Submitted:
20 April 2026
Posted:
21 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Methodology Review
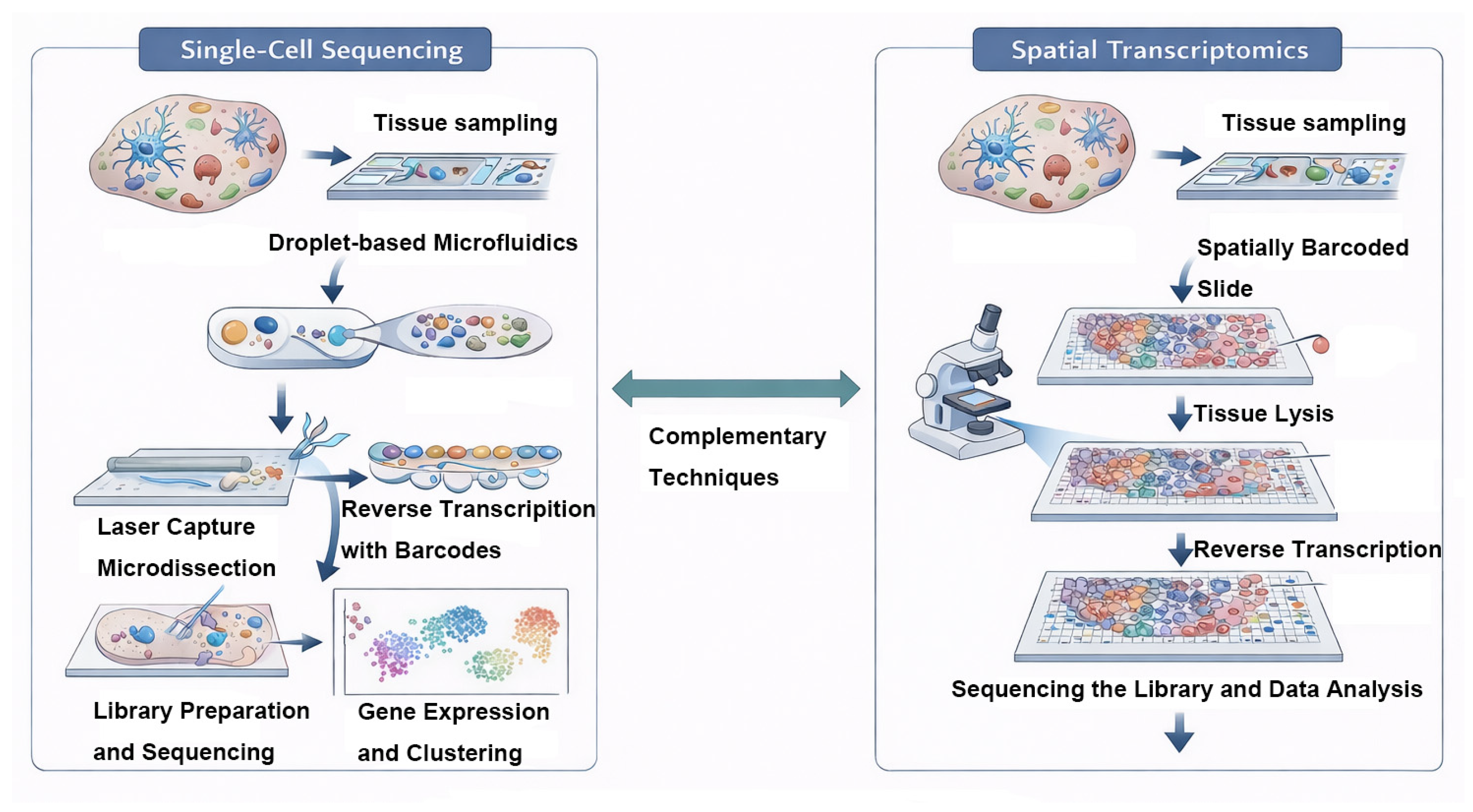
2.1. Single-Cell RNA Sequencing (scRNA-seq/snRNA-seq)
2.2. Spatial Transcriptomics Technology
3. Major Findings
3.1. Animal Model Studies
3.2. Human Studies
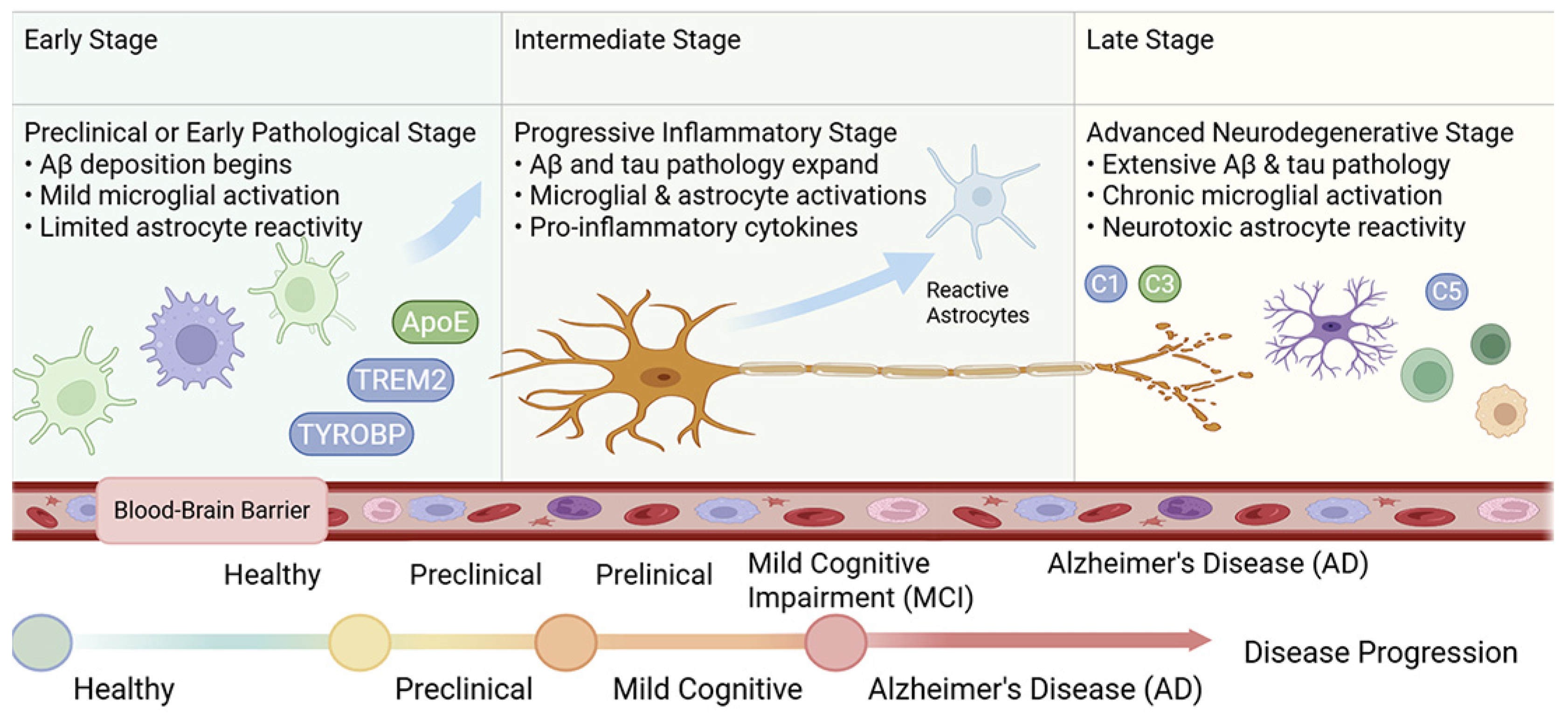
3.3. Comparison Across Disease Stages
3.4. Spatial Transcriptomics Reveals Differences Between Aβ Plaque Proximal and Distal Regions
3.5. Intercellular Interaction Networks and Key Regulatory Factors
4. Technical Limitations and Improvement Needs
5. Clinical Translation Prospects
6. Challenges and Future Directions
7. Conclusion and Outlook
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| AD | Alzheimer’s Disease |
| Aβ | Beta-amyloid |
| NFTs | Intraneuronal neurofibrillary tangles |
| scRNA-seq/snRNA-seq | Single-cell RNA sequencing |
| APP | Amyloid precursor protein |
| GWAS | Genome-wide association studies |
| MG | Microglia |
| AS | Astrocyte |
| DAO | Disease-associated oligodendrocyte |
| FISH | Fluorescence in situ hybridization |
| DAM | Disease-associated microglia |
| DAA | Disease-associated astrocyte |
| DSAD | Down syndrome-associated AD |
| PIGs | Plaque-induced genes |
| TIM | Terminally inflammatory microglia |
| MCI | Mild cognitive impairment |
| SCD | Subjective cognitive decline |
| IFN-I | Type I interferon |
| MGnD | Neurodegenerative microglia |
| CSF | Cerebrospinal fluid |
References
- Mathys, H.; Boix, C. A.; Akay, L. A.; Xia, Z.; Davila-Velderrain, J.; Ng, A. P.; Jiang, X.; Abdelhady, G.; Galani, K.; Mantero, J.; Band, N.; James, B. T.; Babu, S.; Galiana-Melendez, F.; Louderback, K.; Prokopenko, D.; Tanzi, R. E.; Bennett, D. A.; Tsai, L.-H.; Kellis, M. Single-cell multiregion dissection of Alzheimer’s disease. Nature 2024, 632, 858–868. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Global Status Report on the Public Health Response to Dementia, 1st ed.; World Health Organization: Geneva, 2021. [Google Scholar]
- Park, H.; Cho, B.; Kim, H.; Saito, T.; Saido, T. C.; Won, K. J.; Kim, J. Single-cell RNA-sequencing identifies disease-associated oligodendrocytes in male APP NL-G-F and 5XFAD mice. Nat Commun 2023, 14, 802. [Google Scholar] [CrossRef] [PubMed]
- Scheltens, P.; De Strooper, B.; Kivipelto, M.; Holstege, H.; Chetelat, G.; Teunissen, C. E.; Cummings, J.; van der Flier, W. M. Alzheimer’s disease. Lancet 2021, 397, 1577–1590. [Google Scholar] [CrossRef]
- Long, J. M.; Holtzman, D. M. Alzheimer Disease: An Update on Pathobiology and Treatment Strategies. Cell 2019, 179, 312–339. [Google Scholar] [CrossRef] [PubMed]
- Cummings, J.; Zhou, Y.; Lee, G.; Zhong, K.; Fonseca, J.; Cheng, F. Alzheimer’s disease drug development pipeline: 2023. Alzheimers Dement (N Y) 2023, 9, e12385. [Google Scholar] [CrossRef]
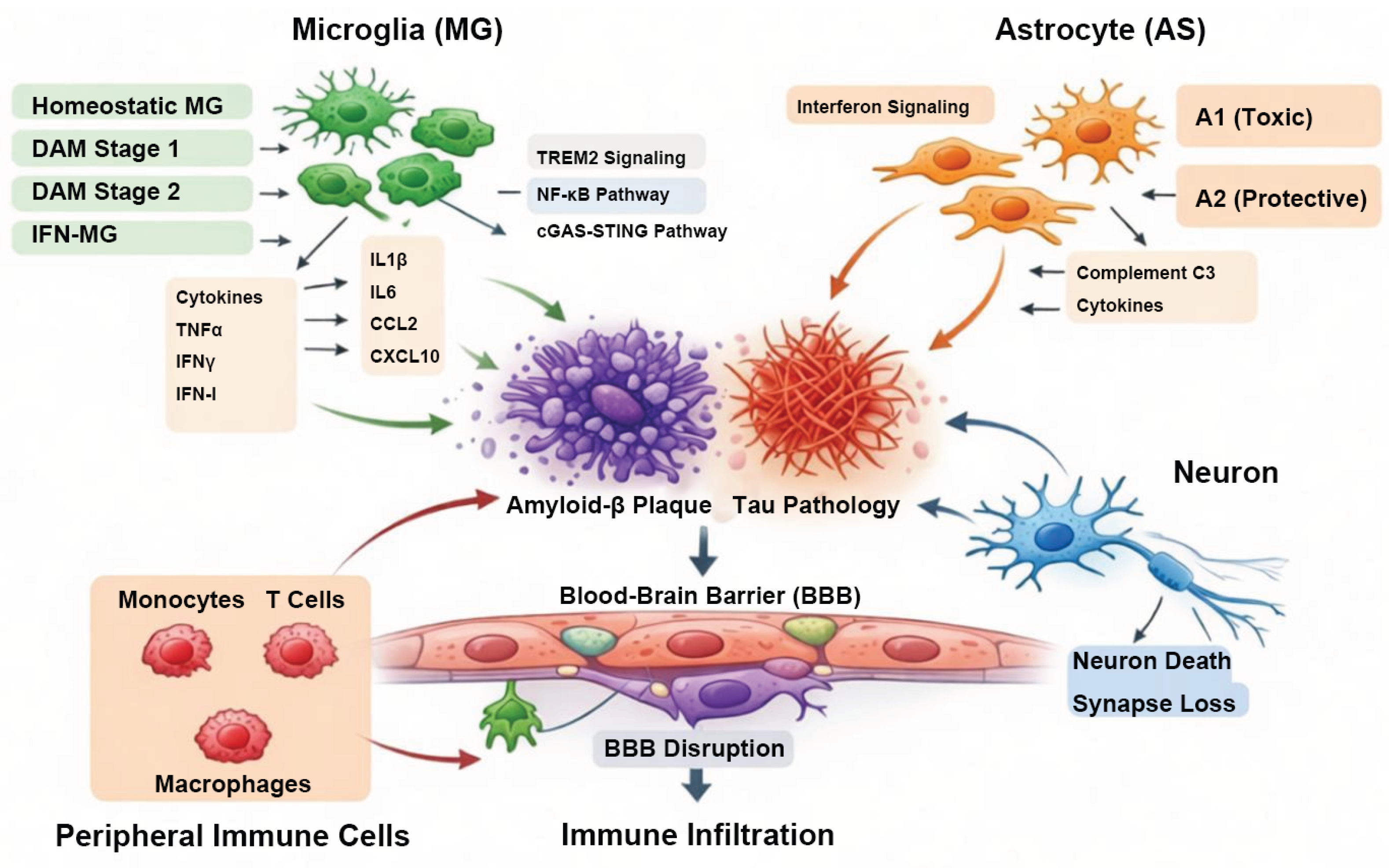
- Singh, D. Astrocytic and microglial cells as the modulators of neuroinflammation in Alzheimer’s disease. J Neuroinflammation 2022, 19, 206. [Google Scholar] [CrossRef]
- Han, J.; Zhang, Z.; Zhang, P.; Yu, Q.; Cheng, Q.; Lu, Z.; Zong, S. The roles of microglia and astrocytes in neuroinflammation of Alzheimer’s disease. Front Neurosci 2025, 19, 1575453. [Google Scholar] [CrossRef]
- Hansen, D. V.; Hanson, J. E.; Sheng, M. Microglia in Alzheimer’s disease. J Cell Biol 2018, 217, 459–472. [Google Scholar] [CrossRef]
- Ising, C.; Venegas, C.; Zhang, S.; Scheiblich, H.; Schmidt, S. V.; Vieira-Saecker, A.; Schwartz, S.; Albasset, S.; McManus, R. M.; Tejera, D.; Griep, A.; Santarelli, F.; Brosseron, F.; Opitz, S.; Stunden, J.; Merten, M.; Kayed, R.; Golenbock, D. T.; Blum, D.; Latz, E.; Buée, L.; Heneka, M. T. NLRP3 inflammasome activation drives tau pathology. Nature 2019, 575, 669–673. [Google Scholar] [CrossRef]
- Kitazawa, M.; Cheng, D.; Tsukamoto, M. R.; Koike, M. A.; Wes, P. D.; Vasilevko, V.; Cribbs, D. H.; LaFerla, F. M. Blocking IL-1 signaling rescues cognition, attenuates tau pathology, and restores neuronal beta-catenin pathway function in an Alzheimer’s disease model. J Immunol 2011, 187, 6539–49. [Google Scholar] [CrossRef]
- Dejanovic, B.; Wu, T.; Tsai, M. C.; Graykowski, D.; Gandham, V. D.; Rose, C. M.; Bakalarski, C. E.; Ngu, H.; Wang, Y.; Pandey, S.; Rezzonico, M. G.; Friedman, B. A.; Edmonds, R.; De Maziere, A.; Rakosi-Schmidt, R.; Singh, T.; Klumperman, J.; Foreman, O.; Chang, M. C.; Xie, L.; Sheng, M.; Hanson, J. E. Complement C1q-dependent excitatory and inhibitory synapse elimination by astrocytes and microglia in Alzheimer’s disease mouse models. Nat Aging 2022, 2, 837–850. [Google Scholar] [CrossRef]
- Abdulkhaliq, A. A.; Alasiri, G.; Kim, B.; Khan, J.; Ajoolabady, A.; Yousof, S. M.; Ren, J.; Tuomilehto, J.; Borai, A.; Alrfaei, B. M.; Pratico, D. TREM2 in neurodegeneration and diseases. Mol Psychiatry 2026. [Google Scholar] [CrossRef]
- Ulland, T. K.; Colonna, M. TREM2 - a key player in microglial biology and Alzheimer disease. Nat Rev Neurol 2018, 14, 667–675. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Huang, Y.; Fan, Y.; Xue, F. Co-regulation of microglial subgroups in Alzheimer’s amyloid pathology: Implications for diagnosis and drug development. PLoS One 2025, 20, e0337741. [Google Scholar] [CrossRef] [PubMed]
- Williams, C. G.; Lee, H. J.; Asatsuma, T.; Vento-Tormo, R.; Haque, A. An introduction to spatial transcriptomics for biomedical research. Genome Med 2022, 14, 68. [Google Scholar] [CrossRef] [PubMed]
- Tang, F.; Barbacioru, C.; Wang, Y.; Nordman, E.; Lee, C.; Xu, N.; Wang, X.; Bodeau, J.; Tuch, B. B.; Siddiqui, A.; Lao, K.; Surani, M. A. mRNA-Seq whole-transcriptome analysis of a single cell. Nat Methods 2009, 6, 377–82. [Google Scholar] [CrossRef]
- Papalexi, E.; Satija, R. Single-cell RNA sequencing to explore immune cell heterogeneity. Nat Rev Immunol 2018, 18, 35–45. [Google Scholar] [CrossRef]
- Mathys, H.; Davila-Velderrain, J.; Peng, Z.; Gao, F.; Mohammadi, S.; Young, J. Z.; Menon, M.; He, L.; Abdurrob, F.; Jiang, X.; Martorell, A. J.; Ransohoff, R. M.; Hafler, B. P.; Bennett, D. A.; Kellis, M.; Tsai, L. H. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature 2019, 570, 332–337. [Google Scholar] [CrossRef]
- Chen, J.; Wang, Y.; Ko, J. Single-cell and spatially resolved omics: Advances and limitations. Journal of Pharmaceutical Analysis 2023, 13, 833–835. [Google Scholar] [CrossRef]
- Yu, Y.; Mai, Y.; Zheng, Y.; Shi, L. Assessing and mitigating batch effects in large-scale omics studies. Genome Biol 2024, 25, 254. [Google Scholar] [CrossRef]
- Stahl, P. L.; Salmen, F.; Vickovic, S.; Lundmark, A.; Navarro, J. F.; Magnusson, J.; Giacomello, S.; Asp, M.; Westholm, J. O.; Huss, M.; Mollbrink, A.; Linnarsson, S.; Codeluppi, S.; Borg, A.; Ponten, F.; Costea, P. I.; Sahlen, P.; Mulder, J.; Bergmann, O.; Lundeberg, J.; Frisen, J. Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science 2016, 353, 78–82. [Google Scholar] [CrossRef] [PubMed]
- Fatemi, M.; Feng, E.; Sharma, C.; Azher, Z.; Goel, T.; Ramwala, O.; Palisoul, S. M.; Barney, R. E.; Perreard, L.; Kolling, F. W.; Salas, L. A.; Christensen, B. C.; Tsongalis, G. J.; Vaickus, L. J.; Levy, J. J. Inferring spatial transcriptomics markers from whole slide images to characterize metastasis-related spatial heterogeneity of colorectal tumors: A pilot study. J Pathol Inform 2023, 14, 100308. [Google Scholar] [CrossRef]
- Rodriques, S. G.; Stickels, R. R.; Goeva, A.; Martin, C. A.; Murray, E.; Vanderburg, C. R.; Welch, J.; Chen, L. M.; Chen, F.; Macosko, E. Z. Slide-seq: A scalable technology for measuring genome-wide expression at high spatial resolution. Science 2019, 363, 1463–1467. [Google Scholar] [CrossRef]
- Wang, G.; Moffitt, J. R.; Zhuang, X. Multiplexed imaging of high-density libraries of RNAs with MERFISH and expansion microscopy. Scientific Reports 2018, 8. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Yang, M.; Deng, Y.; Su, G.; Enninful, A.; Guo, C. C.; Tebaldi, T.; Zhang, D.; Kim, D.; Bai, Z.; Norris, E.; Pan, A.; Li, J.; Xiao, Y.; Halene, S.; Fan, R. High-Spatial-Resolution Multi-Omics Sequencing via Deterministic Barcoding in Tissue. Cell 2020, 183, 1665–1681 e18. [Google Scholar] [CrossRef] [PubMed]
- Keren-Shaul, H.; Spinrad, A.; Weiner, A.; Matcovitch-Natan, O.; Dvir-Szternfeld, R.; Ulland, T. K.; David, E.; Baruch, K.; Lara-Astaiso, D.; Toth, B.; Itzkovitz, S.; Colonna, M.; Schwartz, M.; Amit, I. A Unique Microglia Type Associated with Restricting Development of Alzheimer’s Disease. Cell 2017, 169, 1276–1290 e17. [Google Scholar] [CrossRef]
- Wang, Y.; Cella, M.; Mallinson, K.; Ulrich, J. D.; Young, K. L.; Robinette, M. L.; Gilfillan, S.; Krishnan, G. M.; Sudhakar, S.; Zinselmeyer, B. H.; Holtzman, D. M.; Cirrito, J. R.; Colonna, M. TREM2 lipid sensing sustains the microglial response in an Alzheimer’s disease model. Cell 2015, 160, 1061–71. [Google Scholar] [CrossRef]
- Xue, F.; Du, H. TREM2 mediates microglial anti-inflammatory activations in Alzheimer’s Disease: Lessons learned from transcriptomics. Cells 2021, 10. [Google Scholar] [CrossRef]
- Habib, N.; McCabe, C.; Medina, S.; Varshavsky, M.; Kitsberg, D.; Dvir-Szternfeld, R.; Green, G.; Dionne, D.; Nguyen, L.; Marshall, J. L.; Chen, F.; Zhang, F.; Kaplan, T.; Regev, A.; Schwartz, M. Disease-associated astrocytes in Alzheimer’s disease and aging. Nat Neurosci 2020, 23, 701–706. [Google Scholar] [CrossRef]
- Liddelow, S. A.; Guttenplan, K. A.; Clarke, L. E.; Bennett, F. C.; Bohlen, C. J.; Schirmer, L.; Bennett, M. L.; Munch, A. E.; Chung, W. S.; Peterson, T. C.; Wilton, D. K.; Frouin, A.; Napier, B. A.; Panicker, N.; Kumar, M.; Buckwalter, M. S.; Rowitch, D. H.; Dawson, V. L.; Dawson, T. M.; Stevens, B.; Barres, B. A. Neurotoxic reactive astrocytes are induced by activated microglia. Nature 2017, 541, 481–487. [Google Scholar] [CrossRef]
- Mallach, A.; Zielonka, M.; van Lieshout, V.; An, Y.; Khoo, J. H.; Vanheusden, M.; Chen, W. T.; Moechars, D.; Arancibia-Carcamo, I. L.; Fiers, M.; De Strooper, B. Microglia-astrocyte crosstalk in the amyloid plaque niche of an Alzheimer’s disease mouse model, as revealed by spatial transcriptomics. Cell Rep 2024, 43, 114216. [Google Scholar] [CrossRef]
- Wang, P.; Han, L.; Wang, L.; Tao, Q.; Guo, Z.; Luo, T.; He, Y.; Xu, Z.; Yu, J.; Liu, Y.; Wu, Z.; Xu, B.; Jin, B.; Wei, Y.; Yang, Y.; Cheng, M.; Jiang, Y.; Tian, C.; Zheng, H.; Fan, Z.; Jiang, P.; Gao, Y.; Wu, J.; Wang, S.; Sun, B.; Fang, Z.; Lei, J.; Luo, B.; Wen, H.; Peng, G.; Tang, Y.; Yang, T.; Chen, J.; Zhuang, Z.; Su, X.; Pan, C.; Zhu, K.; Shen, Y.; Liu, S.; Bao, A.; Yao, J.; Wang, J.; Xu, X.; Li, X. M.; Liu, L.; Duan, S.; Zhang, J. Molecular pathways and diagnosis in spatially resolved Alzheimer’s hippocampal atlas. Neuron 2025, 113, 2123–2140 e9. [Google Scholar] [CrossRef]
- Sziraki, A.; Lu, Z.; Lee, J.; Banyai, G.; Anderson, S.; Abdulraouf, A.; Metzner, E.; Liao, A.; Banfelder, J.; Epstein, A.; Schaefer, C.; Xu, Z.; Zhang, Z.; Gan, L.; Nelson, P. T.; Zhou, W.; Cao, J. A global view of aging and Alzheimer’s pathogenesis-associated cell population dynamics and molecular signatures in human and mouse brains. Nat Genet 2023, 55, 2104–2116. [Google Scholar] [CrossRef]
- Miyoshi, E.; Morabito, S.; Henningfield, C. M.; Das, S.; Rahimzadeh, N.; Shabestari, S. K.; Michael, N.; Emerson, N.; Reese, F.; Shi, Z.; Cao, Z.; Srinivasan, S. S.; Scarfone, V. M.; Arreola, M. A.; Lu, J.; Wright, S.; Silva, J.; Leavy, K.; Lott, I. T.; Doran, E.; Yong, W. H.; Shahin, S.; Perez-Rosendahl, M.; Alzheimer’s Biomarkers Consortium-Down, S.; Head, E.; Green, K. N.; Swarup, V. Spatial and single-nucleus transcriptomic analysis of genetic and sporadic forms of Alzheimer’s disease. Nat Genet 2024, 56, 2704–2717. [Google Scholar] [CrossRef]
- Avey, D. R.; Ng, B.; Vialle, R. A.; Kearns, N. A.; de Paiva Lopes, K.; Iatrou, A.; De Tissera, S.; Vyas, H.; Saunders, D. M.; Flood, D. J.; Xu, J.; Tasaki, S.; Gaiteri, C.; Bennett, D. A.; Wang, Y. Uncovering plaque-glia niches in human Alzheimer’s disease brains using spatial transcriptomics. Mol Neurodegener Adv 2025, 1, 2. [Google Scholar] [CrossRef]
- Millet, A.; Ledo, J. H.; Tavazoie, S. F. An exhausted-like microglial population accumulates in aged and APOE4 genotype Alzheimer’s brains. Immunity 2024, 57, 153–170 e6. [Google Scholar] [CrossRef] [PubMed]
- Gerrits, E.; Brouwer, N.; Kooistra, S. M.; Woodbury, M. E.; Vermeiren, Y.; Lambourne, M.; Mulder, J.; Kummer, M.; Moller, T.; Biber, K.; Dunnen, W.; De Deyn, P. P.; Eggen, B. J. L.; Boddeke, E. Distinct amyloid-beta and tau-associated microglia profiles in Alzheimer’s disease. Acta Neuropathol 2021, 141, 681–696. [Google Scholar] [CrossRef]
- Sun, N.; Victor, M. B.; Park, Y. P.; Xiong, X.; Scannail, A. N.; Leary, N.; Prosper, S.; Viswanathan, S.; Luna, X.; Boix, C. A.; James, B. T.; Tanigawa, Y.; Galani, K.; Mathys, H.; Jiang, X.; Ng, A. P.; Bennett, D. A.; Tsai, L. H.; Kellis, M. Human microglial state dynamics in Alzheimer’s disease progression. Cell 2023, 186, 4386–4403 e29. [Google Scholar] [CrossRef]
- Song, L.; Chen, J.; Lo, C. Z.; Guo, Q.; Consortium, Z. I. B.; Feng, J.; Zhao, X. M. Impaired type I interferon signaling activity implicated in the peripheral blood transcriptome of preclinical Alzheimer’s disease. EBioMedicine 2022, 82, 104175. [Google Scholar] [CrossRef] [PubMed]
- Zeng, H.; Huang, J.; Zhou, H.; Meilandt, W. J.; Dejanovic, B.; Zhou, Y.; Bohlen, C. J.; Lee, S. H.; Ren, J.; Liu, A.; Tang, Z.; Sheng, H.; Liu, J.; Sheng, M.; Wang, X. Integrative in situ mapping of single-cell transcriptional states and tissue histopathology in a mouse model of Alzheimer’s disease. Nat Neurosci 2023, 26, 430–446. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Huang, R.; Pan, J. Dynamic Intercellular Networks in the CNS: Mechanisms of Crosstalk from Homeostasis to Neurodegeneration. Int J Mol Sci 2025, 26, 17. [Google Scholar] [CrossRef] [PubMed]
- Guttenplan, K. A.; Weigel, M. K.; Prakash, P.; Wijewardhane, P. R.; Hasel, P.; Rufen-Blanchette, U.; Munch, A. E.; Blum, J. A.; Fine, J.; Neal, M. C.; Bruce, K. D.; Gitler, A. D.; Chopra, G.; Liddelow, S. A.; Barres, B. A. Neurotoxic reactive astrocytes induce cell death via saturated lipids. Nature 2021, 599, 102–107. [Google Scholar] [CrossRef] [PubMed]
- De Strooper, B.; Karran, E. The Cellular Phase of Alzheimer’s Disease. Cell 2016, 164, 603–15. [Google Scholar] [CrossRef] [PubMed]
- Serrano-Pozo, A.; Das, S.; Hyman, B. T. APOE and Alzheimer’s disease: advances in genetics, pathophysiology, and therapeutic approaches. Lancet Neurol 2021, 20, 68–80. [Google Scholar] [CrossRef] [PubMed]
- Clayton, K.; Delpech, J. C.; Herron, S.; Iwahara, N.; Ericsson, M.; Saito, T.; Saido, T. C.; Ikezu, S.; Ikezu, T. Plaque associated microglia hyper-secrete extracellular vesicles and accelerate tau propagation in a humanized APP mouse model. Mol Neurodegener 2021, 16, 18. [Google Scholar] [CrossRef]
- Bakken, T. E.; Hodge, R. D.; Miller, J. A.; Yao, Z.; Nguyen, T. N.; Aevermann, B.; Barkan, E.; Bertagnolli, D.; Casper, T.; Dee, N.; Garren, E.; Goldy, J.; Graybuck, L. T.; Kroll, M.; Lasken, R. S.; Lathia, K.; Parry, S.; Rimorin, C.; Scheuermann, R. H.; Schork, N. J.; Shehata, S. I.; Tieu, M.; Phillips, J. W.; Bernard, A.; Smith, K. A.; Zeng, H.; Lein, E. S.; Tasic, B. Single-nucleus and single-cell transcriptomes compared in matched cortical cell types. PLoS One 2018, 13, e0209648. [Google Scholar] [CrossRef]
- Li, T.; Wang, Z.; Liu, Y.; He, S.; Zou, Q.; Zhang, Y. An overview of computational methods in single-cell transcriptomic cell type annotation. Brief Bioinform 2025, 26. [Google Scholar] [CrossRef]
- Rood, J. E.; Maartens, A.; Hupalowska, A.; Teichmann, S. A.; Regev, A. Impact of the Human Cell Atlas on medicine. Nat Med 2022, 28, 2486–2496. [Google Scholar] [CrossRef] [PubMed]
- Neff, R. A.; Wang, M.; Vatansever, S.; Guo, L.; Ming, C.; Wang, Q.; Wang, E.; Horgusluoglu-Moloch, E.; Song, W. M.; Li, A.; Castranio, E. L.; Tcw, J.; Ho, L.; Goate, A.; Fossati, V.; Noggle, S.; Gandy, S.; Ehrlich, M. E.; Katsel, P.; Schadt, E.; Cai, D.; Brennand, K. J.; Haroutunian, V.; Zhang, B. Molecular subtyping of Alzheimer’s disease using RNA sequencing data reveals novel mechanisms and targets. Sci Adv 2021, 7. [Google Scholar] [CrossRef] [PubMed]
- Narayanaswami, V.; Dahl, K.; Bernard-Gauthier, V.; Josephson, L.; Cumming, P.; Vasdev, N. Emerging PET Radiotracers and Targets for Imaging of Neuroinflammation in Neurodegenerative Diseases: Outlook Beyond TSPO. Mol Imaging 2018, 17, 1536012118792317. [Google Scholar] [CrossRef]
- Benedet, A. L.; Mila-Aloma, M.; Vrillon, A.; Ashton, N. J.; Pascoal, T. A.; Lussier, F.; Karikari, T. K.; Hourregue, C.; Cognat, E.; Dumurgier, J.; Stevenson, J.; Rahmouni, N.; Pallen, V.; Poltronetti, N. M.; Salvado, G.; Shekari, M.; Operto, G.; Gispert, J. D.; Minguillon, C.; Fauria, K.; Kollmorgen, G.; Suridjan, I.; Zimmer, E. R.; Zetterberg, H.; Molinuevo, J. L.; Paquet, C.; Rosa-Neto, P.; Blennow, K.; Suarez-Calvet, M.; Translational Biomarkers in, A.; Dementia study, A. s.; Families, s.; BioCogBank Paris Lariboisiere, c. Differences Between Plasma and Cerebrospinal Fluid Glial Fibrillary Acidic Protein Levels Across the Alzheimer Disease Continuum. JAMA Neurol 2021, 78, 1471–1483. [Google Scholar] [CrossRef]
- Chatterjee, P.; Pedrini, S.; Stoops, E.; Goozee, K.; Villemagne, V. L.; Asih, P. R.; Verberk, I. M. W.; Dave, P.; Taddei, K.; Sohrabi, H. R.; Zetterberg, H.; Blennow, K.; Teunissen, C. E.; Vanderstichele, H. M.; Martins, R. N. Plasma glial fibrillary acidic protein is elevated in cognitively normal older adults at risk of Alzheimer’s disease. Transl Psychiatry 2021, 11, 27. [Google Scholar] [CrossRef]
- Qiu, S.; Joshi, P. S.; Miller, M. I.; Xue, C.; Zhou, X.; Karjadi, C.; Chang, G. H.; Joshi, A. S.; Dwyer, B.; Zhu, S.; Kaku, M.; Zhou, Y.; Alderazi, Y. J.; Swaminathan, A.; Kedar, S.; Saint-Hilaire, M. H.; Auerbach, S. H.; Yuan, J.; Sartor, E. A.; Au, R.; Kolachalama, V. B. Development and validation of an interpretable deep learning framework for Alzheimer’s disease classification. Brain 2020, 143, 1920–1933. [Google Scholar] [CrossRef]
- Lewcock, J. W.; Schlepckow, K.; Di Paolo, G.; Tahirovic, S.; Monroe, K. M.; Haass, C. Emerging Microglia Biology Defines Novel Therapeutic Approaches for Alzheimer’s Disease. Neuron 2020, 108, 801–821. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Mustafa, M.; Yuede, C. M.; Salazar, S. V.; Kong, P.; Long, H.; Ward, M.; Siddiqui, O.; Paul, R.; Gilfillan, S.; Ibrahim, A.; Rhinn, H.; Tassi, I.; Rosenthal, A.; Schwabe, T.; Colonna, M. Anti-human TREM2 induces microglia proliferation and reduces pathology in an Alzheimer’s disease model. J Exp Med 2020, 217. [Google Scholar] [CrossRef] [PubMed]
- Lananna, B. V.; McKee, C. A.; King, M. W.; Del-Aguila, J. L.; Dimitry, J. M.; Farias, F. H. G.; Nadarajah, C. J.; Xiong, D. D.; Guo, C.; Cammack, A. J.; Elias, J. A.; Zhang, J.; Cruchaga, C.; Musiek, E. S. Chi3l1/YKL-40 is controlled by the astrocyte circadian clock and regulates neuroinflammation and Alzheimer’s disease pathogenesis. Sci Transl Med 2020, 12, 574. [Google Scholar] [CrossRef] [PubMed]
- Lian, H.; Litvinchuk, A.; Chiang, A. C.; Aithmitti, N.; Jankowsky, J. L.; Zheng, H. Astrocyte-Microglia Cross Talk through Complement Activation Modulates Amyloid Pathology in Mouse Models of Alzheimer’s Disease. J Neurosci 2016, 36, 577–89. [Google Scholar] [CrossRef]
- Dejanovic, B.; Huntley, M. A.; De Maziere, A.; Meilandt, W. J.; Wu, T.; Srinivasan, K.; Jiang, Z.; Gandham, V.; Friedman, B. A.; Ngu, H.; Foreman, O.; Carano, R. A. D.; Chih, B.; Klumperman, J.; Bakalarski, C.; Hanson, J. E.; Sheng, M. Changes in the Synaptic Proteome in Tauopathy and Rescue of Tau-Induced Synapse Loss by C1q Antibodies. Neuron 2018, 100, 1322–1336 e7. [Google Scholar] [CrossRef]
- Pascoal, T. A.; Benedet, A. L.; Ashton, N. J.; Kang, M. S.; Therriault, J.; Chamoun, M.; Savard, M.; Lussier, F. Z.; Tissot, C.; Karikari, T. K.; Ottoy, J.; Mathotaarachchi, S.; Stevenson, J.; Massarweh, G.; Scholl, M.; de Leon, M. J.; Soucy, J. P.; Edison, P.; Blennow, K.; Zetterberg, H.; Gauthier, S.; Rosa-Neto, P. Microglial activation and tau propagate jointly across Braak stages. Nat Med 2021, 27, 1592–1599. [Google Scholar] [CrossRef]
- Salter, M. W.; Stevens, B. Microglia emerge as central players in brain disease. Nat Med 2017, 23, 1018–1027. [Google Scholar] [CrossRef]
- Lau, S.-F.; Cao, H.; Fu, A. K. Y.; Ip, N. Y. Single-nucleus transcriptome analysis reveals dysregulation of angiogenic endothelial cells and neuroprotective glia in Alzheimer’s disease. Proceedings of the National Academy of Sciences 2020, 117, 25800–25809. [Google Scholar] [CrossRef] [PubMed]
- Ferrari-Souza, J. P.; Povala, G.; Rahmouni, N.; Bellaver, B.; Ferreira, P. C. L.; De Bastiani, M. A.; Leffa, D. T.; Lussier, F. Z.; Aguzzoli, C. S.; Brum, W. S.; Carello-Collar, G.; Borelli, W. V.; Therriault, J.; Macedo, A. C.; Servaes, S.; Stevenson, J.; Pola, I.; Gauthier, S.; Souza, D. O.; Schilling, L. P.; Lourenco, M. V.; Triana-Baltzer, G.; Kolb, H. C.; Benedet, A. L.; Ashton, N. J.; Tudorascu, D. L.; Zetterberg, H.; Blennow, K.; Johnson, S. C.; Pascoal, T. A.; Rosa-Neto, P.; Zimmer, E. R. Microglia modulate Abeta-dependent astrocyte reactivity in Alzheimer’s disease. Nat Neurosci 2026, 29, 81–87. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Firulyova, M.; Manis, M.; Herz, J.; Smirnov, I.; Aladyeva, E.; Wang, C.; Bao, X.; Finn, M. B.; Hu, H.; Shchukina, I.; Kim, M. W.; Yuede, C. M.; Kipnis, J.; Artyomov, M. N.; Ulrich, J. D.; Holtzman, D. M. Microglia-mediated T cell infiltration drives neurodegeneration in tauopathy. Nature 2023, 615, 668–677. [Google Scholar] [CrossRef]
- Padmanabhan, J.; Raja, W. K.; Mungenast, A. E.; Lin, Y.-T.; Ko, T.; Abdurrob, F.; Seo, J.; Tsai, L.-H. Self-Organizing 3D Human Neural Tissue Derived from Induced Pluripotent Stem Cells Recapitulate Alzheimer’s Disease Phenotypes. Plos One 2016, 11, 9. [Google Scholar]
- Yang, A. C.; Vest, R. T.; Kern, F.; Lee, D. P.; Agam, M.; Maat, C. A.; Losada, P. M.; Chen, M. B.; Schaum, N.; Khoury, N.; Toland, A.; Calcuttawala, K.; Shin, H.; Palovics, R.; Shin, A.; Wang, E. Y.; Luo, J.; Gate, D.; Schulz-Schaeffer, W. J.; Chu, P.; Siegenthaler, J. A.; McNerney, M. W.; Keller, A.; Wyss-Coray, T. A human brain vascular atlas reveals diverse mediators of Alzheimer’s risk. Nature 2022, 603, 885–892. [Google Scholar] [CrossRef] [PubMed]
| Plaform/ Technology |
Category | Resolution | Gene Detection Capacity | Throughput | Sample Type | Main Applications/Advantages |
| 10x Chromium (scRNA-snRNA) | Droplet-based (Sequencing) | Single-cell/single-nucleus (~10 μm) | Whole transcriptome (~10,000–20,000 genes) | High (up to 10⁴–10⁵ cells) | Live cells, frozen tissues | Single-cell heterogeneity analysis, high throughput |
| 10x Visium | Spatial capture (Sequencing) | ~55 μm per spot (1–10 cells per spot) | Whole transcriptome (~18,000–20,000 genes) | Moderate (~5,000 spots per section) | Fresh frozen (Visium v1), FFPE (Visium v2) | Whole-transcript spatial mapping, easy operation |
| Slide-seqV2 | Spatial capture (Sequencing) | ~10 μm per bead (near single-cell) | Whole transcriptome | High (millions of beads) | Frozen tissues | Higher spatial resolution, captures more cells |
| Stereo-seq | Spatial capture (Sequencing) | 0.5–3 μm per spot (single-cell and subcellular) | Whole transcriptome | Ultra-high (ultra-dense arrays) | Frozen tissues | Ultra-high resolution, enables subcellular localization |
| MERFISH (MERSCOPE) | In situ imaging (FISH) | Single-cell/subcellular | Scalable to thousands of genes (targeted probes) | Moderate | Fixed tissues | High spatial resolution, near 100% probe capture efficiency |
| seqFISH+ | In situ imaging (FISH) | Single-cell/subcellular | ~10,000 genes | Moderate | Fixed tissues | Ultra-high gene multiplexity, avoids optical crowding via multiple rounds of hybridization |
| GeoMx DSP | Region-of-interest (ROI) capture | Up to cellular level (adjustable ROI) | Targeted panels (hundreds to thousands of genes) | Low | FFPE, frozen tissues | Targeted gene panels, tissue-preserving, highly flexible ROI selection |
| Study (Year) | Sample/Model | Technology | Key Findings | Limitations |
| Keren-Shaul et al. (2017) [27] | 5xFAD mouse brains | scRNA-seq | Identified TREM2-dependent disease-associated microglia (DAM) subpopulation enriched in phagocytosis and inflammation-related genes | Early study, lacks spatial localization information |
| Habib et al. (2020) [30] | Hippocampus of 5xFAD mice | snRNA-seq | Identified disease-associated astrocytes (DAA), whose abundance increases with disease progression | Small sample size, mainly focused on the hippocampus |
| Mallach et al. (2024) [32] | Hippocampus of 5xFAD mice | Spatial transcriptomics (Visium) | Abundant microglial aggregation around plaques disrupts astrocytic signaling; microglial responses are consistent across brain regions, while astrocytic responses show high regional heterogeneity | Mouse model only, limited to the hippocampal region |
| Park et al. (2023) [3] | Male App NL-G-F and 5xFAD mice / human AD brains | scRNA-seq | Identified disease-associated oligodendrocyte (DAO) subpopulation; elevated Erk1/2 signaling activity in DAOs, and Erk inhibition restores myelination and ameliorates AD pathology | Focused on oligodendrocytes, limited coverage of other cell types |
| Study (Year) | Sample Type | Technology | Key Findings | Limitations |
| Mathys et al. (2024) [1] | 283 AD and control brains (6 brain regions) | snRNA-seq | Constructed an atlas of over 100 cell subpopulations; identified impaired neuronal populations in AD and linked Reelin signaling to cognitive resilience | Cross-sectional samples, no integration of spatial information |
| Sziraki et al. (2023) [34] | 1.5 million mouse cells / 118,000 human cells | scRNA-seq (EasySci) | Identified over 300 cell subtypes; revealed AD-associated transcriptional changes in rare cell types | Novel technology, requires further validation |
| Miyoshi et al. (2024) [35] | Brain tissues from sporadic AD (sAD) and Down syndrome-associated AD (DSAD) patients, 5xFAD mice | Spatial transcriptomics + snRNA-seq | Upregulated glial inflammatory programs in superficial cortical layers; validated transcriptional changes near plaques in 5xFAD mice | Differences between DSAD and sAD require further investigation |
| Avey et al. (2025) [36] | Posterior cingulate cortex from 21 AD patients | Spatial transcriptomics (Visium) + IHC | Increased neuronal apoptosis in “low Aβ” plaque regions; upregulated inflammation in “high gliosis” regions; enrichment of DAM/DAA gene modules | Relatively small number of cases |
| Gerrits et al. (2021) [38] | Human AD brain tissues | snRNA-seq | Identified microglial subpopulations specifically associated with Aβ and tau pathologies | Lacks spatial context information |
| Sun et al. (2023) [39] | Human AD brain tissues | snRNA-seq | Identified terminally inflammatory microglia (TIM) subpopulation, which accumulates in aged and APOE4 AD brains and exhibits inflammatory and stress phenotypes | Mainly focused on microglia, limited analysis of other cell types |
| Study | Sample/Model | Technology | Observations in Plaque-Proximal Regions | Observations in Plaque-Distal Regions |
| Avey et al. (2025) [36] | Posterior cingulate cortex of human AD patients | Visium + IHC | “High gliosis” plaque regions: upregulated inflammation and AD-related pathways; enrichment of DAM/DAA gene modules | “Low Aβ” plaque regions: increased neuronal apoptosis markers |
| Mallach et al. (2024) [32] | Hippocampus of 5xFAD mice | High-resolution spatial transcriptomics | Around plaques: dense microglial aggregation, disrupted astrocyte-neuron signaling (synaptic imbalance); elevated CD68 expression | Distal to plaques: weak glial responses, features of normal brain tissue |
| Miyoshi (2024) [35] | Humans and 5xFAD mice | Spatial transcriptomics + snRNA-seq | Near plaques in superficial cortical layers: upregulation of specific inflammatory genes (e.g., IL-1, complement components); transcriptomic profiles correlate with AD risk | Deep cortical layers: relatively downregulated inflammatory genes |
| Limitation Category | Specific Issues | Improvement Strategies |
| Sample Compatibility | Most spatial omics require fresh-frozen samples; scRNA-seq is limited to viable cells | Develop FFPE-compatible spatial technologies; optimize nuclei extraction and tissue fixation techniques |
| Spatial Resolution | Sequencing-based methods have low resolution; imaging-based methods have low throughput and limited coverage | Increase probe density; combine multiple spatial technologies (sequencing + FISH); develop high-throughput imaging protocols |
| Throughput and Coverage | Large-volume samples are difficult to fully cover; long imaging times | Automated microscopy; pre-select plaque regions via regional screening; parallel sequencing of multiple fragments |
| Doublets and Batch Effects | scRNA-seq is prone to doublet formation; batch effects exist across different datasets | Optimize cell concentration control; apply batch integration algorithms (e.g., Harmony); implement strict quality control |
| Cell Type Annotation | A few cell types lack specific markers; ambiguous cell type identification | Utilize multi-omics data (scATAC-seq, protein markers) to provide more phenotypic clues; assist annotation with AI methods |
| Data Integration | Direct comparison and integration of single-cell and spatial data are challenging; limited software tools | Develop cross-modal alignment tools; construct comprehensive databases; share standardized analytical workflows |
| Reproducibility | High tissue heterogeneity makes validation across different samples difficult | Increase sample size; perform multi-center data validation; release open-access data and code |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




