Submitted:
17 April 2026
Posted:
21 April 2026
You are already at the latest version
Abstract
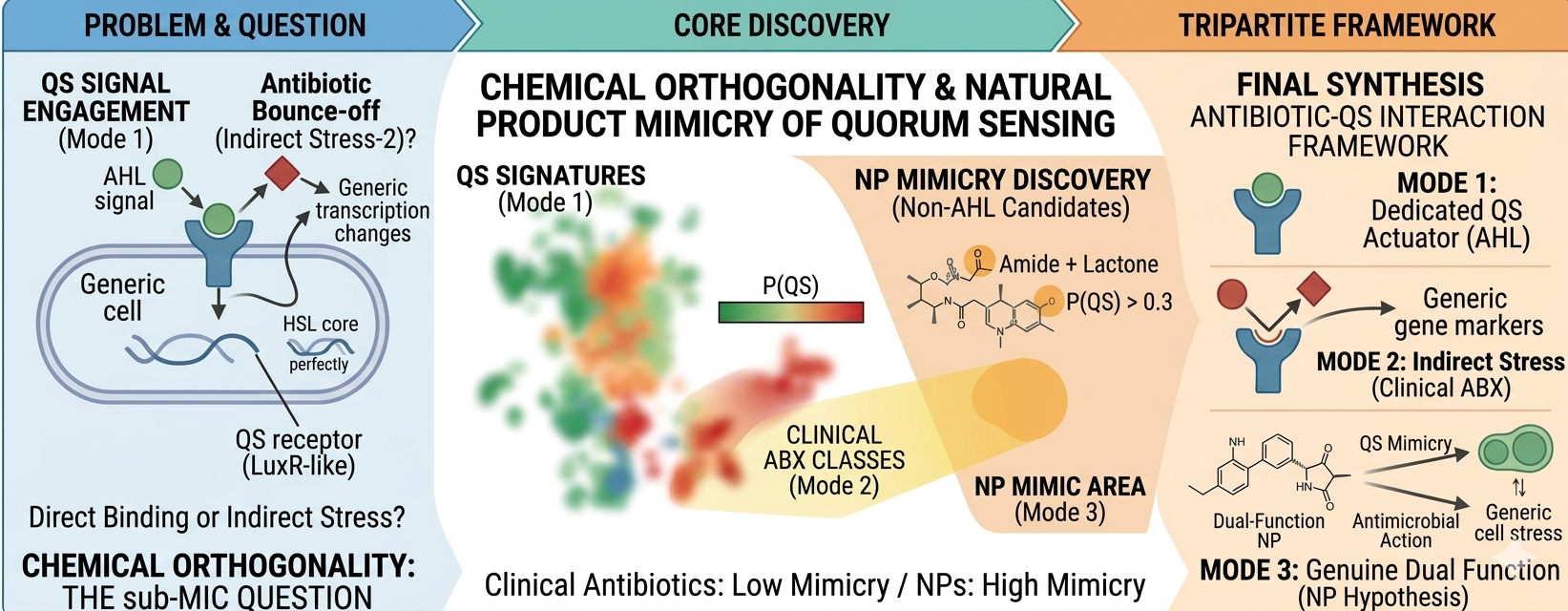
Keywords:
1. Introduction
2. Materials and Methods
2.1. Training Data and Model Architecture
2.2. Validation and Sensitivity Analysis
2.3. Antibacterial Datasets and Natural Product Screening
2.4. Pharmacophore and Statistical Analysis
3. Results and Discussion
3.1. Model Performance and Specificity
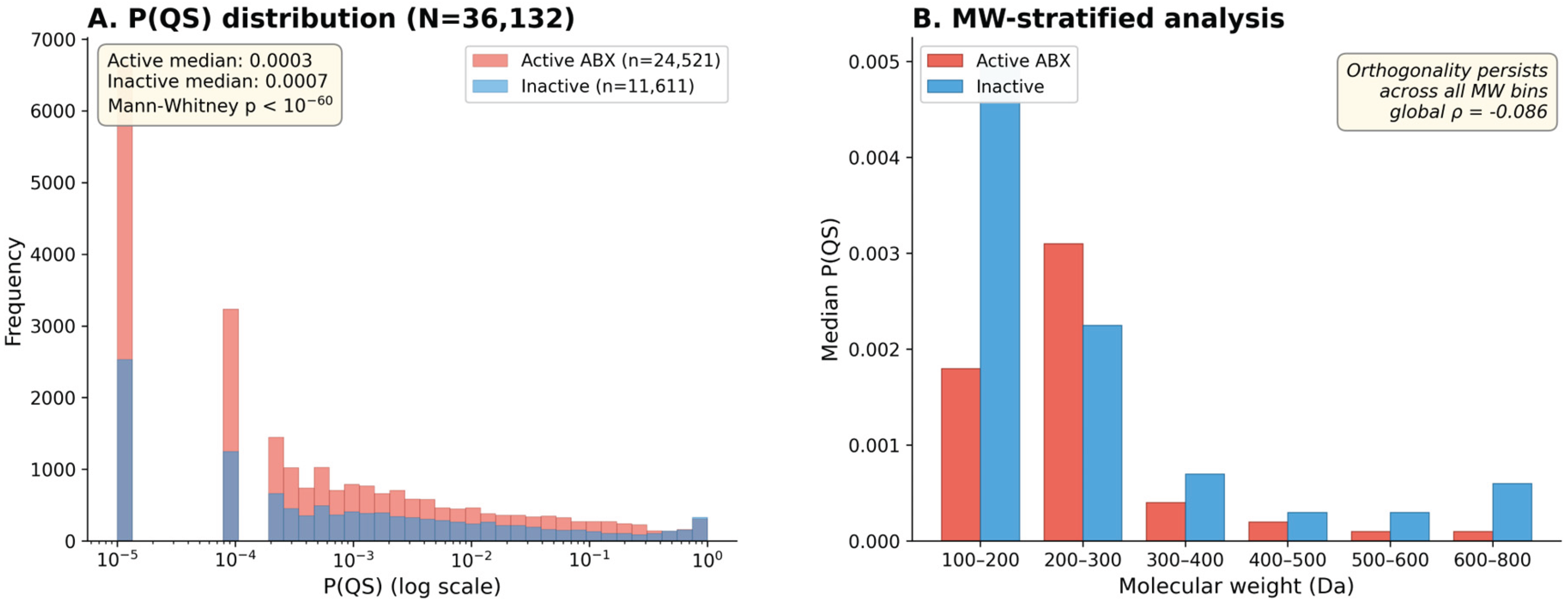
3.2. Chemical Orthogonality of Clinical Antibiotics
3.3. Large-Scale Screening and Natural Product Library
3.4. Non-AHL Candidates: Chemical Profile and Literature Evidence
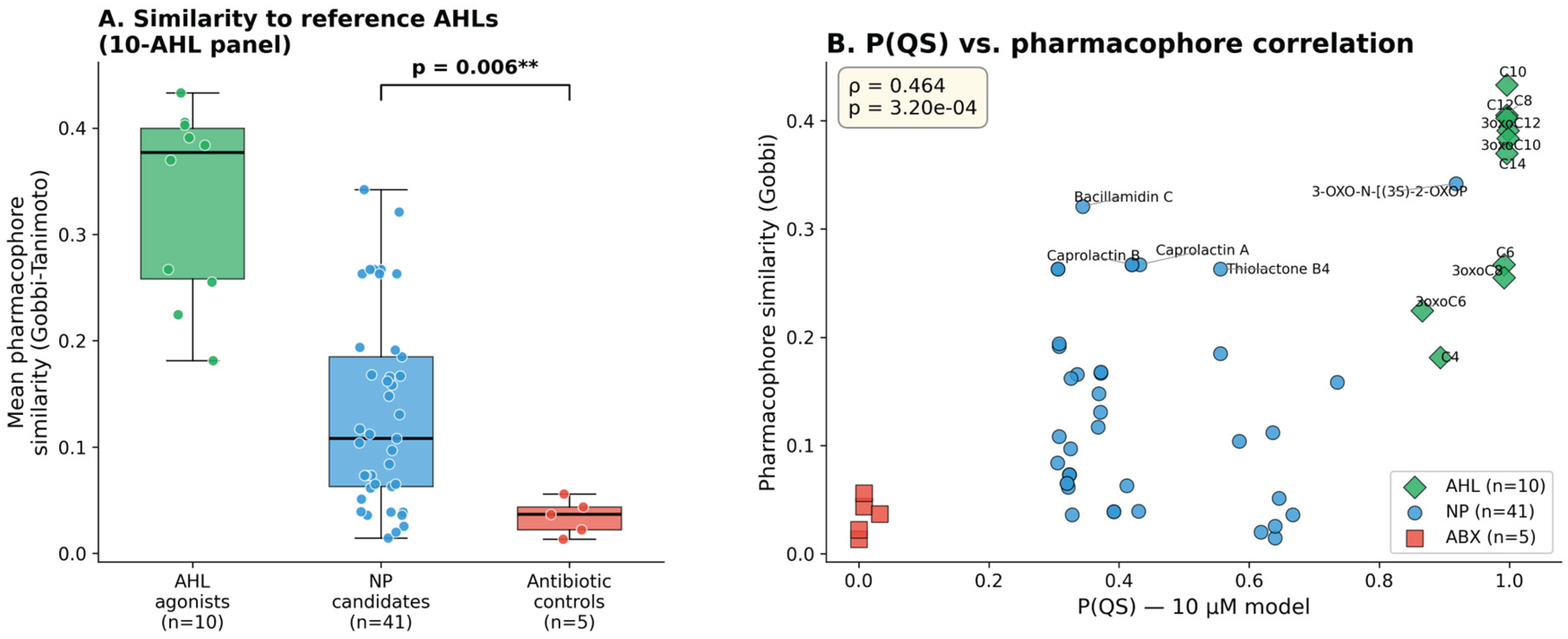
3.5. Independent Pharmacophore Validation
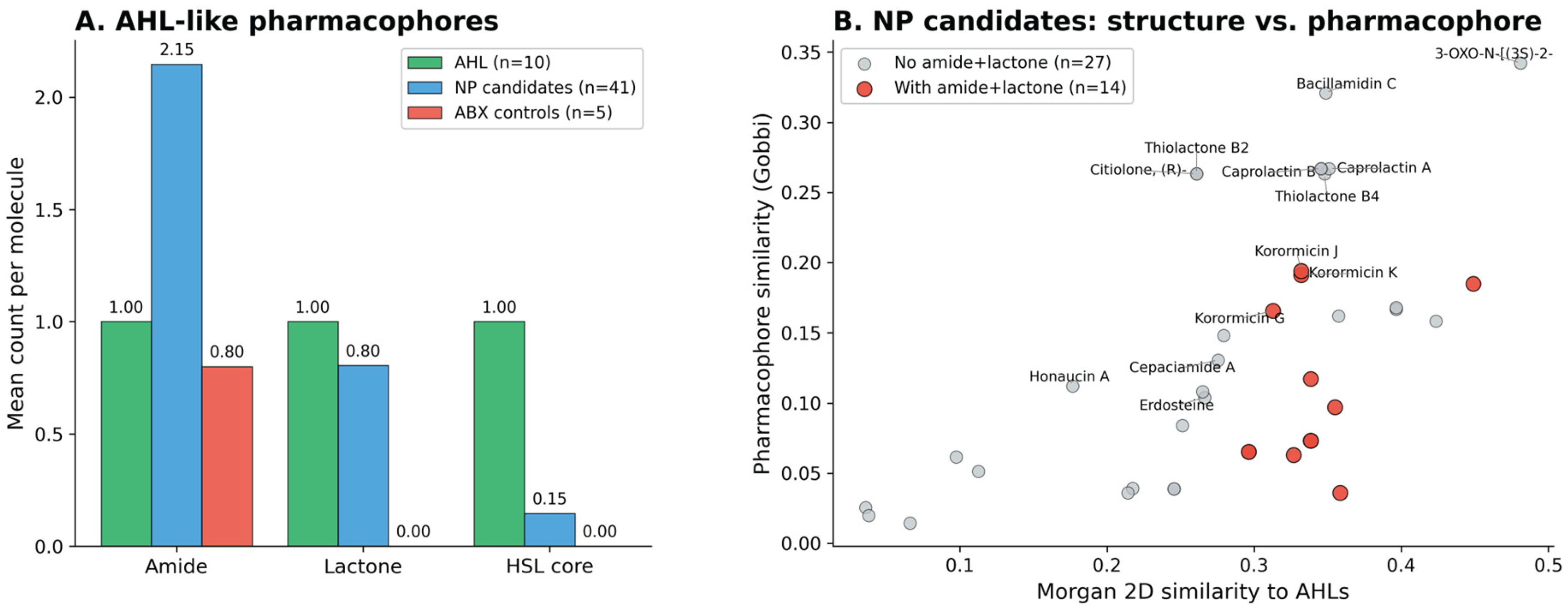
3.6. Sensitivity Analysis
3.7. Mechanistic Synthesis and Implications
3.8. Limitations
4. Conclusions
5. AI Use Declaration
Supplementary Materials
Data Availability Statement
Conflicts of Interest
References
- Davies, J.; Spiegelman, G. B.; Yim, G. The world of subinhibitory antibiotic concentrations. Curr. Opin. Microbiol. 2006, 9, 445–453. [Google Scholar] [CrossRef] [PubMed]
- Romero, D.; Traxler, M. F.; López, D.; Kolter, R. Antibiotics as signal molecules. Chem. Rev. 2011, 111, 5492–5505. [Google Scholar] [CrossRef] [PubMed]
- Goh, E. B.; Yim, G.; Tsui, W.; McClure, J.; Surette, M. G.; Davies, J. Transcriptional modulation of bacterial gene expression by subinhibitory concentrations of antibiotics. Proc. Natl. Acad. Sci. USA 2002, 99, 17025–17030. [Google Scholar]
- Linares, J. F.; Gustafsson, I.; Baquero, F.; Martínez, J. L. Antibiotics as intermicrobial signaling agents instead of weapons. Proc. Natl. Acad. Sci. USA 2006, 103, 19484–19489. [Google Scholar] [PubMed]
- Yim, G.; Wang, H. H.; Davies, J. Antibiotics as signalling molecules. Philos. Trans. R. Soc. B 2007, 362, 1195–1200. [Google Scholar]
- Tateda, K.; Comte, R.; Pechere, J. C.; Köhler, T.; Yamaguchi, K.; Van Delden, C. Azithromycin inhibits quorum sensing in Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 2001, 45, 1930–1933. [Google Scholar] [CrossRef]
- Skindersoe, M. E.; Alhede, M.; Phipps, R.; Yang, L.; Jensen, P. O.; Rasmussen, T. B.; et al. Effects of antibiotics on quorum sensing in Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 2008, 52, 3648–3663. [Google Scholar] [CrossRef]
- Carvalho, R. D.; et al. Aminoglycosides and quorum-sensing regulation in Vibrio cholerae. Microbiol. Res. 2021, 253, 126876. [Google Scholar]
- El-Far, A.; et al. Cephalosporin derivatives inhibit violacein production. J. Mol. Struct. 2023, 1292, 136134. [Google Scholar]
- Grandclément, C.; Tannières, M.; Moréra, S.; Dessaux, Y.; Faure, D. Quorum quenching: role in nature and applied developments. FEMS Microbiol. Rev. 2016, 40, 86–116. [Google Scholar]
- Whiteley, M.; Diggle, S. P.; Greenberg, E. P. Progress in and promise of bacterial quorum sensing research. Nature 2017, 551, 313–320. [Google Scholar] [CrossRef]
- O’Shea, R.; Moser, H. E. Physicochemical properties of antibacterial compounds. J. Med. Chem. 2008, 51, 2871–2878. [Google Scholar] [CrossRef]
- Jonkergouw, C.; et al. Exploration of chemical diversity in intercellular quorum sensing signalling systems. Angew. Chem. Int. Ed. 2024, 63, e202314469. [Google Scholar]
- Mahmoud, A. S.; et al. Machine learning approaches for prediction of quorum sensing modulators. Comput. Biol. Med. 2025, 186, 109612. [Google Scholar]
- Leus, I. V.; Weeks, J. W.; Bonifay, V.; Smith, L.; Richardson, S.; Zgurskaya, H. I. Property space mapping of Pseudomonas aeruginosa permeability to small molecules. Sci. Rep. 2022, 12, 8220. [Google Scholar] [CrossRef] [PubMed]
- Soares, A. C.; et al. COCONUT 2.0: an integrative natural product database. Nucleic Acids Res. 2024, 52, D1302–D1310. [Google Scholar]
- Barker, N. L.; et al. Antibiotic Korormicin A kills bacteria by producing reactive oxygen species. J. Bacteriol. 2019, 201, e00718-18. [Google Scholar] [CrossRef]
- Tebben, J.; Motti, C.; Hay, M. E.; Tapiolas, D. M.; Steinberg, P. D. Five new korormicins from Pseudoalteromonas sp. J010. Mar. Drugs 2014, 12, 2802–2815. [Google Scholar]
- Choi, H.; Mascuch, S. J.; Villa, F. A.; Byrum, T.; Teasdale, M. E.; Smith, J. E.; et al. Honaucins A–C, potent inhibitors of inflammation and bacterial quorum sensing. Chem. Biol. 2012, 19, 589–598. [Google Scholar]
- Mayville, P.; Ji, G.; Beavis, R.; Yang, H.; Goger, M.; Novick, R. P.; Muir, T. W. Structure-activity analysis of synthetic autoinducing thiolactone peptides from Staphylococcus aureus. Proc. Natl. Acad. Sci. USA 1999, 96, 1218–1223. [Google Scholar] [CrossRef]
- Cazzola, M.; Page, C. P.; Matera, M. G. Erdosteine: possible roles in management of biofilms. Pulm. Pharmacol. Ther. 2018, 53, 62–70. [Google Scholar]
- Zhang, H.; et al. Isolation and structure of caprolactins and related piperidine alkaloids from marine bacteria. J. Nat. Prod. 2021. [Google Scholar]
- Huber, B.; Riedel, K.; Hentzer, M.; Heydorn, A.; Gotschlich, A.; Givskov, M.; et al. The cep quorum-sensing system of Burkholderia cepacia. Microbiology 2004. [Google Scholar]
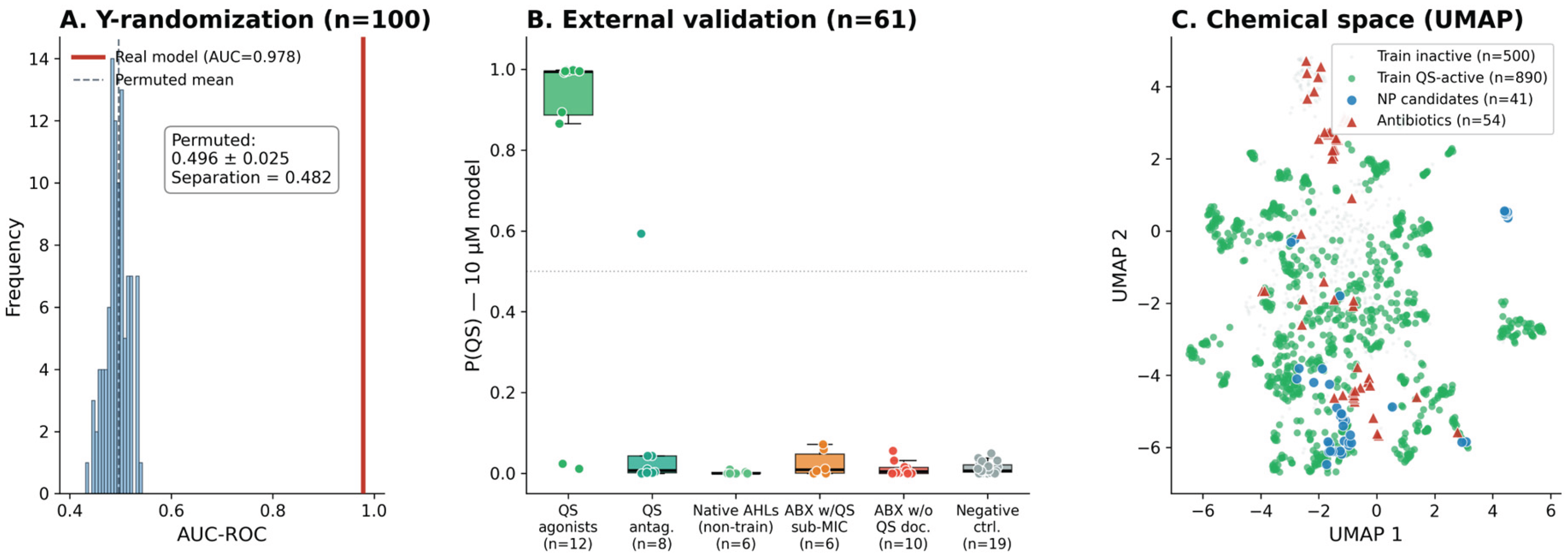
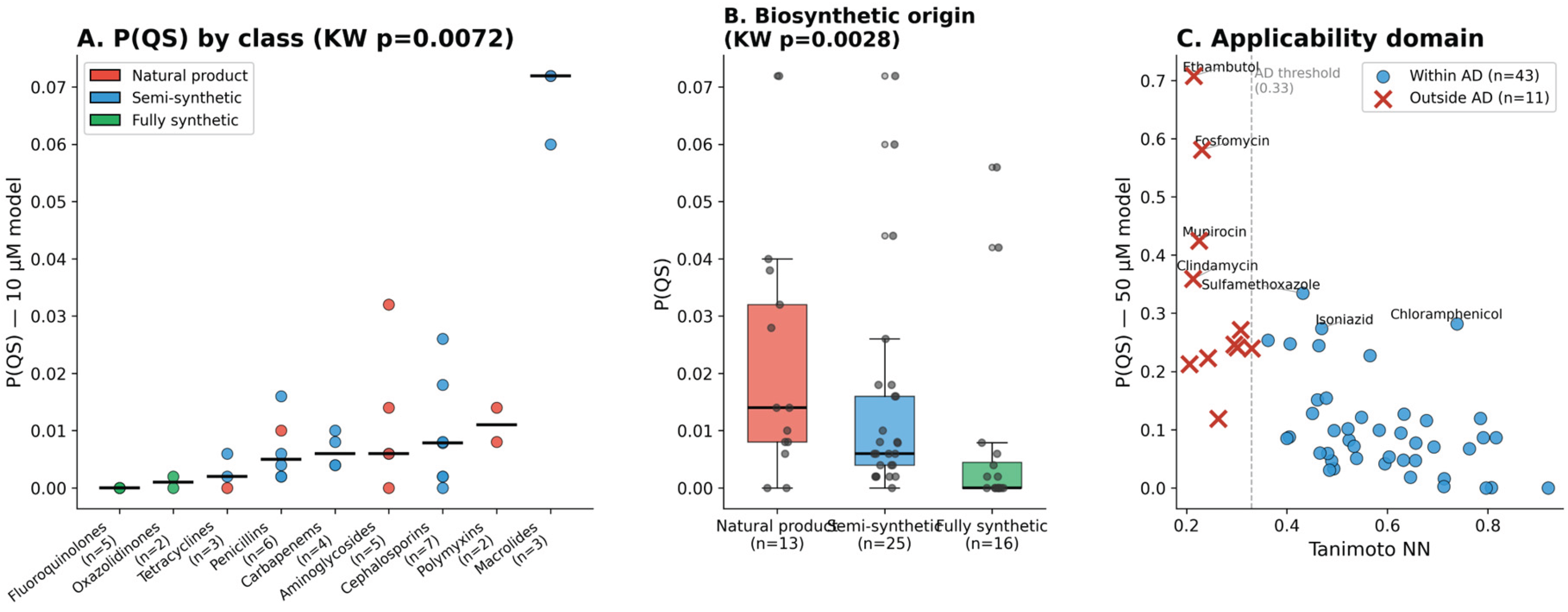
| Compound | P(QS) | Pharm. sim. | NP class | Antibacterial evidence | QS / pharmacophore evidence | Ref. |
|---|---|---|---|---|---|---|
| Honaucin A | 0.636 | 0.155 | Cyanolactone | n.r. | Confirmed QSI (V. harveyi); AHL analog | [19] |
| Erdosteine | 0.585 | 0.127 | Thiolactone | ATB potentiator; anti-biofilm | Thiolactone + amide | [21] |
| Thiolactone B4 | 0.556 | 0.377 | N-acyl amine | Analogs modulate S. aureus | AIP analog (Agr system) | [20] |
| Caprolactin A | 0.432 | 0.180 | Piperid. alkaloid | Antibacterial class (MIC ~5 µg/mL) | Amide + lactam | [22] |
| Caprolactin B | 0.420 | 0.180 | Piperid. alkaloid | Same as A | Amide + lactam | — |
| Triacetylfusigen | 0.412 | 0.051 | Siderophore | Fungal siderophore | Amide + lactone | — |
| Biemamide B | 0.372 | 0.132 | Lipopeptide | Marine ATB class | 5 amides; signal-like features | — |
| Cepaciamide A | 0.371 | 0.094 | Piperid. alkaloid | ATB-producing organism | QS+ org. (CepI/R); amide + lactone | [23] |
| Cladospamide A | 0.368 | 0.115 | Amide | n.r. | Amide + lactone | — |
| Bacillamidin C | 0.344 | 0.202 | Piperid. alkaloid | ATB-producing genus | 3 amides; peptidic scaffold | — |
| Korormicin G | 0.336 | 0.142 | Polyketide | Confirmed ATB (P. aeruginosa MIC ~20 µM) | Amide + lactone; Na+-NQR | [17] |
| Biemamide D | 0.326 | 0.130 | N-acyl amine | Same as Biemamide B | 5 amides | — |
| Korormicin J | 0.308 | 0.223 | Polyketide | Confirmed ATB (Vibrio spp.) | Amide + lactone | [18] |
| Korormicin K | 0.308 | 0.216 | Polyketide | Confirmed ATB (Vibrio spp.) | Amide + lactone | [18] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




