-driven microbial protein production can use land and sunlight more efficiently than conventional crops. Proc. Natl. Acad. Sci. 2021, 118, e2015025118. [Google Scholar] [CrossRef]
- Crandall, B.S.; Harland-Dunaway, M.; Jinkerson, R.E.; Jiao, F. Electro-agriculture: revolutionizing farming for a sustainable future. Joule 2024, 8, 2974–2991. [Google Scholar] [CrossRef]
- Mamaev, D.; Zvyagilskaya, R. Yarrowia lipolytica: a multitalented yeast species of ecological significance. FEMS Yeast Res. 2021, 21, foab008. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



