Submitted:
18 April 2026
Posted:
20 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Leishmania Genomes and Protein-Coding Sequences
2.2. tRNA Gene Annotation
2.3. Analysis of Codon Usage in Protein-Coding Sequences
2.4. Calculation of the tRNA Adaptation Index (tAI)
2.5. Parasite Culture, Protein Samples and Proteomic Data
2.6. Calculation of Relative Protein Abundance
3. Results and Discussion
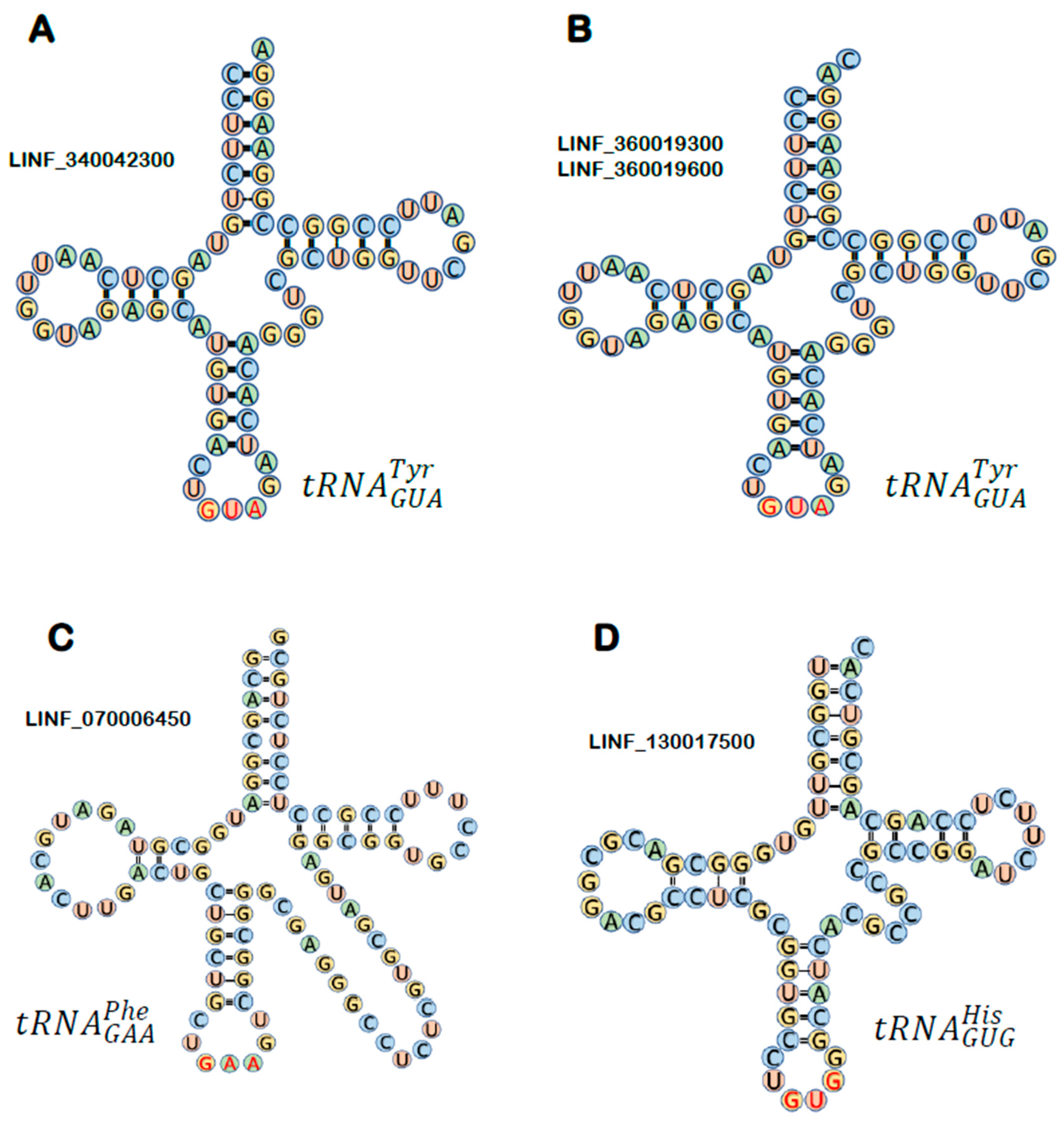
3.1. Annotation of tRNA Genes in the L. infantum Genome
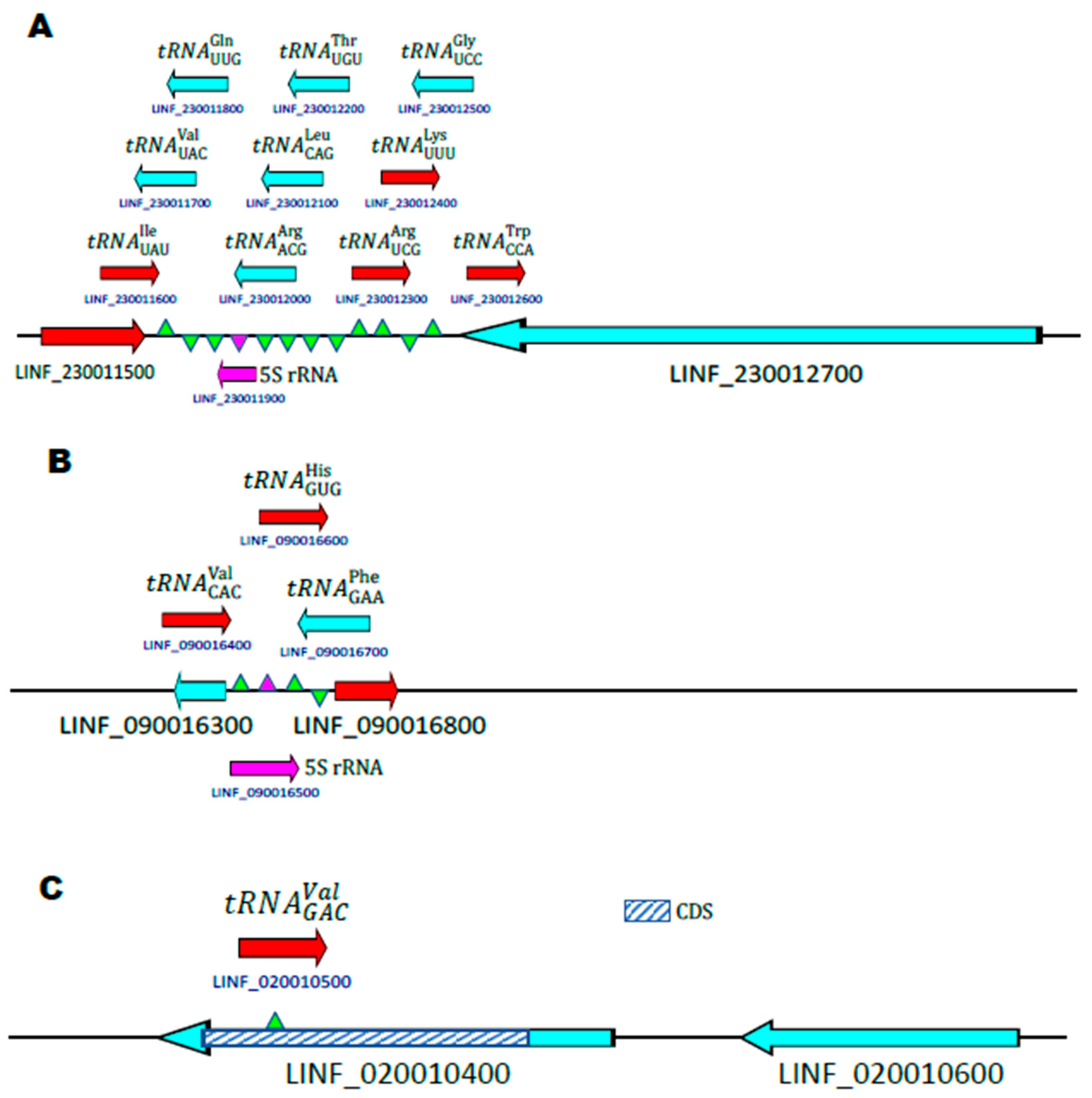
3.2. Genomic Organization of the tRNA Genes
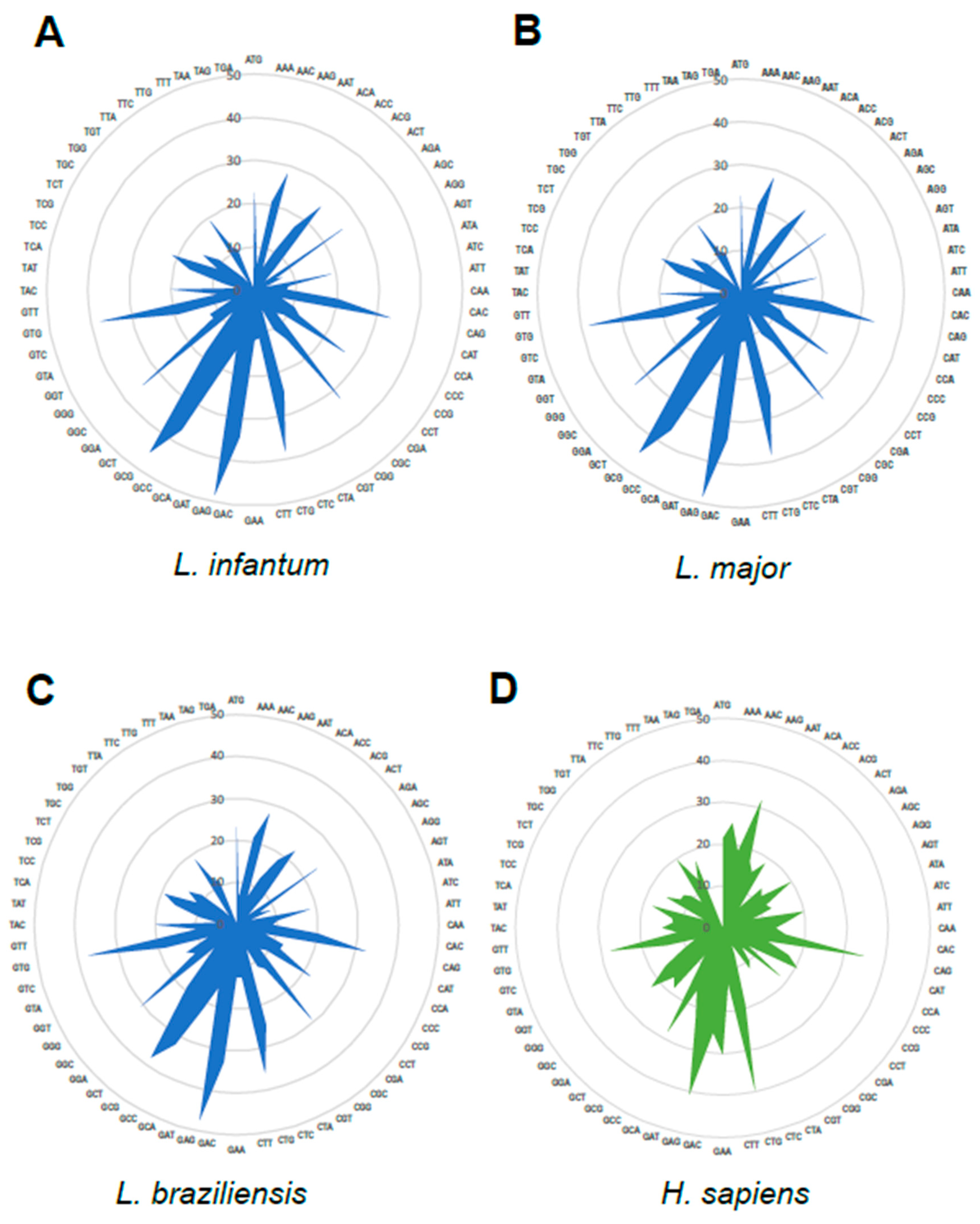
3.3. Analysis of Codon Usage in L. infantum Protein-Coding Sequences
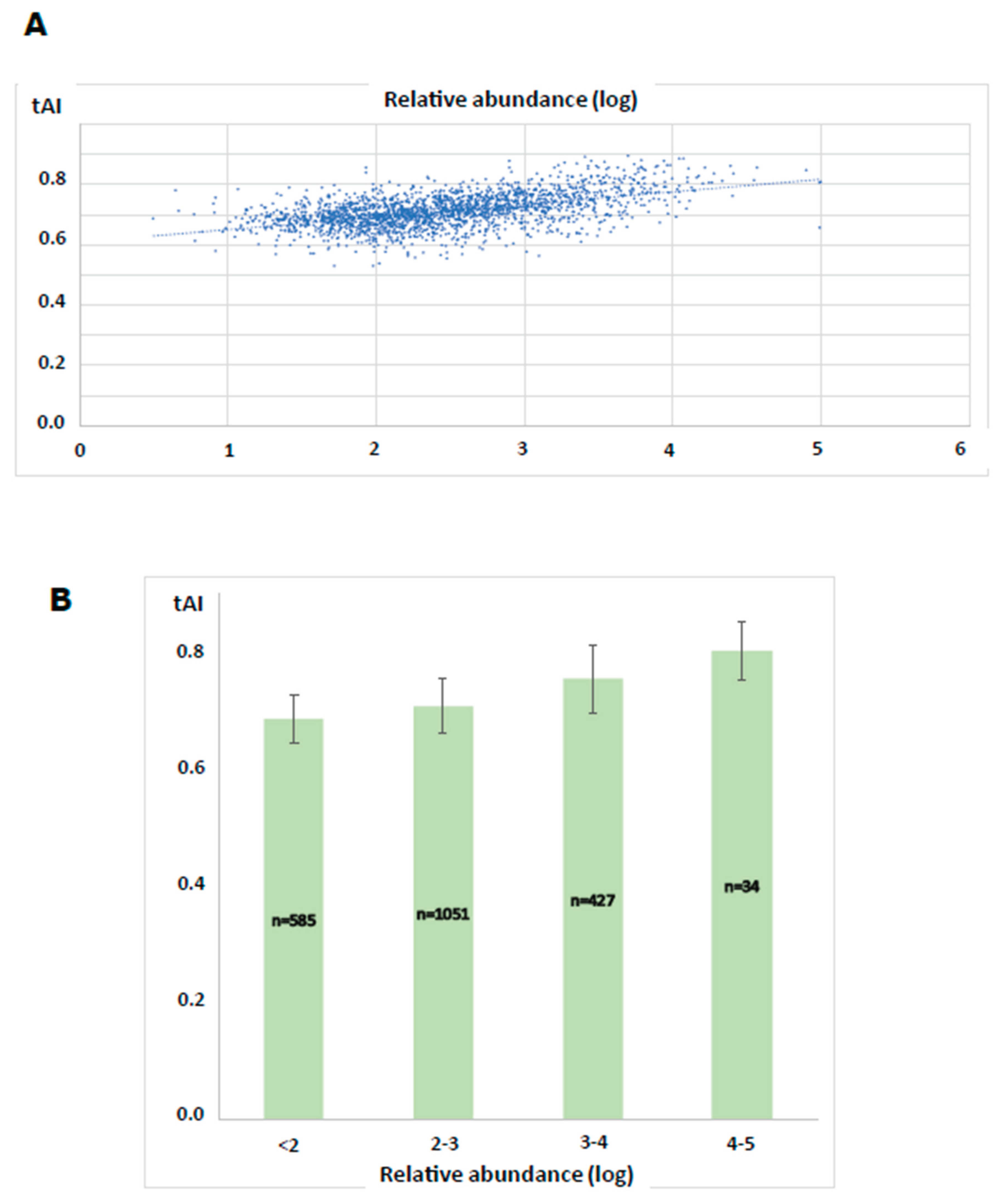
3.4. Determination of the Translational Efficiency of the L. infantum Protein Coding Genes
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Green, R.; Noller, H.F. Ribosomes and Translation. Annu. Rev. Biochem. 1997, 66, 679–716. [Google Scholar] [CrossRef] [PubMed]
- Berg, M.D.; Brandl, C.J. Transfer RNAs: Diversity in Form and Function. RNA Biol. 2021, 18, 316–339. [Google Scholar] [CrossRef]
- Goodenbour, J.M.; Pan, T. Diversity of TRNA Genes in Eukaryotes. Nucleic Acids Res. 2006, 34, 6137–6146. [Google Scholar] [CrossRef]
- Ehrlich, R.; Davyt, M.; López, I.; Chalar, C.; Marín, M. On the Track of the Missing TRNA Genes: A Source of Non-Canonical Functions? Front. Mol. Biosci. 2021, 8, 643701. [Google Scholar] [CrossRef]
- Moreira, D.; Lopez-Garcia, P.; Vickerman, K. An Updated View of Kinetoplastid Phylogeny Using Environmental Sequences and a Closer Outgroup: Proposal for a New Classification of the Class Kinetoplastea. Int J Syst Evol Microbiol 2004, 54, 1861–1875. [Google Scholar] [CrossRef]
- Clayton, C.; Shapira, M. Post-Transcriptional Regulation of Gene Expression in Trypanosomes and Leishmanias. Mol Biochem Parasitol 2007, 156, 93–101. [Google Scholar] [CrossRef]
- Requena, J.M. Lights and Shadows on Gene Organization and Regulation of Gene Expression in Leishmania. Front. Biosci. 2011, 16, 2069–2085. [Google Scholar] [CrossRef]
- Padilla-Mejía, N.E.; Florencio-Martínez, L.E.; Figueroa-Angulo, E.E.; Manning-Cela, R.G.; Hernández-Rivas, R.; Myler, P.J.; Martínez-Calvillo, S. Gene Organization and Sequence Analyses of Transfer RNA Genes in Trypanosomatid Parasites. BMC Genomics 2009, 10, 232. [Google Scholar] [CrossRef]
- Parvathy, S.T.; Udayasuriyan, V.; Bhadana, V. Codon Usage Bias. Mol. Biol. Rep. 2022, 49, 539–565. [Google Scholar] [CrossRef] [PubMed]
- Frumkin, I.; Lajoie, M.J.; Gregg, C.J.; Hornung, G.; Church, G.M.; Pilpel, Y. Codon Usage of Highly Expressed Genes Affects Proteome-Wide Translation Efficiency. Proc. Natl. Acad. Sci. U. S. A 2018, 115, E4940–E4949. [Google Scholar] [CrossRef] [PubMed]
- Davyt, M.; Bharti, N.; Ignatova, Z. Effect of MRNA/TRNA Mutations on Translation Speed: Implications for Human Diseases. J. Biol. Chem. 2023, 299, 105089. [Google Scholar] [CrossRef]
- Rashmi, M.; Swati, D. Comparative Genomics of Trypanosomatid Pathogens Using Codon Usage Bias. Bioinformation 2013, 9, 912–918. [Google Scholar] [CrossRef]
- Gonzalez-de la Fuente, S.; Peiro-Pastor, R.; Rastrojo, A.; Moreno, J.; Carrasco-Ramiro, F.; Requena, J.M.; Aguado, B. Resequencing of the Leishmania Infantum (Strain JPCM5) Genome and de Novo Assembly into 36 Contigs. Sci Rep 2017, 7, 18050. [Google Scholar] [CrossRef]
- Chan, P.P.; Lin, B.Y.; Mak, A.J.; Lowe, T.M. TRNAscan-SE 2.0: Improved Detection and Functional Classification of Transfer RNA Genes. Nucleic Acids Res. 2021, 49, 9077–9096. [Google Scholar] [CrossRef] [PubMed]
- Laslett, D.; Canback, B. ARAGORN, a Program to Detect TRNA Genes and TmRNA Genes in Nucleotide Sequences. Nucleic Acids Res. 2004, 32, 11–16. [Google Scholar] [CrossRef] [PubMed]
- Rich, A.; RajBhandary, U.L. Transfer RNA: Molecular Structure, Sequence, and Properties. Annu. Rev. Biochem. 1976, 45, 805–860. [Google Scholar] [CrossRef]
- dos Reis, M.; Savva, R.; Wernisch, L. Solving the Riddle of Codon Usage Preferences: A Test for Translational Selection. Nucleic Acids Res. 2004, 32, 5036–5044. [Google Scholar] [CrossRef]
- Adán-Jiménez, J.; Sánchez-Salvador, A.; Morato, E.; Solana, J.C.; Aguado, B.; Requena, J.M. A Proteogenomic Approach to Unravel New Proteins Encoded in the Leishmania Donovani (HU3) Genome. Genes (Basel) 2024, 15, 775. [Google Scholar] [CrossRef]
- Tran, N.H.; Qiao, R.; Xin, L.; Chen, X.; Liu, C.; Zhang, X.; Shan, B.; Ghodsi, A.; Li, M. Deep Learning Enables de Novo Peptide Sequencing from Data-Independent-Acquisition Mass Spectrometry. Nat. Methods 2019, 16, 63–66. [Google Scholar] [CrossRef]
- Matzke, M.M.; Brown, J.N.; Gritsenko, M.A.; Metz, T.O.; Pounds, J.G.; Rodland, K.D.; Shukla, A.K.; Smith, R.D.; Waters, K.M.; Mcdermott, J.E.; et al. A Comparative Analysis of Computational Approaches to Relative Protein Quantification Using Peptide Peak Intensities in Label-Free LC-MS Proteomics Experiments. Proteomics 2013, 13, 493–503. [Google Scholar] [CrossRef] [PubMed]
- Cassago, A.; Rodrigues, E.M.; Prieto, E.L.; Gaston, K.W.; Alfonzo, J.D.; Iribar, M.P.; Berry, M.J.; Cruz, A.K.; Thiemann, O.H. Identification of Leishmania Selenoproteins and SECIS Element. Mol Biochem Parasitol 2006, 149, 128–134. [Google Scholar] [CrossRef]
- Díaz-Viraqué, F.; Ehrlich, R.; Robello, C. Genomic Organization of Trypanosoma Cruzi TRNA Genes. Genome Biol. Evol. 2025, 17, evaf108. [Google Scholar] [CrossRef] [PubMed]
- Ranjan Kumar, R.; Jain, R.; Akhtar, S.; Parveen, N.; Ghosh, A.; Sharma, V.; Singh, S. Characterization of Thiamine Pyrophosphokinase of Vitamin B1 Biosynthetic Pathway as a Drug Target of Leishmania Donovani. J. Biomol. Struct. Dyn. 2024, 42, 5669–5685. [Google Scholar] [CrossRef] [PubMed]
- Kaufer, A.; Stark, D.; Ellis, J. Evolutionary Insight into the Trypanosomatidae Using Alignment-Free Phylogenomics of the Kinetoplast. Pathogens 2019, 8, 157. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, A.; Sarkar, R.R. Comparison of Codon Usage Bias across Leishmania and Trypanosomatids to Understand MRNA Secondary Structure, Relative Protein Abundance and Pathway Functions. Genomics 2015, 106, 232–241. [Google Scholar] [CrossRef]
- Silva, J.C.; Gorenstein, M. V.; Li, G.-Z.; Vissers, J.P.C.; Geromanos, S.J. Absolute Quantification of Proteins by LCMSE. Mol. Cell. Proteomics 2006, 5, 144–156. [Google Scholar] [CrossRef]
- Fredrick, K.; Ibba, M. How the Sequence of a Gene Can Tune Its Translation. Cell 2010, 141, 227–229. [Google Scholar] [CrossRef]
| Amino acid | Anticodon | # tRNAs | tRNA gene ID | Wik | Codon | ‰ in CDSa |
| Phe | GAA | 3 | LINF_070006450* LINF_090016700 LINF_310035600 |
1 | TTC | 19.14 |
| AAA | 0 | 0.54 | TTT | 10.32 | ||
| Leu | CAG | 2 | LINF_090013100 LINF_230012100 |
1 | CTG | 38.12 |
| AAG | 3 | LINF_110009900 LINF_110010300 LINF_360074600 |
0.30 | CTT | 11.26 | |
| UAA | 1 | LINF_240022200 | 0.04 | TTA | 1.64 | |
| CAA | 1 | LINF_290025800 | 0.29 | TTG | 10.87 | |
| UAG | 1 | LINF_340017300 | 0.12 | CTA | 4.68 | |
| GAG | 0 | 0.65 | CTC | 24.81 | ||
| Ile | UAU | 1 | LINF_230011600 | 0.14 | ATA | 2.72 |
| AAU | 3 | LINF_240022300 LINF_240022400 LINF_340017100 |
0.44 | ATT | 8.37 | |
| GAU | 0 | 1 | ATC | 18.91 | ||
| Met (i) | CAU | 2 | LINF_090013000 LINF_360074700 |
1 | ATG | 22.65 |
| Met | CAU | 2 | LINF_110015000 LINF_340017200 |
1 | ATG | |
| Val | CAC | 2 | LINF_090016400 LINF_090017100 |
1 | GTG | 37.53 |
| AAC | 2 | LINF_210018800 LINF_340017000 |
0.23 | GTT | 8.73 | |
| UAC | 1 | LINF_230011700 | 0.15 | GTA | 5.45 | |
| GAC | 1 | LINF_020010500 | 0.51 | GTC | 19.16 | |
| Ser | GCU | 2 | LINF_170013500 LINF_210010800 |
1 | AGC | 25.48 |
| CGA | 1 | LINF_290025900 | 0.83 | TCG | 21.18 | |
| AGA | 1 | LINF_310035700 | 0.39 | TCT | 9.97 | |
| UGA | 1 | LINF_340017400 | 0.29 | TCA | 7.37 | |
| ACU | 0 | 0.28 | AGT | 7.23 | ||
| GGA | 0 | 0.64 | TCC | 16.32 | ||
| Pro | AGG | 2 | LINF_210018700 LINF_360033400 |
0.34 | CCT | 8.83 |
| CGG | 2 | LINF_240010900 LINF_340042500 |
1 | CCG | 26.13 | |
| UGG | 1 | LINF_360074800 | 0.40 | CCA | 10.48 | |
| GGG | 0 | 0.47 | CCC | 12.31 | ||
| Thr | UGU | 1 | LINF_230012200 | 0.40 | ACA | 10.02 |
| CGU | 2 | LINF_300026300 LINF_340042400 |
1 | ACG | 25.01 | |
| AGU | 3 | LINF_360019400 LINF_360019500 LINF_360033200 |
0.28 | ACT | 6.97 | |
| GGU | 0 | 0.70 | ACC | 17.47 | ||
| Ala | CGC | 2 | LINF_110010000 LINF_110010200 |
1 | GCG | 45.36 |
| AGC | 2 | LINF_170013600 LINF_310011300 |
0.41 | GCT | 18.51 | |
| UGC | 1 | LINF_330008700 | 0.45 | GCA | 20.40 | |
| GGC | 1 | LINF_290019900 | 0.81 | GCC | 36.95 | |
| Tyr | GUA | 3 | LINF_340042300 LINF_360019300 LINF_360019600 |
1 | TAC | 20.00 |
| AUA | 0 | 0.20 | TAT | 3.97 | ||
| His | GUG | 2 | LINF_090016400 LINF_090017400 LINF_130017500* |
1 | CAC | 20.15 |
| AUG | 0 | 0.33 | CAT | 6.59 | ||
| Gln | CUG | 3 | LINF_160017000 LINF_240022100 LINF_360051500 |
1 | CAG | 33.20 |
| UUG | 1 | LINF_230011800 | 0.23 | CAA | 7.64 | |
| Asn | GUU | 4 | LINF_100020000 LINF_280009850 LINF_340042200 LINF_340042600 |
1 | AAC | 20.81 |
| AUU | 1 | LINF_230016900 | 0.26 | AAT | 5.45 | |
| Lys | CUU | 4 | LINF_030011800 LINF_100019900 LINF_210018900 LINF_330021200 |
1 | AAG | 28.08 |
| UUU | 1 | LINF_230012400 | 0.20 | AAA | 5.64 | |
| Asp | GUC | 3 | LINF_130017500 LINF_170013400 LINF_240024600 |
1 | GAC | 34.26 |
| AUC | 0 | 0.43 | GAT | 14.57 | ||
| Glu | UUC | 1 | LINF_090017100 | 0.24 | GAA | 11.63 |
| CUC | 2 | LINF_150015100 LINF_310019000 |
1 | GAG | 48.53 | |
| Cys | GCA | 1 | LINF_360074500 | 1 | TGC | 14.63 |
| ACA | 0 | 0.27 | TGT | 3.93 | ||
| Trp | CCA | 1 | LINF_230012600 | 1 | TGG | 10.79 |
| Arg | ACG | 4 | LINF_050014900 LINF_070014000 LINF_110015000 LINF_230012000 |
0.32 | CGT | 10.31 |
| CCG | 1 | LINF_090017400 | 0.43 | CGG | 13.87 | |
| UCG | 1 | LINF_230012300 | 0.23 | CGA | 7.54 | |
| UCU | 2 | LINF_180013900 LINF_330008600 |
0.09 | AGA | 2.79 | |
| CCU | 1 | LINF_330008500 | 0.18 | AGG | 5.70 | |
| GCG | 0 | 1 | CGC | 32.50 | ||
| Gly | GCC | 4 | LINF_100020000 LINF_310018900 LINF_360033100 LINF_360033300 |
1 | GGC | 34.75 |
| CCC | 1 | LINF_110010100 | 0.34 | GGG | 11.87 | |
| UCC | 1 | LINF_230012500 | 0.19 | GGA | 6.68 | |
| ACC | 1 | LINF_210019700 | 0.35 | GGT | 12.11 | |
| SeC | UCA | 1 | LINF_060007300 | NA | TGA | NA |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




