Submitted:
17 April 2026
Posted:
21 April 2026
You are already at the latest version
Abstract
Keywords:
Introduction
Methods
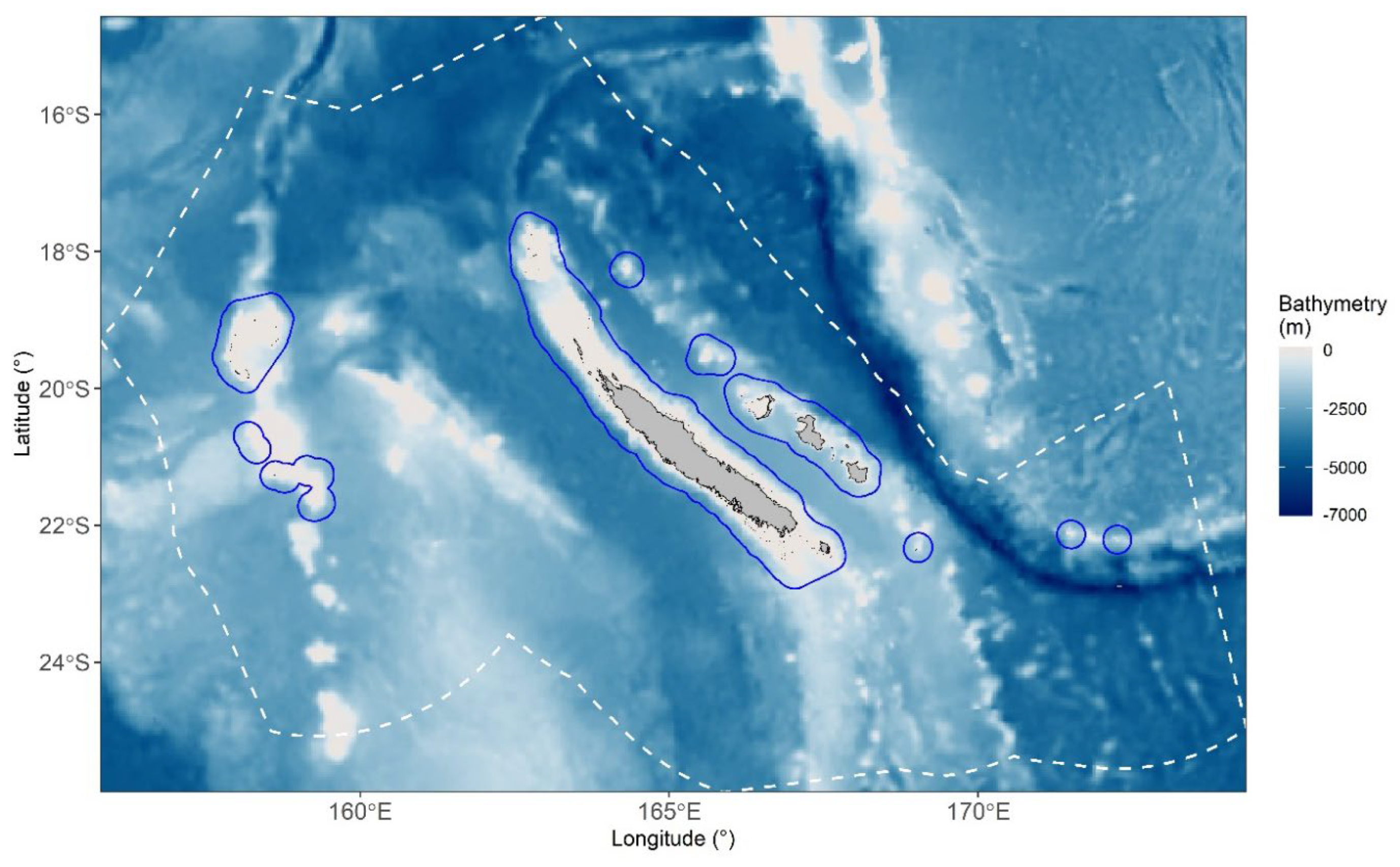
Study Area
Selection of Candidate Species and Probability of Presence in NC
- -
- Absent: no actual observation et no theoretical presence
- -
- Unlikely: species without theoretical presence and with fewer than 10 confirmed observations in total or with theoretical presence and without actual observation.
- -
- Likely: species with at least one confirmed observation in total and theoretical presence
- -
- Present: species with theoretical presence and at least one confirmed observation in each database, with a total of more than 10 confirmed observations in total.
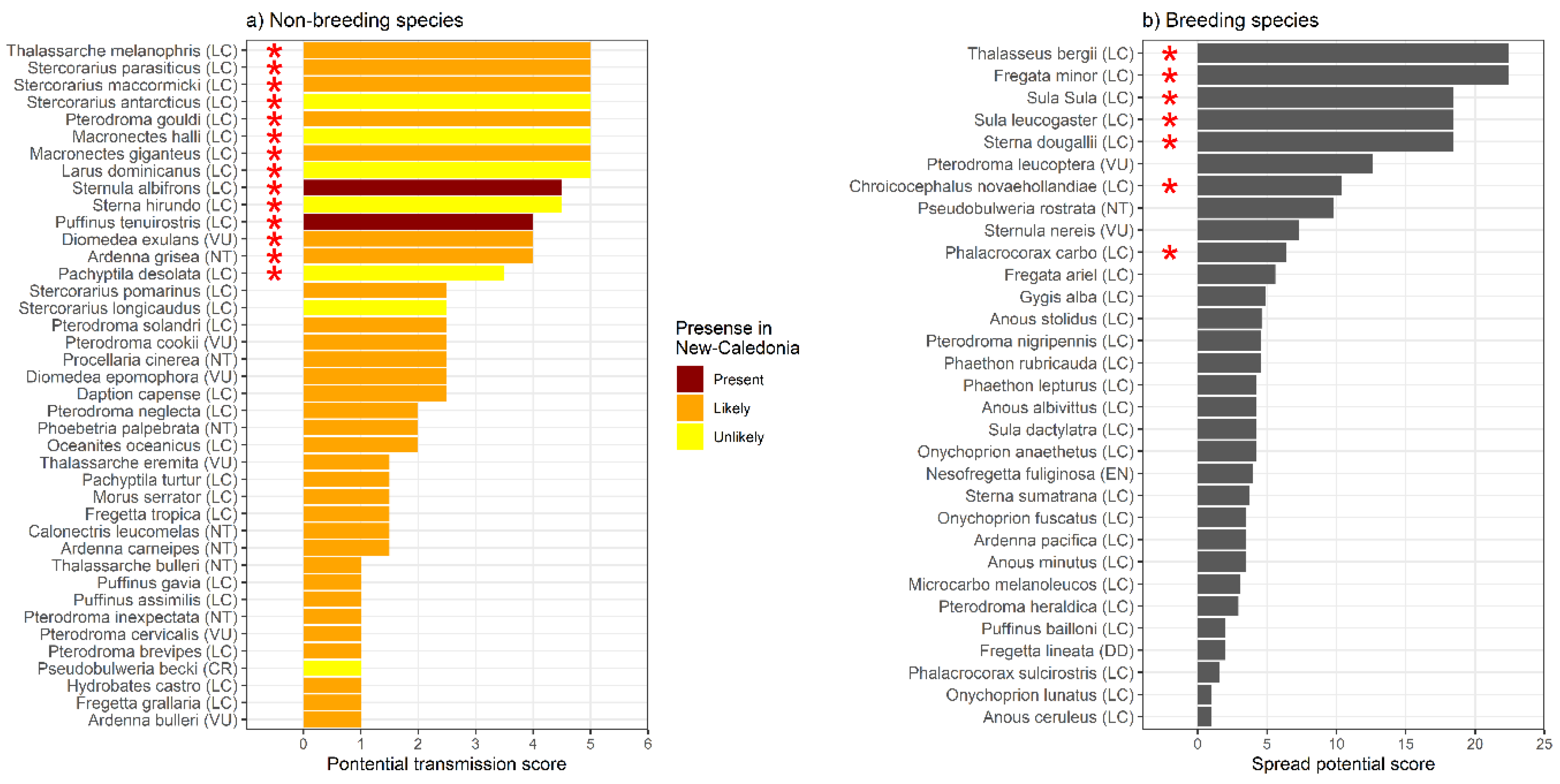
Transmission Potential Score Via Migratory Seabirds and Establishment of HPAIV Within neo-Caledonian Colonies
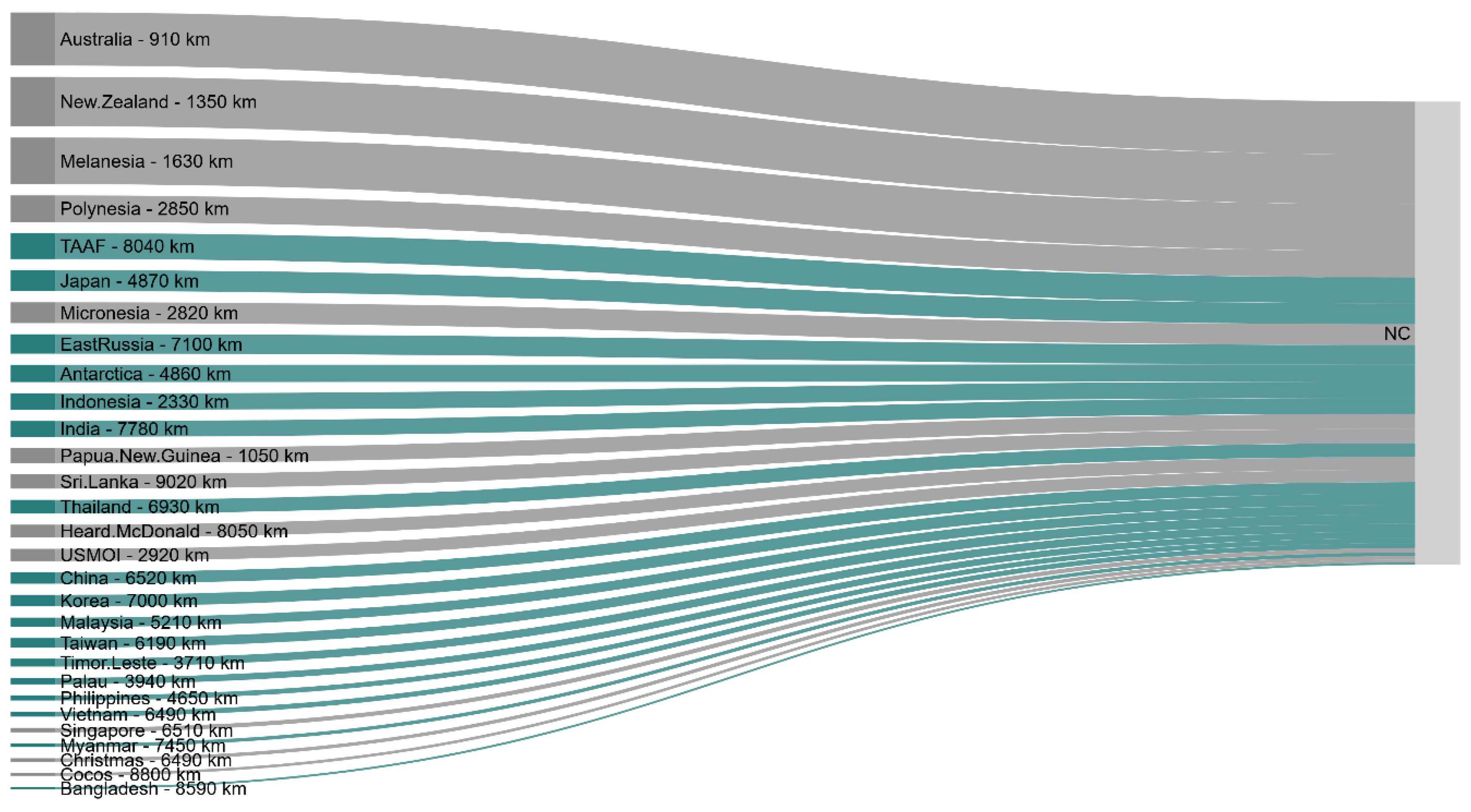
Network of Interactions Between Neighbouring Countries and New Caledonia
- -
- Breeding or resident: 3
- -
- Present during the non-breeding period: 2
- -
- Vagrant, observed, or uncertain origin/presence: 1
Results
Discussion
Which Species May Be Involved in HPAIV Emergence and Spread in New Caledonia?
New Caledonia Still Spared from HPAIV?
Limitations et Perspectives
Data Availability Statement
Acknowledgments
Appendix I
| Score | Justifications | |
|---|---|---|
|
H5N1 (Breeding and non breeding species) |
Affected: 4 Not affected: 1 |
Our hypothesis: A species that has already been affected, regardless of the population concerned, is more likely to introduce H5N1 into NC. For breeding species based on Pearce-Higgins et al.’s framework: Weight = 2 |
|
Diet (Breeding and non breeding species) |
Scavenger / predator: 4 Kleptoparasitism: 3 Ship-following / Surface feeder: 2 Diver: 1 |
These scores are based on potential interactions among individuals. Scavengers and predators feeding directly on other species are the most likely to get infected and spread H5N1, followed by kleptoparasitic species, for which there is a high risk of fluid exchange. Ship-following species are also at risk due to aggregations around fishing vessels and surface-feeding species may engage in competition for food, whereas diving species are generally less likely to come into contact with infected individuals For breeding species based on Pearce-Higgins et al.’s framework: Weight = 2 |
|
Gregariousness (maximum number of individuals per colony in NC) (just for breeding species) |
Occurs in large (1000’s individuals) breeding aggregations: 4 Occurs in moderate (100’s individuals) breeding aggregations: 3 Occurs in small (10’s individuals) breeding aggregations: 2 small (<10) family groups: 1 |
Based on Pearce-Higgins et al.’s framework Weight = 10 |
|
Population size in NC (maximum number of individuals) (just for breeding species) |
Very rare (<1,000 individuals): 4 Rare (1,000–10,000 individuals): 3 Common (10,000–100,000 individuals): 2 Abundant (>100,000 individuals): 1 |
Based on Pearce-Higgins et al.’s framework Weight = 1 |
|
Global conservation status according to the IUCN (just for breeding species) |
Threatened (vulnerable or higher): 4 Near threatened: 3 NA: 2 Least Concern: 1 |
Based on Pearce-Higgins et al.’s framework Weight = 4 |
References
- Airey, M.; Short, K.R. High pathogenicity avian influenza in Australia and beyond: could avian influenza cause the next human pandemic? Microbiol. Aust. 2024, 45, 155–158. [Google Scholar] [CrossRef]
- Alexander, D.; Brown, I. History of highly pathogenic avian influenza. Rev Sci Tech 2009, 28, 19–38. [Google Scholar] [CrossRef]
- Banyard, A.C.; Bennison, A.; Byrne, A.M.P.; Reid, S.M.; Lynton-Jenkins, J.G.; Mollett, B.; De Silva, D.; Peers-Dent, J.; Finlayson, K.; Hall, R.; Blockley, F.; Blyth, M.; Falchieri, M.; Fowler, Z.; Fitzcharles, E.M.; Brown, I.H.; James, J. Detection and spread of high pathogenicity avian influenza virus H5N1 in the Antarctic Region. Nat. Commun. 2024, 15, 7433. [Google Scholar] [CrossRef]
- Barré, N.; Hébert, O.; Aublin, R.; Spaggiari, J.; Chartendrault, V.; Baillon, N. Troisième complément à la liste des oiseaux de Nouvelle-Calédonie. Alauda Dijon 2009, 77, 287–302. [Google Scholar]
- Barré, N.; Villard, P.; Manceau, N.; Monimeau, L.; Ménard, C. Les oiseaux de l’archipel des Loyauté (Nouvelle-Calédonie): Inventaire et éléments d’écologie et de biogéographie. Rev. Décologie 2006, 61, 175–194. [Google Scholar] [CrossRef]
- Bellido-Martín, B.; Rijnink, W.F.; Iervolino, M.; Kuiken, T.; Richard, M.; Fouchier, R.A.M. Evolution, spread and impact of highly pathogenic H5 avian influenza A viruses. Nat. Rev. Microbiol. 2025. [Google Scholar] [CrossRef]
- Bennison, A.; Adlard, S.; Banyard, A.C.; Blockley, F.; Blyth, M.; Browne, E.; Day, G.; Dunn, M.J.; Falchieri, M.; Fitzcharles, E.; Forcada, J.; Forster Davidson, J.; Fox, A.; Hall, R.; Holmes, E.; Hughes, K.; James, J.; Lynton-Jenkins, J.; Marshall, S.; McKenzie, D.; Morley, S.A.; Reid, S.M.; Stubbs, I.; Ratcliffe, N.; Phillips, R.A. A case study of highly pathogenic avian influenza (HPAI) H5N1 at Bird Island, South Georgia: the first documented outbreak in the subantarctic region. Bird Study 2024, 1–12. [Google Scholar] [CrossRef]
- Bernard, A.; Rodrigues, A.S.L.; Cazalis, V.; Grémillet, D. Toward a global strategy for seabird tracking. Conserv. Lett. 2021, 14, e12804. [Google Scholar] [CrossRef]
- Bismuth, D.; Borsa, P. Bilan de l’avifaune des atolls Chesterfield-Bampton et d’Entrecasteaux (Nouvelle-Calédonie). Ornithomedia 15 2022. [Google Scholar]
- Borsa, P.; Philippe, A.; Le Bouteiller, A. Diversité et abondance des oiseaux marins des atolls d’Entrecasteaux (parc naturel de la mer de Corail) estimées à partir de missions ponctuelles effectuées ces deux dernières décennie; Institut de Recherche pour le Développement: Nouméa, Nouvelle-Calédonie, 2021. [Google Scholar]
- Brown, J.D.; Stallknecht, D.E.; Beck, J.R.; Suarez, D.L.; Swayne, D.E. Susceptibility of North American ducks and gulls to H5N1 highly pathogenic avian influenza viruses. Emerg. Infect. Dis. 2006, 12, 1663. [Google Scholar] [CrossRef] [PubMed]
- Clessin, A.; Briand, F.-X.; Tornos, J.; Lejeune, M.; De Pasquale, C.; Fischer, R.; Souchaud, F.; Hirchaud, E.; Hong, S.L.; Bralet, T.; Guinet, C.; McMahon, C.R.; Grasland, B.; Baele, G.; Boulinier, T. Circumpolar spread of avian influenza H5N1 to southern Indian Ocean islands. Nat. Commun. 2025, 16, 8463. [Google Scholar] [CrossRef]
- Couty, M.; Guinat, C.; Fornasiero, D.; Briand, F.-X.; Henry, P.-Y.; Grasland, B.; Palumbo, L.; Le Loc’h, G. The role of wild birds in the global highly pathogenic avian influenza H5 panzootic, 2020–2023. Npj Biodivers. 2026, 5, 1. [Google Scholar] [CrossRef]
- Dalloyau, S. Atlas des oiseaux marins nicheurs des territoires français d’Outre-Mer. In Biotope. ed; Parthénope, 2024. [Google Scholar]
- Duriez, O.; Sassi, Y.; Le Gall-Ladevèze, C.; Giraud, L.; Straughan, R.; Dauverné, L.; Terras, A.; Boulinier, T.; Choquet, R.; Van De Wiele, A.; Hirschinger, J.; Guérin, J.-L.; Le Loc’h, G. Highly pathogenic avian influenza affects vultures’ movements and breeding output. Curr. Biol. 2023, 33, 3766–3774.e3. [Google Scholar] [CrossRef]
- Fernández-Palacios, J.M.; Kreft, H.; Irl, S.D.H.; Norder, S.; Ah-Peng, C.; Borges, P.A.V.; Burns, K.C.; de Nascimento, L.; Meyer, J.-Y.; Montes, E.; Drake, D.R. Scientists’ warning—The outstanding biodiversity of islands is in peril. Glob. Ecol. Conserv. 2021, 31, e01847. [Google Scholar] [CrossRef] [PubMed]
- Gaskin, C.P.; Lukies, K.A.; Whitehead, E.A.; Raine, A.F. Pacific seabird survey and monitoring manual: tools to support seabird conservation across ecosystems in Oceania; Secretariat of the Pacific Regional Environment Programme (SPREP): Apia, Samoa, 2025. [Google Scholar]
- Gorta, S.B.Z.; Berryman, A.J.; Kingsford, R.T.; Klaassen, M.; Clarke, R.H. Kleptoparasitism in seabirds—A potential pathway for global avian influenza virus spread. Conserv. Lett. 2024, 17, e13052. [Google Scholar] [CrossRef]
- Gurney, F. Western Pacific Odyssey tour report 2023. Bird Quest tours. 2023. [Google Scholar]
- Hayakawa, M.; Suzuki-Matsubara, M.; Matsubara, K.; Kanazawa, S.; Fujii, T.; Kitamura, W.; Murofushi, R.A.; Moriyama, A. Population genetic structure of Little Tern (Sternula albifrons) in Japan inferred from nucleotide sequence diversities of the mitochondrial DNA control region. Ornithol. Sci. 2022, 21, 155–163. [Google Scholar] [CrossRef]
- Ismar, S.M.; Phillips, R.A.; Rayner, M.J.; Hauber, M.E. Geolocation tracking of the annual migration of adult Australasian Gannets (Morus serrator) breeding in New Zealand. Wilson J. Ornithol. 2011, 123, 121–125. [Google Scholar] [CrossRef]
- Lane, J.V.; Jeglinski, J.W.E.; Avery-Gomm, S.; Ballstaedt, E.; Banyard, A.C.; Barychka, T.; Brown, I.H.; Brugger, B.; Burt, T.V.; Careen, N.; Castenschiold, J.H.F.; Christensen-Dalsgaard, S.; Clifford, S.; Collins, S.M.; Cunningham, E.; Danielsen, J.; Daunt, F.; D’entremont, K.J.N.; Doiron, P.; Duffy, S.; English, M.D.; Falchieri, M.; Giacinti, J.; Gjerset, B.; Granstad, S.; Grémillet, D.; Guillemette, M.; Hallgrímsson, G.T.; Hamer, K.C.; Hammer, S.; Harrison, K.; Hart, J.D.; Hatsell, C.; Humpidge, R.; James, J.; Jenkinson, A.; Jessopp, M.; Jones, M.E.B.; Lair, S.; Lewis, T.; Malinowska, A.A.; McCluskie, A.; McPhail, G.; Moe, B.; Montevecchi, W.A.; Morgan, G.; Nichol, C.; Nisbet, C.; Olsen, B.; Provencher, J.; Provost, P.; Purdie, A.; Rail, J.-F.; Robertson, G.; Seyer, Y.; Sheddan, M.; Soos, C.; Stephens, N.; Strøm, H.; Svansson, V.; Tierney, T.D.; Tyler, G.; Wade, T.; Wanless, S.; Ward, C.R.E.; Wilhelm, S.I.; Wischnewski, S.; Wright, L.J.; Zonfrillo, B.; Matthiopoulos, J.; Votier, S.C. High pathogenicity avian influenza (H5N. 2024. [Google Scholar]
- Lebarbenchon, C.; Boucher, S.; Feare, C.; Dietrich, M.; Larose, C.; Humeau, L.; Le Corre, M.; Jaeger, A. Migratory patterns of two major influenza virus host species on tropical islands. R. Soc. Open Sci. 2023, 10, 230600. [Google Scholar] [CrossRef] [PubMed]
- Lisovski, S.; Günther, A.; Dewar, M.; Ainley, D.; Aldunate, F.; Arce, R.; Ballard, G.; Bauer, S.; Belliure, J.; Banyard, A.C. Unexpected delayed incursion of highly pathogenic avian influenza H5N1 (clade 2.3. 4.4 b) into the antarctic region. Influenza Other Respir. Viruses 2024, 18, e70010. [Google Scholar] [CrossRef]
- Menkès, C.E.; Allain, V.; Rodier, M.; Gallois, F.; Lebourges-Dhaussy, A.; Hunt, B.P.; Smeti, H.; Pagano, M.; Josse, E.; Daroux, A. Seasonal oceanography from physics to micronekton in the south-west Pacific. Deep Sea Res. Part II Top. Stud. Oceanogr. 2015, 113, 125–144. [Google Scholar] [CrossRef]
- Morten, J.M.; Carneiro, A.P.B.; Beal, M.; Bonnet-Lebrun, A.-S.; Dias, M.P.; Rouyer, M.-M.; Harrison, A.-L.; González-Solís, J.; Jones, V.R.; Garcia Alonso, V.A.; Antolos, M.; Arata, J.A.; Barbraud, C.; Bell, E.A.; Bell, M.; Bose, S.; Broni, S.; de L Brooke, M.; Butchart, S.H.M.; Carlile, N.; Catry, P.; Catry, T.; Charteris, M.; Cherel, Y.; Clark, B.L.; Clay, T.A.; Cole, N.C.; Conners, M.G.; Debski, I.; Delord, K.; Egevang, C.; Elliot, G.; Esefeld, J.; Facer, C.; Fayet, A.L.; Fijn, R.C.; Fischer, J.H.; Franklin, K.A.; Gilg, O.; Gill, J.A.; Granadeiro, J.P.; Guilford, T.; Handley, J.M.; Hanssen, S.A.; Hawkes, L.A.; Hedd, A.; Jaeger, A.; Jones, C.G.; Jones, C.W.; Kopp, M.; Krietsch, J.; Landers, T.J.; Lang, J.; Le Corre, M.; Mallory, M.L.; Masello, J.F.; Maxwell, S.M.; Medrano, F.; Militão, T.; Millar, C.D.; Moe, B.; Montevecchi, W.A.; Navarro-Herrero, L.; Neves, V.C.; Nicholls, D.G.; Nicoll, M.A.C.; Norris, K.; O’Dwyer, T.W.; Parker, G.C.; Peter, H.-U.; Phillips, R.
- Nguyen, T.-Q.; Hutter, C.R.; Markin, A.; Thomas, M.; Lantz, K.; Killian, M.L.; Janzen, G.M.; Vijendran, S.; Wagle, S.; Inderski, B.; Magstadt, D.R.; Li, G.; Diel, D.G.; Frye, E.A.; Dimitrov, K.M.; Swinford, A.K.; Thompson, A.C.; Snekvik, K.R.; Suarez, D.L.; Lakin, S.M.; Schwabenlander, S.; Ahola, S.C.; Johnson, K.R.; Baker, A.L.; Robbe-Austerman, S.; Torchetti, M.K.; Anderson, T.K. Emergence and interstate spread of highly pathogenic avian influenza A(H5N1) in dairy cattle in the United States. Science 2025, 388, eadq0900. [Google Scholar] [CrossRef] [PubMed]
- Peacock, T.P.; Moncla, L.; Dudas, G.; VanInsberghe, D.; Sukhova, K.; Lloyd-Smith, J.O.; Worobey, M.; Lowen, A.C.; Nelson, M.I. The global H5N1 influenza panzootic in mammals. Nature 2025, 637, 304–313. [Google Scholar] [CrossRef] [PubMed]
- Pearce-Higgins, J.W.; Clewley, G.; Bolton, M.; Banyard, A.C.; Falchieri, M.; Lindley, P.; Atkinson, P.W. Assessing the vulnerability of wild bird populations to high pathogenicity avian influenza. Bird Study 2025, 72, 5–19. [Google Scholar] [CrossRef]
- Péron, C.; Grémillet, D. Tracking through life stages: adult, immature and juvenile autumn migration in a long-lived seabird. PLOS ONE 2013, 8, e72713. [Google Scholar] [CrossRef]
- Plaza, P.; Santangeli, A.; Cancellario, T.; Lambertucci, S. Potential Arrival Pathway for Highly Pathogenic Avian Influenza H5N1 to Oceania. Influenza Other Respir. Viruses 2024, 18, e70055. [Google Scholar] [CrossRef]
- Ramírez-Garofalo, J.R.; Lockwood, J.L. Animal vagrancy and the spread of pathogens. Trends Ecol. Evol. 2025, 40, 119–121. [Google Scholar] [CrossRef] [PubMed]
- Rancurel, P. Notes ornithologiques: Liste préliminaire des oiseaux de mer iles et ilots voisins de la Nouvelle Calédonie. Cah. Off. Recherce Sci. Tech. O.-m. Ser. Oceanogr. 1976, 14, 163–168. [Google Scholar]
- Rayner, M.; Hauber, M.; Clout, M.; Seldon, D.; Van Dijken, S.; Bury, S.; Phillips, R. Foraging ecology of the Cook’s petrel Pterodroma cookii during the austral breeding season: a comparison of its two populations. Mar. Ecol. Prog. Ser. 2008, 370, 271–284. [Google Scholar] [CrossRef]
- Rayner, M.J.; Gaskin, C.P.; Taylor, G.A. Seasonal movements, foraging activity, and breeding biology of black-winged petrel (Pterodroma nigripennis) from three New Zealand colonies. Notornis 2023, 70, 111–123. [Google Scholar] [CrossRef]
- Ryding, S.; Ross, T.A.; Klaassen, M. Predicting high pathogenicity avian influenza H5N1 susceptibility in wild birds, with1 special reference to Australia. 2025. [Google Scholar]
- Schreiber, E.A.; Burger, J. Biology of Marine Birds; CRC Press, 2001. [Google Scholar] [CrossRef]
- Soanes, L.M.; Green, J.A.; Bolton, M.; Milligan, G.; Mukhida, F.; Halsey, L.G. Linking foraging and breeding strategies in tropical seabirds. J. Avian Biol. 2021, 52. [Google Scholar] [CrossRef]
- South, A.; Schramm, M.; Massicotte, P. rnaturalearthdata: World Vector Map Data from Natural Earth Used in “rnaturalearth.”. 2024. [Google Scholar]
- Spaggiari, J.; Barré, N.; Baudat-Franceschi, J.; Borsa, P.; Colin, F. New Caledonian seabirds, in: Compendium of Marine Species from New Caledonia 2007a, 415–428.
- Spaggiari, J.; Chartendrault, V.; Barré, N. Zones importantes pour la conservation des oiseaux de Nouvelle-Calédonie; Société calédonienne d’ornithologie—SCO: Nouméa, Nouvelle-Calédonie, 2007b. [Google Scholar]
- Stanislawek, W.L.; Tana, T.; Rawdon, T.G.; Cork, S.C.; Chen, K.; Fatoyinbo, H.; Cogger, N.; Webby, R.J.; Webster, R.G.; Joyce, M.; Tuboltsev, M.A.; Orr, D.; Ohneiser, S.; Watts, J.; Riegen, A.C.; McDougall, M.; Klee, D.; O’Keefe, J.S. Avian influenza viruses in New Zealand wild birds, with an emphasis on subtypes H5 and H7: Their distinctive epidemiology and genomic properties. PLOS ONE 2024, 19, e0303756. [Google Scholar] [CrossRef] [PubMed]
- Talmon, I.; Pekarsky, S.; Bartan, Y.; Thie, N.; Getz, W.M.; Kamath, P.L.; Bowie, R.C.K.; Nathan, R. Using wild-animal tracking for detecting and managing disease outbreaks. Trends Ecol. Evol. 2025, 40, 760–771. [Google Scholar] [CrossRef]
- Weimerskirch, H.; Tarroux, A.; Chastel, O.; Delord, K.; Cherel, Y.; Descamps, S. Population-specific wintering distributions of adult south polar skuas over three oceans. Mar. Ecol. Prog. Ser. 2015, 538, 229–237. [Google Scholar] [CrossRef]
- Wille, M.; Atkinson, R.; Barr, I.G.; Burgoyne, C.; Bond, A.L.; Boyle, D.; Christie, M.; Dewar, M.; Douglas, T.; Fitzwater, T.; Hassell, C.; Jessop, R.; Klaassen, H.; Lavers, J.L.; Leung, K.K.-S.; Ringma, J.; Sutherland, D.R.; Klaassen, M. Long-Distance Avian Migrants Fail to Bring 2.3.4.4b HPAI H5N1 Into Australia for a Second Year in a Row. Influenza Other Respir. Viruses 2024, 18, e13281. [Google Scholar] [CrossRef]
- Wille, M.; Barr, I.G. Resurgence of avian influenza virus. Science 2022, 376, 459–460. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




