Submitted:
17 April 2026
Posted:
21 April 2026
You are already at the latest version
Abstract
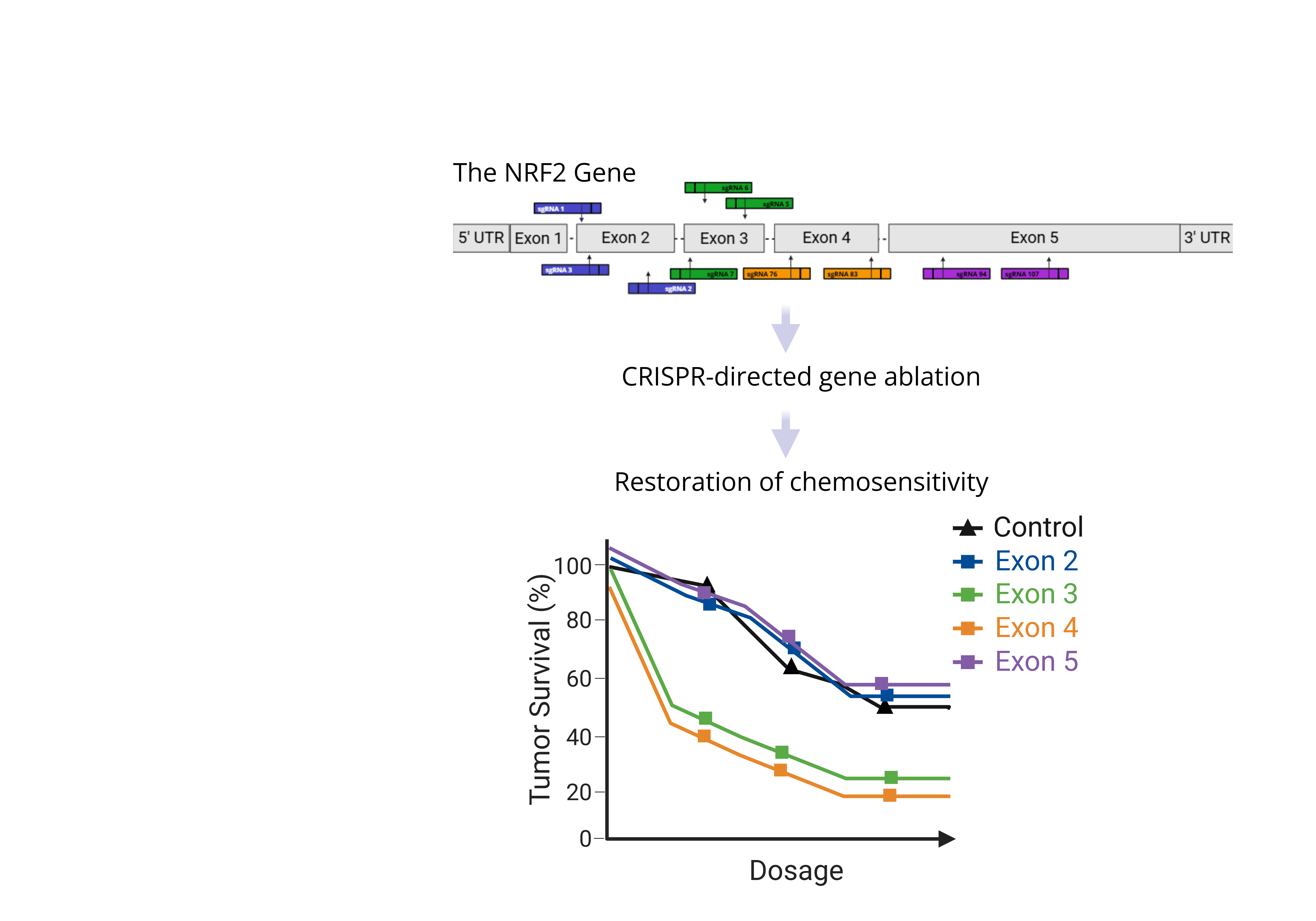
Keywords:
1. Introduction
2. Materials and Methods
2.1. Cell Line and Culture Conditions
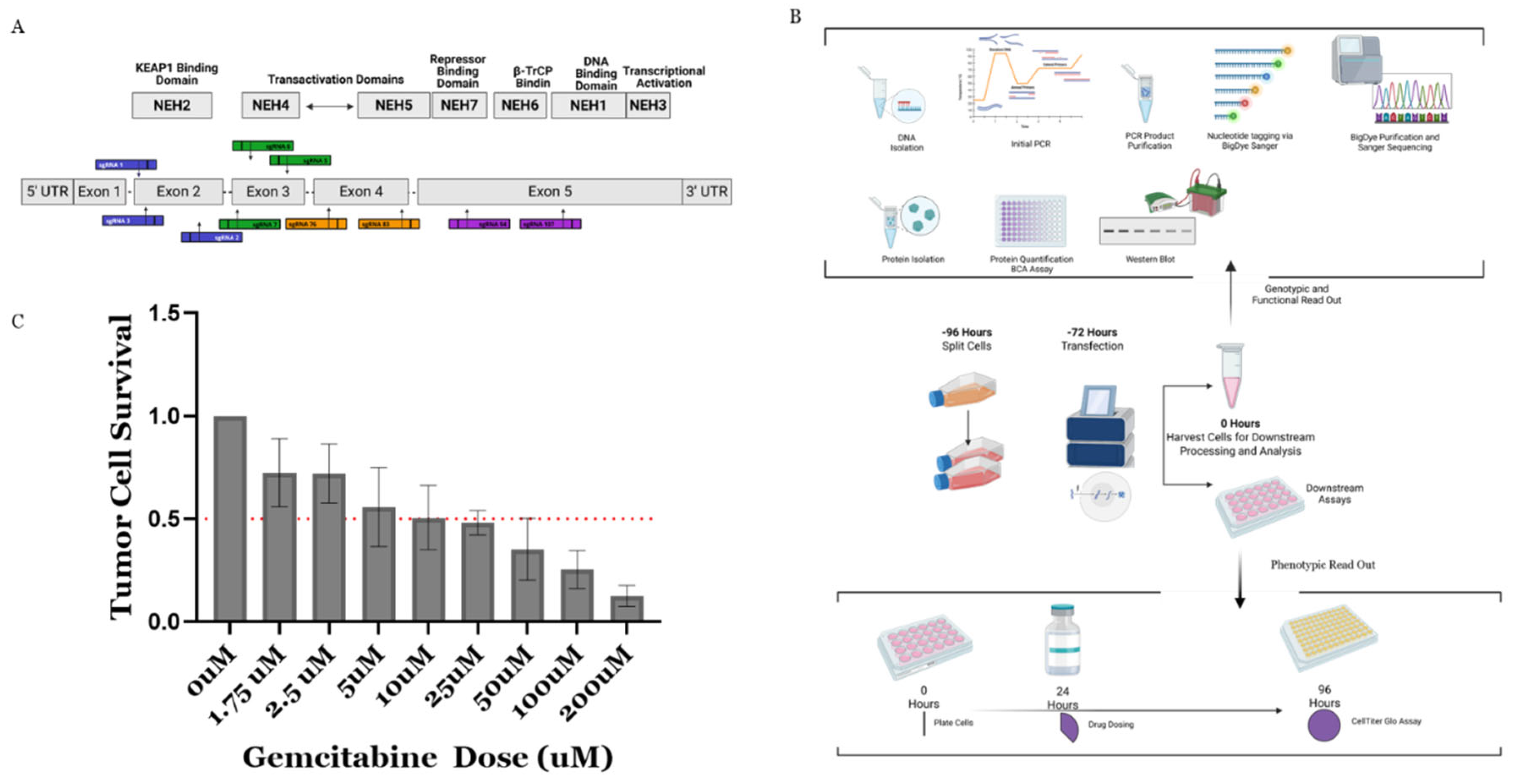
2.2. CRISPR/Cas9 Design
2.3. Nucleofection Transfection
2.4. Primer Design
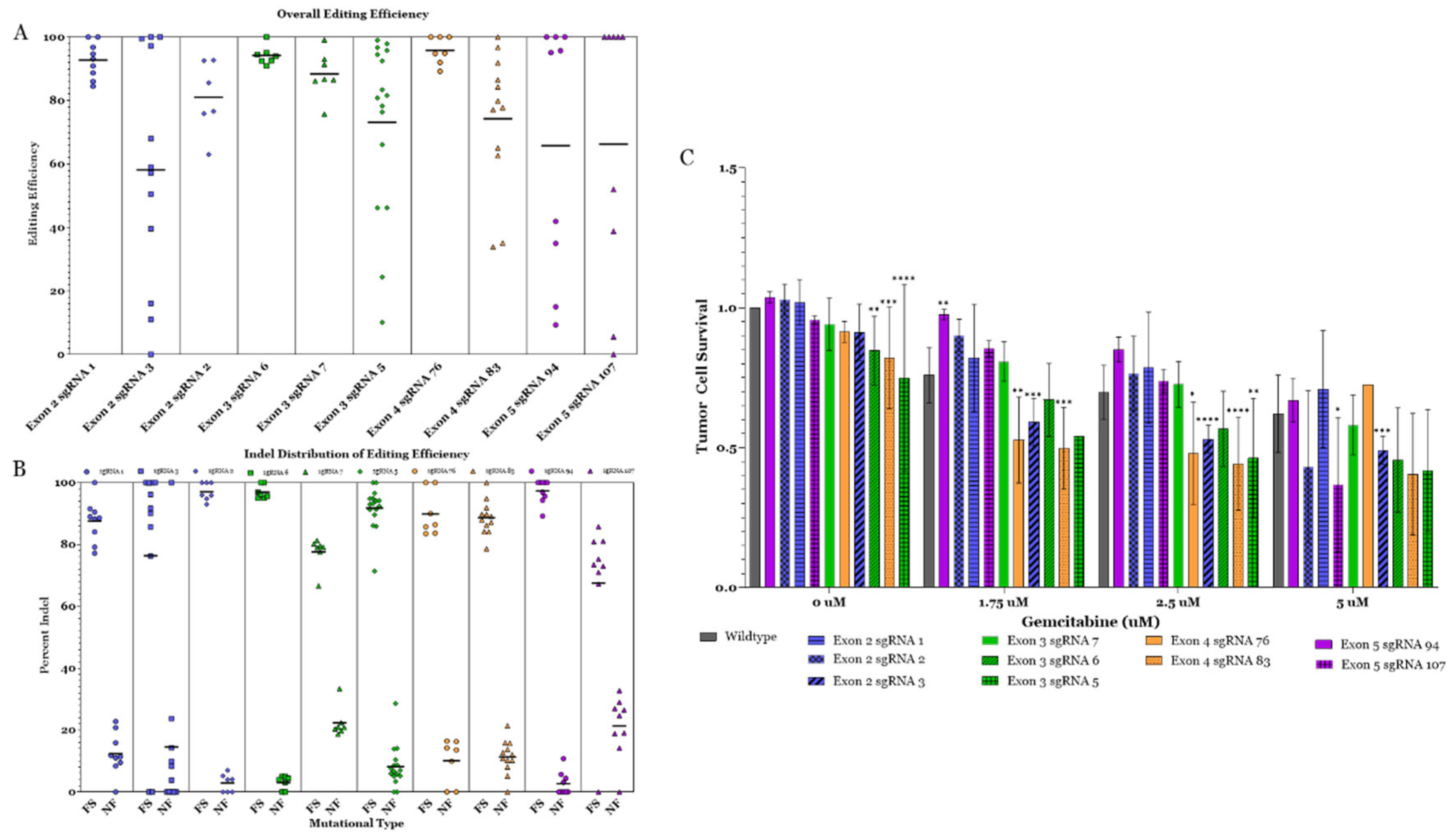
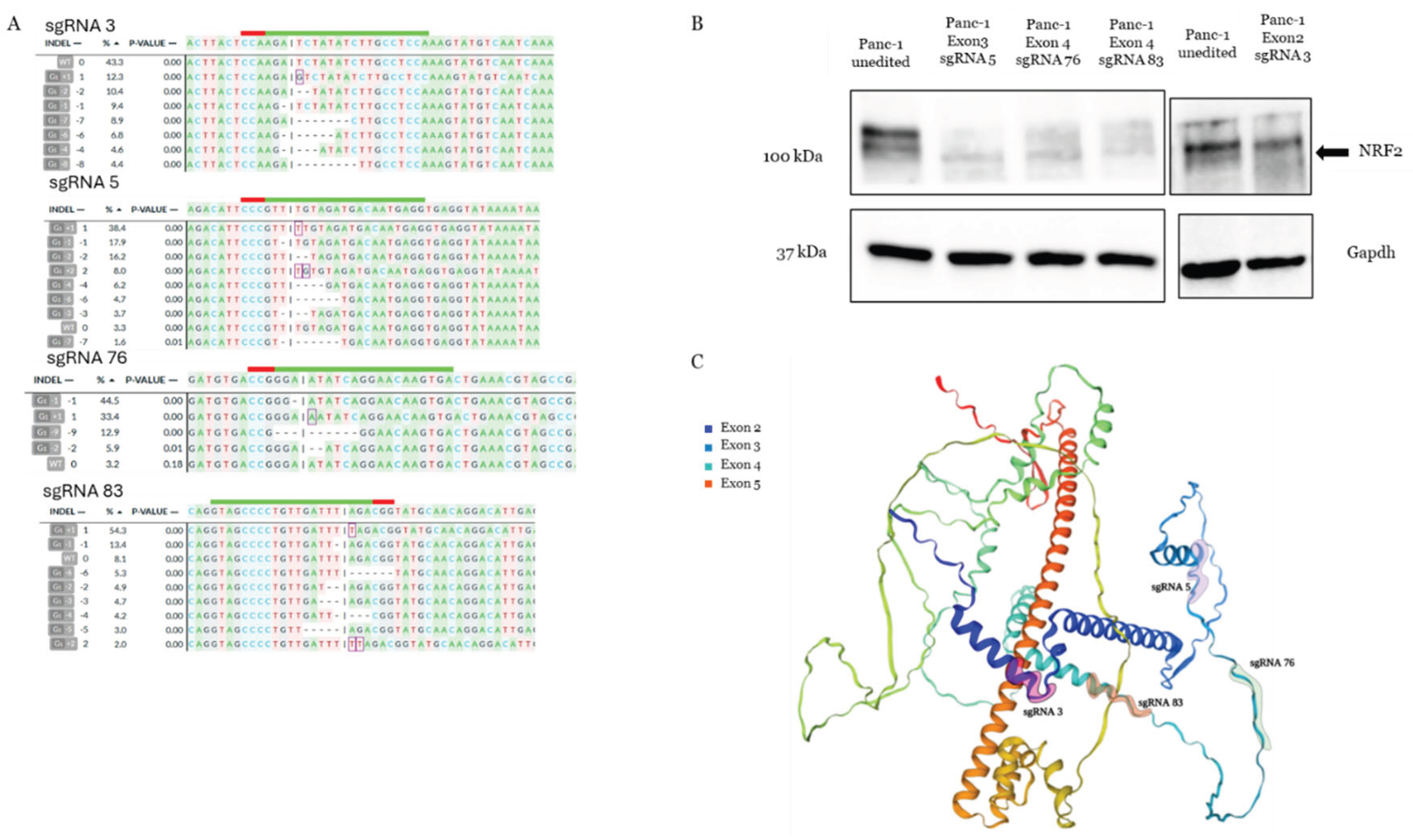
2.5. Sanger Sequencing & Gene Editing Analysis
2.6. Cell Viability
2.7. Western Blot Protein Analysis
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Abbreviations
| CRISPR | clustered regularly interspaced palindromic repeats |
| PDAC | Pancreatic ductal adenocarcinoma |
| sgRNA | Synthetic single guide RNAs |
| INDEL | random insertions and deletions |
Appendix A
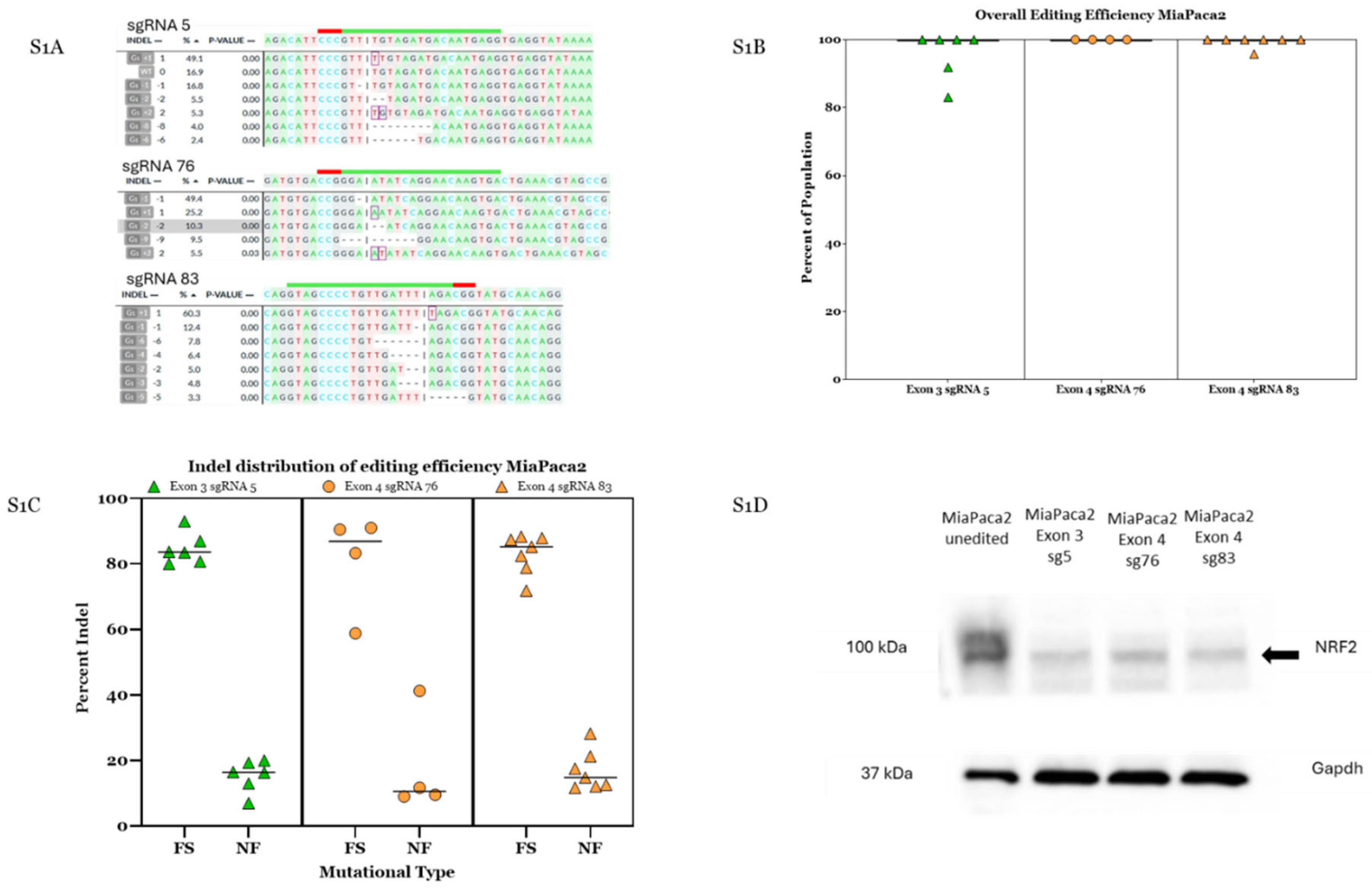

References
- El Jellas, K.; Johansson, B.B.; Fjeld, K.; Antonopoulos, A.; Immervoll, H.; Choi, M.H.; Hoem, D.; Lowe, M.E.; Lombardo, D.; Njølstad, P.R.; et al. The Mucinous Domain of Pancreatic Carboxyl-Ester Lipase (CEL) Contains Core 1/Core 2 O-Glycans That Can Be Modified by ABO Blood Group Determinants. Journal of Biological Chemistry 2018, 293, 19476–19491. [Google Scholar] [CrossRef] [PubMed]
- Rahib, L.; Smith, B.D.; Aizenberg, R.; Rosenzweig, A.B.; Fleshman, J.M.; Matrisian, L.M. Projecting Cancer Incidence and Deaths to 2030: The Unexpected Burden of Thyroid, Liver, and Pancreas Cancers in the United States. Cancer Res. 2014, 74, 2913–2921. [Google Scholar] [CrossRef]
- Park, W.; Chawla, A.; O’Reilly, E.M. Pancreatic Cancer. JAMA 2021, 326, 851. [Google Scholar] [CrossRef]
- American Cancer Society Key Statistics for Pancreatic Cancer.
- National Cancer Institute Cancer Stat Facts: Pancreatic Cancer.
- Blackford, A.L.; Canto, M.I.; Klein, A.P.; Hruban, R.H.; Goggins, M. Recent Trends in the Incidence and Survival of Stage 1A Pancreatic Cancer: A Surveillance, Epidemiology, and End Results Analysis. JNCI: Journal of the National Cancer Institute 2020, 112, 1162–1169. [Google Scholar] [CrossRef]
- Vaziri-Gohar, A.; Hue, J.J.; Abbas, A.; Graor, H.J.; Hajihassani, O.; Zarei, M.; Titomihelakis, G.; Feczko, J.; Rathore, M.; Chelstowska, S.; et al. Increased Glucose Availability Sensitizes Pancreatic Cancer to Chemotherapy. Nat. Commun. 2023, 14, 3823. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Liu, X.; Jin, S.; Chen, Y.; Guo, R. Ferroptosis in Cancer Therapy: A Novel Approach to Reversing Drug Resistance. Mol. Cancer 2022, 21, 47. [Google Scholar] [CrossRef] [PubMed]
- Miess, H.; Dankworth, B.; Gouw, A.M.; Rosenfeldt, M.; Schmitz, W.; Jiang, M.; Saunders, B.; Howell, M.; Downward, J.; Felsher, D.W.; et al. The Glutathione Redox System Is Essential to Prevent Ferroptosis Caused by Impaired Lipid Metabolism in Clear Cell Renal Cell Carcinoma. Oncogene 2018, 37, 5435–5450. [Google Scholar] [CrossRef]
- Waissi, W.; Amé, J.-C.; Mura, C.; Noël, G.; Burckel, H. Gemcitabine-Based Chemoradiotherapy Enhanced by a PARP Inhibitor in Pancreatic Cancer Cell Lines. Int. J. Mol. Sci. 2021, 22, 6825. [Google Scholar] [CrossRef]
- Du, J.; Gu, J.; Li, J. Mechanisms of Drug Resistance of Pancreatic Ductal Adenocarcinoma at Different Levels. Biosci. Rep. 2020, 40. [Google Scholar] [CrossRef]
- Liu, K.; Geng, Y.; Wang, L.; Xu, H.; Zou, M.; Li, Y.; Zhao, Z.; Chen, T.; Xu, F.; Sun, L.; et al. Systematic Exploration of the Underlying Mechanism of Gemcitabine Resistance in Pancreatic Adenocarcinoma. Mol. Oncol. 2022, 16, 3034–3051. [Google Scholar] [CrossRef]
- Stoop, T.F.; Oba, A.; Wu, Y.H.A.; Beaty, L.E.; Colborn, K.L.; Janssen, B. V.; Al-Musawi, M.H.; Franco, S.R.; Sugawara, T.; Franklin, O.; et al. Pathological Complete Response in Patients With Resected Pancreatic Adenocarcinoma After Preoperative Chemotherapy. JAMA Netw. Open 2024, 7, e2417625. [Google Scholar] [CrossRef]
- Dhir, M.; Malhotra, G.K.; Sohal, D.P.S.; Hein, N.A.; Smith, L.M.; O’Reilly, E.M.; Bahary, N.; Are, C. Neoadjuvant Treatment of Pancreatic Adenocarcinoma: A Systematic Review and Meta-Analysis of 5520 Patients. World J. Surg. Oncol. 2017, 15, 183. [Google Scholar] [CrossRef]
- Masuo, H.; Kubota, K.; Shimizu, A.; Notake, T.; Miyazaki, S.; Yoshizawa, T.; Sakai, H.; Hayashi, H.; Soejima, Y. Increased Mitochondria Are Responsible for the Acquisition of Gemcitabine Resistance in Pancreatic Cancer Cell Lines. Cancer Sci. 2023, 114, 4388–4400. [Google Scholar] [CrossRef]
- Siegel, R.L.; Kratzer, T.B.; Giaquinto, A.N.; Sung, H.; Jemal, A. Cancer Statistics, 2025. CA Cancer J. Clin. 2025, 75, 10–45. [Google Scholar] [CrossRef]
- Ferrara, L. Camptothecin Enhances the Frequency of Oligonucleotide-Directed Gene Repair in Mammalian Cells by Inducing DNA Damage and Activating Homologous Recombination. Nucleic Acids Res. 2004, 32, 5239–5248. [Google Scholar] [CrossRef] [PubMed]
- Issa, I.I.; Due, H.; Brøndum, R.F.; Veeravakaran, V.; Haraldsdóttir, H.; Sylvester, C.; Brogaard, A.; Dhanjal, S.; Schmierer, B.; Dybkær, K. CRISPR-Cas9 Knockout Screens Identify DNA Damage Response Pathways and BTK as Essential for Cisplatin Response in Diffuse Large B-Cell Lymphoma. Cancers (Basel). 2024, 16, 2437. [Google Scholar] [CrossRef]
- Engstrom, J.U.; Kmiec, E.B. DNA Replication, Cell Cycle Progression and the Targeted Gene Repair Reaction. Cell Cycle 2008, 7, 1402–1414. [Google Scholar] [CrossRef]
- Bennett, E.P.; Petersen, B.L.; Johansen, I.E.; Niu, Y.; Yang, Z.; Chamberlain, C.A.; Met, Ö; Wandall, H.H.; Frödin, M. INDEL Detection, the ‘Achilles Heel’ of Precise Genome Editing: A Survey of Methods for Accurate Profiling of Gene Editing Induced Indels. Nucleic Acids Res. 2020, 48, 11958–11981. [Google Scholar] [CrossRef]
- Kosicki, M.; Allen, F.; Steward, F.; Tomberg, K.; Pan, Y.; Bradley, A. Cas9-Induced Large Deletions and Small Indels Are Controlled in a Convergent Fashion. Nat. Commun. 2022, 13, 3422. [Google Scholar] [CrossRef] [PubMed]
- Sansbury, B.M.; Hewes, A.M.; Kmiec, E.B. Understanding the Diversity of Genetic Outcomes from CRISPR-Cas Generated Homology-Directed Repair. Commun. Biol. 2019, 2, 458. [Google Scholar] [CrossRef] [PubMed]
- Hsu, P.D.; Lander, E.S.; Zhang, F. Development and Applications of CRISPR-Cas9 for Genome Engineering. Cell 2014, 157, 1262–1278. [Google Scholar] [CrossRef]
- Li, T.; Yang, Y.; Qi, H.; Cui, W.; Zhang, L.; Fu, X.; He, X.; Liu, M.; Li, P.; Yu, T. CRISPR/Cas9 Therapeutics: Progress and Prospects. Signal Transduct. Target. Ther. 2023, 8, 36. [Google Scholar] [CrossRef]
- Ravichandran, M.; Maddalo, D. Applications of CRISPR-Cas9 for Advancing Precision Medicine in Oncology: From Target Discovery to Disease Modeling. Front. Genet. 2023, 14, 1273994. [Google Scholar] [CrossRef] [PubMed]
- Banas, K.H.; Bialk, P.A.; Rivera-Torres, N.; Owens, K.; Reiner, T.N.; Pisarcik, K.M.; Haas, N.; Gielda, E.; Khan, K.; Narra, K.P.; et al. Functional Characterization of Tumor-Specific CRISPR-Directed Gene Editing as a Combinatorial Therapy for the Treatment of Solid Tumors. Molecular Therapy Oncology 2025, 33, 201079. [Google Scholar] [CrossRef]
- Rivera-Torres, N.; Skelly, L.E.; Rogowskyj, J.A.; Aguilar, G.; Banas, K.; Bialk, P.; Kmiec, E.B. Target Choice and Exon Skipping Regulate CRISPR-Directed Gene Editing of NRF2 in Head/Neck and Esophageal Cancer Cells. Molecular Therapy Oncology 2026, 34, 201122. [Google Scholar] [CrossRef] [PubMed]
- Bialk, P.; Wang, Y.; Banas, K.; Kmiec, E.B. Functional Gene Knockout of NRF2 Increases Chemosensitivity of Human Lung Cancer A549 Cells In Vitro and in a Xenograft Mouse Model. Mol. Ther. Oncolytics 2018, 11, 75–89. [Google Scholar] [CrossRef]
- Banas, K.; Modarai, S.; Rivera-Torres, N.; Yoo, B.-C.; Bialk, P.A.; Barrett, C.; Batish, M.; Kmiec, E.B. Exon Skipping Induced by CRISPR-Directed Gene Editing Regulates the Response to Chemotherapy in Non-Small Cell Lung Carcinoma Cells. Gene Ther. 2022, 29, 357–367. [Google Scholar] [CrossRef]
- Wu, S.; Lu, H.; Bai, Y. Nrf2 in Cancers: A Double-Edged Sword. Cancer Med. 2019, 8, 2252–2267. [Google Scholar] [CrossRef]
- Lister, A.; Nedjadi, T.; Kitteringham, N.R.; Campbell, F.; Costello, E.; Lloyd, B.; Copple, I.M.; Williams, S.; Owen, A.; Neoptolemos, J.P.; et al. Nrf2 Is Overexpressed in Pancreatic Cancer: Implications for Cell Proliferation and Therapy. Mol. Cancer 2011, 10, 37. [Google Scholar] [CrossRef] [PubMed]
- Hong, Y. Bin; Kang, H.J.; Kwon, S.Y.; Kim, H.J.; Kwon, K.Y.; Cho, C.H.; Lee, J.-M.; Kallakury, B.V.S.; Bae, I. Nuclear Factor (Erythroid-Derived 2)-Like 2 Regulates Drug Resistance in Pancreatic Cancer Cells. Pancreas 2010, 39, 463–472. [Google Scholar] [CrossRef]
- He, F.; Ru, X.; Wen, T. NRF2, a Transcription Factor for Stress Response and Beyond. Int. J. Mol. Sci. 2020, 21, 4777. [Google Scholar] [CrossRef]
- Grossman, R.; Ram, Z. The Dark Side of Nrf2. World Neurosurg. 2013, 80, 284–286. [Google Scholar] [CrossRef]
- Cykowiak, M.; Krajka-Kuźniak, V. Role of Nrf2 in Pancreatic Cancer. Antioxidants 2021, 11, 98. [Google Scholar] [CrossRef]
- Mukhopadhyay, S.; Goswami, D.; Adiseshaiah, P.P.; Burgan, W.; Yi, M.; Guerin, T.M.; Kozlov, S. V.; Nissley, D. V.; McCormick, F. Undermining Glutaminolysis Bolsters Chemotherapy While NRF2 Promotes Chemoresistance in KRAS-Driven Pancreatic Cancers. Cancer Res. 2020, 80, 1630–1643. [Google Scholar] [CrossRef] [PubMed]
- Xiang, Y.; Ye, W.; Huang, C.; Yu, D.; Chen, H.; Deng, T.; Zhang, F.; Lou, B.; Zhang, J.; Shi, K.; et al. Brusatol Enhances the Chemotherapy Efficacy of Gemcitabine in Pancreatic Cancer via the Nrf2 Signalling Pathway. Oxid. Med. Cell. Longev. 2018, 2018. [Google Scholar] [CrossRef]
- Bae, E.S.; Byun, W.S.; Ock, C.W.; Kim, W.K.; Park, H.J.; Lee, S.K. Periplocin Exerts Antitumor Activity by Regulating Nrf2-Mediated Signaling Pathway in Gemcitabine-Resistant Pancreatic Cancer Cells. Biomedicine & Pharmacotherapy 2023, 157, 114039. [Google Scholar] [CrossRef]
- Hsu, T.; Wang, W.; Chen, A.; Chiu, C.; Liao, P.; Chen, H.; Su, C.; Shen, S.; Tsai, K.; Wang, T.; et al. Nrf2-mediated Adenylosuccinate Lyase Promotes Resistance to Gemcitabine in Pancreatic Ductal Adenocarcinoma Cells through Ferroptosis Escape. J. Cell. Physiol. 2024, 239. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Zhou, Y.; Yang, M.; Wang, K.; Liu, Y.; Zhang, M.; Yang, Y.; Jin, C.; Wang, R.; Hu, R. Digoxin Sensitizes Gemcitabine-Resistant Pancreatic Cancer Cells to Gemcitabine via Inhibiting Nrf2 Signaling Pathway. Redox Biol. 2019, 22, 101131. [Google Scholar] [CrossRef]
- Wang, X.-J.; Sun, Z.; Villeneuve, N.F.; Zhang, S.; Zhao, F.; Li, Y.; Chen, W.; Yi, X.; Zheng, W.; Wondrak, G.T.; et al. Nrf2 Enhances Resistance of Cancer Cells to Chemotherapeutic Drugs, the Dark Side of Nrf2. Carcinogenesis 2008, 29, 1235–1243. [Google Scholar] [CrossRef]
- Lieber, M.; Mazzetta, J.; Nelson-Rees, W.; Kaplan, M.; Todaro, G. Establishment of a Continuous Tumor-cell Line (PANC-1) from a Human Carcinoma of the Exocrine Pancreas. Int. J. Cancer 1975, 15, 741–747. [Google Scholar] [CrossRef]
- Fryer, R.A.; Barlett, B.; Galustian, C.; Dalgleish, A.G. Mechanisms Underlying Gemcitabine Resistance in Pancreatic Cancer and Sensitisation by the IMiDTM Lenalidomide. Anticancer Res. 2011, 31, 3747–3756. [Google Scholar] [PubMed]
- Bloh, K.; Kanchana, R.; Bialk, P.; Banas, K.; Zhang, Z.; Yoo, B.-C.; Kmiec, E.B. Deconvolution of Complex DNA Repair (DECODR): Establishing a Novel Deconvolution Algorithm for Comprehensive Analysis of CRISPR-Edited Sanger Sequencing Data. CRISPR J. 2021, 4, 120–131. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Zhang, J.; Liu, S.; Huang, Y.; Chen, B.; Wang, D. Nrf2 Knockdown by ShRNA Inhibits Tumor Growth and Increases Efficacy of Chemotherapy in Cervical Cancer. Cancer Chemother. Pharmacol. 2012, 69, 485–494. [Google Scholar] [CrossRef] [PubMed]
- Mancini, M.C.S.; Morelli, A.P.; Severino, M.B.; Pavan, I.C.B.; Zambalde, ÉP.; Góis, M.M.; da Silva, L.G.S.; Quintero-Ruiz, N.; Romeiro, C.F.; dos Santos, D.F.G.; et al. Knockout of NRF2 Triggers Prostate Cancer Cells Death through ROS Modulation and Sensitizes to Cisplatin. J. Cell. Biochem. 2022, 123, 2079–2092. [Google Scholar] [CrossRef]
- Prashanth, N.; Meghana, P.; Sandeep Kumar Jain, R.; Rajaput, Pooja S; Satyanarayan, N. D.; Raja Naika, H.; Kumaraswamy, H. M. Nicotine Promotes Epithelial to Mesenchymal Transition and Gemcitabine Resistance via HENT1/RRM1 Signalling in Pancreatic Cancer and Chemosensitizing Effects of Embelin-a Naturally Occurring Benzoquinone. Science of The Total Environment 2024, 914, 169727. [Google Scholar] [CrossRef]
- Waterhouse, A.; Bertoni, M.; Bienert, S.; Studer, G.; Tauriello, G.; Gumienny, R.; Heer, F.T.; de Beer, T.A.P.; Rempfer, C.; Bordoli, L.; et al. SWISS-MODEL: Homology Modelling of Protein Structures and Complexes. Nucleic Acids Res. 2018, 46, W296–W303. [Google Scholar] [CrossRef]
- Brüschweiler, S.; Fuchs, J.E.; Bader, G.; McConnell, D.B.; Konrat, R.; Mayer, M. A Step toward NRF2-DNA Interaction Inhibitors by Fragment-Based NMR Methods. ChemMedChem 2021, 16, 3576–3587. [Google Scholar] [CrossRef]
- Zimta, A.-A.; Cenariu, D.; Irimie, A.; Magdo, L.; Nabavi, S.M.; Atanasov, A.G.; Berindan-Neagoe, I. The Role of Nrf2 Activity in Cancer Development and Progression. Cancers (Basel). 2019, 11, 1755. [Google Scholar] [CrossRef]
- Yagishita, Y.; Fukutomi, T.; Sugawara, A.; Kawamura, H.; Takahashi, T.; Pi, J.; Uruno, A.; Yamamoto, M. Nrf2 Protects Pancreatic β-Cells From Oxidative and Nitrosative Stress in Diabetic Model Mice. Diabetes 2014, 63, 605–618. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



