Submitted:
15 April 2026
Posted:
17 April 2026
You are already at the latest version
Abstract
Keywords:
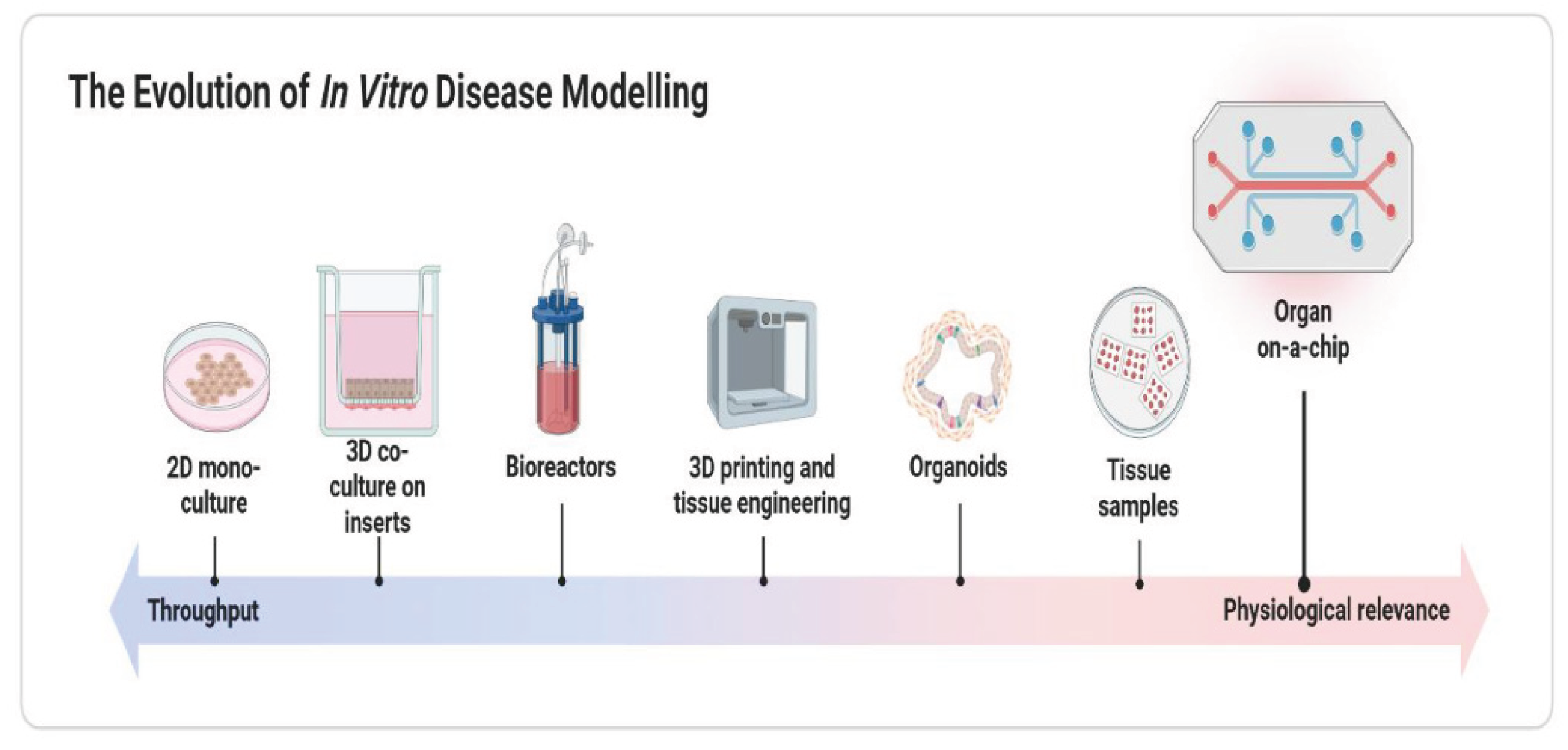
Introduction
Method
SH-SY5Y Cells Expansion
SH-SH5Y Differentiation
Assessment of NLCs Maturity, Purity, and Functional Integrity
Neuronal Purity and Maturity
Functional Assay: Assessment of NLCs Response to NMDA-Induced Excitotoxicity
Modelling Hypoxic-Ischaemic Injury on NLCs
Phase I: Identifying Optimal Hypoxia-Inducing Concentration of Cobalt Chloride
Phase II: Investigating Time-Course of Reoxygenation-Induced Neuronal Injury
Immunocytochemistry
Lactate Dehydrogenase Assay
MTT Assay
Enzyme-Linked Immunosorbent Assay
RNA Extraction
RNA-seq Library Preparation and Sequencing
RNA-seq Data Analysis
Statistical Analysis
Results
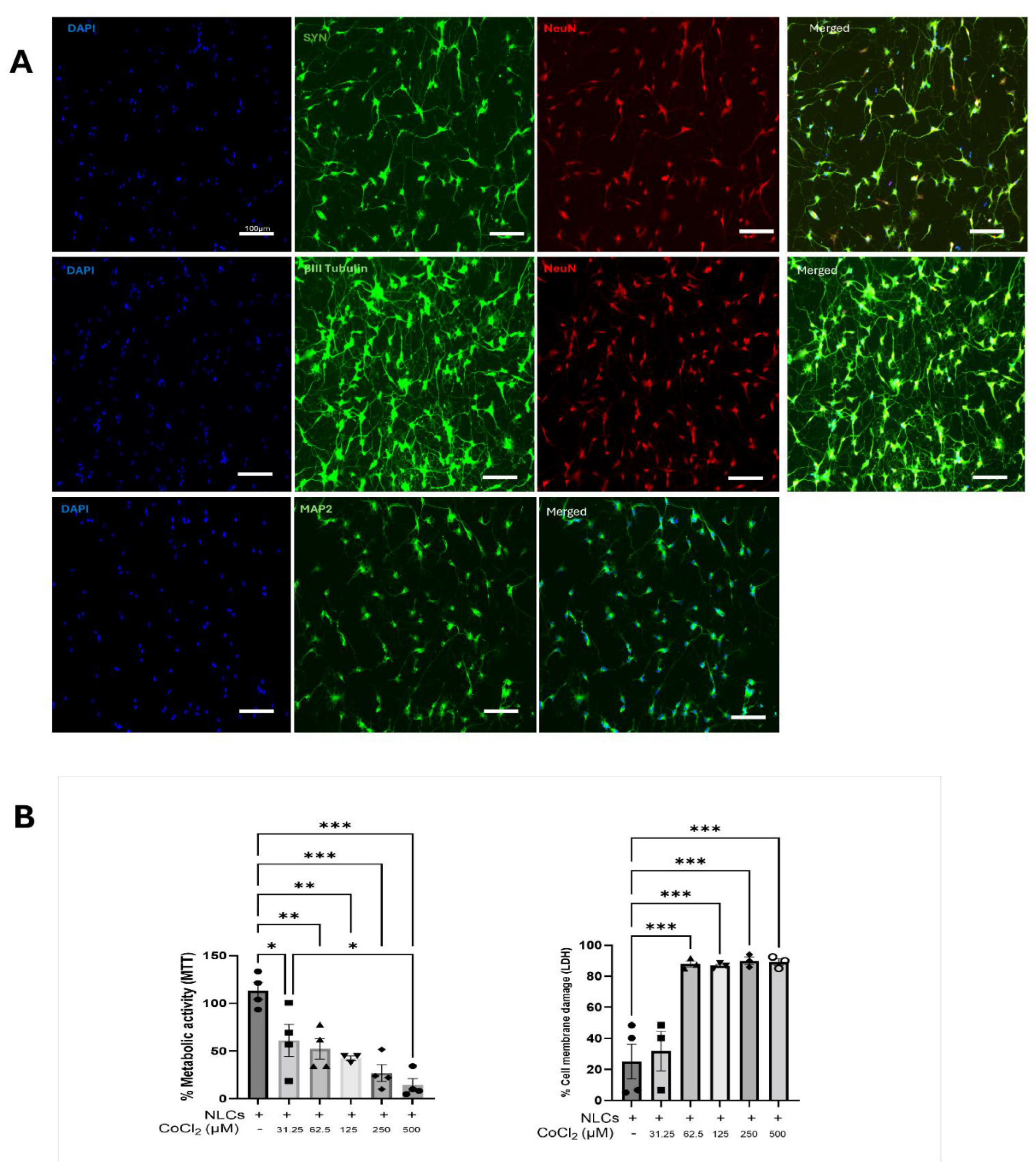
Differentiated SH-SY5Y Cells Have Neuron-Like Characteristics and Express Functional NMDA Receptors
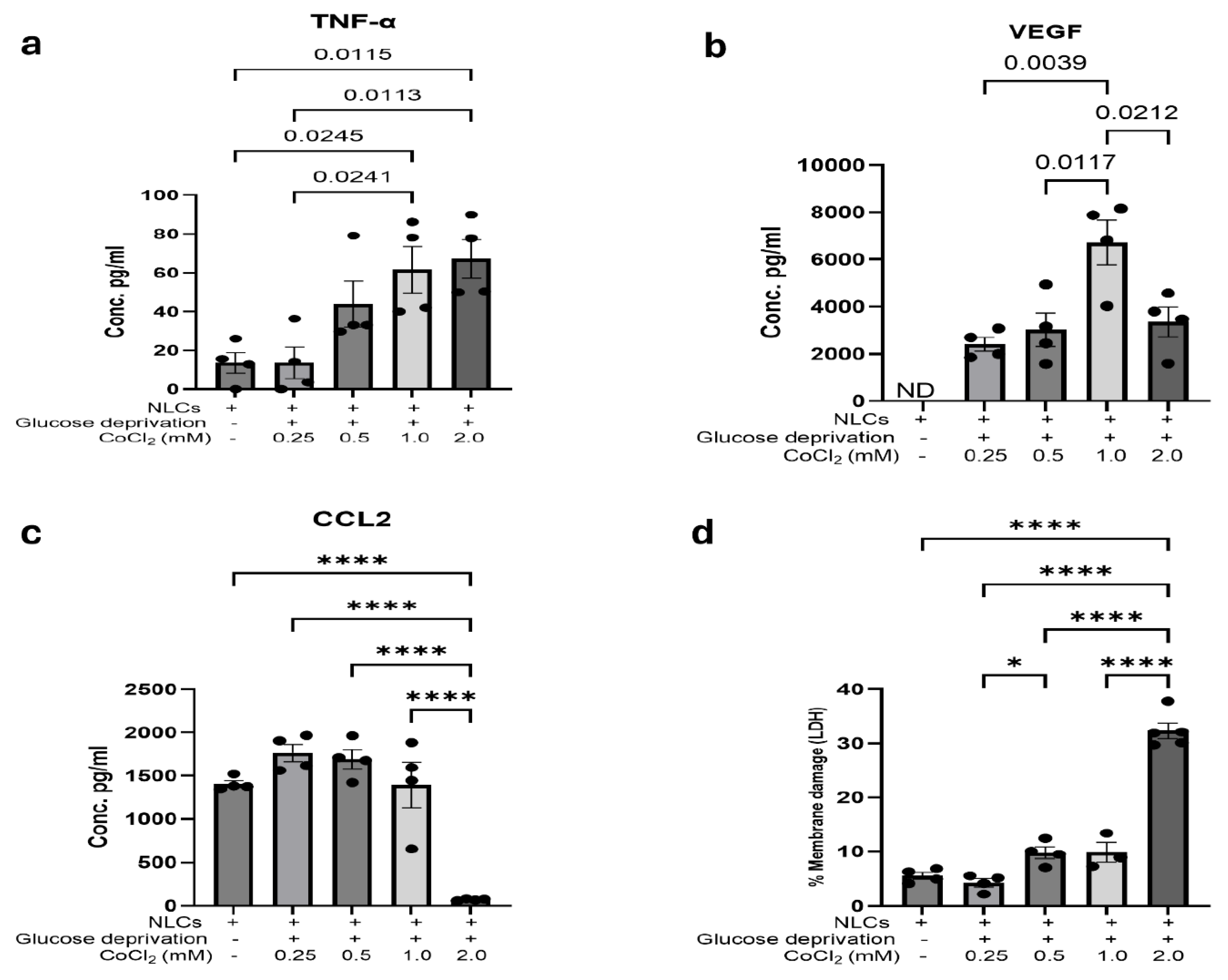
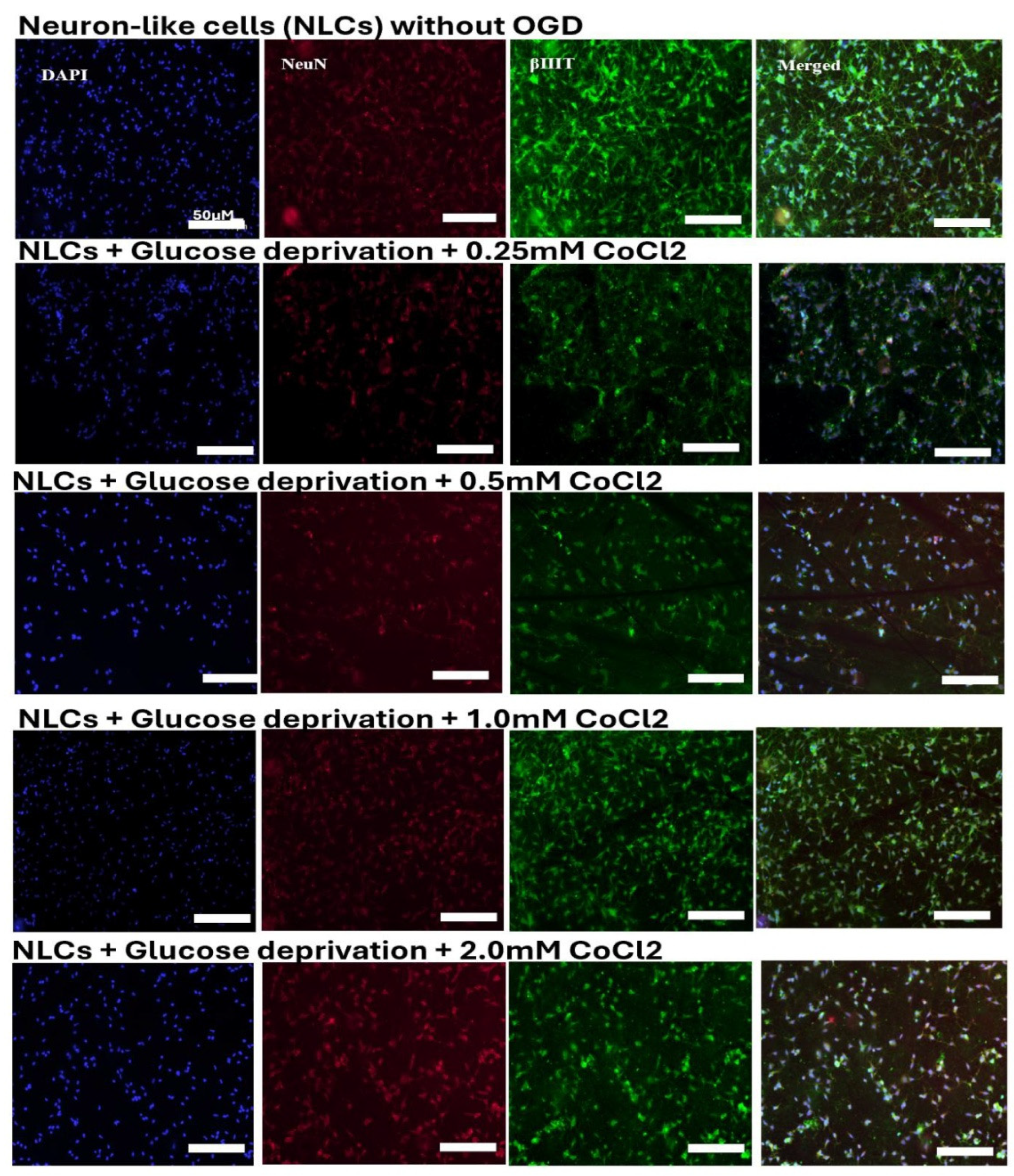
Exposure of NLCs to Glucose Deprivation in the Presence of 1mM CoCl2 Caused Significant Hypoxic Injury
Chemical Hypoxia Accelerates Neuronal Death and Interferes with the Functional and Structural Integrity of Neuron-Like Cells
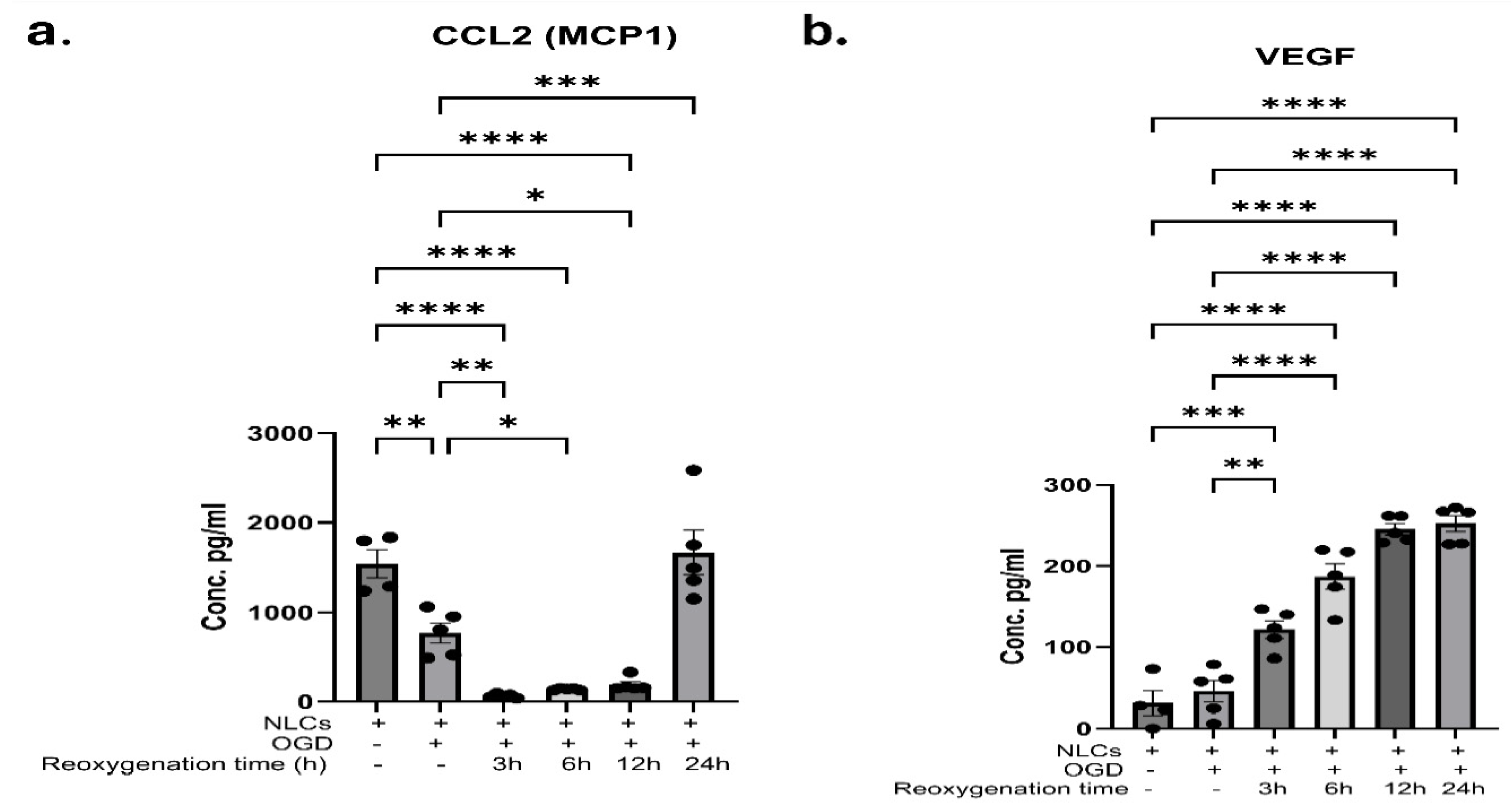
Reoxygenation Following 3 h OGD Resulted in a Temporal Change in Proinflammatory Chemokine and Angiogenesis Factor, VEGF
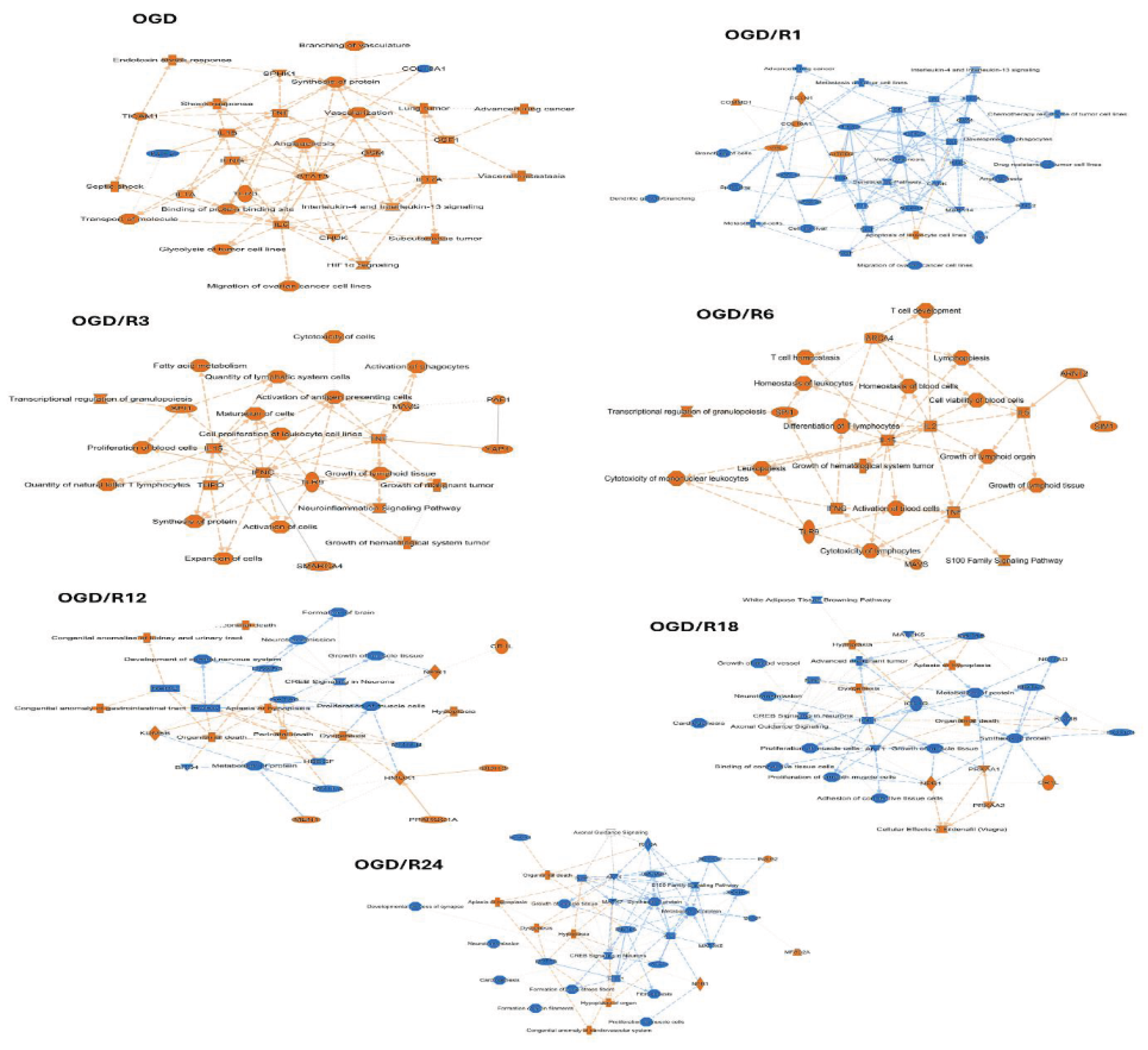
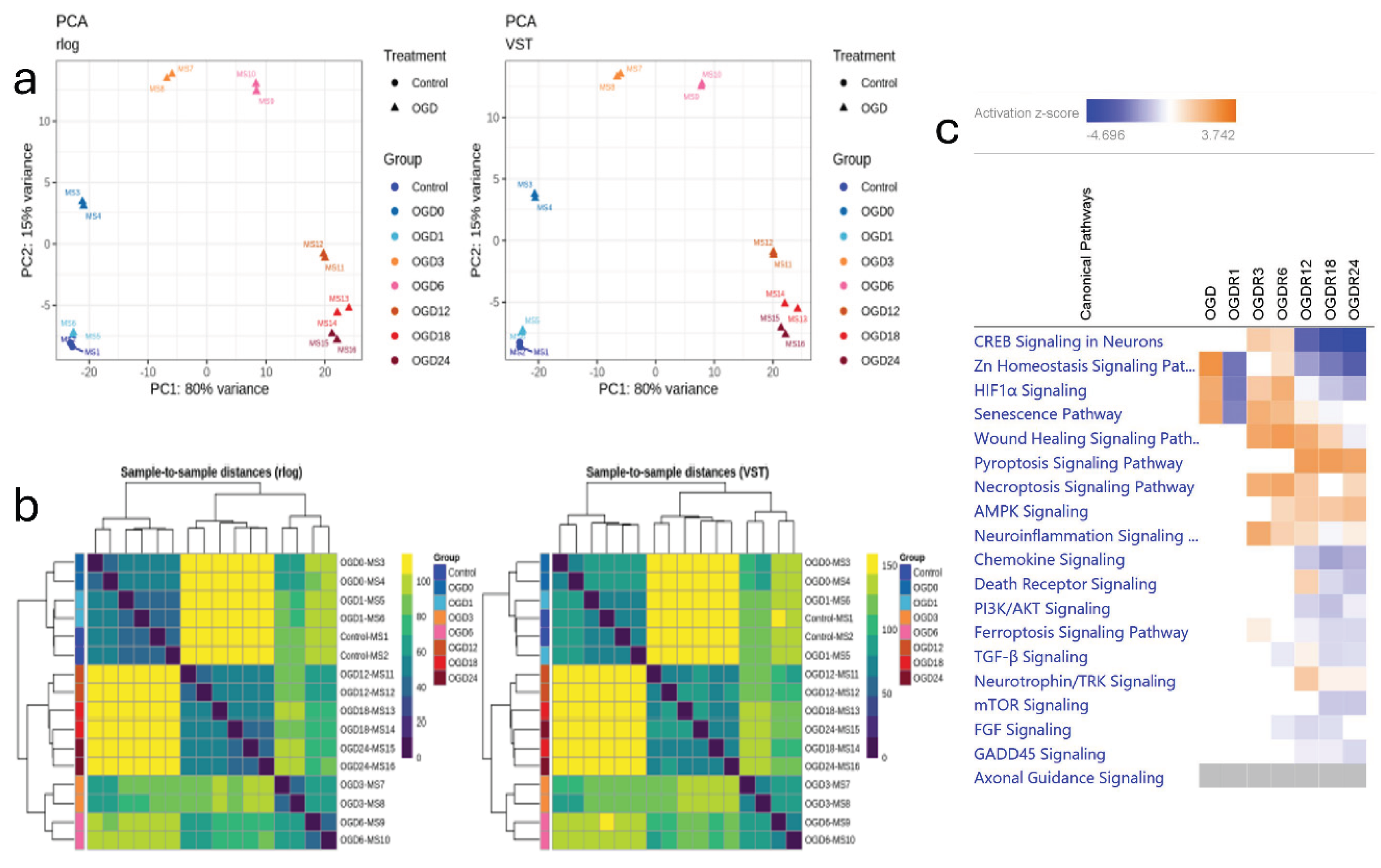
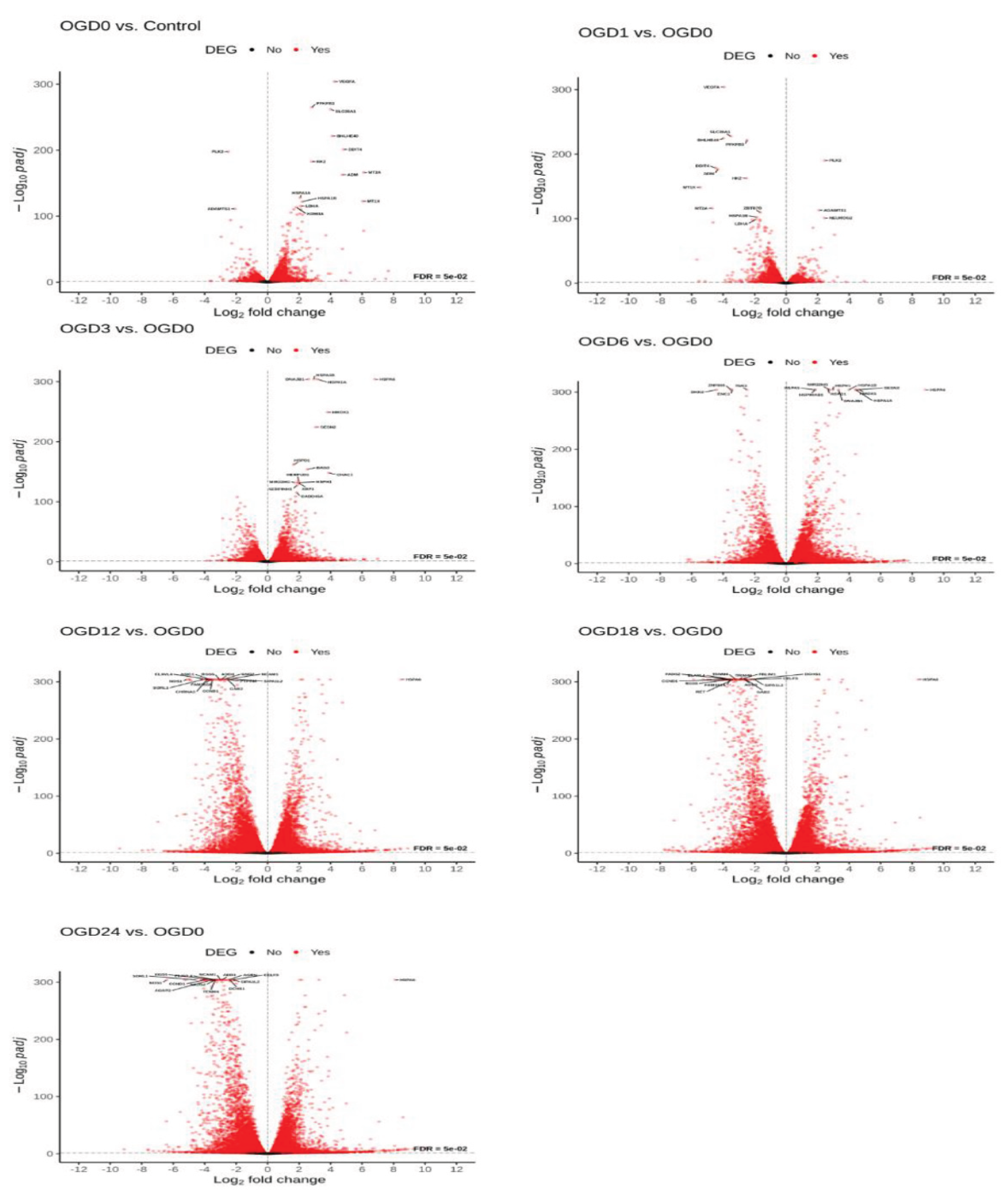
NLCs Undergo Sequential, Multi-Pathway Response Following a 24 h Time-Course OGD/R
Discussion
Conclusions and Future Directions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Pampaloni, F.; Reynaud, E.G.; Stelzer, E.H.K. The third dimension bridges the gap between cell culture and live tissue. Nat. Rev. Mol. Cell Biol. 2007, 8, 839–845. [Google Scholar] [CrossRef] [PubMed]
- Breslin, S.; O’Driscoll, L. Three-dimensional cell culture: the missing link in drug discovery. Drug Discovery Today 2013, 18, 240–249. [Google Scholar] [CrossRef]
- Park, S.; Sharma, H.; Kim, W.; Gwon, Y.; Kim, H.; Choung, Y.-H.; Kim, J. Microengineered organoids: reconstituting organ-level functions in vitro. Organoid 2023, 3. [Google Scholar] [CrossRef]
- Clevers, H. Modeling Development and Disease with Organoids. Cell 2016, 165, 1586–1597. [Google Scholar] [CrossRef] [PubMed]
- Nikolakopoulou, P.; Rauti, R.; Voulgaris, D.; Shlomy, I.; Maoz, B.M.; Herland, A. Recent progress in translational engineered in vitro models of the central nervous system. Brain 2020, 143, 3181–3213. [Google Scholar] [CrossRef]
- Ingber, D.E. Human organs-on-chips for disease modelling, drug development and personalized medicine. Nat. Rev. Genet. 2022, 23, 467–491. [Google Scholar] [CrossRef]
- van Helden, P. The cost of research in developing countries. The EMBO Reports 2012, 13, 395–395. [Google Scholar] [CrossRef]
- Lopez-Suarez, L.; Al Awabdh, S.; Coumoul, X.; Chauvet, C. The SH-SY5Y human neuroblastoma cell line, a relevant in vitro cell model for investigating neurotoxicology in human: Focus on organic pollutants. NeuroToxicology 2022, 92, 131–155. [Google Scholar] [CrossRef]
- Xie, H.-R.; Hu, L.-S.; Li, G.-Y. SH-SY5Y human neuroblastoma cell line: in vitro cell model of dopaminergic neurons in Parkinson's disease. Chin. Med, J. 2010, 123, 1086–92. [Google Scholar]
- Liu, Y.; Eaton, E.D.; Wills, T.E.; McCann, S.K.; Antonic, A.; Howells, D.W. Human Ischaemic Cascade Studies Using SH-SY5Y Cells: a Systematic Review and Meta-Analysis. Transl. Stroke Res. 2018, 9, 564–574. [Google Scholar] [CrossRef]
- Dravid, A.; Raos, B.; Svirskis, D.; O’Carroll, S.J. Optimised techniques for high-throughput screening of differentiated SH-SY5Y cells and application for neurite outgrowth assays. Sci. Rep. 2021, 11, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Bakmiwewa, S.M.; Heng, B.; Guillemin, G.J.; Ball, H.J.; Hunt, N.H. An Effective, Low-cost Method for Achieving and Maintaining Hypoxia During Cell Culture Studies. BioTechniques 2015, 59, 223–229. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Jin, F.; Zhong, H. A novel experimental hypoxia chamber for cell culture. Am. J. Cancer Res. 2014, 4, 53–60. [Google Scholar]
- Rovetta, F.; Stacchiotti, A.; Faggi, F.; Catalani, S.; Apostoli, P.; Fanzani, A.; Aleo, M.F. Cobalt triggers necrotic cell death and atrophy in skeletal C2C12 myotubes. Toxicol. Appl. Pharmacol. 2013, 271, 196–205. [Google Scholar] [CrossRef] [PubMed]
- Muñoz-Sánchez, J.; Chánez-Cárdenas, M.E. The use of cobalt chloride as a chemical hypoxia model. J. Appl. Toxicol. 2019, 39, 556–570. [Google Scholar] [CrossRef]
- Yuan, X.; Li, W.; Yan, Q.; Ou, Y.; Long, Q.; Zhang, P. Biomarkers of mature neuronal differentiation and related diseases. Futur. Sci. OA 2024, 10, 2410146. [Google Scholar] [CrossRef]
- Ankarcrona, M.; Dypbukt, J.M.; Bonfoco, E.; Zhivotovsky, B.; Orrenius, S.; Lipton, S.A.; Nicotera, P. Glutamate-induced neuronal death: A succession of necrosis or apoptosis depending on mitochondrial function. Neuron 1995, 15, 961–973. [Google Scholar] [CrossRef]
- Sims, N.R.; Zaidan, E. Biochemical changes associated with selective neuronal death following short-term cerebral ischaemia. Int J Biochem Cell Biol 1995, 27, 531–50. [Google Scholar] [CrossRef]
- Sun, Y.; Jin, K.; Xie, L.; Childs, J.; Mao, X.O.; Logvinova, A.; Greenberg, D.A. VEGF-induced neuroprotection, neurogenesis, and angiogenesis after focal cerebral ischemia. J. Clin. Investig. 2003, 111, 1843–1851. [Google Scholar] [CrossRef]
- Wu, D.-M.; Liu, J.-P.; Liu, J.; Ge, W.-H.; Wu, S.-Z.; Zeng, C.-J.; Liang, J.; Liu, K.; Lin, Q.; Hong, X.-W.; et al. Immune pathway activation in neurons triggers neural damage after stroke. Cell Rep. 2023, 42, 113368. [Google Scholar] [CrossRef]
- Masoud, G.N.; Li, W. HIF-1α pathway: role, regulation and intervention for cancer therapy. Acta Pharm Sin B 2015, 5, 378–89. [Google Scholar] [CrossRef]
- Beker, M.C.; Evren, E.S.; Özbay, E.; Balaban, B.; Dogan, E.; Yelkenci, H.E.; Ates, N.; Caglayan, A.B.; Doeppner, T.R.; Hermann, D.M.; et al. Circadian clock protein Bmal1 protects against transient focal cerebral ischemia in mice by regulating master signals controlling cell survival and metabolism. Exp. Neurol. 2025, 395, 115481. [Google Scholar] [CrossRef]
- Dong, C.; Li, J.; Tang, Q.; Wang, Y.; Zeng, C.; Du, L.; Sun, Q. Denervation aggravates renal ischemia reperfusion injury via BMAL1-mediated Nrf2/ARE pathway. Arch. Biochem. Biophys. 2023, 746, 109736. [Google Scholar] [CrossRef]
- Zhou, R.; Leng, T.; Yang, T.; Chen, F.; Hu, W.; Xiong, Z.-G. β-Estradiol Protects Against Acidosis-Mediated and Ischemic Neuronal Injury by Promoting ASIC1a (Acid-Sensing Ion Channel 1a) Protein Degradation. Stroke 2019, 50, 2902–2911. [Google Scholar] [CrossRef]
- Schachtrup, C.; Ryu, J.K.; Helmrick, M.J.; Vagena, E.; Galanakis, D.K.; Degen, J.L.; Margolis, R.U.; Akassoglou, K. Fibrinogen Triggers Astrocyte Scar Formation by Promoting the Availability of Active TGF-β after Vascular Damage. J. Neurosci. 2010, 30, 5843–5854. [Google Scholar] [CrossRef]
- Tomolonis, J.A.; Agarwal, S.; Shohet, J.M. Neuroblastoma pathogenesis: Deregulation of embryonic neural crest development. Cell Tissue Res. 2018, 372, 245–262. [Google Scholar] [CrossRef]

| Type of Media | Composition | Note |
|---|---|---|
| Stage I | For every 500ml of high glucose DMEM, add
|
Always add the RA fresh (just before use). Because of its poor aqueous solubility, prepare stocks of 2mM in DMSO (Sigma) and use in the absence of light (Turn off the hood light at the time of use) |
| Stage II |
|
|
| OGD Media |
|
BDNF and B-27 were not included in the media |
| OGD/R Media | 500mL Neurobasal-A media (Gibco 10888-022)
|
BDNF and B-27 were not included in the media |
| Antibodies | Dilutions | Hosts | Manufacturer | |
|---|---|---|---|---|
| Primary Antibodies | β III Tubulin | 1:100 | Mouse | Abcam (Ab78078) |
| Neuronal Nuclei (NeuN) | 1:100 | Rabbit | Proteintech (26975-1-AP) | |
| Growth associated protein 43 (GAP43) | 1:500 | Chicken | Invitrogen (PA5-95660) | |
| Synaptophysin (SYN) | 1:150 | Mouse | Abcam (Ab8049) | |
| Microtubule associated protein 2 (MAP2) | 1:100 | Goat | ||
| Glial fibrillary acidic protein (GFAP)_ | 1:1000 | Chicken | Antibodies (A85307)) | |
| Secondary Antibodies | Alexa 647 | 1:2000 | Chicken | Life technologies (A21449) |
| Alexa 647 | Rabbit | Invitrogen (A31573) | ||
| Alexa 488 | Mouse | Invitrogen (A11001) | ||
| Alexa 488 | Chicken | Invitrogen (A78948) | ||
| Alexa 647 | Anti-mouse | Invitrogen (A31571) | ||
| Alexa 647 | Anti-goat | Invitrogen (A21447) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




