Submitted:
14 April 2026
Posted:
15 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
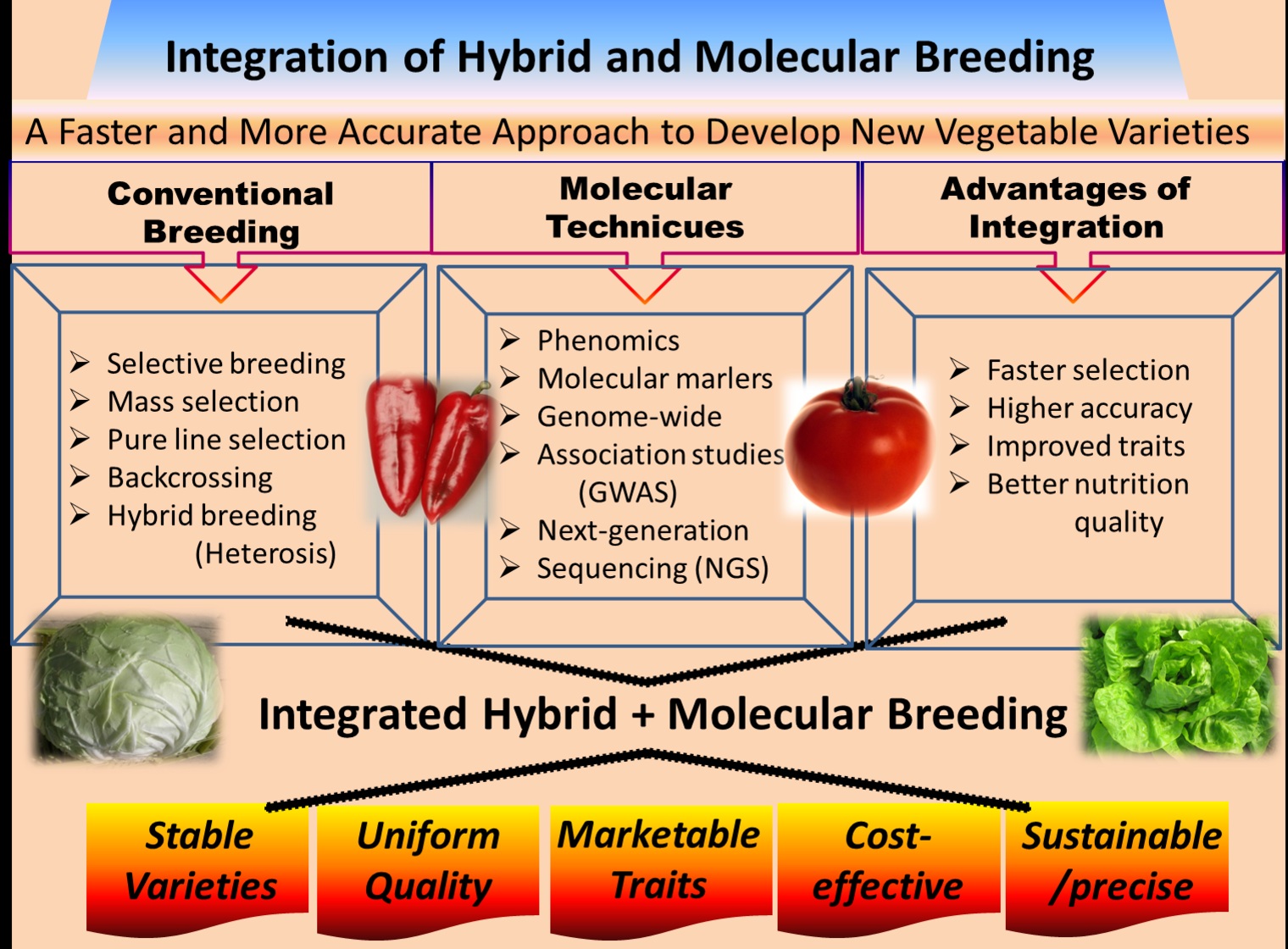
2. Conventional Plant Breeding
3. Genetic Diversity Assessment
4. Marker-Assisted Selection (MAS)
- Tracking favorable alleles (dominant or recessive) through generations;
- Identification of most suitable individuals among the offspring in the separation populations, based on the allelic representation in a part of the genome or the entire genome [64].
4.1. Introgression and Selection of Traits and QTL from Breeding Lines and Wild Relatives
4.2. Marker-Assisted Backcrossing
4.3. Marker-Assisted Recurrent Selection (MARS)
4.4. Cumulative Gene Pyramiding
5. Examples of Marker-Assisted Selection in Vegetable Crops
5.1. Tomato
5.2. Pepper
5.3. Cabbage
5.4. Lettuce
6. Genomic Selection as a Bridge Between Molecular and Hybrid Breeding
7. CRISPR/Cas9 Technology in Vegetable Breeding
7.1. Tomato
7.2. Pepper
7.3. Cabbage
7.4. Cucumber
7.5. Lettuce
8. Advantages of Integrating Hybrid and Molecular Breeding of Vegetables
9. Limitations and Challenges of Integrating Conventional Breeding and Molecular Techniques
10. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| WHO | World Health Organization |
| MAS | Marker-Assisted Selection |
| MABC | Marker-Assisted Backcrossing |
| MARS DNA |
Marker-Assisted Recurrent Selection Deoxyribonucleic Acid |
| RAPD | Random Amplified Polymorphic DNA |
| ISSR | Inter-Simple Sequence Repeats |
| AFLP | Amplified Fragment Length Polymorphism |
| RFLP | Restriction Fragment Length Polymorphism |
| SNP SSR |
Single Nucleotide Polymorphism Simple Sequence Repeat |
| SRAP QTL |
Sequence-Related Amplified Polymorphism Quantitative Trait Loci |
| LD | Linkage Disequilibrium |
| KASP | Kompetitive Allele Specific PCR |
| SCAR | Sequence Characterized Amplified Region |
| CAPS | Cleaved Amplified Polymorphic Sequences |
| PCR dCAPS InDel ERPAR EST-SSR SSAP GS GEBVs GCA SCA DH HTPP AI gLMs sgRNA CRISPR CRISPR/Cas9 |
Polymerase Chain Reaction Derived Cleaved Amplified Polymorphic Sequences Insertion-Deletion Extended Random Primer Amplified Region Expressed Sequence Tag-Simple Sequence Repeats Sequence-specific Amplified Polymorphism Genomic Selection Genomic Estimated Breeding Values General Combining Ability Specific Combining Ability Doubled Haploid High-Throughput Phenotyping Artificial Intelligence Genomic Language Models Single Guide Ribonucleic Acid Clustered Regularly Interspaced Short Palindromic Repeats CRISPR- associated Protein 9 |
| RNP GMOs NHEJ HDR |
Ribonucleoprotein Genetically Modified Organisms Non-Homologous End Joining Homology-Directed Repair |
References
- Abuyusuf, M.; Nath, U. K.; Kim, H.-T.; Islam, M. R.; Park, J.-I.; Nou, I.-S. Molecular Markers Based on Sequence Variation in BoFLC1.C9 for Characterizing Early- and Late-Flowering Cabbage Genotypes. BMC Genet. 2019, 20, 42. [Google Scholar] [CrossRef]
- Dias, J. S. Nutritional Quality and Health Benefits of Vegetables: A Review. Food Nutr. Sci. 2012, 03, 1354–1374. [Google Scholar] [CrossRef]
- Dias, J. C. da S. Dias, J. C. da S. Nutritional Quality and Health Benefits of Vegetables. In Emerging Trends in Disease and Health Research Vol. 4; Book Publisher International (a part of SCIENCEDOMAIN International). 2022, 7–35. [CrossRef]
- Vojnović, Đ; Ilin, Ž; Kostić, A. Ž; Adamović, B.; Kilibarda, S.; Pezo, L.; Popović-Đorđević, J. Phytochemical Composition, Antioxidant Potential, and Bulb Mass of Differently Colored Onions under Distinct Sowing Methods. Qual. Assur. Saf. Crops Foods 2025, 17, 108–126. [Google Scholar] [CrossRef]
- Drewnowski, A.; Rehm, C. D. Energy Intakes of US Children and Adults by Food Purchase Location and by Specific Food Source. Nutr. J. 2013, 12, 59. [Google Scholar] [CrossRef] [PubMed]
- Boye, J.; Zare, F.; Pletch, A. Pulse Proteins: Processing, Characterization, Functional Properties and Applications in Food and Feed. Food Res. Int. 2010, 43, 414–431. [Google Scholar] [CrossRef]
- Food and Agriculture Organization. Pulses: Nutritious Seeds for a Sustainable Future; Food & Agriculture Organization of the United Nations (FAO): Rome, Italy, 2016. [Google Scholar] [CrossRef]
- Slavin, J. L.; Lloyd, B. Health Benefits of Fruits and Vegetables. Adv. Nutr. 2012, 3, 506–516. [Google Scholar] [CrossRef] [PubMed]
- Batoola, U.; Nawaz, R.; Ahmad, S.; Irfan, A.; Irshad, M. A.; Arshad, M.; Salamatullah, A. M.; Shah, A. N.; Dauelbaiti, M.; Butt, A. R.; Bourhiak, M. Physicochemical properties and antioxidant potentials of leafy vegetables grown under different agroecological conditions. International Journal of Food Properties 2024, 2024(27), 1046–1063. [Google Scholar] [CrossRef]
- Soujanya, K.; Kumari, B. A.; Jyothsna, E. Evaluation of Nutritional Quality Characteristics of Selected Uncultivated Green Leafy Vegetables. Ann. Phytomed. 2024, 13. [Google Scholar] [CrossRef]
- Nurzyńska-Wierdak, R. Green Leafy Vegetables (GLVs) as Nutritional and Preventive Agents Supporting Metabolism. Metabolites 2025, 15 502. [Google Scholar] [CrossRef]
- Ramya, V.; Patel, P. Health Benefits of Vegetables. International Journal of Chemical Studies 2019, 7, 82–87. [Google Scholar]
- Ebabhi, A.; Adebayo, R. Nutritional Values of Vegetables. In Vegetable Crops - Health Benefits and Cultivation; IntechOpen, 2022. [CrossRef]
- Devirgiliis, C.; Guberti, E.; Mistura, L.; Raffo, A. Effect of Fruit and Vegetable Consumption on Human Health: An Update of the Literature. Foods 2024, 13, 3149. [Google Scholar] [CrossRef] [PubMed]
- Silva Dias, J. World Importance, Marketing and Trading of Vegetables. Acta Hortic. 2011, 921, 153–169. [Google Scholar] [CrossRef]
- Ramya, V.; Patel, P. Health Benefits of Vegetables. International Journal of Chemical Studies 2019, 7, 82–87. [Google Scholar]
- Ramasamy, S.; Lin, M.-Y.; Wu, W.-J.; Wang, H.-I.; Sotelo-Cardona, P. Evaluating the Potential of Protected Cultivation for Off-Season Leafy Vegetable Production: Prospects for Crop Productivity and Nutritional Improvement. Front. Sustain. Food Syst. 2021, 5. [Google Scholar] [CrossRef]
- Zandalinas, S. I.; Balfagón, D.; Gómez-Cadenas, A.; Mittler, R. Plant Responses to Climate Change: Metabolic Changes under Combined Abiotic Stresses. J. Exp. Bot. 2022, 73, 3339–3354. [Google Scholar] [CrossRef]
- Sánchez-Bermúdez, M.; Del Pozo, J. C.; Pernas, M. Effects of Combined Abiotic Stresses Related to Climate Change on Root Growth in Crops. Front. Plant Sci. 2022, 13, 918537. [Google Scholar] [CrossRef]
- Weiss, J.; Gruda, N. S. Novel Breeding Techniques and Strategies for Enhancing Greenhouse Vegetable Product Quality. Agronomy (Basel) 2025a, 15, 207. [Google Scholar] [CrossRef]
- Bommarco, R.; Kleijn, D.; Potts, S. G. Ecological Intensification: Harnessing Ecosystem Services for Food Security. Trends Ecol. Evol. 2013, 28, 230–238. [Google Scholar] [CrossRef]
- Savary, S.; Willocquet, L.; Pethybridge, S. J.; Esker, P.; McRoberts, N.; Nelson, A. The Global Burden of Pathogens and Pests on Major Food Crops. Nat. Ecol. Evol. 2019, 3, 430–439. [Google Scholar] [CrossRef]
- Tigchelaar, M.; Battisti, D. S.; Naylor, R. L.; Ray, D. K. Future Warming Increases Probability of Globally Synchronized Maize Production Shocks. Proc. Natl. Acad. Sci. U. S. A. 2018, 115, 6644–6649. [Google Scholar] [CrossRef]
- Zörb, C.; Geilfus, C.-M.; Dietz, K.-J. Salinity and Crop Yield. Plant Biol. (Stuttg.) 2019, 21 Suppl 1 (S1), 31–38. [Google Scholar] [CrossRef] [PubMed]
- Mehra, P.; Patel, S. S.; Pansare, U. D. Application of Molecular Markers in Vegetable Improvement : A Review. Plant Arch. 2024, 24. [Google Scholar] [CrossRef]
- Botstein, D.; White, R. L.; Skolnick, M. Construction of a Genetic Linkage Map in Man Using Restriction Fragment Length Pol-Ymorphisms. Am J Hum Genet 1980, 32, 314–331. [Google Scholar] [PubMed]
- Ceccarelli, S. Efficiency of Plant Breeding. Crop Sci. 2015, 55, 87–97. [Google Scholar] [CrossRef]
- He, R.; Ju, J.; Liu, K.; Song, J.; Zhang, S.; Zhang, M.; Hu, Y.; Liu, X.; Li, Y.; Liu, H. Technology of Plant Factory for Vegetable Crop Speed Breeding. Front. Plant Sci. 2024, 15, 1414860. [Google Scholar] [CrossRef]
- Williams, J. G.; Kubelik, A. R.; Livak, K. J.; Rafalski, J. A.; Tingey, S. V. DNA Polymorphisms Amplified by Arbitrary Primers Are Useful as Genetic Markers. Nucleic Acids Res. 1990, 18, 6531–6535. [Google Scholar] [CrossRef]
- Sun, L.; Lai, M.; Ghouri, F.; Nawaz, M. A.; Ali, F.; Baloch, F. S.; Nadeem, M. A.; Aasim, M.; Shahid, M. Q. Modern Plant Breeding Techniques in Crop Improvement and Genetic Diversity: From Molecular Markers and Gene Editing to Artificial Intelligence-A Critical Review. Plants 2024, 13, 2676. [Google Scholar] [CrossRef]
- Vos, P.; Hogers, R.; Bleeker, M.; Reijans, M.; van de Lee, T.; Hornes, M.; Frijters, A.; Pot, J.; Peleman, J.; Kuiper, M. AFLP: A New Technique for DNA Fingerprinting. Nucleic Acids Res. 1995, 23, 4407–4414. [Google Scholar] [CrossRef]
- Singh, A.; Prasad, S. S.; Ingle, K. P.; Das, U.; Ramteke, P. W.; Kurubar, A. R.; Shukla, P. K.; Geethika, P.; Madala, R. S. Molecular Marker-Assisted Selection in Plant Breeding. In Advances in Plant Breeding Strategies; Springer Nature Switzerland: Cham, 2024; pp. 95–111. [Google Scholar] [CrossRef]
- Fetisov, I.; Eizikovich, O.; Charles Diouf, D.; Romanova, E.; Kezimana, P. Advancements in Molecular Breeding Techniques for Soybeans. Plants 2025, 15, 5. [Google Scholar] [CrossRef]
- Charmet, G.; Robert, N.; Perretant, M. R.; Gay, G.; Sourdille, P.; Groos, C.; Bernard, S.; Bernard, M. Marker-Assisted Recurrent Selection for Cumulating Additive and Interactive QTLs in Recombinant Inbred Lines. Theor. Appl. Genet. 1999, 99, 1143–1148. [Google Scholar] [CrossRef]
- Collard, B. C. Y.; Mackill, D. J. Marker-Assisted Selection: An Approach for Precision Plant Breeding in the Twenty-First Century. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2008, 363, 557–572. [Google Scholar] [CrossRef]
- Jiang, G.L. Molecular Markers and Marker-Assisted Breeding in Plants. In Plant Breeding from Laboratories to Fields; InTech, 2013. [CrossRef]
- Thakur, M.; Thakur, H. Vegetable Breeding for Organic Farming: Opportunities and Perspectives. Genetika 2025, 57, 443–456. [Google Scholar] [CrossRef]
- Ebert, A. W. The Role of Vegetable Genetic Resources in Nutrition Security and Vegetable Breeding. Plants 2020, 9, 736. [Google Scholar] [CrossRef]
- Červenski, J.; Slobodan, F.V.; Adamovic, A.B.; Vojnovic, D.D.; Zec, S.S.D. Possibilities of Cabbage Production under Climatic Changes. J. Agric. Sci. Belgrade 2025, 70, 233–247. [Google Scholar] [CrossRef]
- Weiss, J.; Gruda, N. S. Enhancing Nutritional Quality in Vegetables through Breeding and Cultivar Choice in Protected Cultivation. Sci. Hortic. (Amsterdam) 2025b, 339, 113914. [Google Scholar] [CrossRef]
- Tabasum, S.; Nazir, G.; Hussain, K.; Gautam, A.; Mushtaq, F.; Nabi, A.; Ahsan, S. Climate Smart Breeding of Vegetable Crops : A Review. Plant Arch. 2025, 25. [Google Scholar] [CrossRef]
- Ghuge, M. B.; Mirza, A. Vegetable Breeding Strategies. Asian J. Agric. Hortic. Res. 2021, 28–37. [Google Scholar] [CrossRef]
- Singh, H.; Sekhon, B. S.; Kumar, P.; Dhall, R. K.; Devi, R.; Dhillon, T. S.; Sharma, S.; Khar, A.; Yadav, R. K.; Tomar, B. S.; Ntanasi, T.; Sabatino, L.; Ntatsi, G. Genetic Mechanisms for Hybrid Breeding in Vegetable Crops. Plants 2023, 12, 2294. [Google Scholar] [CrossRef]
- Rauf, S.; Basharat, T.; Iqbal, Z. Hybrid Breeding in Vegetable Crops. In Sustainable and Innovative Vegetable Production in times of Climate Change; Springer Nature Singapore: Singapore, 2025; pp. 403–420. [Google Scholar] [CrossRef]
- Riggs, T. J. Breeding F1hybrid Varieties of Vegetables. J. Hortic. Sci. 1988, 63, 369–382. [Google Scholar] [CrossRef]
- Carvalho, F.; Lahlou, R. A.; Silva, L. R. Exploring Bioactive Compounds from Fruit and Vegetable By-Products with Potential for Food and Nutraceutical Applications. Foods 2025, 14, 3884. [Google Scholar] [CrossRef]
- Pidigam, S.; Geetha, A.; Nagaraju, K.; Pandravada, S. R.; Khan, M. S.; Rajasekhar, M.; Sivraj, N.; Vishnukiran, T. Breeding Approaches for the Development of Nutraceutical Vegetables: A Review. Ann. Phytomed. 2022, 11. [Google Scholar] [CrossRef]
- Causse, M.; Saliba-Colombani, V.; Lecomte, L.; Duffé, P.; Rousselle, P.; Buret, M. QTL Analysis of Fruit Quality in Fresh Market Tomato: A Few Chromosome Regions Control the Variation of Sensory and Instrumental Traits. J. Exp. Bot. 2002, 53, 2089–2098. [Google Scholar] [CrossRef] [PubMed]
- Capel, C.; Yuste-Lisbona, F. J.; López-Casado, G.; Angosto, T.; Heredia, A.; Cuartero, J.; Fernández-Muñoz, R.; Lozano, R.; Capel, J. QTL Mapping of Fruit Mineral Contents Provides New Chances for Molecular Breeding of Tomato Nutritional Traits. Theor. Appl. Genet. 2017, 130, 903–913. [Google Scholar] [CrossRef] [PubMed]
- Červenski, J.; Vlajić, S.; Ignjatov, M.; Tamindžić, G.; Zec, S. Agroclimatic Conditions for Cabbage Production. Ratar. Povrt. 2022, 59, 43–50. [Google Scholar] [CrossRef]
- Yu, D.; Gu, X.; Zhang, S.; Dong, S.; Miao, H.; Gebretsadik, K.; Bo, K. Molecular Basis of Heterosis and Related Breeding Strategies Reveal Its Importance in Vegetable Breeding. Hortic. Res. 2021, 8, 120. [Google Scholar] [CrossRef]
- Sahu, D.; Tripathy, S. K.; Sahoo, B.; Nayak, L.; Swarajya, R.; Dalei, D. Role of Molecular Markers in Vegetables Improvement. Journal of Pharmacognosy and Phytochemistry 2019, 8, 456–460. [Google Scholar]
- Singh, J.; Dhall, R. K.; Vikal, Y. Genetic Diversity Studies in Indian Germplasm of Pea (Pisum Sativum L.) Using Morphological and Microsatellite Markers. Genetika 2021, 53, 473–491. [Google Scholar] [CrossRef]
- Kondić-Špika, A.; Kobiljski, B. Biotechnology in modern breeding and agriculture. In Book of the Proceedings, International Conference on BioScience: Biotechnology and Biodiversity,” Step in the Future”. 2012, 201-210. https://fiver.ifvcns.rs/handle/123456789/4822.
- Singh, H. P.; Raigar, O. P.; Chahota, R. K. Estimation of Genetic Diversity and Its Exploitation in Plant Breeding. Bot. Rev. 2022, 88, 413–435. [Google Scholar] [CrossRef]
- Salgotra, R. K.; Chauhan, B. S. Genetic Diversity, Conservation, and Utilization of Plant Genetic Resources. Genes (Basel) 2023, 14, 174. [Google Scholar] [CrossRef]
- Bidyananda, N.; Jamir, I.; Nowakowska, K.; Varte, V.; Vendrame, W. A.; Devi, R. S.; Nongdam, P. Plant Genetic Diversity Studies: Insights from DNA Marker Analyses. Int. J. Plant Biol. 2024, 15, 607–640. [Google Scholar] [CrossRef]
- Vázquez, H.; Jimenez, F. L.; Vico, J. M. C. RAPD Fingerprinting of Pepper (Capsicum Annuum L.) Breeding Lines. Capsicum and Eggplant Newslette 1996, 15, 37–40. [Google Scholar]
- Dijkhuizen, A.; Kennard, W. C.; Havey, M. J.; Staub, J. E. RFLP Variation and Genetic Relationships in Cultivated Cucumber. Euphytica 1996, 90, 79–87. [Google Scholar] [CrossRef]
- Villand, J.; Skroch, P. W.; Lai, T.; Hanson, P.; Kuo, C. G.; Nienhuis, J. Genetic Variation among Tomato Accessions from Primary and Secondary Centers of Diversity. Crop Sci. 1998, 38, 1339–1347. [Google Scholar] [CrossRef]
- Levi, A.; Thomas, C. E.; Newman, M.; Reddy, O. U. K.; Zhang, X.; Xu, Y. ISSR and AFLP Markers Differ among American Watermelon Cultivars with Limited Genetic Diversity. J. Am. Soc. Hortic. Sci. 2004, 129, 553–558. [Google Scholar] [CrossRef]
- Muminović, J.; Merz, A.; Melchinger, A. E.; Lübberstedt, T. 3 Genetic Structure and Diversity among Radish Varieties as Inferred from AFLP and ISSR Analyses. Journal of the American Society for Horticultural Science 2005, 130, 79–87. [Google Scholar] [CrossRef]
- Blanca, J.; Pons, C.; Montero-Pau, J.; Sanchez-Matarredona, D.; Ziarsolo, P.; Fontanet, L.; Fisher, J.; Plazas, M.; Casals, J.; Rambla, J. L.; Riccini, A.; Pombarella, S.; Ruggiero, A.; Sulli, M.; Grillo, S.; Kanellis, A.; Giuliano, G.; Finkers, R.; Cammareri, M.; Grandillo, S.; Mazzucato, A.; Causse, M.; Díez, M. J.; Prohens, J.; Zamir, D.; Cañizares, J.; Monforte, A. J.; Granell, A. European Traditional Tomatoes Galore: A Result of Farmers’ Selection of a Few Diversity-Rich Loci. J. Exp. Bot. 2022, 73, 3431–3445. [Google Scholar] [CrossRef]
- Ibitoye, D.; Akin-Idowu, P. Marker-assisted-selection (MAS): A fast track to increase genetic gain in horticultural crop breeding. African Journal of Biotechnology 2013, 10, 11333–11339. [Google Scholar] [CrossRef]
- Ahmar, S.; Gill, R. A.; Jung, K.-H.; Faheem, A.; Qasim, M. U.; Mubeen, M.; Zhou, W. Conventional and Molecular Techniques from Simple Breeding to Speed Breeding in Crop Plants: Recent Advances and Future Outlook. Int. J. Mol. Sci. 2020, 21, 2590. [Google Scholar] [CrossRef]
- Simko, I.; Jia, M.; Venkatesh, J.; Kang, B.-C.; Weng, Y.; Barcaccia, G.; Lanteri, S.; Bhattarai, G.; Foolad, M. R. Genomics and Marker-Assisted Improvement of Vegetable Crops. CRC Crit. Rev. Plant Sci. 2021, 40, 303–365. [Google Scholar] [CrossRef]
- Lema, M. Marker Assisted Selection in Comparison to Conventional Plant Breeding: Review Article. Agric. Res. Technol. 2018, 14. [Google Scholar] [CrossRef]
- Collard, B. C. Y.; Jahufer, M. Z. Z.; Brouwer, J. B.; Pang, E. C. K. An Introduction to Markers, Quantitative Trait Loci (QTL) Mapping and Marker-Assisted Selection for Crop Improvement: The Basic Concepts. Euphytica 2005, 142, 169–196. [Google Scholar] [CrossRef]
- Guberac, S. Strossmayer University of Osijek, Faculty of Agriculture in Osijek, Kralja Petra Svačića 1d, 31000 Osijek, Croatia; Petrović, S.; Guberac, V.; Marić, S. Molecular Markers in Tomato Breeding. Poljoprivreda 2015, 21, 55–60. [Google Scholar] [CrossRef]
- Xu, Y.; Crouch, J. H. Marker-assisted Selection in Plant Breeding: From Publications to Practice. Crop Sci. 2008, 48, 391–407. [Google Scholar] [CrossRef]
- Bernardo, R. Molecular Markers and Selection for Complex Traits in Plants: Learning from the Last 20 Years. Crop Sci. 2008, 48, 1649–1664. [Google Scholar] [CrossRef]
- Ansari, A. Molecular Markers in Vegetable Improvement. Horticultural Biotechnology Research 2015, 1, 5–10. [Google Scholar] [CrossRef]
- Kalendar, R.; Grob, T.; Regina, M. IRAP and REMAP: two new retrotransposon-based DNA fingerprinting techniques. Theor Appl Genet. 1999, 98, 704–711. [Google Scholar] [CrossRef]
- Jaccoud, D.; Peng, K.; Feinstein, D.; Kilian, A. Diversity Arrays: A Solid State Technology for Sequence Information Independent Genotyping. Nucleic Acids Res. 2001, 29, E25. [Google Scholar] [CrossRef]
- Kulus, D. Genetic Diversity for Breeding Tomato. In Cash Crops; Springer International Publishing: Cham, 2022; pp. 505–521. [Google Scholar] [CrossRef]
- Khazaei, H.; Madduri, A. Role of Tomato Wild Relatives in Breeding Disease-Free Varieties. GenResJ 2022, 3, 64–73. [Google Scholar] [CrossRef]
- Taylor, B. Biosystematics of the tomato. In: Atherton, J. G., and J. Rudich, editors. The Tomato Crop : A Scientific Basis for Improvement. 1st ed. Springer Netherlands, 1986. [CrossRef]
- John Goodstal, F.; Kohler, G. R.; Randall, L. B.; Bloom, A. J.; St Clair, D. A. A Major QTL Introgressed from Wild Lycopersicon Hirsutum Confers Chilling Tolerance to Cultivated Tomato (Lycopersicon Esculentum). Theor. Appl. Genet. 2005, 111, 898–905. [Google Scholar] [CrossRef]
- Mladenov, V.; Boćanski, Ј; Banjac, B. Oplemenjivanje Biljaka: Praktikum; Novi Sad; 2021.1-68.
- Shrestha, S.; Subedi, S.; Jiban, J. Marker-Assisted Selection: A Smart Biotechnological Strategy for Modern Plant Breeding. Peruvian Journal of Agronomy 2020, 4, 104–120. [Google Scholar] [CrossRef]
- Gokidi, Y. U.; Bhanu, A. N.; Singh, M. N. Marker Assisted Recurrent Selection: An Overview. Adv Life Sci 2016, 5, 6493–6499. [Google Scholar]
- Beyene, Y.; Semagn, K.; Mugo, S.; Prasanna, B. M.; Tarekegne, A.; Gakunga, J.; Sehabiague, P.; Meisel, B.; Oikeh, S. O.; Olsen, M.; Crossa, J. Performance and Grain Yield Stability of Maize Populations Developed Using Marker-Assisted Recurrent Selection and Pedigree Selection Procedures. Euphytica 2016, 208, 285–297. [Google Scholar] [CrossRef] [PubMed]
- Cholin, S. S.; Kulkarni, C. C. Prospects of Marker-Assisted Recurrent Selection: Current Insights and Future Implications. Trop. Plant Biol. 2023. [Google Scholar] [CrossRef]
- Na Jinda, A.; Nikornpun, M.; Jeeatid, N.; Thumdee, S.; Thippachote, K.; Pusadee, T.; Kumchai, J. Marker-Assisted Selection of Male-Sterile and Maintainer Line in Chili Improvement by Backcross Breeding. Horticulturae 2023, 9, 357. [Google Scholar] [CrossRef]
- Thomas, W. J. W.; Amas, J. C.; Dolatabadian, A.; Huang, S.; Zhang, F.; Zandberg, J. D.; Neik, T. X.; Edwards, D.; Batley, J. Recent Advances in the Improvement of Genetic Resistance against Disease in Vegetable Crops. Plant Physiol. 2024, 196, 32–46. [Google Scholar] [CrossRef]
- Husaini, A. M. High-Value Pleiotropic Genes for Developing Multiple Stress-Tolerant Biofortified Crops for 21st-Century Challenges. Heredity (Edinb.) 2022, 128, 460–472. [Google Scholar] [CrossRef]
- Suresh, S.; Malathi, D. Gene Pyramiding for Biotic Stress Tolerance In Crop Plants. Wkly. Sci. Res. J 2013, 1, 1–14. [Google Scholar]
- Bai, Y.; Kissoudis, C.; Yan, Z.; Visser, R. G. F.; van der Linden, G. Plant Behaviour under Combined Stress: Tomato Responses to Combined Salinity and Pathogen Stress. Plant J. 2018, 93, 781–793. [Google Scholar] [CrossRef]
- Kage, U.; Kumar, A.; Dhokane, D.; Karre, S.; Kushalappa, A. C. Functional Molecular Markers for Crop Improvement. Crit. Rev. Biotechnol. 2016, 36, 917–930. [Google Scholar] [CrossRef]
- Hasan, M. M.; Rafii, M. Y.; Ismail, M. R.; Mahmood, M.; Rahim, H. A.; Alam, M. A.; Ashkani, S.; Malek, M. A.; Latif, M. A. Marker-Assisted Backcrossing: A Useful Method for Rice Improvement. Biotechnol. Biotechnol. Equip. 2015, 29, 237–254. [Google Scholar] [CrossRef] [PubMed]
- Rana, M.; Sood, A.; Hussain, W.; Kaldate, R.; Sharma, T. R.; Gill, R. K.; Kumar, S.; Singh, S. Chapter 6-Gene Pyramiding and Multiple Character Breeding. In Lentils; Singh, M; Academic Press: Cambridge, MA, USA, 2019, 83–124.
- Dormatey, R.; Sun, C.; Ali, K.; Coulter, J. A.; Bi, Z.; Bai, J. Gene Pyramiding for Sustainable Crop Improvement against Biotic and Abiotic Stresses. Agronomy (Basel) 2020, 10, 1255. [Google Scholar] [CrossRef]
- Verlaan, M. G.; Hutton, S. F.; Ibrahem, R. M.; Kormelink, R.; Visser, R. G. F.; Scott, J. W.; Edwards, J. D.; Bai, Y. The Tomato Yellow Leaf Curl Virus Resistance Genes Ty-1 and Ty-3 Are Allelic and Code for DFDGD-Class RNA-Dependent RNA Polymerases. PLoS Genet. 2013, 9, e1003399. [Google Scholar] [CrossRef] [PubMed]
- Djian-Caporalino, C.; Palloix, A.; Fazari, A.; Marteu, N.; Barbary, A.; Abad, P.; Sage-Palloix, A.-M.; Mateille, T.; Risso, S.; Lanza, R.; Taussig, C.; Castagnone-Sereno, P. Pyramiding, Alternating or Mixing: Comparative Performances of Deployment Strategies of Nematode Resistance Genes to Promote Plant Resistance Efficiency and Durability. BMC Plant Biol. 2014, 14, 53. [Google Scholar] [CrossRef]
- Mapari, A. R.; Mehandi, S. Enhancing Crop Resilience: Advances and Challenges in Marker-Assisted Selection for Disease Resistance. J. Adv. Biol. Biotechnol. 2024, 27, 569–580. [Google Scholar] [CrossRef]
- Ijaz, U.; Sudheesh, S.; Kaur, S.; Sadeque, A.; Bariana, H.; Bansal, U.; Adhikari, K. Mapping of Two New Rust Resistance Genes Uvf-2 and Uvf-3 in Faba Bean. Agronomy (Basel) 2021, 11, 1370. [Google Scholar] [CrossRef]
- Purkaystha, S.; Das, P.; Rashmi, K.; Rout, S.; Nanda, S. Advances in Genetic Mapping of Loci Governing Disease Resistance in Plants. In Biotechnological Advances for Disease Tolerance in Plants; Springer Nature Singapore: Singapore, 2024; pp. 1–27. [Google Scholar]
- Zec, S.; Tamindžić, G.; Azizbekian, S.; Ignjatov, M.; Danojević, D.; Červenski, J.; Vlajić, S.; Vojnović, Đ; Banjac, B. Foliar Application of Ca-Based Fertilizers (Conventional vs. Nanofertilizers): Effects on Fruit Traits, Seed Quality Parameters and Initial Plant Growth of Tomato Genotypes. Horticulturae 2025, 11, 1303. [Google Scholar] [CrossRef]
- Glogovac, S.; Takac, A.; Brbaklic, L.; Trkulja, D.; Cervenski, J.; Gvozdanovic-Varga, J.; Popovic, V. 10.5937/Ratpov50-4344 = Molecular Evaluation of Genetic Variability in Tomato (Lycopersicon Esculentum Mill.) Genotypes by Microsatellite Markers. Ratar. Povrt. 2013, 50, 1–5. [Google Scholar] [CrossRef]
- Ekbiç, E.; Okay, C. Ö. Assessment of Elite Pepper Breeding Lines Using Molecular Markers. Plant Biotechnol. Rep. 2024, 18, 515–524. [Google Scholar] [CrossRef]
- Aktas, H.; Abak, K.; Sensoy, S. Genetic Diversity in Some Turkish Pepper (Capsicum Annuum L.) Genotypes Revealed by AFLP Analyses. African Journal of Biotechnology 2009, 8, 4378–4386. [Google Scholar]
- Bozkalfa, M. K.; Kaygısız Aşcıoğlu, T.; Eşiyok, D. Biber Genotiplerinin Genetik Çeşitliliklerinin Srap Markörleri Kullanılarak Belirlenmesi. ANADOLU J. OF AGRIC. SCI. 2017, 32, 321–329. [Google Scholar] [CrossRef]
- Yildiz, M.; Koçak, M.; Nadeem, M. A.; Cavagnaro, P.; Barboza, K.; Baloch, F. S.; Argün, D.; Keleş, D. Genetic Diversity Analysis in the Turkish Pepper Germplasm Using iPBS Retrotransposon- Based Markers. Turk J. Agric. For. 2020, 44, 1–14. [Google Scholar] [CrossRef]
- Li, X.; Yang, H.; Buirchell, B.; Yan, G. Development of a DNA Marker Tightly Linked to Low-Alkaloid Gene Iucundus in Narrow-Leafed Lupin (Lupinus Angustifolius L.) for Marker-Assisted Selection. Crop Pasture Sci. 2011, 62, 218–224. [Google Scholar] [CrossRef]
- Barka, G. D.; Lee, J. Molecular Marker Development and Gene Cloning for Diverse Disease Resistance in Pepper (Capsicum AnnuumL.): Current Status and Prospects. Plant Breed. Biotechnol. 2020, 8, 89–113. [Google Scholar] [CrossRef]
- Tanaka, Y.; Yoneda, H.; Hosokawa, M.; Miwa, T.; Yazawa, S. Application of Marker-Assisted Selection in Breeding of a New Fresh Pepper Cultivar (Capsicum Annuum) Containing Capsinoids, Low-Pungent Capsaicinoid Analogs. Sci. Hortic. (Amsterdam) 2014, 165, 242–245. [Google Scholar] [CrossRef]
- Paran, I.; van der Voort, J. R.; Lefebvre, V.; Jahn, M.; Landry, L.; van Schriek, M.; Tanyolac, B.; Caranta, C.; Chaim, A. B.; Livingstone, K.; Palloix, A.; Peleman, J. An Integrated Genetic Linkage Map of Pepper (Capsicum Spp.). Mol. Breed. 2004, 13, 251–261. [Google Scholar] [CrossRef]
- Hyeon, N. S.; Woong, Y. J.; Cheorl, K. B.; Byung-Dong, K. Selection of Parental Lines for Hot Pepper Mapping Population Using RFLP and AFLP Analyses. Hortic Environ Biotechnol 1997, 38, 693–697. [Google Scholar]
- Rêgo, E. R.; Rego, M. M.; Farias-Filho, L. P. Genetic Diversity in Pepper (Capsicum Spp.) by Rapd Marker. Acta Hortic. 2011, 918, 341–347. [Google Scholar] [CrossRef]
- Lefebvre, V.; Goffinet, B.; Chauvet, J. C.; Caromel, B.; Signoret, P.; Brand, R.; Palloix, A. Evaluation of Genetic Distances between Pepper Inbred Lines for Cultivar Protection Purposes: Comparison of AFLP, RAPD and Phenotypic Data. Theor. Appl. Genet. 2001, 102, 741–750. [Google Scholar] [CrossRef]
- Tam, S. M.; Mhiri, C.; Vogelaar, A.; Kerkveld, M.; Pearce, S. R.; Grandbastien, M.-A. Comparative Analyses of Genetic Diversities within Tomato and Pepper Collections Detected by Retrotransposon-Based SSAP, AFLP and SSR. Theor. Appl. Genet. 2005, 110, 819–831. [Google Scholar] [CrossRef]
- Dhaliwal, M.S.; Abhay, Y.; S., K. J. Molecular Characterization and Diversity Analysis in Chilli Pepper Using Simple Sequence Repeats (SSR) Markers. Afr. J. Biotechnol. 2014, 13, 3137–3143. [Google Scholar] [CrossRef]
- Villela, J. C. B.; Barbieri, R. L.; Castro, C. M.; Neitzke, R. S.; Vasconcelos, C. S.; Carbonari, T.; Mistura, C. C.; Priori, D. Caracterização Molecular de Variedades Crioulas de Pimentas (Capsicum Baccatum) Com Marcadores Microssatélites. Hortic. Bras. 2014, 32, 131–137. [Google Scholar] [CrossRef]
- Du, H.; Yang, J.; Chen, B.; Zhang, X.; Zhang, J.; Yang, K.; Geng, S.; Wen, C. Target Sequencing Reveals Genetic Diversity, Population Structure, Core-SNP Markers, and Fruit Shape-Associated Loci in Pepper Varieties. BMC Plant Biol. 2019, 19, 578. [Google Scholar] [CrossRef] [PubMed]
- Manivannan, A.; Choi, S.; Jun, T.H.; Yang, E.Y.; Kim, J.H.; Lee, E.S.; Lee, H.E.; Kim, D.S.; Ahn, Y.K. Genotyping by Sequencing-Based Discovery of SNP Markers and Construction of Linkage Map from F5 Population of Pepper with Contrasting Powdery Mildew Resistance Trait. Biomed Res. Int. 2021, 2021, 6673010. [Google Scholar] [CrossRef]
- Xu, X.; Liu, Z.; Lin, X.; Mou, S.; Guan, D.; He, S. Genetic Diversity and Relationship Analysis of Pepper Germplasm Resources Based on Phenotype Traits and SRAP Molecular Markers. J Fujian Agric For Univ 2011, 40, 48–53. [Google Scholar]
- Qiu, Y.H.; Lin, S.T.; Zeng, S.G.; Luo, Y.; Zhu, B.T.; Li, Y.Q. Phenotypic and SRAP genetic diversity analysis of 41 yellow pepper germplasm resources. J South Agric 2021, 52, 2165–2173. [Google Scholar] [CrossRef]
- Maulani, R.; Murti, R. H.; Purwantoro, A. Molecular Diversity in Populations of Chili (Capsicum Annuum l.). SABRAO J. Breed. Genet. 2023, 55, 15–24. [Google Scholar] [CrossRef]
- Danojević, D.; Červenski, J.; Gvozdanović-Varga, J.; Ignjatov, M.; Medić-Pap, S.; Ilić, A.; Bugarski, D.; Takač, A.; Vlajić, S.; Popović, V.; Kiprovski, B.; Bajić, I.; Glogovac, S.; Milošević, D.; Stojanov, N.; Zeremski, T. Nove NS sorte povrtarskih biljnih vrsta, Zbornik referata 56 Savetovanja agronoma i poljoprivrednika Srbije (SAPS) i 2 Savetovanje agronoma Republike Srbije i Republike Srpske. 2022, 121-129, http://fiver.ifvcns.rs/handle/123456789/2548.
- Rani, M.; Jindal, S. K.; Vikal, Y.; Meena, O. P. Genetic Male Sterility Breeding in Heat Tolerant Bell Pepper: Introgression of Ms10 Gene from Hot Pepper through Marker-Assisted Backcrossing. Sci. Hortic. (Amsterdam) 2021, 285, 110172. [Google Scholar] [CrossRef]
- Červenski, J.; Medić-Pap, S.; Danojević, D.; Stojanović, A.; Savić, A. Technological quality of domestic cabbage (Brassica oleracea var. capitata L.) populations and varieties from the Vojvodina Province-Serbia. Contemporary Agriculture. 2014, 63, 473–479, http://fiver.ifvcns.rs/handle/123456789/1305. [Google Scholar]
- Fang, Z.; Liu, Y.; Lou, P.; Liu, G. Current Trends in Cabbage Breeding. J. New Seeds 2004, 6, 75–107. [Google Scholar] [CrossRef]
- Bin, L. J.; Zhang, H. J.; Lin, Y. X.; Wang, S. Y.; Cao, C. S. Molecular Identification of Cytoplasmic Male Sterile (CMS) Type in Common Head Cabbage. Mol. Plant Breed 2009, 7, 1149–1153. [Google Scholar]
- Zhang, Y.; Wang, X. J.; Li, O. C.; Song, H. Y.; Ren, X. S.; Si, J. Molecular Identification of Brassica Oleracea CMS and the Morphologyresponse of Flower to Nuclear Background. Acta Hortic. Sin 2010, 37, 915–922. [Google Scholar]
- Zhang, Y. Y.; Fang, F. Y.; Wang, Q. B.; Liu, Y. M.; Yang, L. M.; Zhuang, M.; Sun, P. T. Molecular Distinction of Two Ogura CMS Sources in Brassica Oleracea Var. Capitata L. Acta Hortic. Sin 2011, 44, 2959–2965. [Google Scholar]
- Chen, C.; Zhuang, M.; Fang, Z.-Y.; Wang, Q.-B.; Zhang, Y.-Y.; Liu, Y.-M.; Yang, L.-M.; Cheng, F. A Co-Dominant Marker BoE332 Applied to Marker-Assisted Selection of Homozygous Male-Sterile Plants in Cabbage (Brassica Oleracea Var. Capitata L.). J. Integr. Agric. 2013, 12, 596–602. [Google Scholar] [CrossRef]
- Yu, H.L.; Li, Z.Y.; Ren, W.J.; Han, F.Q.; Yang, L.M.; Zhuang, M.; Lv, H.H.; Liu, Y.M.; Fang, Z.Y.; Zhang, Y.Y. Creation of Fertility-Restored Materials for Ogura CMS in Brassica Oleracea by Introducing Rfo Gene from Brassica Napus via an Allotriploid Strategy. Theor. Appl. Genet. 2020, 133, 2825–2837. [Google Scholar] [CrossRef]
- Han, F.; Yuan, K.; Kong, C.; Zhang, X.; Yang, L.; Zhuang, M.; Zhang, Y.; Li, Z.; Wang, Y.; Fang, Z.; Lv, H. Fine Mapping and Candidate Gene Identification of the Genic Male-Sterile Gene Ms3 in Cabbage 51S. Theor. Appl. Genet. 2018, 131, 2651–2661. [Google Scholar] [CrossRef]
- Ji, J.L.; Yang, L.M.; Fang, Z.Y.; Zhuang, M.; Zhang, Y.Y.; Lv, H.-H.; Liu, Y.M.; Li, Z.S. Recessive Male Sterility in Cabbage (Brassica Oleracea Var. Capitata) Caused by Loss of Function of BoCYP704B1 Due to the Insertion of a LTR-Retrotransposon. Theor. Appl. Genet. 2017, 130, 1441–1451. [Google Scholar] [CrossRef]
- Wang, X.; Fang, Z.; Sun, P.; Liu, Y.; Yang, L. Identification of a RAPD Marker Linked to a Dominant Male Sterile Gene in Cabbage. Acta Hortic. Sin 1998, 25, 197–198. [Google Scholar]
- Wang, X.; Fang, Z.; Huang, S.; Sun, P.; Liu, Y.; Yang, L.; Zhuang, M.; Qu, D. An Extended Random Primer Amplified Region (ERPAR) Marker Linked to a Dominant Male Sterility Gene in Cabbage (Brassica Oleracea Var. Capitata). Euphytica 2000, 112, 267–273. [Google Scholar] [CrossRef]
- Wang, X.W.; Fang, Z.; Sun, P.; Liu, Y.; Yang, L.; Zhuang, M. A SCAR marker applicable in marker assisted selection of a dominant male sterility gene in cabbage. Acta Hortic. Sin. 2000a, 27, 143–144. [Google Scholar]
- Han, F.; Cui, H.; Zhang, B.; Liu, X.; Yang, L.; Zhuang, M.; Lv, H.; Li, Z.; Wang, Y.; Fang, Z.; Song, J.; Zhang, Y. Map-Based Cloning and Characterization of BoCCD4, a Gene Responsible for White/Yellow Petal Color in B. Oleracea. BMC Genomics 2019a, 20, 242. [Google Scholar] [CrossRef]
- Margalé, E.; Hervé, Y.; Hue, J.; Quiros, C. F. Determination of Genetic Variability by RAPD Markers in Cauliflower, Cabbage and Kale Local Cultivars from France. Genet. Resour. Crop Evol. 1995, 42, 281–289. [Google Scholar] [CrossRef]
- Koutita, O.; Tertivanidis, K.; Koutsos, T. V.; Koutsika-Sotiriou, M.; Skaracis, G. N. Genetic Diversity in Four Cabbage Populations Based on Random Amplified Polymorphic DNA Markers. J. Agric. Sci. 2005, 143, 377–384. [Google Scholar] [CrossRef]
- Saxena, B.; Kaur, R.; Bhardwaj, S. V. Assessment of Genetic Diversity in Cabbage Cultivars Using RAPD and SSR Markers. J. Crop Sci. Biotechnol. 2011, 14, 191–196. [Google Scholar] [CrossRef]
- Li, X.; Yu, H.; Li, Z.; Liu, X.; Fang, Z.; Liu, Y.; Yang, L.; Zhuang, M.; Lv, H.; Zhang, Y. Heterotic Group Classification of 63 Inbred Lines and Hybrid Purity Identification by Using SSR Markers in Winter Cabbage (Brassica Oleracea L. Var. Capitata). Hortic. Plant J 2018a, 4, 158–164. [Google Scholar] [CrossRef]
- Louarn, S.; Torp, A. M.; Holme, I. B.; Andersen, S. B.; Jensen, B. D. Database Derived Microsatellite Markers (SSRs) for Cultivar Differentiation in Brassica Oleracea. Genet. Resour. Crop Evol. 2007, 54, 1717–1725. [Google Scholar] [CrossRef]
- Flannery, M. L.; Mitchell, F. J. G.; Coyne, S.; Kavanagh, T. A.; Burke, J. I.; Salamin, N.; Dowding, P.; Hodkinson, T. R. Plastid Genome Characterisation in Brassica and Brassicaceae Using a New Set of Nine SSRs. Theor. Appl. Genet. 2006, 113, 1221–1231. [Google Scholar] [CrossRef]
- El-Esawi, M. A.; Germaine, K.; Bourke, P.; Malone, R. Genetic Diversity and Population Structure of Brassica Oleracea Germplasm in Ireland Using SSR Markers. C. R. Biol. 2016, 339, 133–140. [Google Scholar] [CrossRef]
- Leroy, X. J.; Leon, K.; Branchard, M. Characterisation of Brassica Oleracea L. by Microsatellite Primers. Plant Syst. Evol 2000, 225, 235–240. [Google Scholar] [CrossRef]
- Faltusová, Z.; Kučera, L.; Ovesná, J. Genetic Diversity of Brassica Oleracea Var. Capitata Gene Bank Accessions Assessed by AFLP. Electron. J. Biotechnol. 2011, 14. [Google Scholar] [CrossRef]
- Van Hintum, T. J. L.; van de Wiel, C. C. M.; Visser, D. L.; van Treuren, R.; Vosman, B. The Distribution of Genetic Diversity in a Brassica Oleracea Gene Bank Collection Related to the Effects on Diversity of Regeneration, as Measured with AFLPs. Theor. Appl. Genet. 2007, 114, 777–786. [Google Scholar] [CrossRef]
- Shapturenko, M. N.; Pechkovskaya, T. V.; Vakula, S. I.; Jakimovich, A. V.; Zabara, Y. M.; Khotyleva, L. V. Informative EST-SSR Markers for Genotyping and Intraspecific Differentiation of Brassica Oleracea Var. Capitata L. Russ. J. Genet. Appl. Res. 2017, 7, 14–20. [Google Scholar] [CrossRef]
- Palmé, A. E.; Hagenblad, J.; Solberg, S. Ø; Aloisi, K.; Artemyeva, A. SNP Markers and Evaluation of Duplicate Holdings of Brassica Oleracea in Two European Genebanks. Plants 2020, 9, 925. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Yu, H.; Li, X.; Zhang, B.; Ren, W.; Liu, X.; Fang, Z.; Yang, L.; Zhuang, M.; Lv, H.; Zhang, Y. Kompetitive Allele-Specific PCR (KASP) Genotyping and Heterotic Group Classification of 244 Inbred Lines in Cabbage (Brassica Oleracea L. Var. Capitata). Euphytica 2020a, 216. [Google Scholar] [CrossRef]
- Crockett, P. A.; Bhalla, P. L.; Lee, C. K.; Singh, M. B. RAPD Analysis of Seed Purity in a Commercial Hybrid Cabbage (Brassica Oleracea Var. Capitata) Cultivar. Genome 2000, 43, 317–321. [Google Scholar] [CrossRef]
- Liu, G.; Liu, L.; Gong, Y.; Wang, Y.; Yu, F.; Shen, H.; Gui, W. Seed Genetic Purity Testing of F1 Hybrid Cabbage (Brassica Oleracea Var. Capitata) with Molecular Marker Analysis. Seed Sci. Technol. 2007, 35, 477–486. [Google Scholar] [CrossRef]
- Ye, S.; Wang, Y.; Huang, D.; Li, J.; Gong, Y.; Xu, L.; Liu, L. Genetic Purity Testing of F1 Hybrid Seed with Molecular Markers in Cabbage (Brassica Oleracea Var. Capitata). Sci. Hortic. (Amsterdam) 2013, 155, 92–96. [Google Scholar] [CrossRef]
- Kawamura, K.; Shimizu, M.; Kawanabe, T.; Pu, Z.; Kodama, T.; Kaji, M.; Osabe, K.; Fujimoto, R.; Okazaki, K. Assessment of DNA Markers for Seed Contamination Testing and Selection of Disease Resistance in Cabbage. Euphytica 2017, 213. [Google Scholar] [CrossRef]
- Cansian, R. L.; Echeverrigaray, S. Discrimination among Cultivars of Cabbage Using Randomly Amplified Polymorphic DNA Markers. HortScience 2000, 35, 1155–1158. [Google Scholar] [CrossRef]
- Li, Z.; Yu, H.; Fang, Z.; Yang, L.; Liu, Y.; Zhuang, M.; Lü, H.; Zhang, Y. Development of SNP Markers in Cabbage and Construction of DNA Fingerprinting of Main Varieties. China J. Agric. Sci 2018b, 51, 2771–2787. [Google Scholar]
- Jo, J.; Kang, M.-Y.; Kim, K. S.; Youk, H. R.; Shim, E.-J.; Kim, H.; Park, J.-S.; Sim, S.-C.; Yu, B. C.; Jung, J.-K. Genome-Wide Analysis-Based Single Nucleotide Polymorphism Marker Sets to Identify Diverse Genotypes in Cabbage Cultivars (Brassica Oleracea Var. Capitata). Sci. Rep. 2022, 12, 20030. [Google Scholar] [CrossRef] [PubMed]
- Han, F.; Zhang, X.; Yuan, K.; Fang, Z.; Yang, L.; Zhuang, M.; Zhang, Y.; Wang, Y.; Liu, Y.; Li, Z.; Lv, H. A User-Friendly KASP Molecular Marker Developed for the DGMS-Based Breeding System in Brassica Oleracea Species. Mol. Breed. 2019b, 39. [Google Scholar] [CrossRef]
- Han, F.Q.; Yang, C.; Fang, Z.Y.; Yang, L.M.; Zhuang, M.; Lv, H.H.; Liu, Y.M.; Li, Z.S.; Liu, B.; Yu, H.L.; Liu, X.P.; Zhang, Y.Y. Inheritance and InDel Markers Closely Linked to Petal Color Gene (Cpc-1) in Brassica Oleracea. Mol. Breed. 2015, 35. [Google Scholar] [CrossRef]
- Liu, D.; Tang, J.; Liu, Z.; Dong, X.; Zhuang, M.; Zhang, Y.; Lv, H.; Sun, P.; Liu, Y.; Li, Z. Fine Mapping of BoGL1, a Gene Controlling the Glossy Green Trait in Cabbage (Brassica Oleracea L. Var. Capitata). Mol. Breed 2017a, 37, 69. [Google Scholar] [CrossRef]
- Liu, Z.; Fang, Z.; Zhuang, M.; Zhang, Y.; Lv, H.; Liu, Y.; Li, Z.; Sun, P.; Tang, J.; Liu, D. Fine-Mapping and Analysis of Cgl1, a Gene Conferring Glossy Trait in Cabbage (Brassica Oleracea L. Var. Capitata). Front. Plant Sci 2017b, 8. [Google Scholar] [CrossRef]
- Liu, D.; Tang, J.; Liu, Z.; Dong, X.; Zhuang, M.; Zhang, Y.; Lv, H.; Sun, P.; Liu, Y.; Li, Z.; Ye, Z.; Fang, Z.; Yang, L. Cgl2 Plays an Essential Role in Cuticular Wax Biosynthesis in Cabbage (Brassica Oleracea L. Var. Capitata). BMC Plant Biol. 2017c, 17, 223. [Google Scholar] [CrossRef]
- Liu, D.; Dong, X.; Liu, Z.; Tang, J.; Zhuang, M.; Zhang, Y.; Lv, H.; Liu, Y.; Li, Z.; Fang, Z.; Yang, L. Fine Mapping and Candidate Gene Identification for Wax Biosynthesis Locus, BoWax1 in Brassica Oleracea L. Var. Capitata. Front. Plant Sci. 2018, 9, 309. [Google Scholar] [CrossRef]
- Ji, J.; Cao, W.; Dong, X.; Liu, Z.; Fang, Z.; Zhuang, M.; Zhang, Y.; Lv, H.; Wang, Y.; Sun, P.; Liu, Y.; Li, Z.; Yang, L. A 252-Bp Insertion in BoCER1 Is Responsible for the Glossy Phenotype in Cabbage (Brassica Oleracea L. Var. Capitata). Mol. Breed. 2018, 38. [Google Scholar] [CrossRef]
- Dong, X.; Ji, J.; Yang, L.; Fang, Z.; Zhuang, M.; Zhang, Y.; Lv, H.; Wang, Y.; Sun, P.; Tang, J.; Liu, D.; Liu, Y.; Li, Z. Fine-Mapping and Transcriptome Analysis of BoGL-3, a Wax-Less Gene in Cabbage (Brassica Oleracea L. Var. Capitata). Mol. Genet. Genomics 2019, 294, 1231–1239. [Google Scholar] [CrossRef]
- Ji, J.; Cao, W.; Tong, L.; Fang, Z.; Zhang, Y.; Zhuang, M.; Wang, Y.; Yang, L.; Lv, H. Identification and Validation of an ECERIFERUM2- LIKE Gene Controlling Cuticular Wax Biosynthesis in Cabbage (Brassica Oleracea L. Var. Capitata L.). Theor. Appl. Genet. 2021, 134, 4055–4066. [Google Scholar] [CrossRef]
- Liu, X.; Yang, C.; Han, F.; Fang, Z.; Yang, L.; Zhuang, M.; Lv, H.; Liu, Y.; Li, Z.; Zhang, Y. Genetics and Fine Mapping of a Yellow-Green Leaf Gene (Ygl-1) in Cabbage (Brassica Oleracea Var. Capitata L.). Mol. Breed 2016. [Google Scholar] [CrossRef]
- Song, H.; Yi, H.; Lee, M.; Han, C.T.; Lee, J.; Kim, H.; Park, J.I.; Nou, I.S.; Kim, S.J.; Hur, Y. Purple Brassica Oleracea Var. Capitata F. Rubra Is Due to the Loss of BoMYBL2-1 Expression. BMC Plant Biol. 2018a, 18, 82. [Google Scholar] [CrossRef] [PubMed]
- Pang, W.; Li, X.; Choi, S. R.; Nguyen, V. D.; Dhandapani, V.; Kim, Y. Y.; Ramchiary, N.; Kim, J. G.; Edwards, D.; Batley, J. Mapping QTLs of Resistance to Head Splitting in Cabbage (Brassica Oleracea Var. Capitata L.). Mol. Breed 2015, 35, 126. [Google Scholar] [CrossRef]
- Zhu, X.; Tai, X.; Ren, Y.; Wang, M.; Chen, J.; Bo, T. QTL-Seq and Marker Development for Resistance to Head Splitting in Cabbage. Euphytica 2022a, 218. [Google Scholar] [CrossRef]
- Zhao, T.; Miao, L.; Zou, M.; Hussain, I.; Yu, H.; Li, J.; Sun, N.; Kong, L.; Wang, S.; Li, J. Development of SSRs Based on the Whole Genome and Screening of Bolting-Resistant SSR Marker in Brassica Oleracea L. Horticulturae 2024, 10. [Google Scholar] [CrossRef]
- Li, Q.; Peng, A.; Yang, J.; Zheng, S.; Li, Z.; Mu, Y.; Chen, L.; Si, J.; Ren, X.; Song, H. A 215-Bp Indel at Intron I of BoFLC2 Affects Flowering Time in Brassica Oleracea Var. Capitata during Vernalization. Theor. Appl. Genet. 2022, 135, 2785–2797. [Google Scholar] [CrossRef]
- Song, H.; Lee, M.; Hwang, B. H.; Han, C. T.; Park, J. I.; Hur, Y. Development and Application of a PCR-Based Molecular Marker for the Identification of High Temperature Tolerant Cabbage (Brassica Oleracea Var. Capitata) Genotypes. Agronomy 2020a, 10. [Google Scholar] [CrossRef]
- Song, H.; Kim, H.; Hwang, B.-H.; Yi, H.; Hur, Y. Natural Variation in Glycine-Rich Region of Brassica Oleracea Cold Shock Domain Protein 5 (BoCSDP5) Is Associated with Low Temperature Tolerance. Genes Genomics 2020b, 42(12), 1407–1417. [Google Scholar] [CrossRef]
- Song, H.; Yi, H.; Han, C.T.; Park, J.I.; Hur, Y. Allelic Variation in Brassica Oleracea CIRCADIAN CLOCK ASSOCIATED 1 (BoCCA1) Is Associated with Freezing Tolerance. Hortic. Environ. Biotechnol. 2018b, 59, 423–434. [Google Scholar] [CrossRef]
- Jiang, M.; Zhao, Y.; Xie, J.; Tian, R.; Chen, Y.; Kang, J. Development of a SCAR Marker for Fusarium Wilt Resistance in Cabbage. Sci. Agric. Sin 2011, 44, 3053–3059. [Google Scholar]
- Pu, Z.J.; Shimizu, M.; Zhang, Y.J.; Nagaoka, T.; Hayashi, T.; Hori, H.; Matsumoto, S.; Fujimoto, R.; Okazaki, K. Genetic Mapping of a Fusarium Wilt Resistance Gene in Brassica Oleracea. Mol. Breed. 2012, 30, 809–818. [Google Scholar] [CrossRef]
- Liu, X.; Han, F.; Kong, C.; Fang, Z.; Yang, L.; Zhang, Y.; Zhuang, M.; Liu, Y.; Li, Z.; Lv, H. Rapid Introgression of the Fusarium Wilt Resistance Gene into an Elite Cabbage Line through the Combined Application of a Microspore Culture, Genome Background Analysis, and Disease Resistance-Specific Marker Assisted Foreground Selection. Front. Plant Sci. 2017, 8, 354. [Google Scholar] [CrossRef]
- Dubina, E. V.; Makukha, Y. A.; Artemeva, A. M.; Fateev, D. A.; Garkusha, S. V.; Gorun, O. L.; Lesnyak, S. A. Molecular Marking in Brassica Oleracea L. Breeding for Resistance to Fusarium Wilt. Russ. J. Genet 2023, 59, 1004–1010. [Google Scholar] [CrossRef]
- Lv, H.H.; Yang, L.M.; Kang, J.G.; Wang, Q.B.; Wang, X.W.; Fang, Z.Y.; Liu, Y.M.; Zhuang, M.; Zhang, Y.Y.; Lin, Y.; Yang, Y.H.; Xie, B.Y.; Liu, B.; Liu, J.S. Development of InDel Markers Linked to Fusarium Wilt Resistance in Cabbage. Mol. Breed. 2013, 32, 961–967. [Google Scholar] [CrossRef]
- Sato, M.; Shimizu, M.; Shea, D. J.; Hoque, M.; Kawanabe, T.; Miyaji, N.; Fujimoto, R.; Fukai, E.; Okazaki, K. Allele Specific DNA Marker for Fusarium Resistance Gene FocBo1 in Brassica Oleracea. Breed. Sci. 2019, 69, 308–315. [Google Scholar] [CrossRef] [PubMed]
- Afrin, K. S.; Rahim, M. A.; Park, J.-I.; Natarajan, S.; Rubel, M. H.; Kim, H.-T.; Nou, I.-S. Screening of Cabbage (Brassica Oleracea L.) Germplasm for Resistance to Black Rot. Plant Breed. Biotechnol. 2018, 6, 30–43. [Google Scholar] [CrossRef]
- Makukha, Y. PCR Identification of Genes of Resistance to Black Rot in White Cabbage Using SSR-Markers. BIO Web Conf. 2020, 21, 00013. [Google Scholar] [CrossRef]
- Hong, J.E.; Afrin, K. S.; Rahim, M. A.; Jung, H.J.; Nou, I.S. Inheritance of Black Rot Resistance and Development of Molecular Marker Linked to Xcc Races 6 and 7 Resistance in Cabbage. Plants 2021, 10, 1940. [Google Scholar] [CrossRef]
- Nomura, K.; Minegishi, Y.; Kimizuka-Takagi, C.; Fujioka, T.; Moriguchi, K.; Shishido, R.; Ikehashi, H. Evaluation of F2 and F3 Plants Introgressed with QTLs for Clubroot Resistance in Cabbage Developed by Using SCAR Markers. Plant Breed. 2005, 124, 371–375. [Google Scholar] [CrossRef]
- Tomita, H.; Shimizu, M.; Asad-Ud Doullah, M. D.; Fujimoto, R.; Okazaki, K. Accumulation of Quantitative Trait Loci Confer-Ring Broad-Spectrum Clubroot Resistance in Brassica Oleracea. Mol. Breed 2013, 32, 889–900. [Google Scholar] [CrossRef]
- Nagaoka, T.; Doullah, M. A. U.; Matsumoto, S.; Kawasaki, S.; Ishikawa, T.; Hori, H.; Okazaki, K. Identification of QTLs That Control Clubroot Resistance in Brassica Oleracea and Comparative Analysis of Clubroot Resistance Genes between B. Rapa and B. Oleracea. Theor. Appl. Genet. 2010, 120, 1335–1346. [Google Scholar] [CrossRef]
- Zhu, M.; Yang, L.; Zhang, Y.; Zhuang, M.; Ji, J.; Hou, X.; Li, Z.; Han, F.; Fang, Z.; Lv, H.; Wang, Y. Introgression of Clubroot Resistant Gene into Brassica Oleracea L. from Brassica Rapa Based on Homoeologous Exchange. Hortic. Res. 2022b, 9, uhac195. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Zhang, B.; Yang, L.; Zhuang, M.; Lv, H.; Wang, Y.; Ji, J.; Hou, X.; Zhang, Y. Fine Mapping and Identification of the Downy Mildew Resistance Gene BoDMR2 in Cabbage (Brassica Oleracea L. Var. Capitata). BMC Plant Biol. 2024, 24, 987. [Google Scholar] [CrossRef] [PubMed]
- Gomez, G.A. Downy Mildew Resistance in B. oleracea. Ph.D. Thesis, The University of East Anglia, Norwich, UK, 2022. [Google Scholar]
- Strembovskiy, I. V.; Kroupin, P. Y. The Current Status and Prospects of Molecular Marker Applications in Head Cabbage (Brassica Oleracea Var. Capitata L.): A Review. Agronomy (Basel) 2025, 15, 2644. [Google Scholar] [CrossRef]
- Shatilov, M. V.; Razin, A. F.; Ivanova, M. I. Analysis of the World Lettuce Market. IOP Conf. Ser. Earth Environ. Sci. 2019, 395, 012053. [Google Scholar] [CrossRef]
- Lebeda, A.; Ryder, E. J.; Grube, R. Lettuce (Asteraceae;Lactuca Spp). In Genetic resources, chromosome engineering and crop improvement; Singh, R. J., Ed.; CRC Press, 2007, 3, 377–472.
- Hartman, Y.; Hooftman, D.; Uwimana, B. Abiotic Stress QTL in Lettuce Crop-Wild Hybrids: Comparing Green-House and Field Experiments. Ecology Evolution 2014, 4, 2395–2409. [Google Scholar] [CrossRef]
- McCabe, M. S.; Garratt, L. C.; Schepers, F.; Jordi, W. J. R. M.; Stoopen, G. M.; Davelaar, E.; van Rhijn, J. H. A.; Power, J. B.; Davey, M. R. Effects of PSAG12-IPT Gene Expression on Development and Senescence in Transgenic Lettuce. Plant Physiol. 2001, 127, 505–516. [Google Scholar] [CrossRef]
- Hassan, M. N.; Mekkawy, S. A.; Mahdy, M.; Salem, K. F. M.; Tawfik, E. Recent Molecular and Breeding Strategies in Lettuce (Lactuca Spp.). Genet. Resour. Crop Evol. 2021, 68, 3055–3079. [Google Scholar] [CrossRef]
- Landry, B. S.; Kesseli, R.; Farrara, B.; Michelmore, R. W. A Genetic Map of Lettuce (Lactuca Sativa L.) with Restriction Frag-Ment Length Polymorphism, Isozymes, Disease Resistance and Morphological Markers. Genetics 1987, 116, 331–337. [Google Scholar] [CrossRef]
- Kesseli, R.; Ochoa, O.; Michelmore, R. Variation at RFLP Loci in Lactuca Spp. and Origin of Cultivated Lettuce (L.Sativa). Genome 1991, 34, 430–436. [Google Scholar] [CrossRef]
- Vermeulen, A.; Desprez, B.; Lancelin, D.; Bannerot, H. Relationship among Cichorium Species and Related Genera as De-Termined by Analysis of Mitochondrial RFLPs. Theor Appl Genet 1994, 88, 159–166. [Google Scholar] [CrossRef]
- Kesseli, R. V.; Paran, I.; Michelmore, R. W. Analysis of a Detailed Genetic Linkage Map of Lactuca Sativa (Lettuce) Con-Structed from RFLP and RAPD Markers. Genetics 1994, 136, 1435–1446. [Google Scholar] [CrossRef] [PubMed]
- El-Esawi, M. Molecular Genetic Markers for Assessing the Genetic Variation and Relationships in Lactuca Germplasm. Annu. Res. Rev. Biol. 2015, 8, 1–13. [Google Scholar] [CrossRef]
- Yamamoto, T.; Nishikawa, A.; Oeda, K. DNA Polymorphisms in Oryza Sativa L. and Lactuca Sativa L. Amplified by Arbitrary Primed PCR. Euphytica 1994, 78, 143–148. [Google Scholar] [CrossRef]
- Tardin, F. D.; Junior, A.; Pereira, M. G. Genetic Diversity and Determination of the Optimum Number of RAPD Markers in Lettuce (Lactuca Sativa L.). Acta Scientiarum: Agronomy Maringa’ 2003, 25, 1–5. [Google Scholar]
- Hill, M.; Witsenboer, H.; Zabeau, M.; Vos, P.; Kesseli, R.; Michelmore, R. PCR-Based Fingerprinting Using AFLPs as a Tool for Studying Genetic Relationships in Lactuca Spp. Theor. Appl. Genet. 1996, 93, 1202–1210. [Google Scholar] [CrossRef]
- Jeuken, M.; Van Wijk, R.; Peleman, J.; Lindhout, P. An Integrated Interspecific AFLP Map of Lettuce (Lactuca) Based on Two L. Sativa 9 L. Saligna F2 Populations. Theor Appl Genet 2001, 103, 638–647. [Google Scholar] [CrossRef]
- Koopman, W.; Zevenbergen, M. J.; Ronald, G.; Van Den Berg, R. G. Species Relationship in Lactuca s. l. (Lactuceae, Asteraceae) Inferred from AFLP Fingerprints. Am J Bot 2001, 88, 1881–1887. [Google Scholar] [CrossRef]
- Van Treuren, R.; Hintum, V. Comparison of Anonymous and Targeted Molecular Markers for the Estimation of Genetic Diversity in Ex-Situ Conserved Lactuca. Theor Appl Genet 2009, 119, 1265–1279. [Google Scholar] [CrossRef]
- Witsenboer, H.; Vogel, J.; Michelmore, R. W. Identification, Genetic Localization, and Allelic Diversity of Selectively Am-Plified Microsatellite Polymorphic Loci in Lettuce and Wild Relatives (Lactuca Spp). Genome 1997, 40, 923–936. [Google Scholar] [CrossRef]
- Van de Wiel, C.; Arens, P.; Vosman, B. Microsatellite Fingerprinting in Lettuce (Lactuca Sativa L.) and Wild Relatives. Plant Cell Rep. 1998, 17, 837–842. [Google Scholar] [CrossRef]
- Van de Wiel, C.; Arens, P.; Vosman, B. Microsatellite Retrieval in Lettuce (Lactuca Sativa L.). Genome 1999, 42, 139–149. [Google Scholar] [CrossRef]
- Rauscher, G.; Simko, I. Development of Genomic SSR Markers for Fingerprinting Lettuce (Lactuca Sativa L.) Cultivars and Mapping Genes. BMC Plant Biol. 2013, 13, 11. [Google Scholar] [CrossRef] [PubMed]
- Rui, S.; Qi, G.; Shuangxi, F.; Chaojie, L.; Yingyan, H. Analysis of Genetic Diversity in Purple Lettuce (Lactuca Sativa L.) by SSR Markers. Pak. J. Bot. 2020, 52. [Google Scholar] [CrossRef] [PubMed]
- Simko, I. Development of EST-SSR Markers for the Study of Population Structure in Lettuce (Lactuca Sativa L.). J. Hered. 2009, 100, 256–262. [Google Scholar] [CrossRef] [PubMed]
- Riar, D. S.; Rustgi, S.; Burke, I. C.; Gill, K. S.; Yenish, J. P. EST-SSR Development from 5 Lactuca Species and Their Use in Studying Genetic Diversity among L. Serriola Biotypes. J. Hered. 2011, 102, 17–28. [Google Scholar] [CrossRef]
- Moreno-Vazquez, S.; Ochoa, O. E.; Faber, N. SNP-Based Codominant Markers for a Recessive Gene Conferring Re-Sistance to Corky Root Rot (Rhizomonas Suberifaciens) in Lettuce (Lactuca Sativa). Genome 2003, 46, 1059–1069. [Google Scholar] [CrossRef]
- Simko, I.; Pechenick, D. A.; McHale, L. K.; Truco, M. J.; Ochoa, O. E.; Michelmore, R. W.; Scheffler, B. E. Development of Molecular Markers for Marker-Assisted Selection of Dieback Disease Resistance in Lettuce (Lactuca Sativa). Acta Hortic. 2010, 859, 401–408. [Google Scholar] [CrossRef]
- Walley, P. G.; Hough, G.; Moore, J. D.; Carder, J.; Elliott, M.; Mead, A.; Jones, J.; Teakle, G.; Barker, G.; Buchanan-Wollaston, V.; Hand, P.; Pink, D.; Collier, R. Towards New Sources of Resistance to the Currant-Lettuce Aphid (Nasonovia Ribisnigri). Mol. Breed. 2017, 37, 4. [Google Scholar] [CrossRef]
- Liu, L.; Liu, Z.; Chen, H.; Zhou, L.; Liu, Y.; Luo, L. SRAP Markers and Morphological Traits Could Be Used in Test of Distinctiveness, Uniformity, and Stability (DUS) of Lettuce (Lactuca Sativa) Varieties. J. Agric. Sci. 2011, 4. [Google Scholar] [CrossRef]
- Salem, K. F. M.; Alghuthaymi, M. A.; Mahdy, M.; Mekkawy, S. A.; Hassan, M. N.; Ibrahim, A. A.; Tawfik, E. Advances in Lettuce (Lactuca Spp.) Molecular Breeding Strategies. In Smart Plant Breeding for Vegetable Crops in Post-genomics Era; Springer Nature Singapore: Singapore, 2023, 251–277. [CrossRef]
- Meuwissen, T. H.; Hayes, B. J.; Goddard, M. E. Prediction of Total Genetic Value Using Genome-Wide Dense Marker Maps. Genetics 2001, 157, 1819–1829. [Google Scholar] [CrossRef] [PubMed]
- Heffner, E. L.; Sorrells, M. E.; Jannink, J.-L. Genomic Selection for Crop Improvement. Crop Sci. 2009, 49, 1–12. [Google Scholar] [CrossRef]
- Varshney, R. K.; Bohra, A.; Yu, J.; Graner, A.; Zhang, Q.; Sorrells, M. E. Designing Future Crops: Genomics-Assisted Breeding Comes of Age. Trends Plant Sci. 2021, 26, 631–649. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Du, S.; Wei, A.; Cheng, Z.; Meng, H.; Han, Y. Hybrid Prediction in Horticulture Crop Breeding: Progress and Challenges. Plants 2024, 13, 2790. [Google Scholar] [CrossRef]
- Kondo, F.; Kumanomido, Y.; D’Andrea, M.; Palombo, V.; Ahmed, N.; Futatsuyama, S.; Nemoto, K.; Matsushima, K. Phenotypic Simulation for Fruit-Related Traits in F1 Progenies of Chili Peppers (Capsicum Annuum) Using Genomic Prediction Based Solely on Parental Information. Mol. Genet. Genomics 2025, 300, 15. [Google Scholar] [CrossRef]
- Zhao, Y.; Li, Z.; Liu, G.; Jiang, Y.; Maurer, H. P.; Würschum, T.; Mock, H.-P.; Matros, A.; Ebmeyer, E.; Schachschneider, R.; Kazman, E.; Schacht, J.; Gowda, M.; Longin, C. F. H.; Reif, J. C. Genome-Based Establishment of a High-Yielding Heterotic Pattern for Hybrid Wheat Breeding. Proc. Natl. Acad. Sci. U. S. A. 2015, 112, 15624–15629. [Google Scholar] [CrossRef]
- Yue, L.; Zhang, S.; Zhang, L.; Liu, Y.; Cheng, F.; Li, G.; Zhang, S.; Zhang, H.; Sun, R.; Li, F. Heterotic Prediction of Hybrid Performance Based on Genome-Wide SNP Markers and the Phenotype of Parental Inbred Lines in Heading Chinese Cabbage (Brassica Rapa L. Ssp. Pekinensis). Sci. Hortic. (Amsterdam) 2022, 296, 110907. [Google Scholar] [CrossRef]
- Crossa, J.; Pérez-Rodríguez, P.; Cuevas, J.; Montesinos-López, O.; Jarquín, D.; de Los Campos, G.; Burgueño, J.; González-Camacho, J. M.; Pérez-Elizalde, S.; Beyene, Y.; Dreisigacker, S.; Singh, R.; Zhang, X.; Gowda, M.; Roorkiwal, M.; Rutkoski, J.; Varshney, R. K. Genomic Selection in Plant Breeding: Methods, Models, and Perspectives. Trends Plant Sci. 2017, 22, 961–975. [Google Scholar] [CrossRef]
- Paspureddy, S. L.; Sasirekhamani, M.; Sneha, P. I.; Deepanshu; Sharma, A.; Harshavardhan, S. S.; Chandravanshi, R.; Sahu, A. Advances in Haploid and Doubled Haploid Technology for Accelerating Breeding Cycles in Crop Improvement. Plant Cell Biotechnol. Mol. Biol. 2025, 26, 61–77. [Google Scholar] [CrossRef]
- Singh, S.; Dey, S. S.; Bhatia, R.; Kumar, R.; Sharma, K.; Behera, T. K. Heterosis and Combining Ability in Cytoplasmic Male Sterile and Doubled Haploid Based Brassica Oleracea Progenies and Prediction of Heterosis Using Microsatellites. PLoS One 2019, 14, e0210772. [Google Scholar] [CrossRef]
- Persa, R.; Ribeiro, P. C. de O.; Jarquin, D. The Use of High-Throughput Phenotyping in Genomic Selection Context. Crop Breed. Appl. Biotechnol. 2021, 21. [Google Scholar] [CrossRef]
- Yoosefzadeh Najafabadi, M.; Hesami, M.; Eskandari, M. Machine Learning-Assisted Approaches in Modernized Plant Breeding Programs. Genes (Basel) 2023, 14, 777. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Yang, F.; Li, J.; Liu, Z.; Han, Y.; Zhang, Q.; Pan, S.; Zhao, X.; Wang, K. Progress and Perspectives on Genomic Selection Models for Crop Breeding. Technology in Agronomy 2025, 5, 0–0. [Google Scholar] [CrossRef]
- Jaganathan, D.; Dutta, B.; Basu, S.; Bag, S.; Srinivasan, R.; Eybishitz, A.; Barchenger, D. W.; Caarls, L.; Simmons, A. M.; Nankar, A. N. Breeding Vegetables for Whitefly Resistance: Past, Present, and Future in the AI Era. Front. Plant Sci. 2025, 16, 1724403. [Google Scholar] [CrossRef] [PubMed]
- Wan, L.; Wang, Z.; Tang, M.; Hong, D.; Sun, Y.; Ren, J.; Zhang, N.; Zeng, H. CRISPR-Cas9 Gene Editing for Fruit and Vegetable Crops: Strategies and Prospects. Horticulturae 2021, 7, 193. [Google Scholar] [CrossRef]
- Li, H.; Yang, X.; Shang, Y.; Zhang, Z.; Huang, S. Vegetable Biology and Breeding in the Genomics Era. Sci. China Life Sci. 2023, 66, 226–250. [Google Scholar] [CrossRef]
- Chaudhuri, A.; Halder, K.; Datta, A. Classification of CRISPR/Cas System and Its Application in Tomato Breeding. Theor. Appl. Genet. 2022, 135, 367–387. [Google Scholar] [CrossRef]
- Brooks, C.; Nekrasov, V.; Lippman, Z. B.; Van Eck, J. Efficient Gene Editing in Tomato in the First Generation Using the Clus-Tered Regularly Interspaced Short Palindromic Repeats/CRISPR-Associated9 System. Plant Physiol 2014, 166, 1292–1297. [Google Scholar] [CrossRef]
- De Toledo Thomazella, P.; Brail, D.; Dahlbeck, Q.; Staskawicz, D. CRISPR-Cas9 Mediated Mutagenesis of a DMR6 Ortholog in Tomato Confers Broad-Spectrum Disease Resistance. BioRxiv 2016. [Google Scholar]
- Nekrasov, V.; Wang, C.; Win, J.; Lanz, C.; Weigel, D.; Kamoun, S. Rapid Generation of a Transgene-Free Powdery Mildew Resistant Tomato by Genome Deletion. Sci. Rep. 2017, 7, 482. [Google Scholar] [CrossRef]
- Prihatna, C.; Barbetti, M. J.; Barker, S. J. A Novel Tomato Fusarium Wilt Tolerance Gene. Front. Microbiol. 2018, 9, 1226. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Wang, L.; Zhao, R.; Yu, W.; Li, R.; Li, Y.; Sheng, J.; Shen, L. Knockout of SlMAPK3 Reduced Disease Resistance to Botrytis Cinerea in Tomato Plants. J. Agric. Food Chem. 2018, 66, 8949–8956. [Google Scholar] [CrossRef] [PubMed]
- Yin, Y.; Qin, K.; Song, X.; Zhang, Q.; Zhou, Y.; Xia, X.; Yu, J. BZR1 Transcription Factor Regulates Heat Stress Tolerance Through FERONIA Receptor-Like Kinase-Mediated Reactive Oxygen Species Signaling in Tomato. Plant Cell Physiol 2018, 59, 2239–2254. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Zhang, L.; Wang, L.; Chen, L.; Zhao, R.; Sheng, J.; Shen, L. Reduction of Tomato-Plant Chilling Tolerance by CRISPR-Cas9-Mediated SlCBF1 Mutagenesis. J. Agric. Food Chem. 2018c, 66, 9042–9051. [Google Scholar] [CrossRef]
- Wang, L.; Chen, L.; Li, R.; Zhao, R.; Yang, M.; Sheng, J.; Shen, L. Reduced Drought Tolerance by CRISPR/Cas9-Mediated SlMAPK3 Mutagenesis in Tomato Plants. J. Agric. Food Chem. 2017, 65, 8674–8682. [Google Scholar] [CrossRef]
- Butler, N. M.; Atkins, P. A.; Voytas, D. F.; Douches, D. S. Generation and Inheritance of Targeted Mutations in Potato (Sola-Num Tuberosum L.) Using the CRISPR/Cas System. PLoS ONE 2015, 10. [Google Scholar] [CrossRef]
- Bari, V. K.; Nassar, J. A.; Kheredin, S. M.; Gal-On, A.; Ron, M.; Britt, A.; Steele, D.; Yoder, J.; Aly, R. CRISPR/Cas9-Mediated Mutagenesis of CAROTENOID CLEAVAGE DIOXYGENASE 8 in Tomato Provides Resistance against the Parasitic Weed Phelipanche Aegyptiaca. Sci. Rep. 2019, 9, 11438. [Google Scholar] [CrossRef]
- Bari, V. K.; Nassar, J. A.; Aly, R. CRISPR/Cas9 Mediated Mutagenesis of MORE AXILLARY GROWTH 1 in Tomato Confers Resistance to Root Parasitic Weed Phelipanche Aegyptiaca. Sci. Rep. 2021, 11, 3905. [Google Scholar] [CrossRef]
- Li, T.; Yang, X.; Yu, Y.; Si, X.; Zhai, X.; Zhang, H.; Dong, W.; Gao, C.; Xu, C. Domestication of Wild Tomato Is Accelerated by Genome Editing. Nat. Biotechnol. 2018d, 36, 1160–1163. [Google Scholar] [CrossRef]
- Rodríguez-Leal, D.; Lemmon, Z. H.; Man, J.; Bartlett, M. E.; Lippman, Z. B. Engineering Quantitative Trait Variation for Crop Improvement by Genome Editing. Cell 2017, 171, 470–480.e8. [Google Scholar] [CrossRef]
- Čermák, T.; Baltes, N. J.; Čegan, R.; Zhang, Y.; Voytas, D. F. High-Frequency, Precise Modification of the Tomato Genome. Genome Biol. 2015, 16(1), 232. [Google Scholar] [CrossRef]
- Deng, L.; Wang, H.; Sun, C.; Li, Q.; Jiang, H.; Du, M.; Li, C.B.; Li, C. Efficient Generation of Pink-Fruited Tomatoes Using CRISPR/Cas9 System. J. Genet. Genomics 2018, 45, 51–54. [Google Scholar] [CrossRef] [PubMed]
- Filler Hayut, S.; Melamed Bessudo, C.; Levy, A. A. Targeted Recombination between Homologous Chromosomes for Precisebreeding in Tomato. Nat. Commun 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Yang, D.; Zhang, Y.; Wu, L.; Zhang, Y.; Ye, L.; Pan, C.; He, Y.; Huang, L.; Ruan, Y.L.; Lu, G. Evidence for a Specific and Critical Role of Mitogen-Activated Protein Kinase 20 in Uni-to-Binucleate Transition of Microgametogenesis in Tomato. New Phytol. 2018, 219, 176–194. [Google Scholar] [CrossRef] [PubMed]
- Nonaka, S.; Arai, C.; Takayama, M.; Matsukura, C.; Ezura, H. Efficient Increase of Ɣ-Aminobutyric Acid (GABA) Content in Tomato Fruits by Targeted Mutagenesis. Sci. Rep. 2017, 7, 7057. [Google Scholar] [CrossRef]
- Li, X.; Wang, Y.; Chen, S.; Tian, H.; Fu, D.; Zhu, B.; Luo, Y.; Zhu, H. Lycopene Is Enriched in Tomato Fruit by CRISPR/Cas9- Mediated Multiplex Genome Editing. Front. Plant Sci 2018e, 9. [Google Scholar] [CrossRef]
- Ito, Y.; Nishizawa-Yokoi, A.; Endo, M.; Mikami, M.; Toki, S. CRISPR/Cas9-Mediated Mutagenesis of the RIN Locus That Regulates Tomato Fruit Ripening. Biochem. Biophys. Res. Commun. 2015, 467, 76–82. [Google Scholar] [CrossRef]
- Lang, Z.; Wang, Y.; Tang, K.; Tang, D.; Datsenka, T.; Cheng, J.; Zhang, Y.; Handa, A. K.; Zhu, J.K. Critical Roles of DNA Demethylation in the Activation of Ripening-Induced Genes and Inhibition of Ripening-Repressed Genes in Tomato Fruit. Proc. Natl. Acad. Sci. U. S. A. 2017, 114, E4511–E4519. [Google Scholar] [CrossRef]
- Wang, D.; Samsulrizal, N. H.; Yan, C.; Allcock, N. S.; Craigon, J.; Blanco-Ulate, B.; Ortega-Salazar, I.; Marcus, S. E.; Bagheri, H. M.; Perez Fons, L.; Fraser, P. D.; Foster, T.; Fray, R.; Knox, J. P.; Seymour, G. B. Characterization of CRISPR Mutants Targeting Genes Modulating Pectin Degradation in Ripening Tomato. Plant Physiol. 2019, 179, 544–557. [Google Scholar] [CrossRef]
- Yu, Q.H.; Wang, B.; Li, N.; Tang, Y.; Yang, S.; Yang, T.; Xu, J.; Guo, C.; Yan, P.; Wang, Q.; Asmutola, P. CRISPR/Cas9-Induced Targeted Mutagenesis and Gene Replacement to Generate Long-Shelf Life Tomato Lines. Sci. Rep. 2017, 7, 11874. [Google Scholar] [CrossRef]
- Roldan, M. V. G.; Périlleux, C.; Morin, H.; Huerga-Fernandez, S.; Latrasse, D.; Benhamed, M.; Bendahmane, A. Natural and Induced Loss of Function Mutations in SlMBP21 MADS-Box Gene Led to Jointless-2 Phenotype in Tomato. Sci. Rep. 2017, 7, 4402. [Google Scholar] [CrossRef] [PubMed]
- Klap, C.; Yeshayahou, E.; Bolger, A. M.; Arazi, T.; Gupta, S. K.; Shabtai, S.; Usadel, B.; Salts, Y.; Barg, R. Tomato Facultative Parthenocarpy Results from SlAGAMOUS-LIKE 6 Loss of Function. Plant Biotechnol. J. 2017, 15, 634–647. [Google Scholar] [CrossRef] [PubMed]
- Ueta, R.; Abe, C.; Watanabe, T.; Sugano, S. S.; Ishihara, R.; Ezura, H.; Osakabe, Y.; Osakabe, K. Rapid Breeding of Parthenocarpic Tomato Plants Using CRISPR/Cas9. Sci. Rep. 2017, 7, 507. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.; Alon, I.; Naomi, O.; Sun, T. P. DELLA-ARF/IAA Interaction Mediates Crosstalk between Gibberellin and Auxin Signaling in Controlling Fruit Initiation in Solanum Lycopersicum. Plant Cell 2018, 30, 1710–1728. [Google Scholar] [CrossRef]
- Xu, C.; Park, S. J.; Van Eck, J.; Lippman, Z. B. Control of Inflorescence Architecture in Tomato by BTB/POZ Transcriptional Regulators. Genes Dev. 2016, 30, 2048–2061. [Google Scholar] [CrossRef]
- Soyk, S.; Müller, N. A.; Park, S. J.; Schmalenbach, I.; Jiang, K.; Hayama, R.; Zhang, L.; Van Eck, J.; Jiménez-Gómez, J. M.; Lippman, Z. B. Variation in the Flowering Gene SELF PRUNING 5G Promotes Day-Neutrality and Early Yield in Tomato. Nat. Genet. 2017, 49, 162–168. [Google Scholar] [CrossRef]
- Nie, H.; Shi, Y.; Geng, X.; Xing, G. CRISRP/Cas9-Mediated Targeted Mutagenesis of Tomato Polygalacturonase Gene (SlPG) Delays Fruit Softening. Front. Plant Sci. 2022, 13, 729128. [Google Scholar] [CrossRef]
- Yang, Q.; Cai, L.; Wang, M.; Gan, G.; Li, W.; Li, W.; Jiang, Y.; Yuan, Q.; Qin, C.; Yu, C.; Wang, Y. CRISPR/Cas9 Allows for the Quick Improvement of Tomato Firmness Breeding. Curr. Issues Mol. Biol. 2024, 47, 9. [Google Scholar] [CrossRef]
- Ahmad, S.; Tang, L.; Shahzad, R.; Mawia, A. M.; Rao, G. S.; Jamil, S.; Wei, C.; Sheng, Z.; Shao, G.; Wei, X.; Hu, P.; Mahfouz, M. M.; Hu, S.; Tang, S. CRISPR-Based Crop Improvements: A Way Forward to Achieve Zero Hunger. J. Agric. Food Chem. 2021, 69, 8307–8323. [Google Scholar] [CrossRef]
- Chaudhary, N.; Sandhu, R. A Comprehensive Review on Speed Breeding Methods and Applications. Euphytica 2024, 220(3). [Google Scholar] [CrossRef]
- Ullah, I.; Zhang, H.; Khan, M. R.; Mateen, A.; Pei, Y.; Shakeel, A.; Bhat, A. H.; Fu, C.; Chen, R. Molecular Mecha-Nisms and Genomic Strategies for Enhancing Stress Resilience in Pepper Crop. Scientia Horticulturae 2025, 352. [Google Scholar]
- Mishra, R.; Mohanty, J. N.; Mahanty, B.; Joshi, R. K. A Single Transcript CRISPR/Cas9 Mediated Mutagenesis of CaERF28 Confers Anthracnose Resistance in Chilli Pepper (Capsicum Annuum L.). Planta 2021, 254, 5. [Google Scholar] [CrossRef] [PubMed]
- Bulle, M.; Venkatapuram, A. K.; Abbagani, S.; Kirti, P. B. CRISPR/Cas9 Based Genome Editing of Phytoene Desaturase (PDS) Gene in Chilli Pepper (Capsicum Annuum L.). J. Genet. Eng. Biotechnol. 2024, 22, 100380. [Google Scholar] [CrossRef] [PubMed]
- Choi, H.; Shin, H.; Kim, C. Y.; Park, J.; Kim, H. Highly Efficient CRISPR/Cas9-RNP Mediated CaPAD1 Editing in Protoplasts of Three Pepper (Capsicum Annuum L.) Cultivars. Plant Signal. Behav. 2024, 19, 2383822. [Google Scholar] [CrossRef]
- Park, S.-I.; Kim, H.-B.; Jeon, H.-J.; Kim, H. Agrobacterium-Mediated Capsicum Annuum Gene Editing in Two Cultivars, Hot Pepper CM334 and Bell Pepper Dempsey. Int. J. Mol. Sci. 2021, 22, 3921. [Google Scholar] [CrossRef]
- Hou, X.M.; Zhang, H.F.; Liu, S.Y.; Wang, X.K.; Zhang, Y.M.; Meng, Y.C.; Luo, D.; Chen, R.G. The NAC Transcription Factor CaNAC064 Is a Regulator of Cold Stress Tolerance in Peppers. Plant Sci. 2020, 291, 110346. [Google Scholar] [CrossRef]
- Kong, X.M.; Zhou, Q.; Zhou, X.; Wei, B.D.; Ji, S.J. Transcription Factor CaNAC1 Regulates Low-Temperature-Induced Phospholipid Degradation in Green Bell Pepper. J. Exp. Bot. 2020, 71, 1078–1091. [Google Scholar] [CrossRef]
- Ma, X.; Yu, Y.N.; Jia, J.H.; Li, Q.H.; Gong, Z.H. The Pepper MYB Transcription Factor CaMYB306 Accelerates Fruit Coloration and Negatively Regulates Cold Resistance. Sci. Hortic. (Amsterdam) 2022, 295, 110892. [Google Scholar] [CrossRef]
- Zhang, H.X.; Zhu, W.C.; Feng, X.H.; Wei, J. J.H.; Gong, A.M. Transcription Factor CaSBP12 Negatively Regulates Salt Stress Tolerance in Pepper (Capsicum Annuum L). Int J Mol Sci 2020b, 21. [Google Scholar] [CrossRef]
- Zhang, H.; Pei, Y.; Zhu, F.; He, Q.; Zhou, Y.; Ma, B.; Chen, X.; Guo, J.; Khan, A.; Jahangir, M. CaSnRK2. 4-Mediated Phosphorylation of CaNAC035 Regulates Abscisic Acid Synthesis in Pepper (Capsicum Annuum L.) Responding to Cold Stress. Plant J. 2024a, 117, 1377–1391. [Google Scholar] [CrossRef]
- Ma, J.; Wang, L.Y.; Dai, J.X.; Wang, Y.; Lin, D. The NAC-Type Transcription Factor CaNAC46 Regulates the Salt and Drought Tolerance of Transgenic Arabidopsis Thaliana. BMC Plant Biol 2021, 21, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Lim, J.; Lim, C. W.; Lee, S. C. Role of Pepper MYB Transcription Factor CaDIM1 in Regulation of the Drought Response. Front. Plant Sci. 2022, 13, 1028392. [Google Scholar] [CrossRef] [PubMed]
- Oh, D.; Lim, C. W.; Lee, S. C. GRAS-Type Transcription Factor CaGRAS1 Functions as a Positive Regulator of the Drought Response in Capsicum Annuum. Environ. Exp. Bot. 2022, 198, 104853. [Google Scholar] [CrossRef]
- Guo, W.L.; Wang, S.B.; Chen, R.G.; Chen, B.H.; Du, X.H.; Yin, Y.X.; Gong, Z.H.; Zhang, Y.Y. Characterization and Expression Profile of CaNAC2 Pepper Gene. Front. Plant Sci. 2015b, 6, 755. [Google Scholar] [CrossRef]
- Wang, Y.; Dang, F.; Liu, Z.; Wang, X.; Eulgem, T.; Lai, Y.; Yu, L.; She, J.; Shi, Y.; Lin, J. CaWRKY 58, Encoding a Group I WRKY Transcription Factor of Capsicum Annuum, Negatively Regulates Resistance to Ralstonia Solanacearum Infection. Mol Plant Pathol 2013, 14, 131–144. [Google Scholar] [CrossRef]
- Noman, A.; Hussain, A.; Adnan, M.; Khan, M. I.; Ashraf, M. F.; Zainab, M.; Khan, K. A.; Ghramh, H. A.; He, S. A Novel MYB Transcription Factor CaPHL8 Provide Clues about Evolution of Pepper Immunity Againstsoil Borne Pathogen. Microb. Pathog. 2019, 137, 103758. [Google Scholar] [CrossRef]
- Cai, W.; Yang, S.; Wu, R.; Cao, J.; Shen, L.; Guan, D.; Shuilin, H. Pepper NAC-Type Transcription Factor NAC2c Balances the Trade-off between Growth and Defense Responses. Plant Physiol. 2021, 186, 2169–2189. [Google Scholar] [CrossRef]
- Hussain, A.; Qayyum, A.; Farooq, S.; Almutairi, S. M.; Rasheed, R. A.; Qadir, M.; Vyhnánek, T.; Sun, Y. Pepper Immunity against Ralstonia Solanacearum Is Positively Regulated by CaWRKY3 through Modulation of Different WRKY Transcription Factors. BMC Plant Biol. 2024, 24, 522. [Google Scholar] [CrossRef]
- Oh, S.K.; Baek, K.H.; Park, J. M.; Yi, S. Y.; Yu, S. H.; Kamoun, S.; Choi, D. Capsicum Annuum WRKY Protein CaWRKY1 Is a Negative Regulator of Pathogen Defense. New Phytol. 2008, 177, 977–989. [Google Scholar] [CrossRef]
- Dang, F.; Wang, Y.; She, J.; Lei, Y.; Liu, Z.; Eulgem, T.; Lai, Y.; Lin, J.; Yu, L.; Lei, D. Overexpression of CaWRKY27, a Subgroup Iie WRKY Transcription Factor of Capsicum Annuum, Positively Regulates Tobacco Resistance to Ralstonia Solanacearum in-Fection. Physiol Plant 2014, 150, 397–411. [Google Scholar] [CrossRef]
- Hussain, A.; Li, X.; Weng, Y.; Liu, Z.; Ashraf, M. F.; Noman, A.; Yang, S.; Ifnan, M.; Qiu, S.; Yang, Y.; Guan, D.; He, S. CaWRKY22 Acts as a Positive Regulator in Pepper Response to RalstoniaSolanacearum by Constituting Networks with CaWRKY6, CaWRKY27, CaWRKY40, and CaWRKY58. Int. J. Mol. Sci. 2018, 19, 1426. [Google Scholar] [CrossRef]
- Guo, M.; Zhai, Y.F.; Lu, J.P.; Chai, L.; Chai, W.G.; Gong, Z.H.; Lu, M.H. Characterization of CaHsp70-1, a Pepper Heat-Shock Protein Gene in Response to Heat Stress and Some Regulation Exogenous Substances in Capsicum Annuum L. Int. J. Mol. Sci. 2014, 15, 19741–19759. [Google Scholar] [CrossRef]
- Guo, M.; Lu, J.P.; Zhai, Y.F.; Chai, W.G.; Gong, Z.H.; Lu, M.H. Genome-Wide Analysis, Expression Profile of Heat Shock Factor Gene Family (CaHsfs) and Characterisation of CaHsfA2 in Pepper (Capsicum Annuum L.). BMC Plant Biol. 2015a, 15, 151. [Google Scholar] [CrossRef]
- Dang, F.; Lin, J.; Xue, B.; Chen, Y.; Guan, D.; Wang, Y.; He, S. CaWRKY27 Negatively Regulates H2O2-Mediated Thermotol-Erance in Pepper (Capsicum Annuum). Front Plant Sci 2018, 9. [Google Scholar] [CrossRef]
- Jingyuan, Z.; Xuexiao, Z.; Zhenchuan, M.; Bingyan, X. A Novel Pepper (Capsicum Annuum L.) WRKY Gene, CaWRKY30, Is Involved in Pathogen Stress Responses. J. Plant Biol. 2011, 54, 329–337. [Google Scholar] [CrossRef]
- Hussain, A.; Khan, M. I.; Albaqami, M.; Mahpara, S.; Noorka, I. R.; Ahmed, M. A. A.; Aljuaid, B. S.; El-Shehawi, A. M.; Liu, Z.; Farooq, S.; Zuan, A. T. K. CaWRKY30 Positively Regulates Pepper Immunity by Targeting CaWRKY40 against Ralstonia Solanacearum Inoculation through Modulating Defense-Related Genes. Int. J. Mol. Sci. 2021, 22, 12091. [Google Scholar] [CrossRef] [PubMed]
- Dang, F.F.; Wang, Y.N.; Yu, L.; Eulgem, T.; Lai, Y.; Liu, Z.Q.; Wang, X.; Qiu, A.L.; Zhang, T.X.; Lin, J.; Chen, Y.S.; Guan, D.Y.; Cai, H.Y.; Mou, S.L.; He, S.L. CaWRKY40, a WRKY Protein of Pepper, Plays an Important Role in the Regulation of Tolerance to Heat Stress and Resistance to Ralstonia Solanacearum Infection: CaWRKY40 in Heat andR. Solanacearuminfection. Plant Cell Environ. 2013, 36, 757–774. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Ma, X.; Xiao, L.-D.; Yu, Y.-N.; Yan, H.-L.; Gong, Z.-H. CaWRKY5 Acts as a Negative Regulator in Response to Colleto-Trichumscovillei Infection in Pepper. Plants 2023, 12(10). [Google Scholar]
- Cheng, W.; Wang, N.; Li, Y.; Zhou, X.; Bai, X.; Liu, L.; Ma, X.; Wang, S.; Li, X.; Gong, B.; Jiang, Y.; Azeem, M.; Zhu, L.; Chen, L.; Wang, H.; Chu, M. CaWRKY01-10 and CaWRKY08-4 Confer Pepper’s Resistance to Phytophthora Capsici Infection by Directly Activating a Cluster of Defense-Related Genes. J. Agric. Food Chem. 2024, 72, 11682–11693. [Google Scholar] [CrossRef]
- Feng, X.H.; Zhang, H.X.; Gai, A. M.; Cheng, W.X.; Li, Y. S.B.; Gong, X.X. A Small Heat Shock Protein CaHsp25. 9 Positively Regulates Heat, Salt, and Drought Stress Tolerance in Pepper (Capsicum Annuum L.). Plant Physiol Biochem 2019, 142, 151–162. [Google Scholar] [CrossRef]
- Liu, R.; Song, J.; Liu, S.; Chen, C.; Zhang, S.; Wang, J.; Xiao, Y.; Cao, B.; Lei, J.; Zhu, Z. Genome-Wide Identification of the Capsicum bHLH Transcription Factor Family: Discovery of a Candidate Regulator Involved in the Regulation of Species-Specific Bioactive Metabolites. BMC Plant Biol 2021a, 21. [Google Scholar] [CrossRef]
- Liu, Y.L.; Liu, S.; Xiao, J.J.; Cheng, G.X.; Ul; H. S. Gong, Z.H. CaHSP18. 1a, a Small Heat Shock Protein from Pepper (Capsicum Annuum L) Positively Responds to Heat Drought and Salt Tolerance. Horticulturae 2021a, 7. [Google Scholar] [CrossRef]
- Lee, S. C.; Choi, H. W.; Hwang, I. S.; Choi, D. S.; Hwang, B. K. Functional Roles of the Pepper Pathogen-Induced bZIP Transcription Factor, CAbZIP1, in Enhanced Resistance to Pathogen Infection and Environmental Stresses. Planta 2006, 224, 1209–1225. [Google Scholar] [CrossRef] [PubMed]
- Lim, C. W.; Baek, W.; Lim, S.; Han, S.-W.; Lee, S. C. Expression and Functional Roles of the Pepper Pathogen-Induced bZIP Transcription Factor CabZIP2 in Enhanced Disease Resistance to Bacterial Pathogen Infection. Mol. Plant. Microbe. Interact. 2015, 28, 825–833. [Google Scholar] [CrossRef] [PubMed]
- Gai, W.X.; Ma, X.; Qiao, Y.M.; Shi, B.H.; Ul Haq, S.; Li, Q.H.; Wei, A.M.; Liu, K.K.; Gong, Z.H. Characterization of the bZIP Transcription Factor Family in Pepper (Capsicum Annuum L.): CabZIP25 Positively Modulates the Salt Tolerance. Front. Plant Sci. 2020, 11, 139. [Google Scholar] [CrossRef]
- Hong, J.P.; Kim, W. T. Isolation and Functional Characterization of the Ca-DREBLP1 Gene Encoding a Dehydra-Tion-Responsive Element Binding-Factor-like Protein 1 in Hot Pepper (Capsicum Annuum L. Cv. Pukang). Planta 2005, 220, 875–888. [Google Scholar] [CrossRef]
- Parkash, C.; Kumar, S.; Thakur, N.; Singh, S.; Sharma, B. B. Cabbage: Breeding and Genomics. isvs 2023, 50 (Special), 231–243. [Google Scholar] [CrossRef]
- Jansson, S. Gene-Edited Plants on the Plate: The “CRISPR Cabbage Story.”. Physiol. Plant. 2018, 164, 396–405. [Google Scholar] [CrossRef]
- Ma, C.; Zhu, C.; Zheng, M.; Liu, M.; Zhang, D.; Liu, B.; Li, Q.; Si, J.; Ren, X.; Song, H. CRISPR/Cas9-Mediated Multiple Gene Editing in Brassica Oleracea Var. Capitata Using the Endogenous tRNA-Processing System. Hortic. Res. 2019a, 6, 20. [Google Scholar] [CrossRef]
- Ma, C.; Liu, M.; Li, Q.; Si, J.; Ren, X.; Song, H. Efficient BoPDS Gene Editing in Cabbage by the CRISPR/Cas9 System. Hortic. Plant J. 2019b, 5, 164–169. [Google Scholar] [CrossRef]
- Murovec, J.; Guček, K.; Bohanec, B.; Avbelj, M.; Jerala, R. DNA-Free Genome Editing of Brassica Oleracea and B. Rapa Protoplasts Using CRISPR-Cas9 Ribonucleoprotein Complexes. Front. Plant Sci. 2018, 9, 1594. [Google Scholar] [CrossRef]
- Kaur, C.; Lee, M.; Yao, J.; Nam, D.; Lee, G. J. CRISPR/Cas12-Mediated Detection of Xanthomonas Campestris Pv. Campestris (Race 1 and 4) for Improved Pathogen Diagnosis in Cabbage. Horticulturae Sci. Technol 2023a, 41, 338. [Google Scholar]
- Vicente, J. G.; Holub, E. B. X Anthomonas Campestris Pv. Campestris (Cause of Black Rot of Crucifers) in the Genomic Era Is Still a Worldwide Threat to Brassica Crops. Mol. Plant Pathol. 2013, 14, 2–18. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.; Li, S.; Chen, W.; Zhang, J.; Zhang, L.; Sun, W.; Wang, Z. Plant Dehydrins: Expression, Regulatory Networks, and Protective Roles in Plants Challenged by Abiotic Stress. Int. J. Mol. Sci. 2021, 22, 12619. [Google Scholar] [CrossRef] [PubMed]
- Kaur, C.; Lee, M.; Jeon, Y.; Lee, G. J. Enhancing Black-Rot Resistance in Cabbage Using CRISPR/Cas9 System. Horticulturae Sci. Technol. 2023b, 41, 423. [Google Scholar]
- Zhao, Z.; Qi, Y.; Yang, Z.; Cheng, L.; Sharif, R.; Raza, A.; Chen, P.; Hou, D.; Li, Y. Exploring the Agrobacterium-Mediated Transformation with CRISPR/Cas9 in Cucumber (Cucumis Sativus L.). Mol. Biol. Rep. 2022, 49, 11481–11490. [Google Scholar] [CrossRef]
- Feng, J.; Wang, N.; Li, Y.; Wang, H.; Zhang, W.; Wang, H.; Chai, S. Recent Progress in Genetic Transformation and Gene Editing Technology in Cucurbit Crops. Agronomy (Basel) 2023, 13, 755. [Google Scholar] [CrossRef]
- Hu, B.; Li, D.; Liu, X.; Qi, J.; Gao, D.; Zhao, S.; Huang, S.; Sun, J.; Yang, L. Engineering Non-Transgenic Gynoecious Cucumber Using an Improved Transformation Protocol and Optimized CRISPR/Cas9 System. Mol. Plant 2017, 10(12), 1575–1578. [Google Scholar] [CrossRef]
- Feng, S.; Zhang, J.; Mu, Z.; Wang, Y.; Wen, C.; Wu, T.; Yu, C.; Li, Z.; Wang, H. Recent Progress on the Molecular Breeding of Cucumis Sativus L. in China. Theor. Appl. Genet. 2020, 133, 1777–1790. [Google Scholar] [CrossRef]
- Ma, Z.; Ma, L.; Zhou, J. Applications of CRISPR/Cas Genome Editing in Economically Important Fruit Crops: Recent Advances and Future Directions. Mol. Hortic. 2023, 3, 1. [Google Scholar] [CrossRef]
- Pawełkowicz, M.; Zieniuk, B.; Staszek, P.; Przybysz, A. From Sequencing to Genome Editing in Cucurbitaceae: Application of Modern Genomic Techniques to Enhance Plant Traits. Agriculture 2024, 14, 90. [Google Scholar] [CrossRef]
- Chandrasekaran, J.; Brumin, M.; Wolf, D.; Leibman, D.; Klap, C.; Pearlsman, M.; Sherman, A.; Arazi, T.; Gal-On, A. Development of Broad Virus Resistance in Non-Transgenic Cucumber Using CRISPR/Cas9 Technology: Virus Resistance in Cucumber Using CRISPR/Cas9. Mol. Plant Pathol. 2016, 17, 1140–1153. [Google Scholar] [CrossRef] [PubMed]
- Dey, S. S.; Singh, S.; Munshi, A. D.; Behera, T. K. Classical Genetics and Traditional Breeding. In Compendium of Plant Genomes; Springer International Publishing: Cham, 2022; pp. 159–183. [Google Scholar] [CrossRef]
- Riu, Y.-S.; Kim, G. H.; Chung, K. W.; Kong, S.-G. Enhancement of the CRISPR/Cas9-Based Genome Editing System in Lettuce (Lactuca Sativa L.) Using the Endogenous U6 Promoter. Plants 2023, 12, 878. [Google Scholar] [CrossRef] [PubMed]
- Bertier, L. D.; Ron, M.; Huo, H.; Bradford, K. J.; Britt, A. B.; Michelmore, R. W. High-Resolution Analysis of the Efficiency, Heritability, and Editing Outcomes of CRISPR/Cas9-Induced Modifications of NCED4 in Lettuce (Lactuca Sativa). G3 (Bethesda) 2018, 8, 1513–1521. [Google Scholar] [CrossRef] [PubMed]
- Luo, C.; Wang, S.; Ning, K.; Chen, Z.; Wang, Y.; Yang, J.; Wang, Q. LsAP2 Regulates Leaf Morphology by Inhibiting CIN-like TCP Transcription Factors and Repressing LsKAN2 in Lettuce. Hortic. Res. 2021, 8, 184. [Google Scholar] [CrossRef]
- Nguyen, C. D.; Li, J.; Mou, B.; Gong, H.; Huo, H. A Case Study of Using an Efficient CRISPR/Cas9 System to Develop Variegated Lettuce. Vegetable Research 2021, 1, 1–10. [Google Scholar] [CrossRef]
- Choi, S. H.; Ahn, W. S.; Jie, E. Y.; Cho, H.-S.; Kim, S. W. Development of Late-Bolting Plants by CRISPR/Cas9-Mediated Genome Editing from Mesophyll Protoplasts of Lettuce. Plant Cell Rep. 2022, 41, 1627–1630. [Google Scholar] [CrossRef]
- Ahmad, F.; Akram, A.; Farman, K.; Abbas, T.; Bibi, A.; Khalid, S.; Waseem, M. Molecular Markers and Marker Assisted Plant Breeding: Current Status and Their Applications in Agricultural Development. Journal of Environmental and Agricultural Sciences 2017, 11, 35–50. [Google Scholar]
- Pejić, I.; Šatović, Z. Konvencionalno i Molekularno Oplemenjivanje Bilja / Molekularno Oplemenjivanje Bilja, Agronomski Fakultet, Zagreb - Pregled Integracije Molekularnih i Klasičnih Metoda. 2022, 3–20.
- Lamichhane, S.; Thapa, S. Advances from Conventional to Modern Plant Breeding Methodologies. Plant Breed. Biotechnol. 2022, 10, 1–14. [Google Scholar] [CrossRef]
- Wanga, M. A.; Shimelis, H.; Mashilo, J.; Laing, M. D. Opportunities and Challenges of Speed Breeding: A Review. Plant Breed. 2021, 140, 185–194. [Google Scholar] [CrossRef]
- Datta, A. Genetic Engineering for Improving Quality and Productivity of Crops. Agric. & Food Secur. 2013, 2, 15. [Google Scholar] [CrossRef]
- Singh, R. K.; Jamir, S. Advancements in vegetable breeding techniques for quality enhancement. Plant Archives 2025, 25, 26–35. [Google Scholar] [CrossRef]
- Xu, Y.; Zhang, X.; Li, H.; Zheng, H.; Zhang, J.; Olsen, M. S.; Varshney, R. K.; Prasanna, B. M.; Qian, Q. Smart Breeding Driven by Big Data, Artificial Intelligence, and Integrated Genomic-Enviromic Prediction. Mol. Plant 2022, 15, 1664–1695. [Google Scholar] [CrossRef] [PubMed]
- Sharma, A.; Namdeo, A.; Mahadik, K. Molecular Markers: New Prospects in Plant Genome Analysis. Pharmacognosy reviews 2008, 2. [Google Scholar]
- Nadeem, M. A.; Nawaz, M. A.; Shahid, M. Q.; Doğan, Y.; Comertpay, G.; Yıldız, M.; Hatipoğlu, R.; Ahmad, F.; Alsaleh, A.; Labhane, N.; Özkan, H.; Chung, G.; Baloch, F. S. DNA Molecular Markers in Plant Breeding: Current Status and Recent Advancements in Genomic Selection and Genome Editing. Biotechnol. Biotechnol. Equip. 2018, 32, 261–285. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, Y. Advances in Molecular Breeding of Vegetable Crops. Horticulturae 2022, 8, 821. [Google Scholar] [CrossRef]
- Kumar, J.; Choudhary, A. K.; Solanki, R. K.; Pratap, A. Towards Marker-Assisted Selection in Pulses: A Review: Towards Marker-Assisted Selection in Pulses. Plant Breed. 2011, 130, 297–313. [Google Scholar] [CrossRef]
- Fu, Y.B.; Dong, Y.B. Genetic Erosion under Modern Plant Breeding: Case Studies in Canadian Crop Gene Pools. In Sustainable Development and Biodiversity; Springer International Publishing: Cham, 2015; pp. 89–104. [Google Scholar]
- Behera, T. K.; Devi, J.; Tiwari, J. K.; Singh, B. K. Vegetable Breeding: Status and Strategies. isvs 2023, 50 (Special), 131–145. [Google Scholar] [CrossRef]
- Song, S.; Ong, E. J. K.; Lee, A. M. J.; Chew, F. T. How Crop Breeding Programs Can Improve Plant Factories’ Business and Environmental Sustainability: Insights from a Farm Level Analysis. Sustain. Prod. Consum. 2024, 44, 298–311. [Google Scholar] [CrossRef]
- Zhang, S.; Zhu, H. Development and Prospect of Gene-Edited Fruits and Vegetables. Food Qual. Saf. 2024, 8, fyad045. [Google Scholar] [CrossRef]
- Brennan, J. P.; Martin, P. J.; Brennan, J. P.; Martin, P. J. Developing Cost Functions for a Wheat Breeding Program. Unknown 2006. [CrossRef]
- Morris, M. L.; Bellon, M. R. Participatory Plant Breeding Research: Opportunities and Challenges for the International Crop Improvement System. Euphytica 2004, 136, 21–35. [Google Scholar] [CrossRef]
- Shu, Q.; Wu, D. Rice: Breeding. In Encyclopedia of Food Grains; Elsevier, 2016, 304–310.
- Sharma, R.; Yang, C. J.; Rossi, N.; Irving, E.; Tuffin, A.; Aliki, H.; Powell, W.; Dawson, I. K. Integrating Molecular Genetics with Plant Breeding to Deliver Impact. Plant Physiol. 2025, 198. [Google Scholar] [CrossRef]
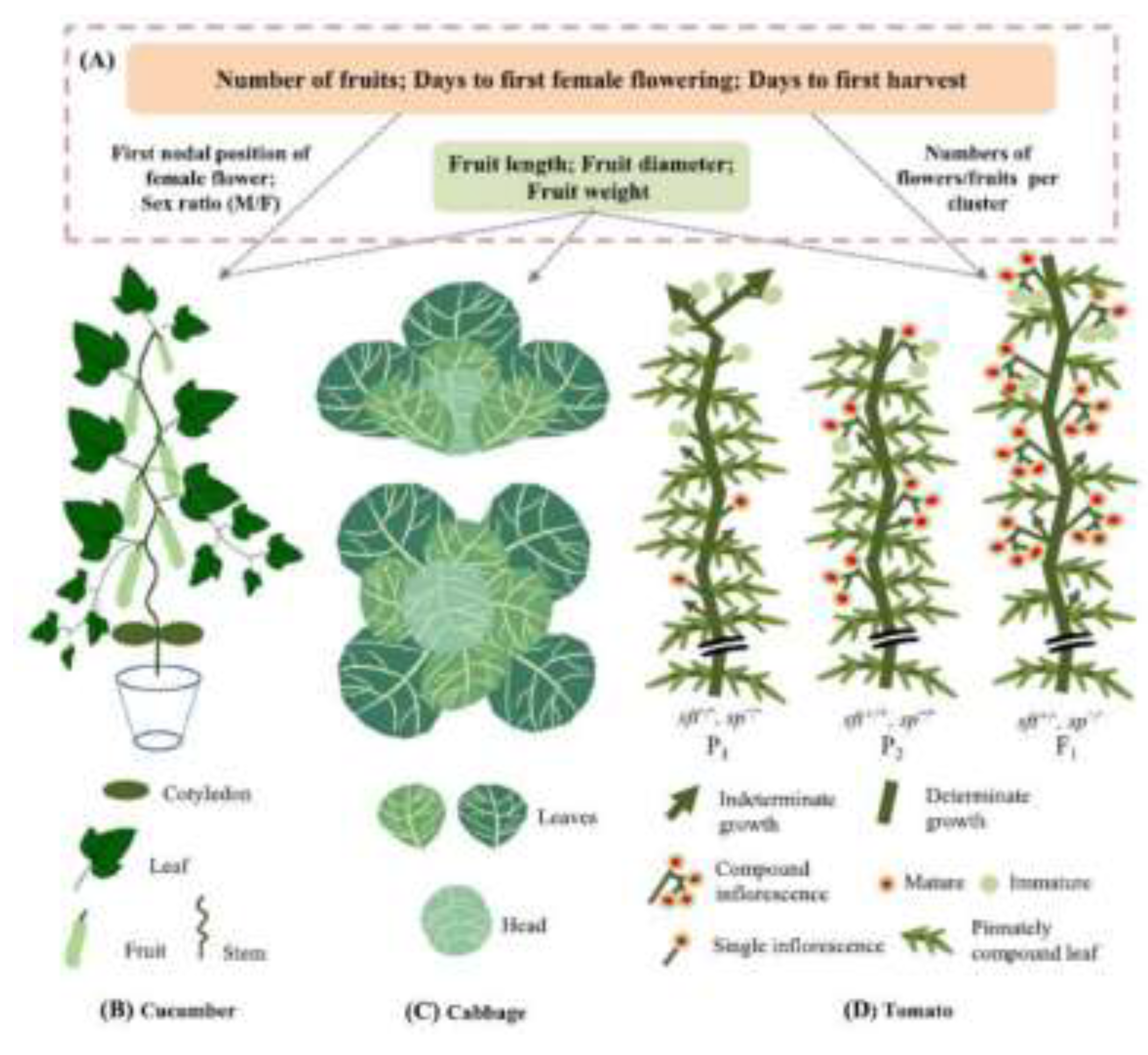
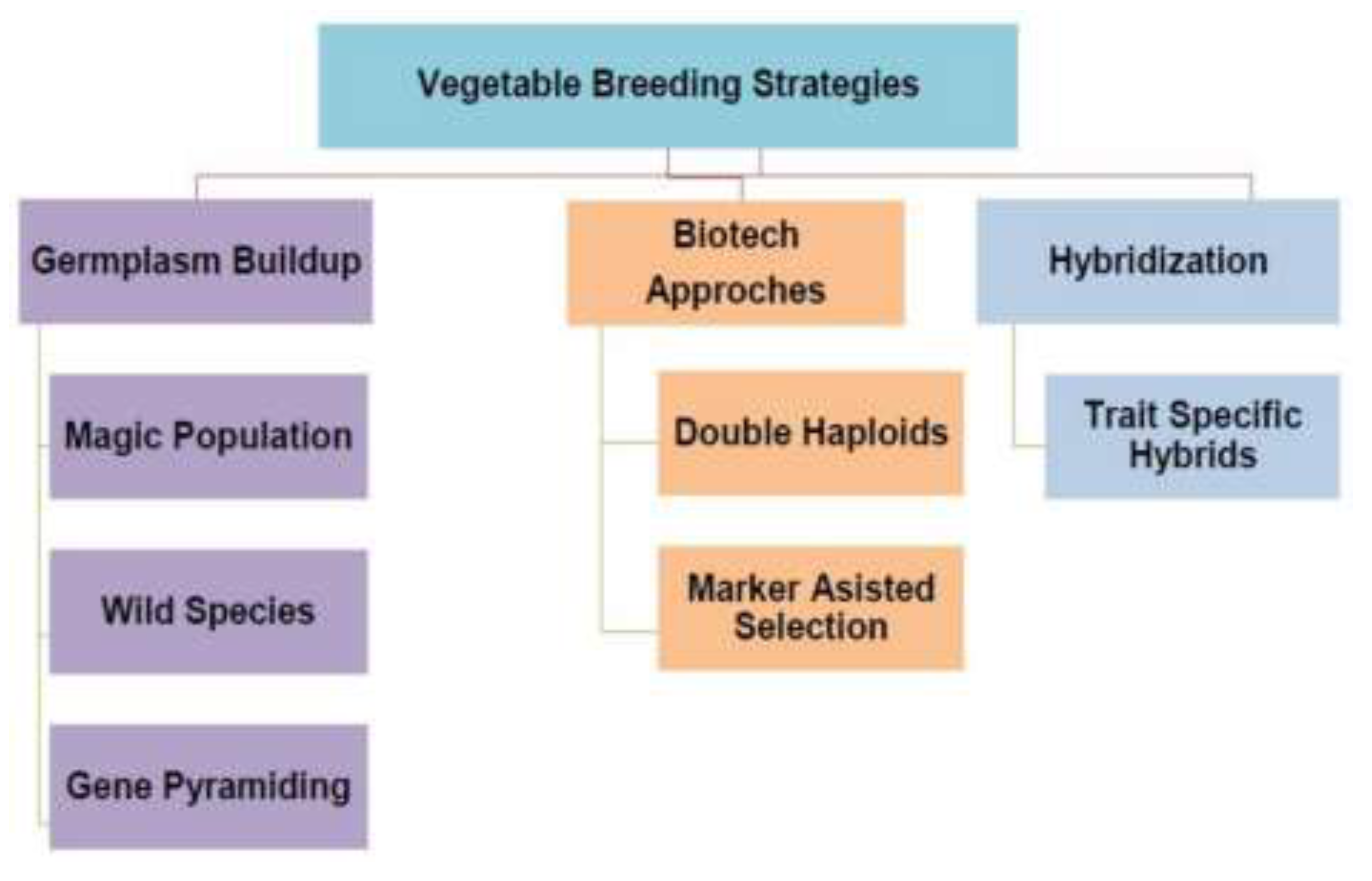
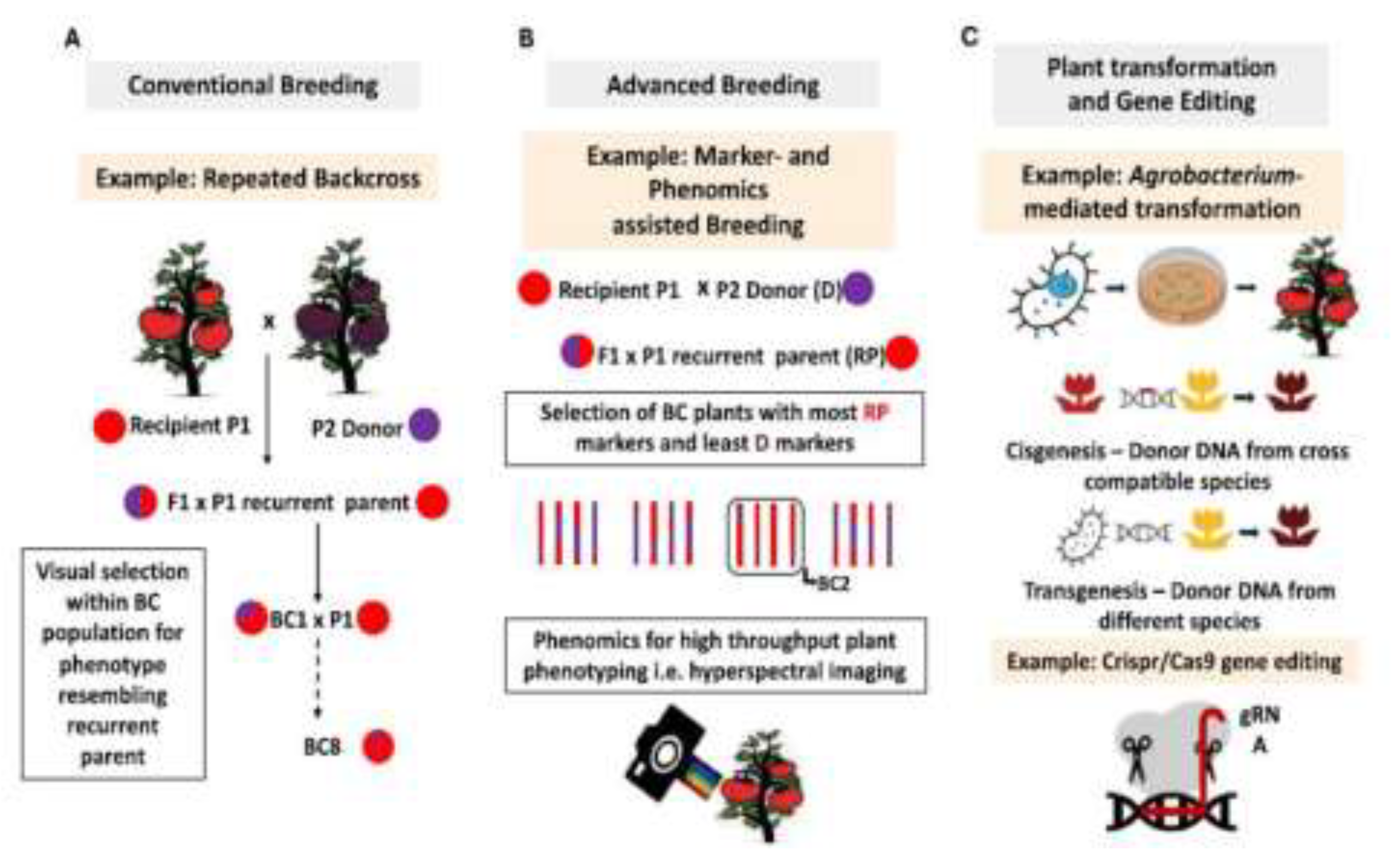
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



