Submitted:
17 April 2026
Posted:
20 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
1.1. Iranian Population and the Zoroastrians as an Ethnic and Religious Minority
1.2. Iran and Its Diverse Geography
1.3. Genetic Studies
2. Materials and Methods
3. Results
4. Discussion
5. Conclusion
References
- Organization, W.H. Health data overview for the Islamic Republic of Iran. Available online: https://data.who.int/countries/364 (accessed on.
- Ashrafian-Bonab, M.; Lawson Handley, L.J.; Balloux, F. Is urbanization scrambling the genetic structure of human populations? A case study. Heredity 2007, 98, 151-156. [CrossRef]
- Behnam, B.; Zakeri, M. Genetics and genomic medicine in Iran. Mol Genet Genomic Med 2019, 7, e00606. [CrossRef]
- State, U.S.D.o. IRAN 2022 INTERNATIONAL RELIGIOUS FREEDOM REPORT. Available online: https://www.state.gov/reports/2022-report-on-international-religious-freedom/iran (accessed on.
- Jahanian, D. The History of Zoroastrians after Arab Invasion; Alien in Their Homeland. Available online: https://www.cais-soas.com/CAIS/History/Post-Sasanian/zoroastrians_after_arab_invasion.htm (accessed on.
- Chaubey, G.; Ayub, Q.; Rai, N.; Prakash, S.; Mushrif-Tripathy, V.; Mezzavilla, M.; Pathak, A.K.; Tamang, R.; Firasat, S.; Reidla, M.; et al. “Like sugar in milk”: reconstructing the genetic history of the Parsi population. Genome Biol 2017, 18, 110. [CrossRef]
- Quintana-Murci, L.; Krausz, C.; Zerjal, T.; Sayar, S.H.; Hammer, M.F.; Mehdi, S.Q.; Ayub, Q.; Qamar, R.; Mohyuddin, A.; Radhakrishna, U.; et al. Y-chromosome lineages trace diffusion of people and languages in southwestern Asia. Am J Hum Genet 2001, 68, 537-542. [CrossRef]
- Qamar, R.; Ayub, Q.; Mohyuddin, A.; Helgason, A.; Mazhar, K.; Mansoor, A.; Zerjal, T.; Tyler-Smith, C.; Mehdi, S.Q. Y-Chromosomal DNA Variation in Pakistan. The American Journal of Human Genetics 2002, 70, 1107-1124. [CrossRef]
- Dhalla, M.N. History of Zoroastrianism; AMS Press: 1977.
- Cavalli-Sforza, L.L.; Menozzi, P.; Piazza, A. The history and geography of human genes; Princeton university press: 1994.
- Shah, B. Rise and fall of Ottoman Empire-Seljuk Turks of central Asia. 2015.
- Blair, J.B.a.S. Islam: A Thousand Years of Faith and Power 1st ed.; Yale University Press: 2002; p. 304.
- Daryaee, T. The Oxford Handbook of Iranian History, 1st ed.; Oxford University Press: UK, 2012; p. 436.
- Bosworth, C.E. The Ghaznavids: Their Empire in Afghanistan and Eastern Iran; Hassell Street Press: 2021; p. 352.
- V, J. Zoroaster, Zarathushtra. Available online: https://www.hinduwebsite.com/zoroastrianism/zoroaster.asp (accessed on 02/01/2026).
- Fisher, W.B. The Cambridge History of Iran: The Land of Iran: Volume 1; Cambridge University Press: United Kingdom, 1968; p. 804.
- Soucek, V.V.B.a.S. An Historical Geography of Iran; Princeton University Press: USA, 2017; p. 310.
- POLK, W.R. Understanding Iran: Everything You Need to Know, from Persia to the Islamic Republic, from Cyrus to Ahmadinejad; Griffin: 2011; p. 288.
- Khoshsirat, Z. Toward a Linguistic Anthropology of Iran: Essays on Language, Culture, and Society; Springer: 2026; p. 273.
- Wellman, B. History of Iran: An Enthralling Overview of Iranian History, Starting from the Ancient Persians through the Persian Empire to Modern Iran; Billy Wellman: 2023; p. 160.
- Speth, J.D. Modern human origins: Do we know what we’re looking for? Quaternary Environments and Humans 2026, 4, 100102. [CrossRef]
- Gomes, C.L. Chapter 20 - Archaeogenetics of human history. In DNA in Health, Identity, and Ancestry, Edinur, H.A., Kofi, A.E., Lahiri, D., Eds.; Academic Press: 2026; pp. 333-355.
- Ammerman, A.J.; Cavalli-Sforza, L.L. The Neolithic transition and the genetics of populations in Europe. 2014.
- Ruffo, P.; Traynor, B.J.; Conforti, F.L. Unveiling the regulatory potential of the non-coding genome: Insights from the human genome project to precision medicine. Genes & Diseases 2025, 12, 101652. [CrossRef]
- Achilli, A.; Rengo, C.; Battaglia, V.; Pala, M.; Olivieri, A.; Fornarino, S.; Magri, C.; Scozzari, R.; Babudri, N.; Santachiara-Benerecetti, A.S.; et al. Saami and Berbers--an unexpected mitochondrial DNA link. Am J Hum Genet 2005, 76, 883-886. [CrossRef]
- Katara, A.; Chand, S.; Chaudhary, H.; Chaudhry, V.; Chandra, H.; Dubey, R.C. Evolution and applications of Next Generation Sequencing and its intricate relations with chromatographic and spectrometric techniques in modern day sciences. Journal of Chromatography Open 2024, 5, 100121. [CrossRef]
- Luo, H.; Wang, Y.; Hou, H.; Yang, J.; Liu, Y.-X. Advances and applications in sequencing-based pathogen surveillance. aBIOTECH 2026, 7, 100004. [CrossRef]
- Jiménez-Arroyo, C.; Molinero, N.; del Campo, R.; Delgado, S.; Moreno-Arribas, M.V. Human gut microbiome study through metagenomics: Recent advances and challenges for clinical implementation. Enfermedades infecciosas y microbiologia clinica (English ed.) 2025, 43, 698-708. [CrossRef]
- Tyler-Smith, C.; Zerjal, T.; Xue, Y.; Wells, R.S.; Bao, W.; Zhu, S.; Qamar, R.; Ayub, Q.; Mohyuddin, A.; Fu, S.; et al. Y-chromosomal DNA variation and human population history. International Congress Series 2003, 1239, 281-282. [CrossRef]
- Soh, P.X.Y.; Adams, A.; Bornman, M.S.R.; Jiang, J.; Stricker, P.D.; Mutambirwa, S.B.A.; Jaratlerdsiri, W.; Hayes, V.M. Y chromosome variation and prostate cancer ancestral disparities. iScience 2025, 28, 112437. [CrossRef]
- Hashiyada, M.; Umetsu, K.; Yuasa, I.; Tamura, A.; Matsusue, A.; Suzuki, K.; Kashimura, S.; Funayama, M. Population genetics of 17 Y-chromosomal STR loci in Japanese. Forensic Science International: Genetics 2008, 2, e69-e70. [CrossRef]
- Livni, J.; Skorecki, K. Distinguishing between founder and host population mtDNA lineages in the Ashkenazi population. Human Gene 2025, 45, 201445. [CrossRef]
- Castillo, A.; Gomes, V.; Ossa, H.; Ribeiro, B.; Prata, M.J.; Rondón, F.; Simão, F.; Gusmão, L. An overview of the mtDNA diversity across the Colombian Andean region. Forensic Science International: Genetics 2025, 78, 103288. [CrossRef]
- Colucci, M.; Wetton, J.H.; Rolf, B.; Sheehan, N.; Jobling, M.A. Evaluating genome-wide and targeted forensic sequencing approaches to kinship determination. Forensic Science International: Genetics 2025, 76, 103228. [CrossRef]
- Jobling, M.A. The impact of recent events on human genetic diversity. Philosophical Transactions of the Royal Society B: Biological Sciences 2012, 367, 793-799. [CrossRef]
- Al-Zahery, N.; Semino, O.; Benuzzi, G.; Magri, C.; Passarino, G.; Torroni, A.; Santachiara-Benerecetti, A.S. Y-chromosome and mtDNA polymorphisms in Iraq, a crossroad of the early human dispersal and of post-Neolithic migrations. Mol Phylogenet Evol 2003, 28, 458-472. [CrossRef]
- Kumar, L.; Ahlawat, B.; Kumar, S.; Mushrif-Tripathy, V.; Rai, N. Maternal ancestry of first Parsi settlers of India using ancient mitogenome. Mitochondrion 2023, 71, 104-111. [CrossRef]
- López, S.; Thomas, M.G.; van Dorp, L.; Ansari-Pour, N.; Stewart, S.; Jones, A.L.; Jelinek, E.; Chikhi, L.; Parfitt, T.; Bradman, N.; et al. The Genetic Legacy of Zoroastrianism in Iran and India: Insights into Population Structure, Gene Flow, and Selection. The American Journal of Human Genetics 2017, 101, 353-368. [CrossRef]
- Ahlawat, B.; Dwivedi, A.; Konar, S.; Gupta, S.; Rai, N. Chapter 12 - Applications of genetics in archaeological investigations: exploring human migration and genetic evolution in Indian subcontinent. In Investigative and Predictive DNA Testing, Dash, H.R., Ed.; Academic Press: 2026; pp. 173-188.
- Morawala-Patell, V.; Pasha, N.; Krishnasamy, K.; Mittal, B.; Gopalakrishnan, C.; Mugasimangalam, R.; Sharma, N.; Khanna-Gupta, A.; Bhote-Patell, P.; Rao, S.; Jain, R. The first complete Zoroastrian-Parsi mitochondrial reference genome and genetic signatures of an endogamous non-smoking population. Meta Gene 2021, 28, 100882. [CrossRef]
- Suarez-Trujillo, F.; Amirzargar, A.; Hadinedoushan, H.; Juarez, I.; Nikbin, B.; Gil-Martin, R.; Vaquero-Yuste, C.; Arnaiz-Villena, A. HLA study in Iranian desert Yazd province inhabitants. Human Immunology 2023, 84, 311-312. [CrossRef]
- Quintana-Murci, L.; Chaix, R.; Wells, R.S.; Behar, D.M.; Sayar, H.; Scozzari, R.; Rengo, C.; Al-Zahery, N.; Semino, O.; Santachiara-Benerecetti, A.S.; et al. Where West Meets East: The Complex mtDNA Landscape of the Southwest and Central Asian Corridor. The American Journal of Human Genetics 2004, 74, 827-845. [CrossRef]
- Stausberg, M. From Power to Powerlessness: Zoroastrianism in Iranian History. 2012; pp. 171-193.
- Lashgary, Z.; Khodadadi, A.; Singh, Y.; Houshmand, S.M.; Mahjoubi, F.; Sharma, P.; Singh, S.; Seyedin, M.; Srivastava, A.; Ataei, M.; et al. Y chromosome diversity among the Iranian religious groups: A reservoir of genetic variation. Annals of Human Biology 2011, 38, 364-371. [CrossRef]
- Jonnalagadda, M., Ozarkar, S. and Mushrif-Tripathy, V. . Population affinities of Parsis in the Indian subcontinent. International Journal of Osteoarchaeology 2011, 103-110. [CrossRef]
- Torroni, A.; Bandelt, H.J.; Macaulay, V.; Richards, M.; Cruciani, F.; Rengo, C.; Martinez-Cabrera, V.; Villems, R.; Kivisild, T.; Metspalu, E.; et al. A signal, from human mtDNA, of postglacial recolonization in Europe. Am J Hum Genet 2001, 69, 844-852. [CrossRef]
- Syndercombe Court, D. Mitochondrial DNA in forensic use. Emerg Top Life Sci 2021, 5, 415-426. [CrossRef]
- Salas, A.; Bandelt, H.J.; Macaulay, V.; Richards, M.B. Phylogeographic investigations: the role of trees in forensic genetics. Forensic Sci Int 2007, 168, 1-13. [CrossRef]
- Yoshizawa, N.; Saigo, T.; Suzuki, T. A novel restriction fragment length polymorphism (RFLP) -based method for rapid genotyping of bovine coronaviruses. J Virol Methods 2025, 337, 115188. [CrossRef]
- Butler, J.M.; Schoske, R.; Vallone, P.M.; Kline, M.C.; Redd, A.J.; Hammer, M.F. A novel multiplex for simultaneous amplification of 20 Y chromosome STR markers. Forensic Sci Int 2002, 129, 10-24. [CrossRef]
- Parkin, E.J.; Kraayenbrink, T.; van Driem, G.L.; Tshering Of Gaselô, K.; de Knijff, P.; Jobling, M.A. 26-Locus Y-STR typing in a Bhutanese population sample. Forensic Sci Int 2006, 161, 1-7. [CrossRef]
- Consortium, T.Y.C. The Human Y Chromosomal Haplogroup Tree. Available online: https://web.archive.org/web/20120716130221/http://ycc.biosci.arizona.edu/ (accessed on.
- Emmerova, B.; Ehler, E.; Comas, D.; Votrubova, J.; Vanek, D. Comparison of Y-chromosomal haplogroup predictors. Forensic Science International: Genetics Supplement Series 2017, 6, e145-e147. [CrossRef]
- Behnam, B.; Modarressi, M.H.; Conti, V.; Taylor, K.E.; Puliti, A.; Wolfe, J. Expression of Tsga10 sperm tail protein in embryogenesis and neural development: from cilium to cell division. Biochem Biophys Res Commun 2006, 344, 1102-1110. [CrossRef]
- Xiang, M.; Wang, Y.; Xu, W.; Zheng, N.; Zhang, J.; Duan, Z.; Zha, X.; Shi, X.; Wang, F.; Cao, Y.; Zhu, F. Pathogenesis of acephalic spermatozoa syndrome caused by splicing mutation and de novo deletion in TSGA10. J Assist Reprod Genet 2021, 38, 2791-2799. [CrossRef]

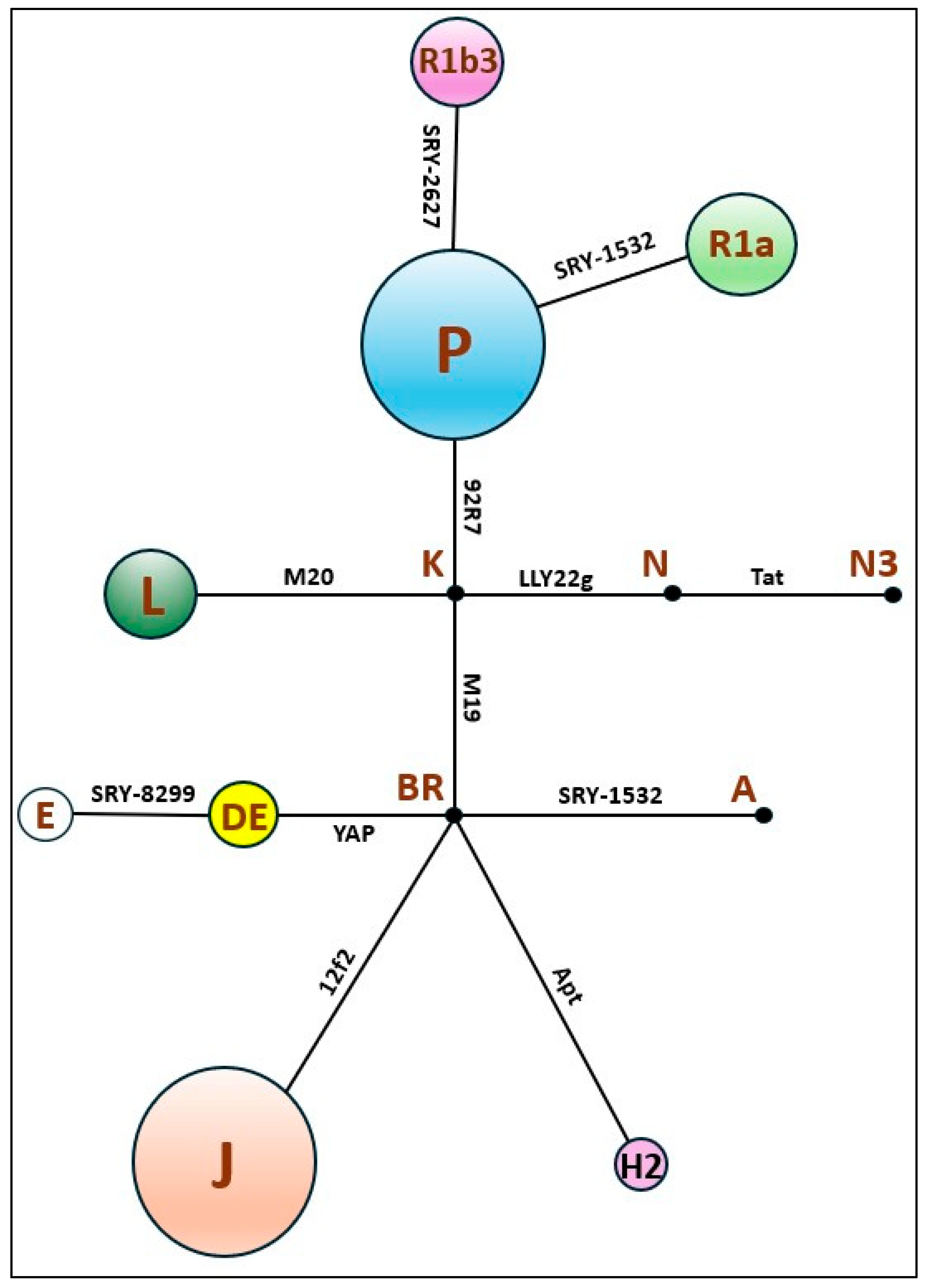
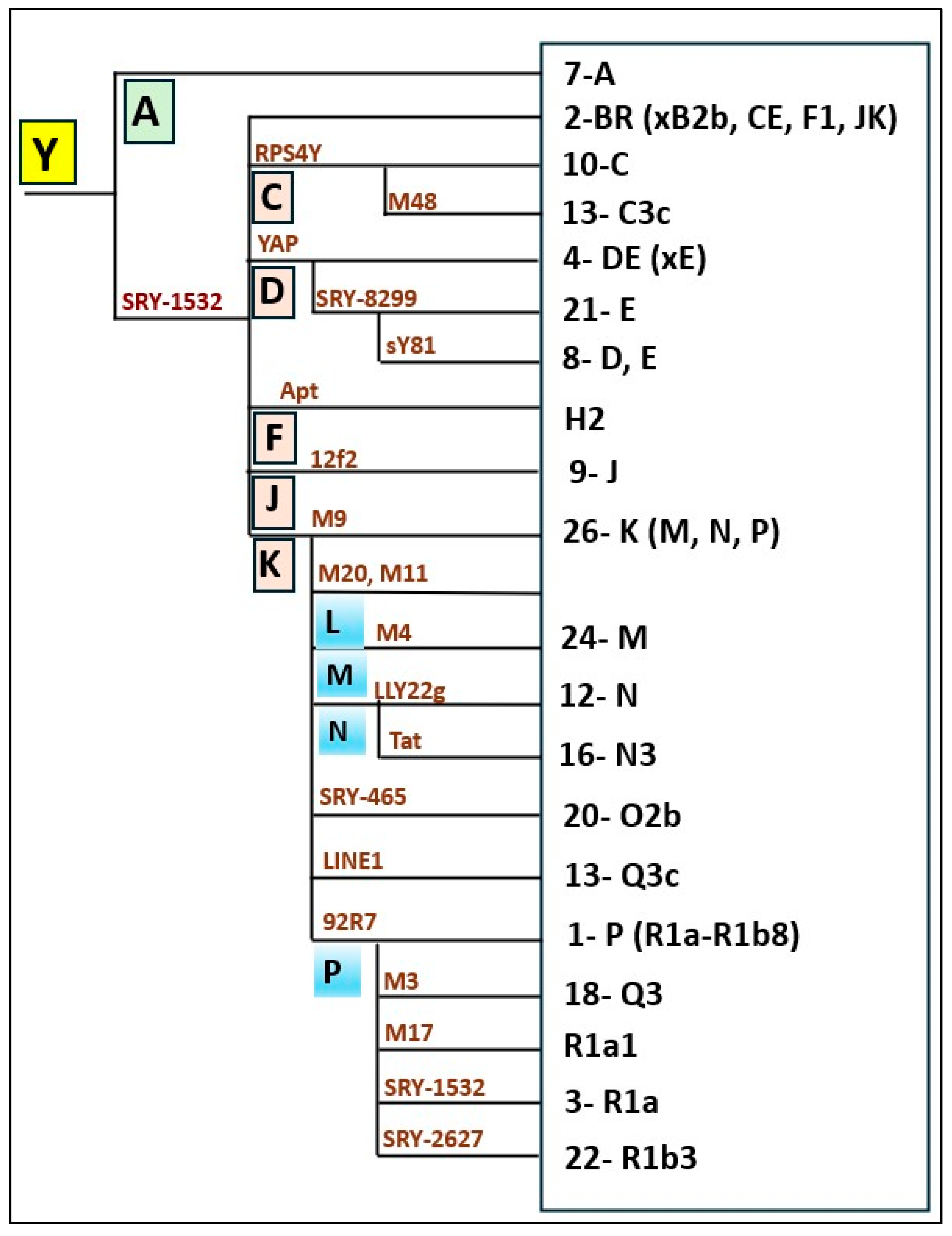
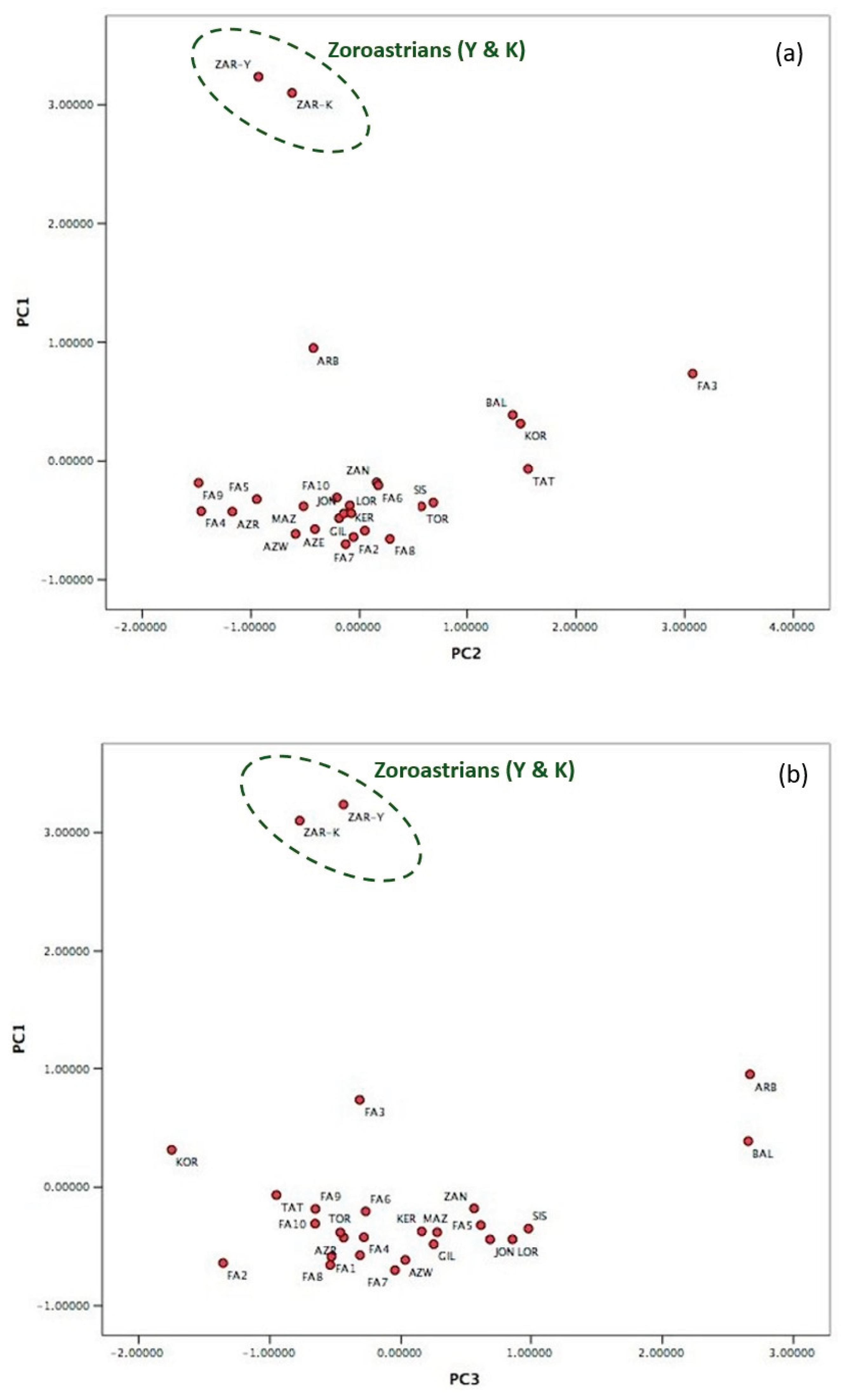
| Haplogroups | Derived state at | FA9 & FA10 | ZAR-Y | ZAR-K | AZE & AZW | KOR | Parsi mean (Qamar et al., 2002) | Pakistani mean (Qamar et al., 2002) | Iranian mean (Quintana-Murci et al., 2001) | European mean (Rosser et al., 2000) |
| No. | 94 | 40 | 54 | 94 | 94 | 90 | 628 | 401 | 3677 | |
| P (xR1b8, R1a, Q3) | 92R7 | 38.3 | 28.4 | 32.2 | 33.4 | 29.2 | 26 | 14.7 | - | 38 |
| BR-xB2b, CE, F1, H and JK | SRY10831a | - | - | - | - | - | 3.3 | 10.9 | - | 22 |
| R1a1 | M17 | 16.9 | 12.9 | 13.2 | 16.3 | 17.9 | 7.7 | 35.9 | 14.6 | 14 |
| DE (xE) | YAP | 7.2 | 8.8 | 9.1 | 9.2 | 18.4 | - | - | - | 0 |
| E3a | M2 | - | - | - | - | - | 0 | 1.2 | - | 0.2 |
| J2 | 12f2 | 21 | 34.8 | 36.1 | 27 | 28.3 | 38.8 | 15.4 | 43 | 8 |
| N | LLY22g | 3.1 | 1.2 | 1.4 | 2.1 | 0 | 0 | 1.5 | - | 0.8 |
| O3c | LINE1 | 0 | 0 | 0 | 0 | 0.2 | 0 | 1.5 | - | - |
| F1 | Apt | 4.8 | 5.2 | 5.7 | 7.1 | 4.8 | - | - | - | - |
| N3 | Tat | 1.6 | 0.8 | 0.8 | 0.2 | 0 | - | - | - | 6 |
| O2b | SRY+465 | 0 | 0 | 0 | 0.4 | 0 | - | - | - | - |
| E | SRY4064 | 5.2 | 6.4 | 7.2 | 8.5 | 14.8 | 5.5 | 1.9 | - | 9 |
| R1b8 | SRY-2627 | 14.4 | 10.2 | 9.4 | 9.3 | 3.8 | - | - | - | 23 |
| M | M4 | 0 | 0 | 0 | 0 | 0 | - | - | - | - |
| K | M9 | 2.7 | 0.9 | 1.1 | 3.1 | 5.1 | 0 | 1.9 | - | 1.5 |
| L | M20 | 9.2 | 9.6 | 9.8 | 4.2 | 2.3 | 17.7 | 12.8 | - | - |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




