Submitted:
26 April 2026
Posted:
27 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Materials and Methods
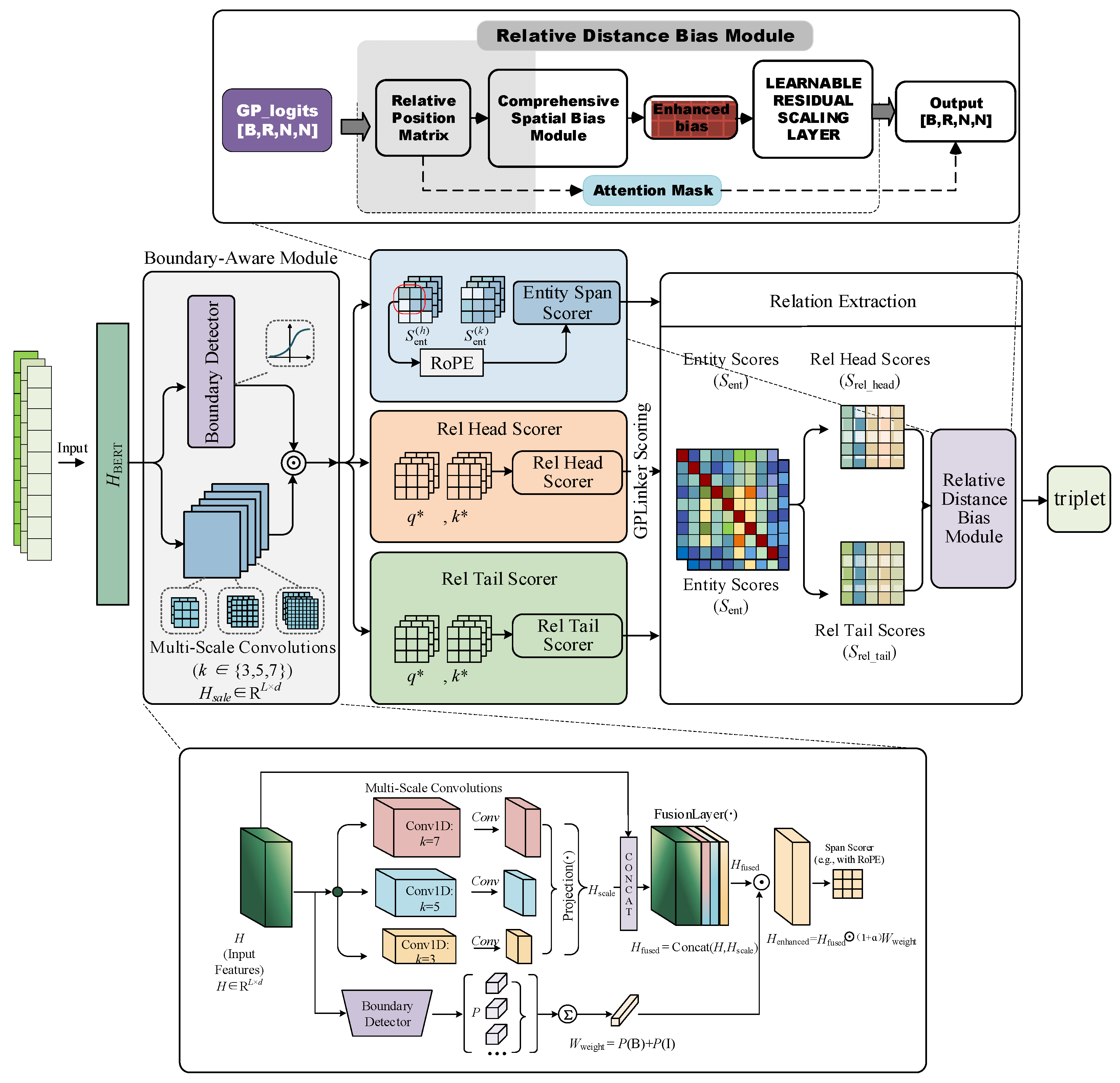
2.2. Model
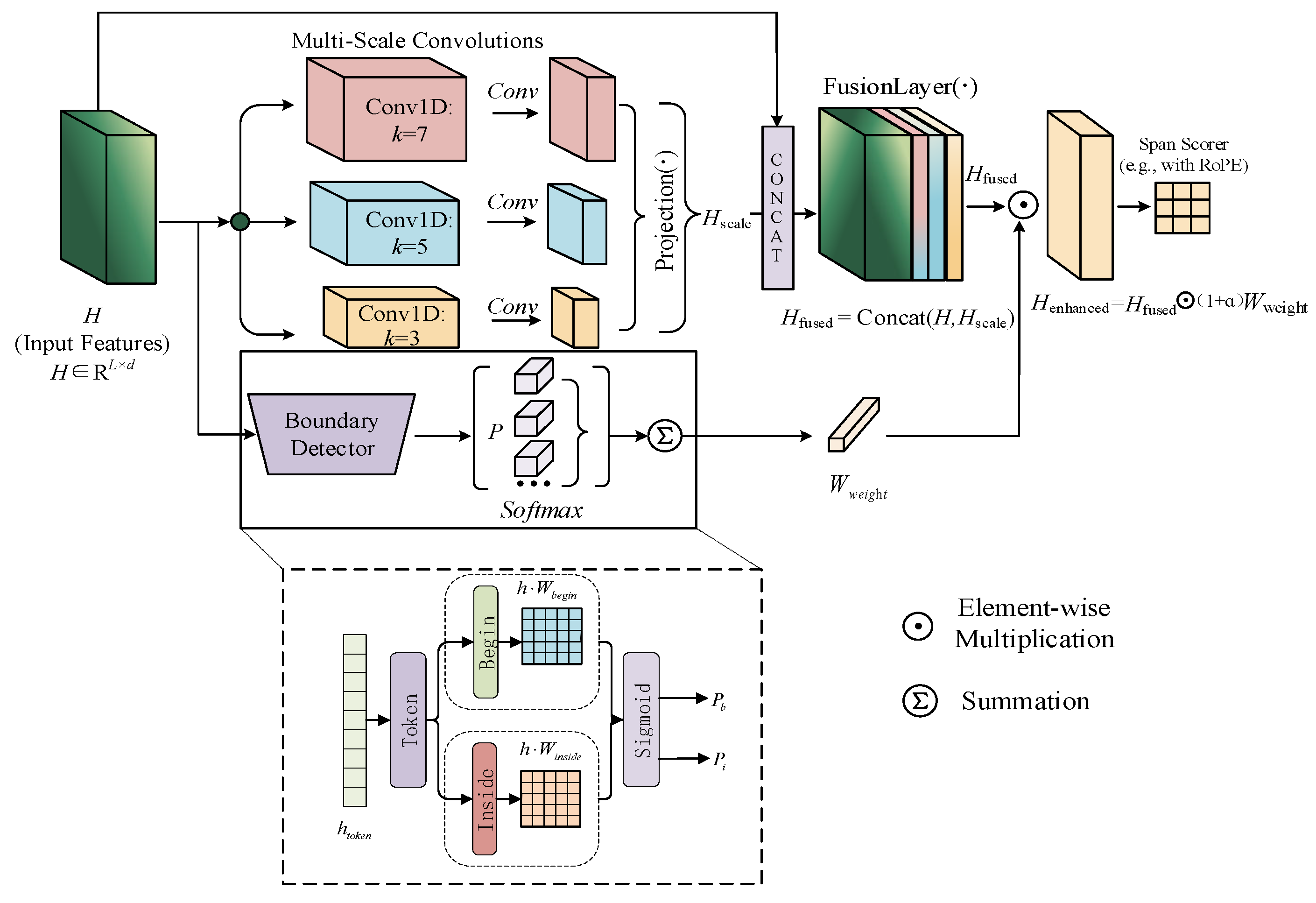
2.2.1. Boundary-Aware Module
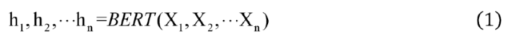
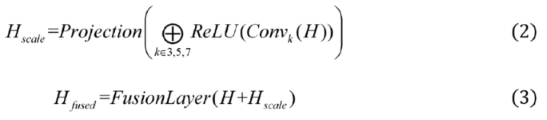
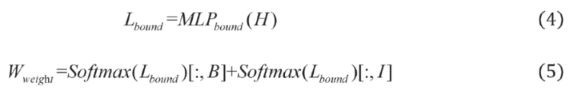
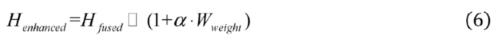
2.2.2. Encoding Layer

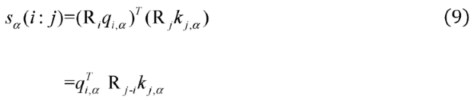
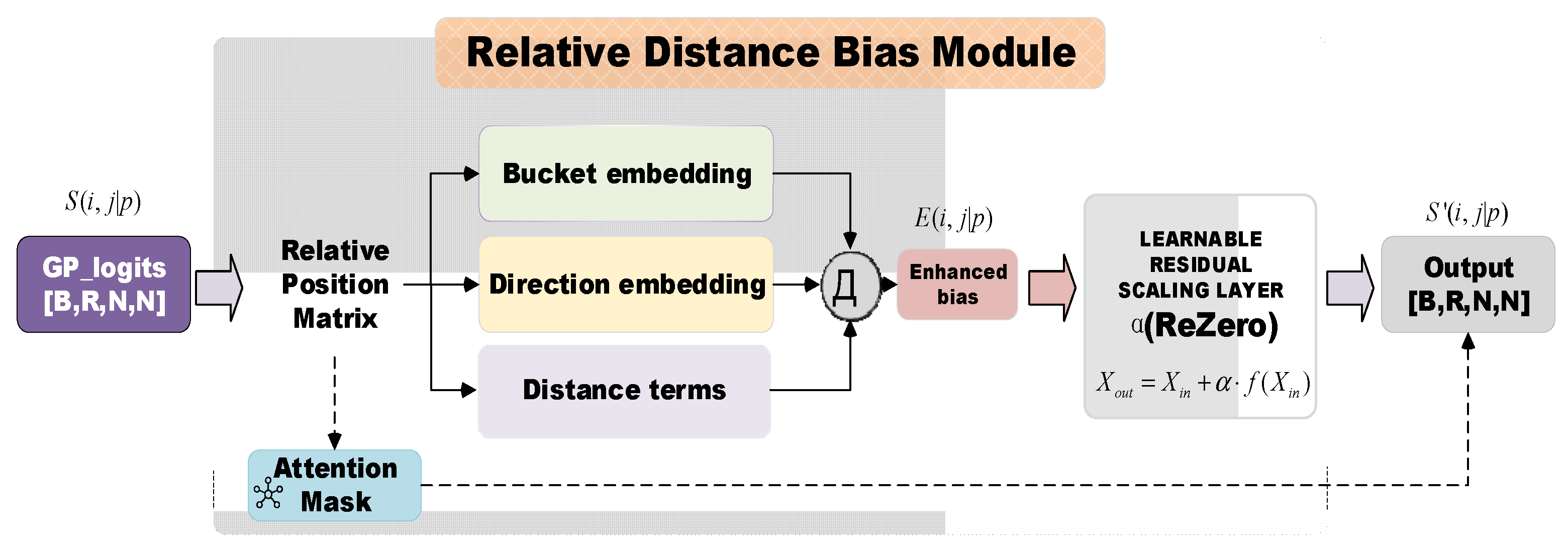
2.2.3. Relative Distance Bias Module
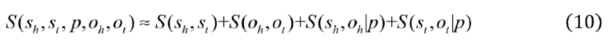
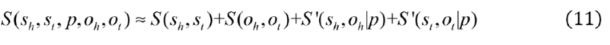
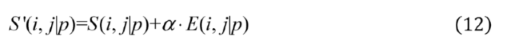
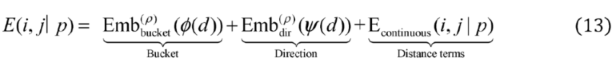
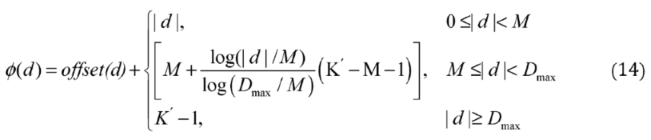

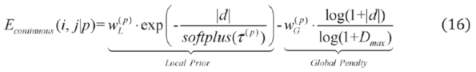
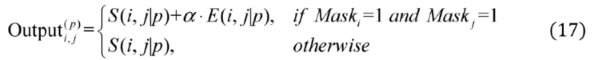
2.2.3. Weighted Sparse Multi-Label Cross-Entropy
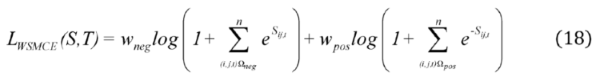

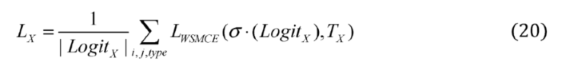
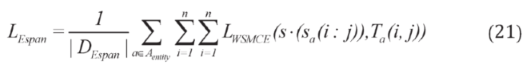
2.3. Experimental Setup
2.3.1. Parameters

2.3.2. Models
3. Results
3.1. Recognition Performances of Different Models
3.2. Ablation Study
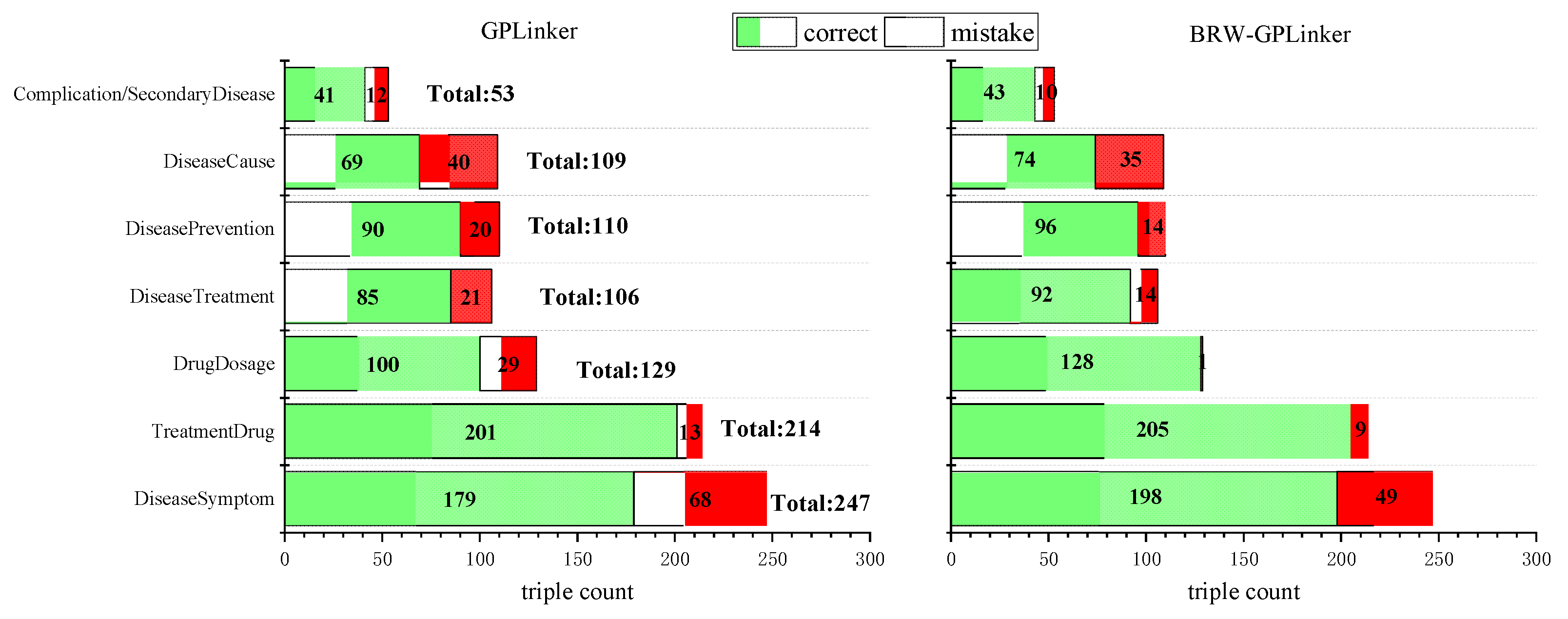
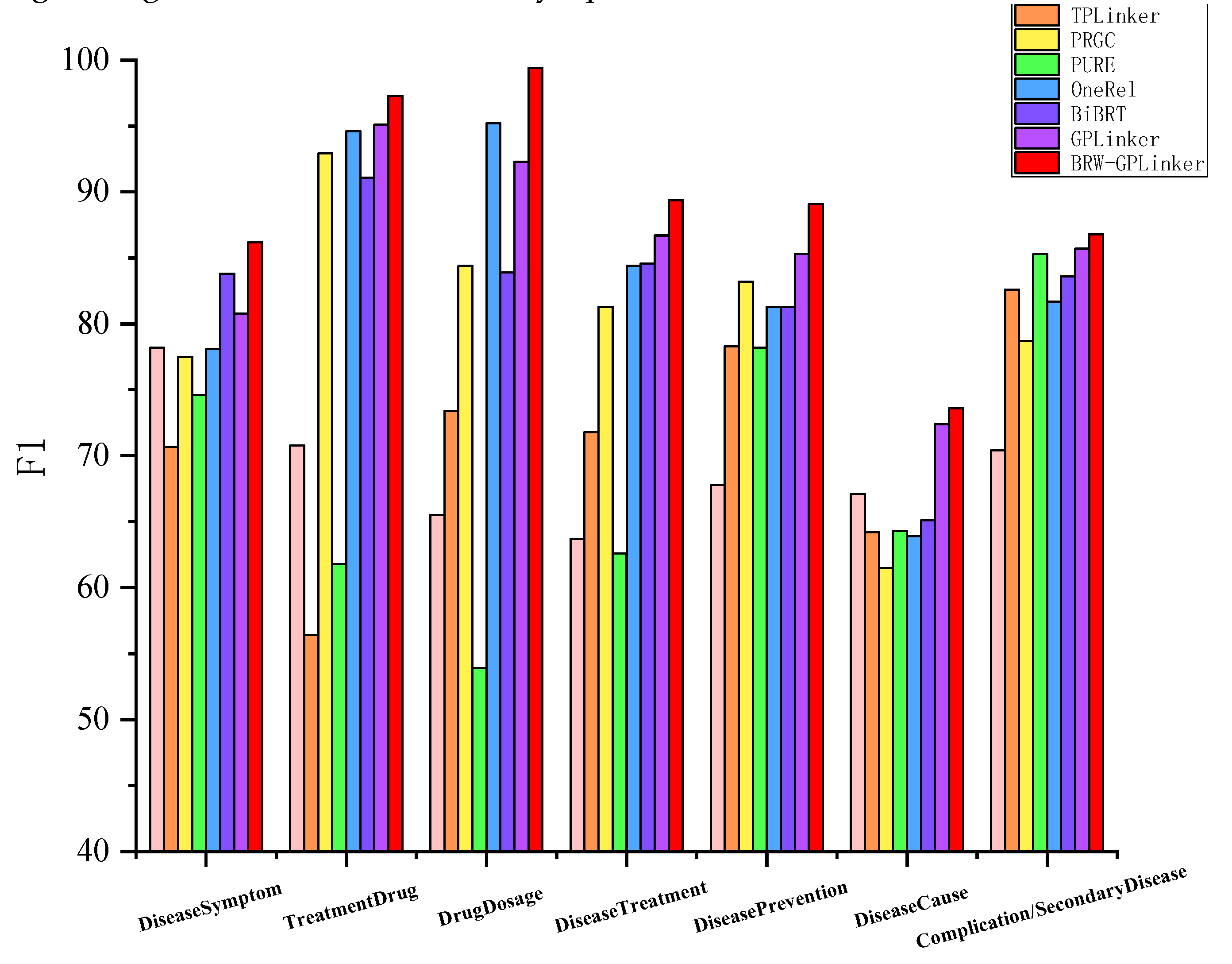
3.3. Performance on Different Triples by Relation Category
3.4. Performance on Different Triples by Relation Category
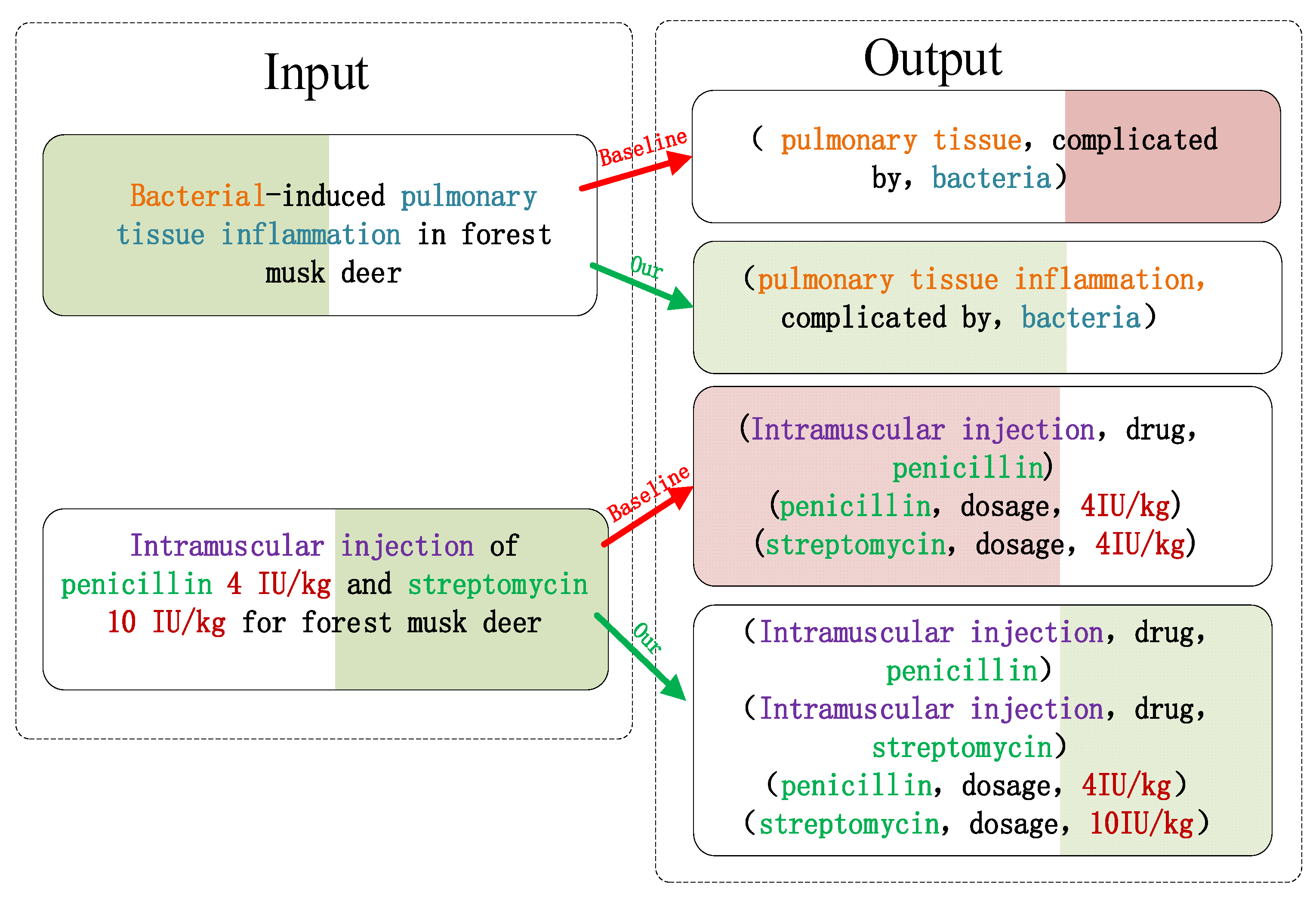
3.5. Qualitative Analysis
4. Discussion
4.1. Model Performance
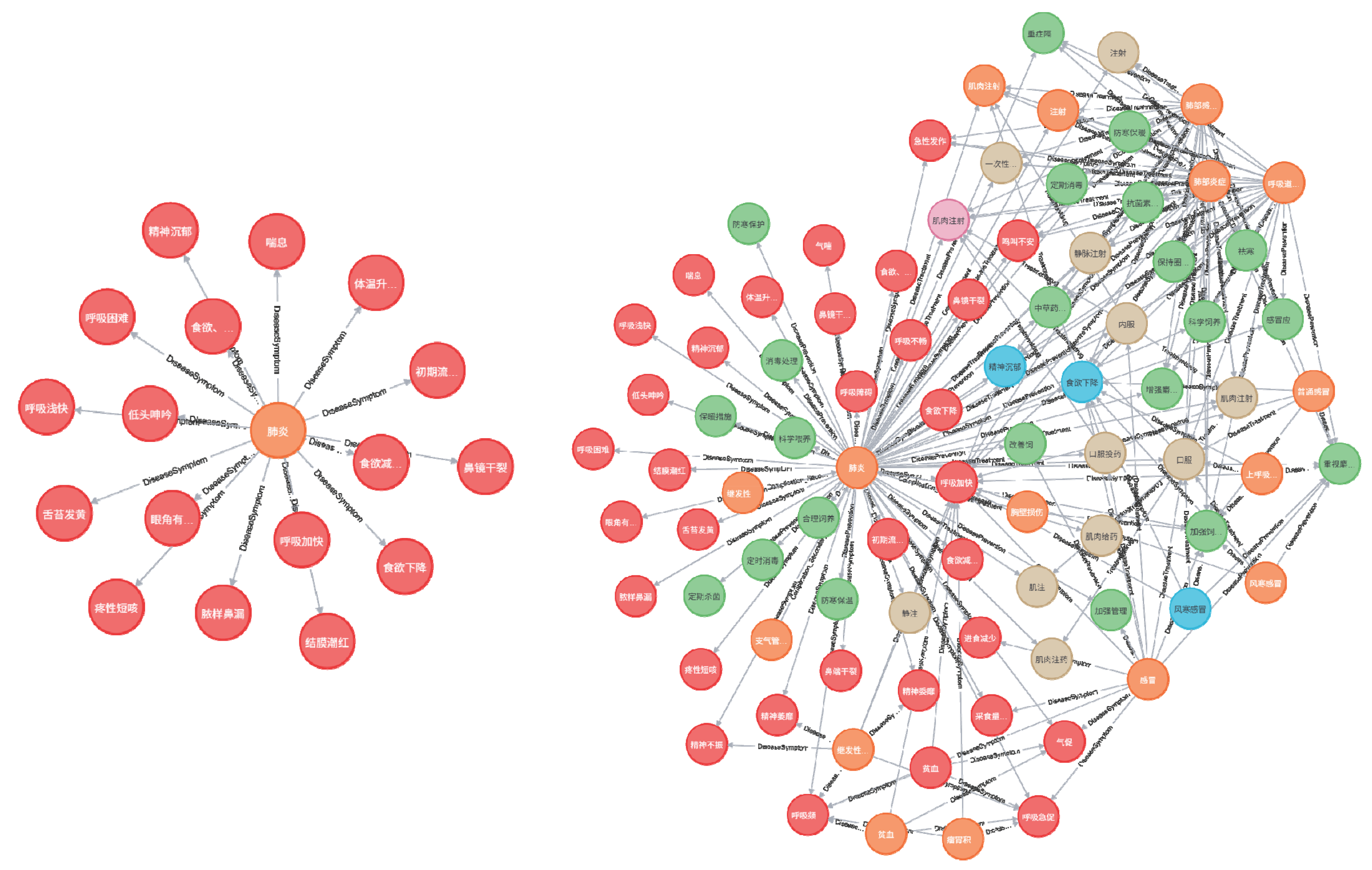
4.2. Partial Construction of the Forest Musk Deer Disease Knowledge Graph
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lu, X.; Sheng, Y.; Ye, H.; Yang, Z.; Meng, X. Development of Stereotypic Behaviors and Personality Traits of Captive Male Forest Musk Deer and Relationships with Musk Secretion. Vet. Sci. 2026, 13, 261. [Google Scholar] [CrossRef]
- Liu, K.; Xie, L.; Deng, M.; Zhang, X.; Luo, J.; Li, X. Zoology, chemical composition, pharmacology, quality control and future perspective of Musk (Moschus): A review. Chin. Med. 2021, 16, 46. [Google Scholar] [CrossRef]
- Liu, H.; Xiao, L.; Liu, Z.; Deng, Y.; Zhu, J.; Yang, C.; Liu, Q.; Tian, D.; Cui, X.; Peng, J. Impacts of Captive Domestication and Geographical Divergence on the Gut Microbiome of Endangered Forest Musk Deer. Animals 2025, 15, 1954. [Google Scholar] [CrossRef]
- Hu, J.; An, X.; Yang, P.; Tan, R.; Chen, T.; Chen, J.; Tao, Y.; Li, X.; Sun, K.; Zhang, S.; et al. Analysis of the Primary Pathogenic Bacteria in Abscess Disease of Musk Deer Using Metagenomic Approaches. Animals 2025, 15, 1105. [Google Scholar] [CrossRef]
- Yang, N.; Li, H.; Yang, X.; Wu, Y.; Lv, Z.; Zhang, Z.; Ma, X.; Zhou, X.; Zhang, X.; Zhao, K.; et al. Furazolidone reduces the pathogenesis of Trueperella pyogenes and Pseudomonas aeruginosa co-infection in a mouse model. Heliyon 2024, 10. [Google Scholar] [CrossRef] [PubMed]
- Lu, G.; Wang, Z.; Zhang, B.; Zhou, Z.; Hu, D.; Zhang, D. Detecting forest musk deer abscess disease pathogens using 16S rRNA high-throughput sequencing technology. Animals 2023, 13, 3142. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Wang, R.; Hu, Q.; Lv, N.; Zhang, L.; Yang, Z.; Zhou, Y.; Wang, X. Characterization of Pseudomonas aeruginosa bacteriophages and control hemorrhagic pneumonia on a mice model. Front. Microbiol. 2024, 15, 1396774. [Google Scholar] [CrossRef] [PubMed]
- Zhao, W.; Ren, Z.; Luo, Y.; Cheng, J.; Wang, J.; Wang, Y.; Yang, Z.; Yao, X.; Zhong, Z.; Yang, W.; et al. Metagenomics analysis of the gut microbiome in healthy and bacterial pneumonia forest musk deer. Genes. Genom. 2021, 43, 43–53. [Google Scholar] [CrossRef]
- Achhami, B.; Gurung, S.; Deshar, S.; Khaiju, S.; Thapa, L.K.; Gurung, S. Silent predators: Revealing the parasites of Himalayan musk deer (Moschus leucogaster) in Manaslu Conservation Area, Nepal. Int. J. Parasitol. Parasites Wildl. 2025, 101119. [Google Scholar] [CrossRef]
- Gao, Y.; Fu, Y.; Yan, L.; Hu, D.; Jiang, B.; Zhang, D. First record of traumatic myiasis obtained from forest musk deer (Moschus berezovskii). Int. J. Parasitol. Parasites Wildl. 2021, 16, 70–74. [Google Scholar] [CrossRef]
- Deng, L.; Chen, S.; Meng, W.; Zhou, Z.; Liu, H.; Zhong, Z.; Fu, H.; Shen, L.; Cao, S.; Tan, K.S.; et al. Changes in gut microbiota composition associated with the presence of enteric Protist Blastocystis in captive Forest musk deer (Moschus Berezovskii). Microbiol. Spectr. 2022, 10, e02269-21. [Google Scholar] [CrossRef]
- Shi, X.; Cheng, Z.; Zheng, C.; Wang, K.; Yang, J.; Jie, H.; Li, Y.; Zhang, M. The blood transcriptome of musk deer under heat stress condition reveals the regulatory mechanism of genes to maintain homeostasis metabolism. BMC Genom. 2025, 26, 400. [Google Scholar] [CrossRef]
- Yang, C.; Wang, H.; Sai, J.; Zhang, P.; Yan, M. Construction of knowledge graph of forest musk deer based on BiLSTM-CRF model and DPA method. In Proceedings of the Proceedings of the IOP Conference Series: Earth and Environmental Science, Chengdu, China, 21–23 August 2020; Vol. 615, p. 012008. [Google Scholar]
- Hao, Y.; Wang, H.; Zhang, D. Chinese Named Entity Recognition Method for Musk Deer Domain Based on Cross-Attention Enhanced Lexicon Features. Comput. Mater. Contin. 2025, 83, 2989. [Google Scholar] [CrossRef]
- Feng, H.; Wang, L.; Cao, F.; Ma, J.; Tang, J.; Feng, C.; Su, Z. Forest musk deer (Moschus berezovskii) in China: Research and protection. J. Vertebr. Biol. 2023, 72, 22067-1. [Google Scholar] [CrossRef]
- Yuan, W.; Wang, H.; Pe, L.; Zhang, T.; Hao, Y.; Lu, J.; Yan, W. Research on Entity and Relationship Extraction with Small Training Samples for Cotton Pests and Diseases. Agriculture 2024, 14, 457. [Google Scholar] [CrossRef]
- Huang, J.; Hao, X.; Wang, Y.; Song, R.; Mu, Z.; Niu, G.; Guo, X. Joint topic entity and intent recognition model for multimodal agricultural diseases and pests question answering. Comput. Electron. Agric. 2026, 241, 111253. [Google Scholar] [CrossRef]
- Gao, Y.; Li, X.; Wang, M.; Li, S. SemSynJE: Semantic and syntactic enhanced joint extraction of entity and relation for sheep disease prevention and treatment. Comput. Electron. Agric. 2025, 237, 110607. [Google Scholar] [CrossRef]
- De, S.; Sanyal, D.K.; Mukherjee, I. Fine-tuned encoder models with data augmentation beat ChatGPT in agricultural named entity recognition and relation extraction. Expert Syst. Appl. 2025, 277, 127126. [Google Scholar] [CrossRef]
- Wei, Z.; Su, J.; Wang, Y.; Tian, Y.; Chang, Y. A novel cascade binary tagging framework for relational triple extraction. In Proceedings of the Proceedings of the 58th Annual Meeting of the Association for Computational Linguistics (ACL), Seattle, USA, 5–10 July 2020; pp. 1476–1488. [Google Scholar]
- Wang, Y.; Yu, B.; Zhang, Y.; Liu; Tingwen bit Zhu, H.; Sun, L. TPLinker: Single-stage joint extraction of entities and relations through token pair linking. In Proceedings of the 28th International Conference on Computational Linguistics (COLING), Barcelona, Spain, 8–13 December 2020; pp. 1572–1582. [Google Scholar]
- Zheng, H.; Wen, R.; Chen, X.; Yang, Y.; Zhang, Y.; Zhang, Z.; Zhang, N.; Qin, B.; Ming, X.; Zheng, Y. PRCC: Potential relation and global correspondence based joint relational triple extraction. In Proceedings of the 59th Annual Meeting of the Association for Computational Linguistics and the 11th International Joint Conference on Natural Language Processing (ACL-IJCNLP), Bangkok, Thailand, 1–6 August 2021; pp. 6225–6235. [Google Scholar]
- Shang, Y.M.; Huang, K.; Mao, X. Onerel: Joint entity and relation extraction with one module in one step. In Proceedings of the 36th AAAI Conference on Artificial Intelligence (AAAI), Vancouver, Canada, 22 February–1 March 2022; Vol. 36, pp. 11285–11293. [Google Scholar]
- Ren, F.; Zhang, L.; Zhao, X.; Yin, S.; Liu, S.; Li, B. A simple but effective bidirectional framework for relational triple extraction. In Proceedings of the Proceedings of the 15th ACM International Conference on Web Search and Data Mining (WSDM), Phoenix, USA, 21–25 February 2022; pp. 824–832. [Google Scholar]
- Lou, Y.; Gao, M.; Zhang, S.; Yang, F.; Wang, S.; He, Y.; Yang, J.e.a. Chinese Named Entity Recognition for Dairy Cow Diseases by Fusion of Multi-Semantic Features Using Self-Attention-Based Deep Learning. Animals 2025, 15, 822. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Zhang, L.; Wu, H.; Wang, C.; Chen, C.; Zhu, H.; Liang, F. Chinese named entity recognition for agricultural diseases based on entity-related visual prompts injection. Comput. Electron. Agric. 2024, 227, 109493. [Google Scholar] [CrossRef]
- Wang, Y.; Tong, H.; Zhu, Z.; Hou, F.; Li, Y. Enhancing biomedical named entity recognition with parallel boundary detection and category classification. BMC Bioinform. 2025, 26, 63. [Google Scholar] [CrossRef]
- Guo, Q.; Dong, Y.; Tian, L.; Kang, Z.; Zhang, Y.; Wang, S. BANER: Boundary-aware LLMs for few-shot named entity recognition. In Proceedings of the Proceedings of the 31st International Conference on Computational Linguistics (COLING), Abu Dhabi, UAE, 19–24 January 2025; pp. 10375–10389. [Google Scholar]
- Zhu, W.; Fu, Y. Biomedical relation extraction based on a cascade binary tagging framework with varied decoders. Expert Syst. Appl. 2026, 295, 128838. [Google Scholar] [CrossRef]
- Su, J.; Ahmed, M.; Lu, Y.; Pan, S.; Bo, W.; Liu, Y. Roformer: Enhanced transformer with rotary position embedding. Neurocomputing 2024, 568, 127063. [Google Scholar] [CrossRef]
- Raffel, C.; Shazeer, N.; Roberts, A.; Lee, K.; Narang, S.; Matena, M.; Zhou, Y.; Li, W.; Liu, P.J. Exploring the limits of transfer learning with a unified text-to-text transformer. J. Mach. Learn. Res. 2020, 21, 1–67. [Google Scholar]
- Su, J.; Murtadha, A.; Pan, S.; Hou, J.; Sun, J.; Huang, W.; Wen, B.; Liu, Y. Global pointer: Novel efficient span-based approach for named entity recognition. arXiv 2022, arXiv:2208.03054. [Google Scholar] [CrossRef]
- Guan, T.; Zan, H.; Zhou, X.; Xu, H.; Zhang, K. CMelE: Construction and evaluation of Chinese medical information extraction dataset. In Proceedings of the Proceedings of the 9th CCF International Conference on Natural Language Processing and Chinese Computing (NLPCC), Zhengzhou, China, 14–18 October 2020; pp. 270–282. [Google Scholar]
- Devlin, J.; Chang, M.W.; Lee, K.; Toutanova, K. Bert: Pre-training of deep bidirectional transformers for language understanding. In Proceedings of the Proceedings of the 2019 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies (NAACL-HLT), Minneapolis, MN, USA, 2–7 June 2019; pp. 4171–4186. [Google Scholar]
- Tang, R.; Chen, Y.; Qin, Y.; Huang, R.; Zheng, Q. Boundary regression model for joint entity and relation extraction. Expert Syst. Appl. 2023, 229, 120441. [Google Scholar] [CrossRef]
- Bachlechner, T.; Majumder, B.P.; Mao, H.; Cottrell, G.; McAuley, J. Rezero is all you need: Fast convergence at large depth. In Proceedings of the Proceedings of the 37th Conference on Uncertainty in Artificial Intelligence (UAI), Mexico City, Mexico, 27–30 July 2021; pp. 1352–1361. [Google Scholar]
- Wang, L.; Wu, F.; Liu, X.; Cao, J.; Ma, M.; Qu, Z. Relationship extraction between entities with long distance dependencies and noise based on semantic and syntactic features. Sci. Rep. 2025, 15, 15750. [Google Scholar] [CrossRef]
- Long, T.; Yu, R.; You, X.; Shen, W.; Wei, X.; Gu, Z. FSCA-YOLO: An Enhanced YOLO-Based Model for Multi-Target Dairy Cow Behavior Recognition. Animals 2025, 15, 2631. [Google Scholar] [CrossRef]
- Zhong, Z.; Chen, D. A frustratingly easy approach for entity and relation extraction. In Proceedings of the Proceedings of the 2021 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies (NAACL-HLT), Toronto, ON, Canada, 6–11 June 2021; pp. 50–61. [Google Scholar]
| Entity Type | Definition | Relation Type | Definition of Relation | Example (Triplet) |
|---|---|---|---|---|
| Disease | Pathological conditions of forest musk deer. | DiseaseCause | The pathological or environmental origin of a disease. | (Gastroenteritis, DiseaseCause, Bacteria) |
| Symptoms | Clinical signs of a specific disease. | DiseaseSymptom | The correlation between a disease and its clinical manifestations. | (Pneumonia, DiseaseSymptom, Dyspnea) |
| Drug | Agents used for clinical intervention prescribed for a condition. | TreatmentDrug | Medication prescribed for a condition. | (Abscess, TreatmentDrug, Penicillin) |
| Dosage | Amount or frequency of medication for a drug. | DrugDosage | Administration details for a drug. | (Amoxicillin, DrugDosage, 20mg/kg) |
| Treatment | Therapeutic procedures or strategies approach to treat a disease. | DiseaseTreatment | Strategies for prophylaxis and health management. | (Fracture, DiseaseTreatment, Splinting) |
| Prevention | Actions taken to inhibit disease spread. | DiseasePrevention | Strategies for prophylaxis and health management. | (Parasitism, DiseasePrevention, Deworming) |
| Causes | Clinical causes of disease. | Complication/ SecondaryDisease |
Secondary conditions arising from a primary disease. | (Pneumonia, Complication/Secondary Disease, Laryngitis) |
| MS-data | CMeIE-V2 | ||
|---|---|---|---|
| Parameters | Value | Parameters | Value |
| Batch size | 8 | Batch size | 16 |
| Learning rate | 2e-5 | Learning rate | 2e-5 |
| Bert dim | 768 | Bert dim | 768 |
| epochs | 100 | epochs | 50 |
| hidden_size | 64 | hidden_size | 64 |
| Optimization algorithm | Adam | Optimization algorithm | Adam |
| Model | MS-data | CMeIE-V2 | ||||
|---|---|---|---|---|---|---|
| F1 | Re | P | F1 | Re | P | |
| CasRel | 0.702 | 0.654 | 0.764 | 0.482 | 0.477 | 0.488 |
| TPLinker | 0.781 | 0.776 | 0.798 | 0.502 | 0.498 | 0.507 |
| PRGC | 0.836 | 0.846 | 0.826 | 0.579 | 0.572 | 0.586 |
| PURE | 0.743 | 0.753 | 0.724 | 0.532 | 0.548 | 0.527 |
| Onerel | 0.842 | 0.808 | 0.879 | 0.585 | 0.586 | 0.583 |
| BiBRT | 0.844 | 0.824 | 0.864 | 0.539 | 0.504 | 0.579 |
| GPlinker | 0.867 | 0.833 | 0.903 | 0.583 | 0.525 | 0.630 |
| Our | 0.887 | 0.861 | 0.918 | 0.590 | 0.551 | 0.633 |
| Version | BAM | WSMCE | RDBM | F1 | Re | P |
|---|---|---|---|---|---|---|
| GPLinker | 0.867 | 0.833 | 0.903 | |||
| Version1 | √ | 0.875 | 0.849 | 0.898 | ||
| Version2 | √ | 0.877 | 0.843 | 0.904 | ||
| Version3 | √ | 0.876 | 0.838 | 0.907 | ||
| Version4 | √ | √ | 0.878 | 0.855 | 0.905 | |
| Version5 | √ | √ | 0.878 | 0.823 | 0.914 | |
| Version6 | √ | √ | 0.881 | 0.854 | 0.910 | |
| BRW-GPLinker | √ | √ | √ | 0.887 | 0.861 | 0.916 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




