Submitted:
14 April 2026
Posted:
15 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
1.1. Heterogeneity of Autism Spectrum Disorder and the Rationale for a Metabolic Subtype
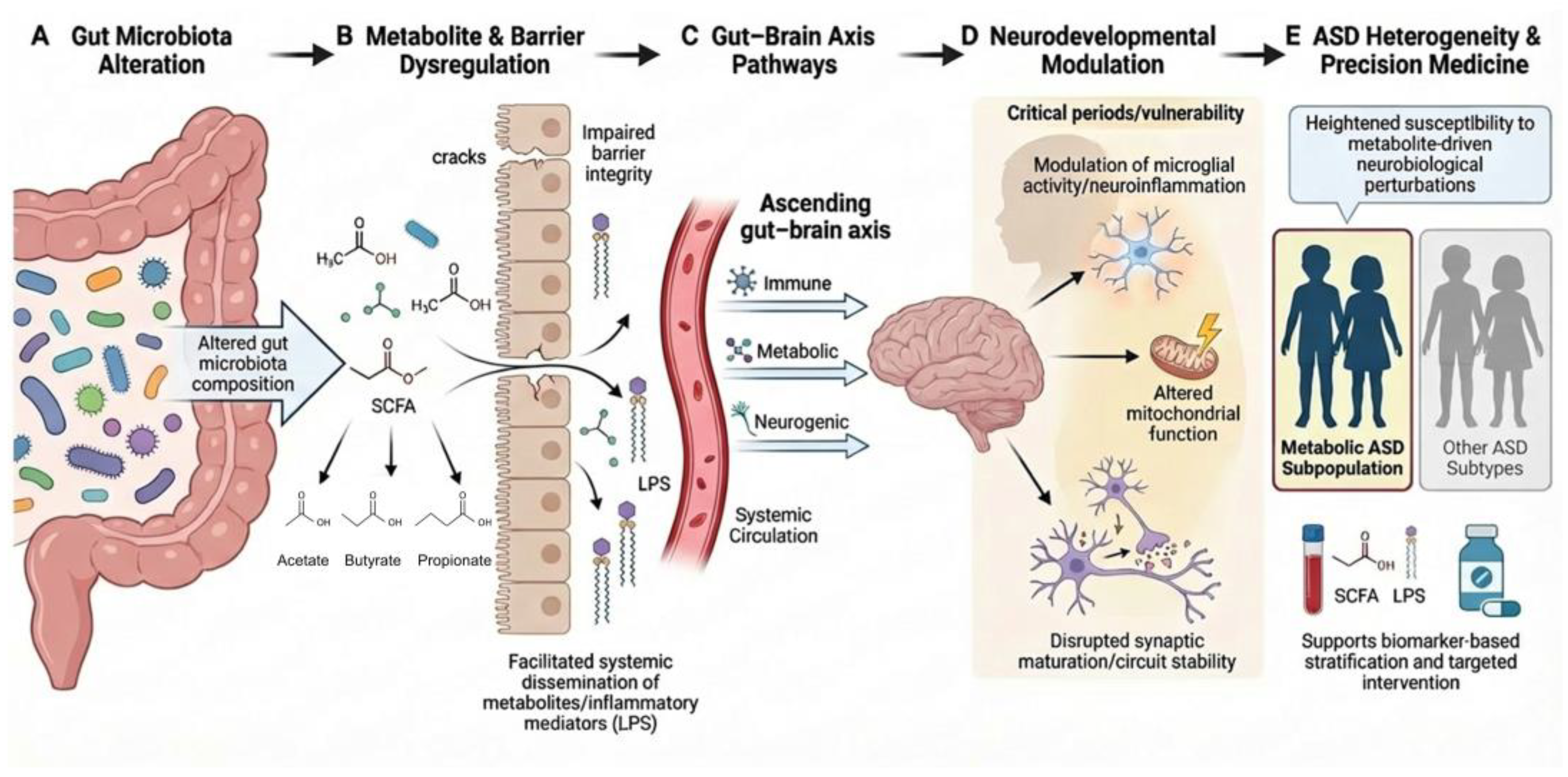
1.2. The Expanded Concept of the Gut–Brain Axis and the Pathophysiological Significance of Gut-Derived Metabolites
1.3. Research Trends on Short-Chain Fatty Acids and the Scope of This Review
2. Physiological Roles of Gut Microbiota and Short-Chain Fatty Acids
2.1. Composition of the Gut Microbiota and Mechanisms of Short-Chain Fatty Acid Production
2.2. Physiological Functions of Major Short-Chain Fatty Acids and Gut–Immune–Neural Homeostasis
2.3. SCFA Signaling Mechanisms: Receptor-Mediated Pathways and Epigenetic Regulation
2.4. Developmental Stage– and Context-Dependent Actions of SCFAs
3. Gut Microbiota and Short-Chain Fatty Acid Imbalance in Autism Spectrum Disorder
3.1. Alterations in Gut Microbiota Composition in Individuals with ASD and Their Functional Implications
3.2. Clinical Heterogeneity of Reported Short-Chain Fatty Acid Alterations in Individuals with ASD
3.3. Selective Associations Between Propionate Alterations and ASD Phenotypes
4. Neuro-pathophysiological Implications of Short-Chain Fatty Acids with a Focus on Propionate
4.1. Absorption, Systemic Distribution, and Central Nervous System Accessibility of Propionate
4.3. Mitochondrial Dysfunction and Energy Metabolic Stress
4.4. Distinguishing Direct Effects from Axis-Mediated Indirect Effects
5. Retrograde Signaling via the Gut–Brain Axis
5.1. Conceptual Framework of Gut Metabolite–Based Bottom-Up Signaling
5.2. Signal Amplification Mediated by Intestinal Barrier Dysfunction and Systemic Inflammation
5.3. Integration of Neuro–Immune–Metabolic Pathways and Central Nervous System Responses
5.4. Limitations of Preclinical Evidence and Clinical Implications
6. Gut-Derived Metabolites and ASD: Integration of Clinical Evidence, Boundaries of Interpretation, and a Research Roadmap for Precision Interventions
6.1. Integrative Interpretation of Clinical Evidence: Signals Are Present, but Not Universal Markers
6.2. Guarding Against Premature Causal Inference: Reframing “Limitations” as Design Requirements
6.3. Therapeutic Implications of the Metabolic ASD Concept: Defining the Target and the Endpoints, Not the Magnitude of Effect
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Choi, J.; Lee, S.; Won, J.; Jin, Y.; Hong, Y.; Hur, T.Y.; Kim, J.H.; Lee, S.R.; Hong, Y. Pathophysiological and neurobehavioral characteristics of a propionic acid-mediated autism-like rat model. PLoS One 2018, 13(2), e0192925. [Google Scholar] [CrossRef]
- Tsai, C.H.; Chen, K.L.; Li, H.J.; Chen, K.H.; Hsu, C.W.; Lu, C.H.; Hsieh, K.Y.; Huang, C.Y. The symptoms of autism including social communication deficits and repetitive and restricted behaviors are associated with different emotional and behavioral problems. Sci Rep. 2020, 10(1), 20509. [Google Scholar] [CrossRef] [PubMed]
- Delehanty, A.; Wetherby, A.M. Coordinated social communication in toddlers with and without autism spectrum disorder during a home observation. Autism Dev Lang Impair 2022, 7, 23969415221121089. [Google Scholar] [CrossRef] [PubMed]
- Lombardo, M.V.; Lai, M.C.; Baron-Cohen, S. Big data approaches to decomposing heterogeneity across the autism spectrum. Mol Psychiatry 2019, 24(10), 1435–1450. [Google Scholar] [CrossRef]
- Reichow, B.; Hume, K.; Barton, E.E.; Boyd, B.A. Early intensive behavioral intervention (EIBI) for young children with autism spectrum disorders (ASD). Cochrane Database Syst Rev 2018, 5(5), CD009260. [Google Scholar] [CrossRef]
- Jobin, A. Varied treatment response in young children with autism: A relative comparison of structured and naturalistic behavioral approaches. Autism 2020, 24(2), 338–351. [Google Scholar] [CrossRef]
- Al-Beltagi, M.; Saeed, N.K.; Bediwy, A.S.; Bediwy, E.A.; Elbeltagi, R. Decoding the genetic landscape of autism: A comprehensive review. World J Clin Pediatr 2024, 13(3), 98468. [Google Scholar] [CrossRef]
- Satterstrom FK, Kosmicki JA, Wang J, Breen MS, De Rubeis S, An JY, Peng M, Collins R, Grove J, Klei L, Stevens C, Reichert J, Mulhern MS, Artomov M, Gerges S, Sheppard B, Xu X, Bhaduri A, Norman U, Brand H, Schwartz G, Nguyen R, Guerrero EE, Dias C; Autism Sequencing Consortium; iPSYCH-Broad Consortium; Betancur C, Cook EH, Gallagher L, Gill M, Sutcliffe JS, Thurm A, Zwick ME, Børglum AD, State MW, Cicek AE, Talkowski ME, Cutler DJ, Devlin B, Sanders SJ, Roeder K, Daly MJ, Buxbaum JD. Large-Scale Exome Sequencing Study Implicates Both Developmental and Functional Changes in the Neurobiology of Autism. Cell. 2020 Feb 6;180(3):568-584.e23. [CrossRef]
- Fu, J.M.; Satterstrom, F.K.; Peng, M.; Brand, H.; Collins, R.L.; Dong, S.; Wamsley, B.; Klei, L.; Wang, L.; Hao, S.P.; Stevens, C.R.; Cusick, C.; Babadi, M.; Banks, E.; Collins, B.; Dodge, S.; Gabriel, S.B.; Gauthier, L.; Lee, S.K.; Liang, L.; Ljungdahl, A.; Mahjani, B.; Sloofman, L.; Smirnov, A.N.; Barbosa, M.; Betancur, C.; Brusco, A.; Chung, B.H.Y.; Cook, E.H.; Cuccaro, M.L.; Domenici, E.; Ferrero, G.B.; Gargus, J.J.; Herman, G.E.; Hertz-Picciotto, I.; Maciel, P.; Manoach, D.S.; Passos-Bueno, M.R.; Persico, A.M.; Renieri, A.; Sutcliffe, J.S.; Tassone, F.; Trabetti, E.; Campos, G.; Cardaropoli, S.; Carli, D.; Chan, M.C.Y.; Fallerini, C.; Giorgio, E.; Girardi, A.C.; Hansen-Kiss, E.; Lee, S.L.; Lintas, C.; Ludena, Y.; Nguyen, R.; Pavinato, L.; Pericak-Vance, M.; Pessah, I.N.; Schmidt, R.J.; Smith, M.; Costa, C.I.S.; Trajkova, S.; Wang, J.Y.T.; Yu, M.H.C.; Cutler, D.J.; De Rubeis, S.; Buxbaum, J.D.; Daly, M.J.; Devlin, B.; Roeder, K.; Sanders, S.J.; Talkowski, M.E.; Autism Sequencing Consortium (ASC); Broad Institute Center for Common Disease Genomics (Broad-CCDG); iPSYCH-BROAD Consortium. Rare coding variation provides insight into the genetic architecture and phenotypic context of autism. Nat Genet. 2022, 54(9), 1320–1331. [Google Scholar] [CrossRef]
- De Rubeis S, He X, Goldberg AP, Poultney CS, Samocha K, Cicek AE, Kou Y, Liu L, Fromer M, Walker S, Singh T, Klei L, Kosmicki J, Shih-Chen F, Aleksic B, Biscaldi M, Bolton PF, Brownfeld JM, Cai J, Campbell NG, Carracedo A, Chahrour MH, Chiocchetti AG, Coon H, Crawford EL, Curran SR, Dawson G, Duketis E, Fernandez BA, Gallagher L, Geller E, Guter SJ, Hill RS, Ionita-Laza J, Jimenz Gonzalez P, Kilpinen H, Klauck SM, Kolevzon A, Lee I, Lei I, Lei J, Lehtimäki T, Lin CF, Ma'ayan A, Marshall CR, McInnes AL, Neale B, Owen MJ, Ozaki N, Parellada M, Parr JR, Purcell S, Puura K, Rajagopalan D, Rehnström K, Reichenberg A, Sabo A, Sachse M, Sanders SJ, Schafer C, Schulte-Rüther M, Skuse D, Stevens C, Szatmari P, Tammimies K, Valladares O, Voran A, Li-San W, Weiss LA, Willsey AJ, Yu TW, Yuen RK; DDD Study; Homozygosity Mapping Collaborative for Autism; UK10K Consortium; Cook EH, Freitag CM, Gill M, Hultman CM, Lehner T, Palotie A, Schellenberg GD, Sklar P, State MW, Sutcliffe JS, Walsh CA, Scherer SW, Zwick ME, Barett JC, Cutler DJ, Roeder K, Devlin B, Daly MJ, Buxbaum JD. Synaptic, transcriptional and chromatin genes disrupted in autism. Nature. 2014 Nov 13;515(7526):209-15. [CrossRef]
- Liu, S.; Xi, H.; Xue, X.; Sun, X.; Huang, H.; Fu, D.; Mi, Y.; He, Y.; Yang, P.; Tang, Y.; Zheng, P. Clostridium butyricum regulates intestinal barrier function via trek1 to improve behavioral abnormalities in mice with autism spectrum disorder. Cell Biosci. 2024, 14(1), 95. [Google Scholar] [CrossRef]
- Nithianantharajah, J.; Balasuriya, G.K.; Franks, A.E.; Hill-Yardin, E.L. Using Animal Models to Study the Role of the Gut-Brain Axis in Autism. Curr Dev Disord Rep. Epub 2017 May 12. 2017, 4(2), 28–36. [Google Scholar] [CrossRef]
- Hsiao, E.Y.; McBride, S.W.; Chow, J.; Mazmanian, S.K.; Patterson, P.H. Modeling an autism risk factor in mice leads to permanent immune dysregulation. Proc Natl Acad Sci U S A 2012, 109(31), 12776–81. [Google Scholar] [CrossRef] [PubMed]
- Choi, G.B.; Yim, Y.S.; Wong, H.; Kim, S.; Kim, H.; Kim, S.V.; Hoeffer, C.A.; Littman, D.R.; Huh, J.R. The maternal interleukin-17a pathway in mice promotes autism-like phenotypes in offspring. Science 2016, 351(6276), 933–9. [Google Scholar] [CrossRef] [PubMed]
- Yardeni, T.; Cristancho, A.G.; McCoy, A.J.; Schaefer, P.M.; McManus, M.J.; Marsh, E.D.; Wallace, D.C. An mtDNA mutant mouse demonstrates that mitochondrial deficiency can result in autism endophenotypes. Proc Natl Acad Sci U S A 2021, 118(6), e2021429118. [Google Scholar] [CrossRef]
- Ahn, Y.; Sabouny, R.; Villa, B.R.; Yee, N.C.; Mychasiuk, R.; Uddin, G.M.; Rho, J.M.; Shutt, T.E. Aberrant Mitochondrial Morphology and Function in the BTBR Mouse Model of Autism Is Improved by Two Weeks of Ketogenic Diet. Int J Mol Sci 2020, 21(9), 3266. [Google Scholar] [CrossRef] [PubMed]
- Leader, G.; Abberton, C.; Cunningham, S.; Gilmartin, K.; Grudzien, M.; Higgins, E.; Joshi, L.; Whelan, S.; Mannion, A. Gastrointestinal Symptoms in Autism Spectrum Disorder: A Systematic Review. Nutrients 2022, 14(7), 1471. [Google Scholar] [CrossRef]
- Cao, X.; Liu, K.; Liu, J.; Liu, Y.W.; Xu, L.; Wang, H.; Zhu, Y.; Wang, P.; Li, Z.; Wen, J.; Shen, C.; Li, M.; Nie, Z.; Kong, X.J. Dysbiotic Gut Microbiota and Dysregulation of Cytokine Profile in Children and Teens With Autism Spectrum Disorder. Front Neurosci 2021, 15, 635925. [Google Scholar] [CrossRef]
- Arteaga-Henríquez, G.; Gisbert, L.; Ramos-Quiroga, J.A. Immunoregulatory and/or Anti-inflammatory Agents for the Management of Core and Associated Symptoms in Individuals with Autism Spectrum Disorder: A Narrative Review of Randomized, Placebo-Controlled Trials. CNS Drugs 2023, 37(3), 215–229. [Google Scholar] [CrossRef]
- Chen, L.; Shi, X.J.; Liu, H.; Mao, X.; Gui, L.N.; Wang, H.; Cheng, Y. Oxidative stress marker aberrations in children with autism spectrum disorder: a systematic review and meta-analysis of 87 studies (N = 9109). Transl Psychiatry 2021, 11(1), 15. [Google Scholar] [CrossRef]
- Main, P.A.; Angley, M.T.; O'Doherty, C.E.; Thomas, P.; Fenech, M. The potential role of the antioxidant and detoxification properties of glutathione in autism spectrum disorders: a systematic review and meta-analysis. Nutr Metab (Lond) 2012, 9, 35. [Google Scholar] [CrossRef]
- Weissman, J.R.; Kelley, R.I.; Bauman, M.L.; Cohen, B.H.; Murray, K.F.; Mitchell, R.L.; Kern, R.L.; Natowicz, M.R. Mitochondrial disease in autism spectrum disorder patients: a cohort analysis. PLoS One 2008, 3(11), e3815. [Google Scholar] [CrossRef]
- Rose, S.; Niyazov, D.M.; Rossignol, D.A.; Goldenthal, M.; Kahler, S.G.; Frye, R.E. Clinical and Molecular Characteristics of Mitochondrial Dysfunction in Autism Spectrum Disorder. Mol Diagn Ther 2018, 22(5), 571–593. [Google Scholar] [CrossRef]
- Carabotti, M.; Scirocco, A.; Maselli, M.A.; Severi, C. The gut-brain axis: interactions between enteric microbiota, central and enteric nervous systems. Ann Gastroenterol 2015, 28(2), 203–209. [Google Scholar]
- Guo, C.; Huo, Y.J.; Li, Y.; Han, Y.; Zhou, D. Gut-brain axis: Focus on gut metabolites short-chain fatty acids. World J Clin Cases 2022, 10(6), 1754–1763. [Google Scholar] [CrossRef]
- Li, C.; Liang, Y.; Qiao, Y. Messengers From the Gut: Gut Microbiota-Derived Metabolites on Host Regulation. Front Microbiol 2022, 13, 863407. [Google Scholar] [CrossRef] [PubMed]
- Deng, W.; Yi, P.; Xiong, Y.; Ying, J.; Lin, Y.; Dong, Y.; Wei, G.; Wang, X.; Hua, F. Gut Metabolites Acting on the Gut-Brain Axis: Regulating the Functional State of Microglia. Aging Dis. 2024, 15(2), 480–502. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, H.; Leyrolle, Q.; Koistinen, V.; Kärkkäinen, O.; Layé, S.; Delzenne, N.; Hanhineva, K. Microbiota-derived metabolites as drivers of gut-brain communication. Gut Microbes 2022, 14(1), 2102878. [Google Scholar] [CrossRef] [PubMed]
- Ang, Z.; Xiong, D.; Wu, M.; Ding, J.L. FFAR2-FFAR3 receptor heteromerization modulates short-chain fatty acid sensing. FASEB J 2018, 32(1), 289–303. [Google Scholar] [CrossRef]
- Tolhurst, G.; Heffron, H.; Lam, Y.S.; Parker, H.E.; Habib, A.M.; Diakogiannaki, E.; Cameron, J.; Grosse, J.; Reimann, F.; Gribble, F.M. Short-chain fatty acids stimulate glucagon-like peptide-1 secretion via the G-protein-coupled receptor FFAR2. Diabetes 2012, 61(2), 364–71. [Google Scholar] [CrossRef]
- Thomas, S.P.; Denu, J.M. Short-chain fatty acids activate acetyltransferase p300. Elife 2021, 10, e72171. [Google Scholar] [CrossRef]
- Yan, H.; Ajuwon, K.M. Butyrate modifies intestinal barrier function in IPEC-J2 cells through a selective upregulation of tight junction proteins and activation of the Akt signaling pathway. PLoS One 2017, 12(6), e0179586. [Google Scholar] [CrossRef]
- Pang, S.; Ren, Z.; Ding, H.; Chan, P. Short-chain fatty acids mediate enteric and central nervous system homeostasis in Parkinson's disease: Innovative therapies and their translation. Neural Regen Res. 2026, 21(3), 938–956. [Google Scholar] [CrossRef]
- Singh, N.; Gurav, A.; Sivaprakasam, S.; Brady, E.; Padia, R.; Shi, H.; Thangaraju, M.; Prasad, P.D.; Manicassamy, S.; Munn, D.H.; Lee, J.R.; Offermanns, S.; Ganapathy, V. Activation of Gpr109a, receptor for niacin and the commensal metabolite butyrate, suppresses colonic inflammation and carcinogenesis. Immunity Epub 2014 Jan 9. 2014, 40(1), 128–39. [Google Scholar] [CrossRef]
- Kaisar, M.M.M.; Pelgrom, L.R.; van der Ham, A.J.; Yazdanbakhsh, M.; Everts, B. Butyrate Conditions Human Dendritic Cells to Prime Type 1 Regulatory T Cells via both Histone Deacetylase Inhibition and G Protein-Coupled Receptor 109A Signaling. Front Immunol 2017, 8, 1429. [Google Scholar] [CrossRef]
- Ren, T.; Li, D.; Sun, F.; Pan, L.; Wang, A.; Li, X.; Bao, Y.; Zhang, M.; Zheng, F.; Yue, H. Indole Propionic Acid Regulates Gut Immunity: Mechanisms of Metabolite-Driven Immunomodulation and Barrier Integrity. J Microbiol Biotechnol. 2025, 35, e2503045. [Google Scholar] [CrossRef]
- Baranwal, G.; Goodlett, B.L.; Arenaz, C.M.; Creed, H.A.; Navaneethabalakrishnan, S.; Rutkowski, J.M.; Alaniz, R.C.; Mitchell, B.M. Indole Propionic Acid Increases T Regulatory Cells and Decreases T Helper 17 Cells and Blood Pressure in Mice with Salt-Sensitive Hypertension. Int J Mol Sci. 2023, 24(11), 9192. [Google Scholar] [CrossRef]
- Hoyles, L.; Snelling, T.; Umlai, U.K.; Nicholson, J.K.; Carding, S.R.; Glen, R.C.; McArthur, S. Microbiome-host systems interactions: protective effects of propionate upon the blood-brain barrier. Microbiome 2018, 6(1), 55. [Google Scholar] [CrossRef] [PubMed]
- Fock, E.; Parnova, R. Mechanisms of Blood-Brain Barrier Protection by Microbiota-Derived Short-Chain Fatty Acids. Cells 2023, 12(4), 657. [Google Scholar] [CrossRef] [PubMed]
- Lagod, P.P.; Naser, S.A. The Role of Short-Chain Fatty Acids and Altered Microbiota Composition in Autism Spectrum Disorder: A Comprehensive Literature Review. Int J Mol Sci. 2023, 24(24), 17432. [Google Scholar] [CrossRef] [PubMed]
- Boets, E.; Deroover, L.; Houben, E.; Vermeulen, K.; Gomand, S.V.; Delcour, J.A.; Verbeke, K. Quantification of in Vivo Colonic Short Chain Fatty Acid Production from Inulin. Nutrients 2015, 7(11), 8916–29. [Google Scholar] [CrossRef]
- Morrison, D.J.; Preston, T. Formation of short chain fatty acids by the gut microbiota and their impact on human metabolism. Gut Microbes 2016, 7(3), 189–200. [Google Scholar] [CrossRef]
- Peng, L.; Li, Z.R.; Green, R.S.; Holzman, I.R.; Lin, J. Butyrate enhances the intestinal barrier by facilitating tight junction assembly via activation of AMP-activated protein kinase in Caco-2 cell monolayers. J Nutr. 2009, 139(9), 1619–25. [Google Scholar] [CrossRef] [PubMed]
- Kim, C.H.; Park, J.; Kim, M. Gut microbiota-derived short-chain Fatty acids, T cells, and inflammation. Immune Netw 2014, 14(6), 277–88. [Google Scholar] [CrossRef] [PubMed]
- Kimura, I.; Inoue, D.; Hirano, K.; Tsujimoto, G. The SCFA Receptor GPR43 and Energy Metabolism. Front Endocrinol (Lausanne) 2014, 5, 85. [Google Scholar] [CrossRef] [PubMed]
- Colombo, A.V.; Sadler, R.K.; Llovera, G.; Singh, V.; Roth, S.; Heindl, S.; Sebastian Monasor, L.; Verhoeven, A.; Peters, F.; Parhizkar, S.; Kamp, F.; Gomez de Aguero, M.; MacPherson, A.J.; Winkler, E.; Herms, J.; Benakis, C.; Dichgans, M.; Steiner, H.; Giera, M.; Haass, C.; Tahirovic, S.; Liesz, A. Microbiota-derived short chain fatty acids modulate microglia and promote Aβ plaque deposition. Elife 2021, 10, e59826. [Google Scholar] [CrossRef]
- Wang, J.; Pan, J.; Chen, H.; Li, Y.; Amakye, W.K.; Liang, J.; Ma, B.; Chu, X.; Mao, L.; Zhang, Z. Fecal Short-Chain Fatty Acids Levels Were Not Associated With Autism Spectrum Disorders in Chinese Children: A Case-Control Study. Front Neurosci 2019, 13, 1216. [Google Scholar] [CrossRef]
- Liu, S.; Li, E.; Sun, Z.; Fu, D.; Duan, G.; Jiang, M.; Yu, Y.; Mei, L.; Yang, P.; Tang, Y.; Zheng, P. Altered gut microbiota and short chain fatty acids in Chinese children with autism spectrum disorder. Sci Rep. 2019, 9(1), 287. [Google Scholar] [CrossRef]
- MacFabe, D.F. Short-chain fatty acid fermentation products of the gut microbiome: implications in autism spectrum disorders. Microb Ecol Health Dis 2012, 23. [Google Scholar] [CrossRef]
- Claesson, M.J.; Cusack, S.; O'Sullivan, O.; Greene-Diniz, R.; de Weerd, H.; Flannery, E.; Marchesi, J.R.; Falush, D.; Dinan, T.; Fitzgerald, G.; Stanton, C.; van Sinderen, D.; O'Connor, M.; Harnedy, N.; O'Connor, K.; Henry, C.; O'Mahony, D.; Fitzgerald, A.P.; Shanahan, F.; Twomey, C.; Hill, C.; Ross, R.P.; O'Toole, P.W. Composition, variability, and temporal stability of the intestinal microbiota of the elderly. Proc Natl Acad Sci U S A 2011, 108 Suppl 1(Suppl 1), 4586–91. [Google Scholar] [CrossRef]
- Rinninella, E.; Raoul, P.; Cintoni, M.; Franceschi, F.; Miggiano, G.A.D.; Gasbarrini, A.; Mele, M.C. What is the Healthy Gut Microbiota Composition? A Changing Ecosystem across Age, Environment, Diet, and Diseases. Microorganisms 2019, 7(1), 14. [Google Scholar] [CrossRef]
- den Besten, G.; van Eunen, K.; Groen, A.K.; Venema, K.; Reijngoud, D.J.; Bakker, B.M. The role of short-chain fatty acids in the interplay between diet, gut microbiota, and host energy metabolism. J Lipid Res. 2013, 54(9), 2325–40. [Google Scholar] [CrossRef]
- Kasubuchi, M.; Hasegawa, S.; Hiramatsu, T.; Ichimura, A.; Kimura, I. Dietary gut microbial metabolites, short-chain fatty acids, and host metabolic regulation. Nutrients 2015, 7(4), 2839–49. [Google Scholar] [CrossRef] [PubMed]
- Price, C.E.; Valls, R.A.; Ramsey, A.R.; Loeven, N.A.; Jones, J.T.; Barrack, K.E.; Schwartzman, J.D.; Royce, D.B.; Cramer, R.A.; Madan, J.C.; Ross, B.D.; Bliska, J.; O'Toole, G.A. Intestinal Bacteroides modulates inflammation, systemic cytokines, and microbial ecology via propionate in a mouse model of cystic fibrosis. mBio 2024, 15(2), e0314423. [Google Scholar] [CrossRef] [PubMed]
- Singh, V.; Lee, G.; Son, H.; Koh, H.; Kim, E.S.; Unno, T.; Shin, J.H. Butyrate producers, "The Sentinel of Gut": Their intestinal significance with and beyond butyrate, and prospective use as microbial therapeutics. Front Microbiol. 2023, 13, 1103836. [Google Scholar] [CrossRef] [PubMed]
- Tsukuda, N.; Yahagi, K.; Hara, T.; Watanabe, Y.; Matsumoto, H.; Mori, H.; Higashi, K.; Tsuji, H.; Matsumoto, S.; Kurokawa, K.; Matsuki, T. Key bacterial taxa and metabolic pathways affecting gut short-chain fatty acid profiles in early life. ISME J 2021, 15(9), 2574–2590. [Google Scholar] [CrossRef]
- Culp, E.J.; Goodman, A.L. Cross-feeding in the gut microbiome: Ecology and mechanisms. Cell Host Microbe 2023, 31(4), 485–499. [Google Scholar] [CrossRef]
- Boets, E.; Gomand, S.V.; Deroover, L.; Preston, T.; Vermeulen, K.; De Preter, V.; Hamer, H.M.; Van den Mooter, G.; De Vuyst, L.; Courtin, C.M.; Annaert, P.; Delcour, J.A.; Verbeke, K.A. Systemic availability and metabolism of colonic-derived short-chain fatty acids in healthy subjects: a stable isotope study. J Physiol. 2017, 595(2), 541–555. [Google Scholar] [CrossRef]
- Cummings, J.H.; Pomare, E.W.; Branch, W.J.; Naylor, C.P.; Macfarlane, G.T. Short chain fatty acids in human large intestine, portal, hepatic and venous blood. Gut 1987, 28(10), 1221–7. [Google Scholar] [CrossRef]
- Yang, W.; Yu, T.; Huang, X.; Bilotta, A.J.; Xu, L.; Lu, Y.; Sun, J.; Pan, F.; Zhou, J.; Zhang, W.; Yao, S.; Maynard, C.L.; Singh, N.; Dann, S.M.; Liu, Z.; Cong, Y. Intestinal microbiota-derived short-chain fatty acids regulation of immune cell IL-22 production and gut immunity. Nat Commun. 2020, 11(1), 4457. [Google Scholar] [CrossRef]
- Silva, Y.P.; Bernardi, A.; Frozza, R.L. The Role of Short-Chain Fatty Acids From Gut Microbiota in Gut-Brain Communication. Front Endocrinol (Lausanne) 2020, 11, 25. [Google Scholar] [CrossRef]
- Yamashita, H.; Maruta, H.; Jozuka, M.; Kimura, R.; Iwabuchi, H.; Yamato, M.; Saito, T.; Fujisawa, K.; Takahashi, Y.; Kimoto, M.; Hiemori, M.; Tsuji, H. Effects of acetate on lipid metabolism in muscles and adipose tissues of type 2 diabetic Otsuka Long-Evans Tokushima Fatty (OLETF) rats. Biosci Biotechnol Biochem. 2009, 73(3), 570–6. [Google Scholar] [CrossRef]
- Frost, G.; Sleeth, M.L.; Sahuri-Arisoylu, M.; Lizarbe, B.; Cerdan, S.; Brody, L.; Anastasovska, J.; Ghourab, S.; Hankir, M.; Zhang, S.; Carling, D.; Swann, J.R.; Gibson, G.; Viardot, A.; Morrison, D.; Louise Thomas, E.; Bell, J.D. The short-chain fatty acid acetate reduces appetite via a central homeostatic mechanism. Nat Commun. 2014, 5, 3611. [Google Scholar] [CrossRef] [PubMed]
- Forte, N.; Marfella, B.; Nicois, A.; Palomba, L.; Paris, D.; Motta, A.; Pina Mollica, M.; Di Marzo, V.; Cristino, L. The short-chain fatty acid acetate modulates orexin/hypocretin neurons: A novel mechanism in gut-brain axis regulation of energy homeostasis and feeding. Biochem Pharmacol Epub 2024 Jun 20. 2024, 226, 116383. [Google Scholar] [CrossRef] [PubMed]
- Donohoe, D.R.; Garge, N.; Zhang, X.; Sun, W.; O'Connell, T.M.; Bunger, M.K.; Bultman, S.J. The microbiome and butyrate regulate energy metabolism and autophagy in the mammalian colon. Cell Metab. 2011, 13(5), 517–26. [Google Scholar] [CrossRef] [PubMed]
- Siavoshian, S.; Segain, J.P.; Kornprobst, M.; Bonnet, C.; Cherbut, C.; Galmiche, J.P.; Blottière, H.M. Butyrate and trichostatin A effects on the proliferation/differentiation of human intestinal epithelial cells: induction of cyclin D3 and p21 expression. Gut 2000, 46(4), 507–14. [Google Scholar] [CrossRef]
- Hu, M.; Alashkar Alhamwe, B.; Santner-Nanan, B.; Miethe, S.; Harb, H.; Renz, H.; Potaczek, D.P.; Nanan, R.K. Short-Chain Fatty Acids Augment Differentiation and Function of Human Induced Regulatory T Cells. Int J Mol Sci 2022, 23(10), 5740. [Google Scholar] [CrossRef]
- Henao-Mejia, J.; Elinav, E.; Jin, C.; Hao, L.; Mehal, W.Z.; Strowig, T.; Thaiss, C.A.; Kau, A.L.; Eisenbarth, S.C.; Jurczak, M.J.; Camporez, J.P.; Shulman, G.I.; Gordon, J.I.; Hoffman, H.M.; Flavell, R.A. Inflammasome-mediated dysbiosis regulates progression of NAFLD and obesity. Nature 2012, 482(7384), 179–85. [Google Scholar] [CrossRef]
- Perry, R.J.; Borders, C.B.; Cline, G.W.; Zhang, X.M.; Alves, T.C.; Petersen, K.F.; Rothman, D.L.; Kibbey, R.G.; Shulman, G.I. Propionate Increases Hepatic Pyruvate Cycling and Anaplerosis and Alters Mitochondrial Metabolism. J Biol Chem. 2016, 291(23), 12161–70. [Google Scholar] [CrossRef]
- Yoshida, H.; Ishii, M.; Akagawa, M. Propionate suppresses hepatic gluconeogenesis via GPR43/AMPK signaling pathway. Arch Biochem Biophys 2019, 672, 108057. [Google Scholar] [CrossRef]
- Heimann, E.; Nyman, M.; Degerman, E. Propionic acid and butyric acid inhibit lipolysis and de novo lipogenesis and increase insulin-stimulated glucose uptake in primary rat adipocytes. Adipocyte 2014, 4(2), 81–8. [Google Scholar] [CrossRef]
- Weitkunat, K.; Schumann, S.; Nickel, D.; Kappo, K.A.; Petzke, K.J.; Kipp, A.P.; Blaut, M.; Klaus, S. Importance of propionate for the repression of hepatic lipogenesis and improvement of insulin sensitivity in high-fat diet-induced obesity. Mol Nutr Food Res. 2016, 60(12), 2611–2621. [Google Scholar] [CrossRef]
- Smith, P.M.; Howitt, M.R.; Panikov, N.; Michaud, M.; Gallini, C.A.; Bohlooly-Y, M.; Glickman, J.N.; Garrett, W.S. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 2013, 341(6145), 569–73. [Google Scholar] [CrossRef] [PubMed]
- Haghikia, A.; Zimmermann, F.; Schumann, P.; Jasina, A.; Roessler, J.; Schmidt, D.; Heinze, P.; Kaisler, J.; Nageswaran, V.; Aigner, A.; Ceglarek, U.; Cineus, R.; Hegazy, A.N.; van der Vorst, E.P.C.; Döring, Y.; Strauch, C.M.; Nemet, I.; Tremaroli, V.; Dwibedi, C.; Kränkel, N.; Leistner, D.M.; Heimesaat, M.M.; Bereswill, S.; Rauch, G.; Seeland, U.; Soehnlein, O.; Müller, D.N.; Gold, R.; Bäckhed, F.; Hazen, S.L.; Haghikia, A.; Landmesser, U. Propionate attenuates atherosclerosis by immune-dependent regulation of intestinal cholesterol metabolism. Eur Heart J 2022, 43(6), 518–533. [Google Scholar] [CrossRef] [PubMed]
- Kimura, I.; Inoue, D.; Maeda, T.; Hara, T.; Ichimura, A.; Miyauchi, S.; Kobayashi, M.; Hirasawa, A.; Tsujimoto, G. Short-chain fatty acids and ketones directly regulate sympathetic nervous system via G protein-coupled receptor 41 (GPR41). Proc Natl Acad Sci U S A 2011, 108(19), 8030–5. [Google Scholar] [CrossRef] [PubMed]
- Ang, Z.; Er, J.Z.; Tan, N.S.; Lu, J.; Liou, Y.C.; Grosse, J.; Ding, J.L. Human and mouse monocytes display distinct signalling and cytokine profiles upon stimulation with FFAR2/FFAR3 short-chain fatty acid receptor agonists. Sci Rep. 2016, 6, 34145. [Google Scholar] [CrossRef]
- Lavoie, S.; Chun, E.; Bae, S.; Brennan, C.A.; Gallini Comeau, C.A.; Lang, J.K.; Michaud, M.; Hoveyda, H.R.; Fraser, G.L.; Fuller, M.H.; Layden, B.T.; Glickman, J.N.; Garrett, W.S. Expression of Free Fatty Acid Receptor 2 by Dendritic Cells Prevents Their Expression of Interleukin 27 and Is Required for Maintenance of Mucosal Barrier and Immune Response Against Colorectal Tumors in Mice. Gastroenterology 2020, 158(5), 1359–1372.e9. [Google Scholar] [CrossRef]
- Licciardi, P.V.; Ververis, K.; Karagiannis, T.C. Histone deacetylase inhibition and dietary short-chain Fatty acids. ISRN Allergy 2011, 2011, 869647. [Google Scholar] [CrossRef]
- Folkerts, J.; Redegeld, F.; Folkerts, G.; Blokhuis, B.; van den Berg, M.P.M.; de Bruijn, M.J.W.; van IJcken, W.F.J.; Junt, T.; Tam, S.Y.; Galli, S.J.; Hendriks, R.W.; Stadhouders, R.; Maurer, M. Butyrate inhibits human mast cell activation via epigenetic regulation of FcεRI-mediated signaling. Allergy 2020, 75(8), 1966–1978. [Google Scholar] [CrossRef]
- Park, J.; Kim, M.; Kang, S.G.; Jannasch, A.H.; Cooper, B.; Patterson, J.; Kim, C.H. Short-chain fatty acids induce both effector and regulatory T cells by suppression of histone deacetylases and regulation of the mTOR-S6K pathway. Mucosal Immunol 2015, 8(1), 80–93. [Google Scholar] [CrossRef]
- Lin, M.Y.; de Zoete, M.R.; van Putten, J.P.; Strijbis, K. Redirection of Epithelial Immune Responses by Short-Chain Fatty Acids through Inhibition of Histone Deacetylases. Front Immunol 2015, 6, 554. [Google Scholar] [CrossRef]
- Otten, B.M.J.; Sthijns, M.M.J.P.E.; Troost, F.J. A Combination of Acetate, Propionate, and Butyrate Increases Glucose Uptake in C2C12 Myotubes. Nutrients 2023, 15(4), 946. [Google Scholar] [CrossRef]
- Saleri, R.; Borghetti, P.; Ravanetti, F.; Cavalli, V.; Ferrari, L.; De Angelis, E.; Andrani, M.; Martelli, P. Effects of different short-chain fatty acids (SCFA) on gene expression of proteins involved in barrier function in IPEC-J2. Porcine Health Manag 2022, 8(1), 21. [Google Scholar] [CrossRef] [PubMed]
- Park J, Wang Q, Wu Q, Mao-Draayer Y, Kim CH. Bidirectional regulatory potentials of short-chain fatty acids and their G-protein-coupled receptors in autoimmune neuroinflammation. Sci Rep. 2019 Jun 20;9(1):8837. https://doi.org/10.1038/s41598-019-45311-y. Erratum in: Sci Rep. 2019 Nov 20;9(1):17511. [CrossRef]
- Corrêa-Oliveira, R.; Fachi, J.L.; Vieira, A.; Sato, F.T.; Vinolo, M.A. Regulation of immune cell function by short-chain fatty acids. Clin Transl Immunology 2016, 5(4), e73. [Google Scholar] [CrossRef] [PubMed]
- Cusick, S.E.; Georgieff, M.K. The Role of Nutrition in Brain Development: The Golden Opportunity of the "First 1000 Days. J Pediatr. 2016, 175, 16–21. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.; Lee-Sarwar, K.; Kelly, R.S.; Lasky-Su, J.A.; Litonjua, A.A.; Weiss, S.T.; Liu, Y.Y. Identifying the critical time window for the association of the early-life gut microbiome and metabolome with childhood neurodevelopment. medRxiv 2022. [Google Scholar] [CrossRef]
- Kasani, P.H.; Yun, C.H.; Cho, K.H.; Jeong, S.J. Neonatal gut microbiota stratification and identification of SCFA-associated microbial subgroups using unsupervised clustering and machine learning classification. Front Microbiol 2025, 16, 1668451. [Google Scholar] [CrossRef]
- Ruohtula, T.; de Goffau, M.C.; Nieminen, J.K.; Honkanen, J.; Siljander, H.; Hämäläinen, A.M.; Peet, A.; Tillmann, V.; Ilonen, J.; Niemelä, O.; Welling, G.W.; Knip, M.; Harmsen, H.J.; Vaarala, O. Maturation of Gut Microbiota and Circulating Regulatory T Cells and Development of IgE Sensitization in Early Life. Front Immunol 2019, 10, 2494. [Google Scholar] [CrossRef]
- Beghetti, I.; Barone, M.; Brigidi, P.; Sansavini, A.; Corvaglia, L.; Aceti, A.; Turroni, S. Early-life gut microbiota and neurodevelopment in preterm infants: a narrative review. Front Nutr. 2023, 10, 1241303. [Google Scholar] [CrossRef]
- Kwon, H.K.; Choi, G.B.; Huh, J.R. Maternal inflammation and its ramifications on fetal neurodevelopment. Trends Immunol. 2022, 43(3), 230–244. [Google Scholar] [CrossRef]
- Hsu, C.Y.; Khachatryan, L.G.; Younis, N.K.; Mustafa, M.A.; Ahmad, N.; Athab, Z.H.; Polyanskaya, A.V.; Kasanave, E.V.; Mirzaei, R.; Karampoor, S. Microbiota-derived short chain fatty acids in pediatric health and diseases: from gut development to neuroprotection. Front Microbiol 2024, 15, 1456793. [Google Scholar] [CrossRef]
- Hernández-Martínez, C.; Canals, J.; Voltas, N.; Martín-Luján, F.; Arija, V. Circulating Levels of Short-Chain Fatty Acids during Pregnancy and Infant Neurodevelopment. Nutrients 2022, 14(19), 3946. [Google Scholar] [CrossRef]
- MacFabe, D.F. Enteric short-chain fatty acids: microbial messengers of metabolism, mitochondria, and mind: implications in autism spectrum disorders. Microb Ecol Health Dis. 2015, 26, 28177. [Google Scholar] [CrossRef]
- Zhang, D.; Zhou, M.; Qiu, Y.; Xu, H.; Liu, H.; Liu, Y.; Xie, L. Cross-generational mechanisms of maternal gut microbiota in modulating offspring autism spectrum disorder risk: from the gut-brain axis to translational challenges in precision interventions. Front Aging Neurosci 2025, 17, 1642240. [Google Scholar] [CrossRef] [PubMed]
- Cao, Q.; Shen, M.; Li, R.; Liu, Y.; Zeng, Z.; Zhou, J.; Niu, D.; Zhang, Q.; Wang, R.; Yao, J.; Zhang, G. Elucidating the specific mechanisms of the gut-brain axis: the short-chain fatty acids-microglia pathway. J Neuroinflammation 2025, 22(1), 133. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Guo, W.; Li, S.; Sun, B.; Li, N.; Xie, D.; Dong, Z.; Luo, D.; Chen, W.; Fu, W.; Zheng, J.; Zhu, J. Alteration of the gut microbiota profile in children with autism spectrum disorder in China. Front Microbiol 2024, 14, 1326870. [Google Scholar] [CrossRef] [PubMed]
- Lou, M.; Cao, A.; Jin, C.; Mi, K.; Xiong, X.; Zeng, Z.; Pan, X.; Qie, J.; Qiu, S.; Niu, Y.; Liang, H.; Liu, Y.; Chen, L.; Liu, Z.; Zhao, Q.; Qiu, X.; Jin, Y.; Sheng, X.; Hu, Z.; Jin, G.; Liu, J.; Liu, X.; Wang, Y. Deviated and early unsustainable stunted development of gut microbiota in children with autism spectrum disorder. Gut 2022, 71(8), 1588–1599. [Google Scholar] [CrossRef]
- Chen, Q.; Xu, J.; Yang, J.; Qin, X.; Fan, J.; Ke, H.; Yang, Z.; Zheng, W.; Li, X.; Huang, L.; Ning, W. Gut microbiota analysis in children with autism spectrum disorder and their family members. Sci Rep. 2025, 15(1), 44282. [Google Scholar] [CrossRef]
- Iglesias-Vázquez, L.; Van Ginkel Riba, G.; Arija, V.; Canals, J. Composition of Gut Microbiota in Children with Autism Spectrum Disorder: A Systematic Review and Meta-Analysis. Nutrients 2020, 12(3), 792. [Google Scholar] [CrossRef]
- Li, H.; Guo, W.; Li, S.; Sun, B.; Li, N.; Xie, D.; Dong, Z.; Luo, D.; Chen, W.; Fu, W.; Zheng, J.; Zhu, J. Alteration of the gut microbiota profile in children with autism spectrum disorder in China. Front Microbiol 2024, 14, 1326870. [Google Scholar] [CrossRef]
- Kang, D.W.; Park, J.G.; Ilhan, Z.E.; Wallstrom, G.; Labaer, J.; Adams, J.B.; Krajmalnik-Brown, R. Reduced incidence of Prevotella and other fermenters in intestinal microflora of autistic children. PLoS One 2013, 8(7), e68322. [Google Scholar] [CrossRef]
- Retuerto, M.; Al-Shakhshir, H.; Herrada, J.; McCormick, T.S.; Ghannoum, M.A. Analysis of Gut Bacterial and Fungal Microbiota in Children with Autism Spectrum Disorder and Their Non-Autistic Siblings. Nutrients 2024, 16(17), 3004. [Google Scholar] [CrossRef]
- Coretti L, Paparo L, Riccio MP, Amato F, Cuomo M, Natale A, Borrelli L, Corrado G, Comegna M, Buommino E, Castaldo G, Bravaccio C, Chiariotti L, Berni Canani R, Lembo F. Gut Microbiota Features in Young Children With Autism Spectrum Disorders. Front Microbiol. 2018 Dec 19;9:3146. https://doi.org/10.3389/fmicb.2018.03146. Erratum in: Front Microbiol. 2019 May 03;10:920. [CrossRef]
- Darwesh, M.K.; Bakr, W.; Omar, T.E.I.; El-Kholy, M.A.; Azzam, N.F. Unraveling the relative abundance of psychobiotic bacteria in children with Autism Spectrum Disorder. Sci Rep. 2024, 14(1), 24321. [Google Scholar] [CrossRef]
- Xiang, F.; Zhang, M.; Wei, X.; Chang, J. Gut microbiota composition and phylogenetic analysis in autism spectrum disorder: a comparative study. Front Psychiatry 2025, 16, 1609638. [Google Scholar] [CrossRef]
- Yap CX, Henders AK, Alvares GA, Wood DLA, Krause L, Tyson GW, Restuadi R, Wallace L, McLaren T, Hansell NK, Cleary D, Grove R, Hafekost C, Harun A, Holdsworth H, Jellett R, Khan F, Lawson LP, Leslie J, Frenk ML, Masi A, Mathew NE, Muniandy M, Nothard M, Miller JL, Nunn L, Holtmann G, Strike LT, de Zubicaray GI, Thompson PM, McMahon KL, Wright MJ, Visscher PM, Dawson PA, Dissanayake C, Eapen V, Heussler HS, McRae AF, Whitehouse AJO, Wray NR, Gratten J. Autism-related dietary preferences mediate autism-gut microbiome associations. Cell. 2021 Nov 24;184(24):5916-5931.e17. https://doi.org/10.1016/j.cell.2021.10.015. Epub 2021 Nov 11. Erratum in: Cell. 2024 Jan 18;187(2):495-510. Cell. [CrossRef]
- Li, H.; Liu, C.; Huang, S.; Wang, X.; Cao, M.; Gu, T.; Ou, X.; Pan, S.; Lin, Z.; Wang, X.; Zhu, Y.; Jing, J. Multi-omics analyses demonstrate the modulating role of gut microbiota on the associations of unbalanced dietary intake with gastrointestinal symptoms in children with autism spectrum disorder. Gut Microbes 2023, 15(2), 2281350. [Google Scholar] [CrossRef] [PubMed]
- Chavira, A.; Wang, E.H.J.; Mills, R.H. Meta-analysis of the autism gut microbiome identifies factors influencing study discrepancies and machine learning classification. bioRxiv. 2022.
- Vernocchi, P.; Ristori, M.V.; Guerrera, S.; Guarrasi, V.; Conte, F.; Russo, A.; Lupi, E.; Albitar-Nehme, S.; Gardini, S.; Paci, P.; Ianiro, G.; Vicari, S.; Gasbarrini, A.; Putignani, L. Gut Microbiota Ecology and Inferred Functions in Children With ASD Compared to Neurotypical Subjects. Front Microbiol 2022, 13, 871086. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Gong, X.; Hu, B.; Lin, L.; Lin, X.; Gong, W.; Zhang, B.; Cao, M.; Xu, Y.; Xia, R.; Zheng, G.; Wu, S.; Zhang, Y. Altered Gut Microbiota and Short-chain Fatty Acids in Chinese Children with Constipated Autism Spectrum Disorder. Sci Rep. 2023, 13(1), 19103. [Google Scholar] [CrossRef] [PubMed]
- Deng, W.; Wang, S.; Li, F.; Wang, F.; Xing, Y.P.; Li, Y.; Lv, Y.; Ke, H.; Li, Z.; Lv, P.J.; Hao, H.; Chen, Y.; Xiao, X. Gastrointestinal symptoms have a minor impact on autism spectrum disorder and associations with gut microbiota and short-chain fatty acids. Front Microbiol 2022, 13, 1000419. [Google Scholar] [CrossRef]
- Manokasemsan, W.; Jariyasopit, N.; Poungsombat, P.; Kaewnarin, K.; Wanichthanarak, K.; Kurilung, A.; Duangkumpha, K.; Limjiasahapong, S.; Pomyen, Y.; Chaiteerakij, R.; Tansawat, R.; Srisawat, C.; Sirivatanauksorn, Y.; Sirivatanauksorn, V.; Khoomrung, S. Quantifying fecal and plasma short-chain fatty acids in healthy Thai individuals. Comput Struct Biotechnol J 2024, 23, 2163–2172. [Google Scholar] [CrossRef]
- Needham, B.D.; Adame, M.D.; Serena, G.; Rose, D.R.; Preston, G.M.; Conrad, M.C.; Campbell, A.S.; Donabedian, D.H.; Fasano, A.; Ashwood, P.; Mazmanian, S.K. Plasma and Fecal Metabolite Profiles in Autism Spectrum Disorder. Biol Psychiatry 2021, 89(5), 451–462. [Google Scholar] [CrossRef]
- Kang, D.W.; Adams, J.B.; Vargason, T.; Santiago, M.; Hahn, J.; Krajmalnik-Brown, R. Distinct Fecal and Plasma Metabolites in Children with Autism Spectrum Disorders and Their Modulation after Microbiota Transfer Therapy. mSphere 2020, 5(5), e00314-20. [Google Scholar] [CrossRef]
- Trivedi, N.; Erickson, H.E.; Bala, V.; Chhonker, Y.S.; Murry, D.J. A Concise Review of Liquid Chromatography-Mass Spectrometry-Based Quantification Methods for Short Chain Fatty Acids as Endogenous Biomarkers. Int J Mol Sci. 2022, 23(21), 13486. [Google Scholar] [CrossRef]
- Rohde, J.K.; Fuh, M.M.; Evangelakos, I.; Pauly, M.J.; Schaltenberg, N.; Siracusa, F.; Gagliani, N.; Tödter, K.; Heeren, J.; Worthmann, A. A Gas Chromatography Mass Spectrometry-Based Method for the Quantification of Short Chain Fatty Acids. Metabolites 2022, 12(2), 170. [Google Scholar] [CrossRef] [PubMed]
- Scortichini, S.; Boarelli, M.C.; Silvi, S.; Fiorini, D. Development and validation of a GC-FID method for the analysis of short chain fatty acids in rat and human faeces and in fermentation fluids. J Chromatogr B Analyt Technol Biomed Life Sci. 2020, 1143, 121972. [Google Scholar] [CrossRef] [PubMed]
- Liebisch, G.; Ecker, J.; Roth, S.; Schweizer, S.; Öttl, V.; Schött, H.F.; Yoon, H.; Haller, D.; Holler, E.; Burkhardt, R.; Matysik, S. Quantification of Fecal Short Chain Fatty Acids by Liquid Chromatography Tandem Mass Spectrometry-Investigation of Pre-Analytic Stability. Biomolecules 2019, 9(4), 121. [Google Scholar] [CrossRef]
- Levkova, M.; Chervenkov, T.; Pancheva, R. Genus-Level Analysis of Gut Microbiota in Children with Autism Spectrum Disorder: A Mini Review. Children (Basel) 2023, 10(7), 1103. [Google Scholar] [CrossRef] [PubMed]
- Abdelli, L.S.; Samsam, A.; Naser, S.A. Propionic Acid Induces Gliosis and Neuro-inflammation through Modulation of PTEN/AKT Pathway in Autism Spectrum Disorder. Sci Rep. 2019, 9(1), 8824. [Google Scholar] [CrossRef]
- Thomas, R.H.; Meeking, M.M.; Mepham, J.R.; Tichenoff, L.; Possmayer, F.; Liu, S.; MacFabe, D.F. The enteric bacterial metabolite propionic acid alters brain and plasma phospholipid molecular species: further development of a rodent model of autism spectrum disorders. J Neuroinflammation 2012, 9, 153. [Google Scholar] [CrossRef]
- Saunders, N.R.; Liddelow, S.A.; Dziegielewska, K.M. Barrier mechanisms in the developing brain. Front Pharmacol 2012, 3, 46. [Google Scholar] [CrossRef]
- Saunders, N.R.; Dziegielewska, K.M.; Møllgård, K.; Habgood, M.D. Physiology and molecular biology of barrier mechanisms in the fetal and neonatal brain. J Physiol. 2018, 596(23), 5723–5756. [Google Scholar] [CrossRef]
- Fleming, A.; Diekmann, H.; Goldsmith, P. Functional characterisation of the maturation of the blood-brain barrier in larval zebrafish. PLoS One 2013, 8(10), e77548. [Google Scholar] [CrossRef]
- Verscheijden, L.F.M.; van Hattem, A.C.; Pertijs, J.C.L.M.; de Jongh, C.A.; Verdijk, R.M.; Smeets, B.; Koenderink, J.B.; Russel, F.G.M.; de Wildt, S.N. Developmental patterns in human blood-brain barrier and blood-cerebrospinal fluid barrier ABC drug transporter expression. Histochem Cell Biol. 2020, 154(3), 265–273. [Google Scholar] [CrossRef]
- Wang, C.; He, T.; Qin, J.; Jiao, J.; Ji, F. The roles of immune factors in neurodevelopment. Front Cell Neurosci 2025, 19, 1451889. [Google Scholar] [CrossRef]
- Maffezzini, C.; Calvo-Garrido, J.; Wredenberg, A.; Freyer, C. Metabolic regulation of neurodifferentiation in the adult brain. Cell Mol Life Sci 2020, 77(13), 2483–2496. [Google Scholar] [CrossRef]
- Wang, Q.; Yang, Q.; Liu, X. The microbiota-gut-brain axis and neurodevelopmental disorders. Protein Cell 2023, 14(10), 762–775. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.L.; Millischer, V.; Rodin, S.; MacFabe, D.F.; Villaescusa, J.C.; Lavebratt, C. Enteric short-chain fatty acids promote proliferation of human neural progenitor cells. J Neurochem. 2020, 154(6), 635–646. [Google Scholar] [CrossRef] [PubMed]
- Erny, D.; Hrabě de Angelis, A.L.; Jaitin, D.; Wieghofer, P.; Staszewski, O.; David, E.; Keren-Shaul, H.; Mahlakoiv, T.; Jakobshagen, K.; Buch, T.; Schwierzeck, V.; Utermöhlen, O.; Chun, E.; Garrett, W.S.; McCoy, K.D.; Diefenbach, A.; Staeheli, P.; Stecher, B.; Amit, I.; Prinz, M. Host microbiota constantly control maturation and function of microglia in the CNS. Nat Neurosci 2015, 18(7), 965–77. [Google Scholar] [CrossRef] [PubMed]
- Kimura, I.; Miyamoto, J.; Ohue-Kitano, R.; Watanabe, K.; Yamada, T.; Onuki, M.; Aoki, R.; Isobe, Y.; Kashihara, D.; Inoue, D.; Inaba, A.; Takamura, Y.; Taira, S.; Kumaki, S.; Watanabe, M.; Ito, M.; Nakagawa, F.; Irie, J.; Kakuta, H.; Shinohara, M.; Iwatsuki, K.; Tsujimoto, G.; Ohno, H.; Arita, M.; Itoh, H.; Hase, K. Maternal gut microbiota in pregnancy influences offspring metabolic phenotype in mice. Science 2020, 367(6481), eaaw8429. [Google Scholar] [CrossRef]
- Liu, X.; Li, X.; Xia, B.; Jin, X.; Zeng, Z.; Yan, S.; Li, L.; Yuan, S.; Zhao, S.; Dai, X.; Yin, F.; Cadenas, E.; Liu, R.H.; Zhao, B.; Hou, M.; Liu, Z.; Liu, X. Gut microbiota mediates high-diet alleviation of maternal obesity-induced cognitive and social deficits in offspring. bioRxiv 2020. [Google Scholar]
- Schafer, D.P.; Lehrman, E.K.; Kautzman, A.G.; Koyama, R.; Mardinly, A.R.; Yamasaki, R.; Ransohoff, R.M.; Greenberg, M.E.; Barres, B.A.; Stevens, B. Microglia sculpt postnatal neural circuits in an activity and complement-dependent manner. Neuron 2012, 74(4), 691–705. [Google Scholar] [CrossRef]
- Schafer, D.P.; Lehrman, E.K.; Stevens, B. The "quad-partite" synapse: microglia-synapse interactions in the developing and mature CNS. Glia 2013, 61(1), 24–36. [Google Scholar] [CrossRef]
- Lobzhanidze, G.; Japaridze, N.; Lordkipanidze, T.; Rzayev, F.; MacFabe, D.; Zhvania, M. Behavioural and brain ultrastructural changes following the systemic administration of propionic acid in adolescent male rats. Further development of a rodent model of autism. Int J Dev Neurosci 2020, 80(2), 139–156. [Google Scholar] [CrossRef]
- Zhvania, M.G.; Lobzhanidze, G.; Pochkhidze, N.; Japaridze, N.; Tchelidze, P.; Rzayev, F.; Gasimov, E. Propionic acid affects the synaptic architecture of rat hippocampus and prefrontal cortex. Micron 2024, 181, 103624. [Google Scholar] [CrossRef] [PubMed]
- Lagod, P.P.; Abdelli, L.S.; Naser, S.A. An In Vivo Model of Propionic Acid-Rich Diet-Induced Gliosis and Neuro-Inflammation in Mice (FVB/N-Tg(GFAPGFP)14Mes/J): A Potential Link to Autism Spectrum Disorder. Int J Mol Sci. 2024, 25(15), 8093. [Google Scholar] [CrossRef] [PubMed]
- Mehan, S.; Rahi, S.; Tiwari, A.; Kapoor, T.; Rajdev, K.; Sharma, R.; Khera, H.; Kosey, S.; Kukkar, U.; Dudi, R. Adenylate cyclase activator forskolin alleviates intracerebroventricular propionic acid-induced mitochondrial dysfunction of autistic rats. Neural Regen Res. 2020, 15(6), 1140–1149. [Google Scholar] [CrossRef] [PubMed]
- Frye, R.E.; Rose, S.; Chacko, J.; Wynne, R.; Bennuri, S.C.; Slattery, J.C.; Tippett, M.; Delhey, L.; Melnyk, S.; Kahler, S.G.; MacFabe, D.F. Modulation of mitochondrial function by the microbiome metabolite propionic acid in autism and control cell lines. Transl Psychiatry 2016, 6(10), e927. [Google Scholar] [CrossRef]
- Schwab, M.A.; Sauer, S.W.; Okun, J.G.; Nijtmans, L.G.; Rodenburg, R.J.; van den Heuvel, L.P.; Dröse, S.; Brandt, U.; Hoffmann, G.F.; Ter Laak, H.; Kölker, S.; Smeitink, J.A. Secondary mitochondrial dysfunction in propionic aciduria: a pathogenic role for endogenous mitochondrial toxins. Biochem J 2006, 398(1), 107–12. [Google Scholar] [CrossRef]
- Chen Q, Qiao Y, Xu XJ, You X, Tao Y. Urine Organic Acids as Potential Biomarkers for Autism-Spectrum Disorder in Chinese Children. Front Cell Neurosci. 2019 Apr 30;13:150. https://doi.org/10.3389/fncel.2019.00150. Erratum in: Front Cell Neurosci. 2019 Aug 22;13:388. [CrossRef]
- Kurochkin, I.; Khrameeva, E.; Tkachev, A.; Stepanova, V.; Vanyushkina, A.; Stekolshchikova, E.; Li, Q.; Zubkov, D.; Shichkova, P.; Halene, T.; Willmitzer, L.; Giavalisco, P.; Akbarian, S.; Khaitovich, P. Metabolome signature of autism in the human prefrontal cortex. Commun Biol. 2019, 2, 234. [Google Scholar] [CrossRef]
- Oh, M.; Kim, S.A.; Yoo, H.J. Higher Lactate Level and Lactate-to-Pyruvate Ratio in Autism Spectrum Disorder. Exp Neurobiol 2020, 29(4), 314–322. [Google Scholar] [CrossRef]
- Rossignol, D.A.; Frye, R.E. Mitochondrial dysfunction in autism spectrum disorders: a systematic review and meta-analysis. Mol Psychiatry Epub 2011 Jan 25. 2012, 17(3), 290–314. [Google Scholar] [CrossRef]
- Giulivi, C.; Zhang, Y.F.; Omanska-Klusek, A.; Ross-Inta, C.; Wong, S.; Hertz-Picciotto, I.; Tassone, F.; Pessah, I.N. Mitochondrial dysfunction in autism. JAMA 2010, 304(21), 2389–96. [Google Scholar] [CrossRef]
- Féron, F.; Caillol, D.; Fourel, L.; Leimkuhler, S.; Iranzo, O.; Gepner, B.; Guiraudie-Capraz, G. An impaired glycolysis induces ATP deficiency and reduced cell respiration in stem cells of patients with autism spectrum disorders. Sci Rep. 2025, 15(1), 38353. [Google Scholar] [CrossRef]
- Longo, N.; Price, L.B.; Gappmaier, E.; Cantor, N.L.; Ernst, S.L.; Bailey, C.; Pasquali, M. Anaplerotic therapy in propionic acidemia. Mol Genet Metab 2017, 122(1-2), 51–59. [Google Scholar] [CrossRef]
- Go S, Kramer TT, Verhoeven AJ, Oude Elferink RPJ, Chang JC. The extracellular lactate-to-pyruvate ratio modulates the sensitivity to oxidative stress-induced apoptosis via the cytosolic NADH/NAD+ redox state. Apoptosis. 2021 Feb;26(1-2):38-51. https://doi.org/10.1007/s10495-020-01648-8. Epub 2020 Nov 23. Erratum in: Apoptosis. 2024 Dec;29(11-12):1858-1859. [CrossRef]
- Loh, J.S.; Mak, W.Q.; Tan, L.K.S.; Ng, C.X.; Chan, H.H.; Yeow, S.H.; Foo, J.B.; Ong, Y.S.; How, C.W.; Khaw, K.Y. Microbiota-gut-brain axis and its therapeutic applications in neurodegenerative diseases. Signal Transduct Target Ther 2024, 9(1), 37. [Google Scholar] [CrossRef] [PubMed]
- Munir, M.U.; Ali, S.A.; Chung, K.H.K.; Kakinen, A.; Javed, I.; Davis, T.P. Reverse engineering the Gut-Brain Axis and microbiome-metabolomics for symbiotic/pathogenic balance in neurodegenerative diseases. Gut Microbes 2024, 16(1), 2422468. [Google Scholar] [CrossRef] [PubMed]
- Bonaz, B.; Bazin, T.; Pellissier, S. The Vagus Nerve at the Interface of the Microbiota-Gut-Brain Axis. Front Neurosci 2018, 12, 49. [Google Scholar] [CrossRef]
- Jameson KG, Kazmi SA, Son C, Mazdeyasnan D, Leshan E, Vuong HE, Paramo J, Lopez-Romero A, Yang L, Schweizer FE, Hsiao EY. Vagal interoception of microbial metabolites from the small intestinal lumen. bioRxiv [Preprint]. 2023 Dec 19:2023.12.18.572257. doi: 10.1101/2023.12.18.572257. Update in: iScience. 2024 Dec 27;28(2):111699. [CrossRef]
- Huo, J.Y.; Jiang, W.Y.; Yin, T.; Xu, H.; Lyu, Y.T.; Chen, Y.Y.; Chen, M.; Geng, J.; Jiang, Z.X.; Shan, Q.J. Intestinal Barrier Dysfunction Exacerbates Neuroinflammation via the TLR4 Pathway in Mice With Heart Failure. Front Physiol. 2021, 12, 712338. [Google Scholar] [CrossRef]
- Barbara G, Barbaro MR, Fuschi D, Palombo M, Falangone F, Cremon C, Marasco G, Stanghellini V. Inflammatory and Microbiota-Related Regulation of the Intestinal Epithelial Barrier. Front Nutr. 2021 Sep 13;8:718356. https://doi.org/10.3389/fnut.2021.718356. Erratum in: Front Nutr. 2021 Nov 01;8:790387. [CrossRef]
- Yu, L.C.; Wang, J.T.; Wei, S.C.; Ni, Y.H. Host-microbial interactions and regulation of intestinal epithelial barrier function: From physiology to pathology. World J Gastrointest Pathophysiol 2012, 3(1), 27–43. [Google Scholar] [CrossRef]
- Kang, G.G.; Trevaskis, N.L.; Murphy, A.J.; Febbraio, M.A. Diet-induced gut dysbiosis and inflammation: Key drivers of obesity-driven NASH. iScience 2022, 26(1), 105905. [Google Scholar] [CrossRef]
- Crawford, C.K.; Lopez Cervantes, V.; Quilici, M.L.; Armién, A.G.; Questa, M.; Matloob, M.S.; Huynh, L.D.; Beltran, A.; Karchemskiy, S.J.; Crakes, K.R.; Kol, A. Inflammatory cytokines directly disrupt the bovine intestinal epithelial barrier. Sci Rep. 2022, 12(1), 14578. [Google Scholar] [CrossRef]
- Ghosh, S.S.; Wang, J.; Yannie, P.J.; Ghosh, S. Intestinal Barrier Dysfunction, LPS Translocation, and Disease Development. J Endocr Soc. 2020, 4(2), bvz039. [Google Scholar] [CrossRef]
- Thaiss, C.A.; Levy, M.; Grosheva, I.; Zheng, D.; Soffer, E.; Blacher, E.; Braverman, S.; Tengeler, A.C.; Barak, O.; Elazar, M.; Ben-Zeev, R.; Lehavi-Regev, D.; Katz, M.N.; Pevsner-Fischer, M.; Gertler, A.; Halpern, Z.; Harmelin, A.; Aamar, S.; Serradas, P.; Grosfeld, A.; Shapiro, H.; Geiger, B.; Elinav, E. Hyperglycemia drives intestinal barrier dysfunction and risk for enteric infection. Science 2018, 359(6382), 1376–1383. [Google Scholar] [CrossRef] [PubMed]
- Cani, P.D.; Osto, M.; Geurts, L.; Everard, A. Involvement of gut microbiota in the development of low-grade inflammation and type 2 diabetes associated with obesity. Gut Microbes 2012, 3(4), 279–88. [Google Scholar] [CrossRef] [PubMed]
- Mohammad, S.; Thiemermann, C. Role of Metabolic Endotoxemia in Systemic Inflammation and Potential Interventions. Front Immunol 2021, 11, 594150. [Google Scholar] [CrossRef] [PubMed]
- Yin, F.; Sancheti, H.; Patil, I.; Cadenas, E. Energy metabolism and inflammation in brain aging and Alzheimer's disease. Free Radic Biol Med 2016, 100, 108–122. [Google Scholar] [CrossRef]
- Qin, L.; Wu, X.; Block, M.L.; Liu, Y.; Breese, G.R.; Hong, J.S.; Knapp, D.J.; Crews, F.T. Systemic LPS causes chronic neuroinflammation and progressive neurodegeneration. Glia 2007, 55(5), 453–62. [Google Scholar] [CrossRef]
- Riazi, K.; Galic, M.A.; Kuzmiski, J.B.; Ho, W.; Sharkey, K.A.; Pittman, Q.J. Microglial activation and TNFalpha production mediate altered CNS excitability following peripheral inflammation. Proc Natl Acad Sci U S A 2008, 105(44), 17151–6. [Google Scholar] [CrossRef]
- De Felice, F.G.; Lourenco, M.V. Brain metabolic stress and neuroinflammation at the basis of cognitive impairment in Alzheimer's disease. Front Aging Neurosci 2015, 7, 94. [Google Scholar] [CrossRef]
- Camandola, S.; Mattson, M.P. Brain metabolism in health, aging, and neurodegeneration. EMBO J 2017, 36(11), 1474–1492. [Google Scholar] [CrossRef]
- Lauro, C.; Limatola, C. Metabolic Reprograming of Microglia in the Regulation of the Innate Inflammatory Response. Front Immunol 2020, 11, 493. [Google Scholar] [CrossRef]
- Sangineto, M.; Ciarnelli, M.; Cassano, T.; Radesco, A.; Moola, A.; Bukke, V.N.; Romano, A.; Villani, R.; Kanwal, H.; Capitanio, N.; Duda, L.; Avolio, C.; Serviddio, G. Metabolic reprogramming in inflammatory microglia indicates a potential way of targeting inflammation in Alzheimer's disease. Redox Biol. Epub 2023 Aug 9. 2023, 66, 102846. [Google Scholar] [CrossRef]
- Adams, J.B.; Audhya, T.; Geis, E.; Gehn, E.; Fimbres, V.; Pollard, E.L.; Mitchell, J.; Ingram, J.; Hellmers, R.; Laake, D.; Matthews, J.S.; Li, K.; Naviaux, J.C.; Naviaux, R.K.; Adams, R.L.; Coleman, D.M.; Quig, D.W. Comprehensive Nutritional and Dietary Intervention for Autism Spectrum Disorder-A Randomized, Controlled 12-Month Trial. Nutrients 2018, 10(3), 369. [Google Scholar] [CrossRef] [PubMed]
- Grimaldi, R.; Gibson, G.R.; Vulevic, J.; Giallourou, N.; Castro-Mejía, J.L.; Hansen, L.H.; Leigh Gibson, E.; Nielsen, D.S.; Costabile, A. A prebiotic intervention study in children with autism spectrum disorders (ASDs). Microbiome 2018, 6(1), 133. [Google Scholar] [CrossRef]
- Kang, D.W.; Adams, J.B.; Gregory, A.C.; Borody, T.; Chittick, L.; Fasano, A.; Khoruts, A.; Geis, E.; Maldonado, J.; McDonough-Means, S.; Pollard, E.L.; Roux, S.; Sadowsky, M.J.; Lipson, K.S.; Sullivan, M.B.; Caporaso, J.G.; Krajmalnik-Brown, R. Microbiota Transfer Therapy alters gut ecosystem and improves gastrointestinal and autism symptoms: an open-label study. Microbiome 2017, 5(1), 10. [Google Scholar] [CrossRef]
| Study | Model | Sample Type | Age/Stage | ASD Characteristics | ASD patients | ||
|---|---|---|---|---|---|---|---|
| Acetate | Propionate | Butyrate | |||||
| Wang et al., 2019 [47] | Human | Feces | Children | ASD vs TD controls | ↔ | ↔ | ↔ |
| Liu et al., 2019 [48] | ↓ | NS | ↓ | ||||
| Study | Model | Sample type | Increased taxa | Decreased taxa | Functional implication |
|---|---|---|---|---|---|
| Liu et al., 2019 [48] | Human (ASD) | Feces | Propionate- and acetate-producing taxa; Desulfovibrio (G) | Butyrate-associated taxa; Ruminococcaceae (F), Eubacterium (G), Lachnospiraceae (F), Erysipelotrichaceae (F) | Altered SCFA-related microbial balance |
| Li et al., 2024 [97] | Human (ASD) | Fusobacteria (P), Firmicutes (P), Verrucomicrobia (P), | Lentisphaerae (P), Bacteroidetes (P), Euryarchaeota (P), Patescibacteria (P) | Altered microbial community structure | |
| Lou et al., 2022 [98] | Human (ASD) |
Bifidobacterium (G), Veillonella (G), Enterobacteriaceae (F), Lachnospiraceae (F) |
Clostridium (G), Veillonella ratti (S) |
Early-life cohort-specific microbial deviation | |
| Iglesias-Vázquez et al., 2020 [100] | Human (ASD) | Bacteroides (G), Parabacteroides (G), Clostridium (G), Faecalibacterium (G), Phascolarctobacterium (G) | Coprococcus (G), Bifidobacterium (G) | Context-dependent lack of a consistent ASD microbial signature | |
| Kang et al., 2013 [102] | Human (ASD) | NR | Prevotella (G), Coprococcus (G), unclassified Veillonellaceae (F) | Disrupted microbial interactions | |
| Coretti et al., 2018 [104] | Human (ASD) | Bacteroidetes (P), Proteobacteria (P) | Actinobacteria (P) | Pro-inflammatory microbial shift |
| Source of heterogeneity | Categories / examples | Impact on SCFA readouts | Implications for interpretation |
|---|---|---|---|
| Sample source | Feces vs plasma/serum | Captures luminal fermentation output vs absorbed/systemically available SCFAs, influenced by host metabolism (e.g., hepatic clearance) | Fecal and circulating SCFAs are not interchangeable |
| GI symptom stratification | Constipation, diarrhea, abdominal pain | Alters intestinal transit time, fermentation kinetics, absorption efficiency, and stool water content | Null findings may reflect signal dilution in unstratified cohorts |
| Developmental stage | Infancy, childhood, adolescence | Age-dependent microbiota maturation and diet transitions shift baseline SCFA profiles | Age mismatch limits cross-study comparability |
| Diet and fiber intake | High vs low fermentable fiber; dietary patterns | Directly modulates microbial substrate availability and SCFA production | Dietary control is critical for stronger causal inference |
| Medication exposure | Antibiotics, probiotics/prebiotics, laxatives, psychotropics, PPIs | Reshapes microbial composition and metabolic output; may interact with GI status | Medication history should be systematically captured and adjusted for |
| Analytical workflow / platform | GC-FID, GC–MS, LC–MS/MS (with derivatization and internal standards) | Differences in sensitivity, specificity, and quantification range across platforms | Methodological heterogeneity contributes substantially to variability |
| Reporting units / normalization | Absolute concentration vs relative proportion; wet vs dry weight; different unit scales | Limits quantitative comparability and may invert apparent group differences | Directional interpretation is frequently more appropriate than magnitude alone |
| Experimental design / model | Human cohorts vs rodent models; exposure-based vs endogenous metabolism | Exposure paradigms may not recapitulate endogenous SCFA dynamics in humans | Animal findings should not be directly generalized to human ASD |
| Developmental stage | SCFA-related context | Biological processes potentially affected | Observed or reported outcomes | Representative references |
|---|---|---|---|---|
| Pregnancy (maternal) | Circulating maternal SCFAs reflecting gut fermentation | Maternal metabolic–immune milieu influencing fetal neurodevelopmental susceptibility | Associations with offspring neurodevelopmental measures in cohort studies | [93] |
| Prenatal period (maternal immune activation) | SCFA–immune crosstalk potentially intersecting with maternal inflammation | Immune signaling at the maternal–fetal interface | Increased vulnerability to neurodevelopmental alterations in offspring | [14,95] |
| Early postnatal (infancy) | Infant gut fermentation–derived SCFAs with limited systemic availability | Microglial maturation and synaptic pruning (preclinical support) | Atypical early developmental trajectories in early-life microbiome studies | [90,92] |
| Childhood | Diet-modulated gut and plasma metabolomic profiles | Systemic metabolic shifts linked to microbiota modulation | Context-dependent associations with autism-related traits | [107,115] |
| Juvenile (experimental animal models) | Experimental SCFA exposure (e.g., propionate/PPA) | Neuroinflammation and mitochondrial stress pathways | ASD-like behavioral phenotypes in specific paradigms | [1,94] |
| Biomarker domain | Candidate markers | Measurement source | Biological rationale |
|---|---|---|---|
| Gut-derived metabolites | Relative SCFA patterns (acetate, propionate, butyrate) | Feces / plasma | Reflects microbial fermentation patterns and systemic availability |
| Gastrointestinal integrity | Lipopolysaccharide-binding protein (LBP) | Serum | Indicates barrier-related metabolite translocation |
| Immune activation | Representative inflammatory cytokines (e.g., IL-6, IL-17A) | Serum | Captures inflammatory tone linked to metabolic signaling |
| Oxidative stress / redox imbalance | Redox-related markers | Blood | Reflects oxidative imbalance relevant to neuronal vulnerability |
| Mitochondrial / energy metabolism | Energy metabolism–related indicators | Blood | Reflects altered cellular energy metabolism |
| Microbiota structure | Relative depletion of fermentative taxa | Fecal sequencing | Reflects reduced SCFA-producing capacity |
| Developmental timing | Perinatal/early-life exposures | Clinical history | Identifies sensitive developmental windows |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




