Submitted:
13 April 2026
Posted:
15 April 2026
You are already at the latest version
Abstract
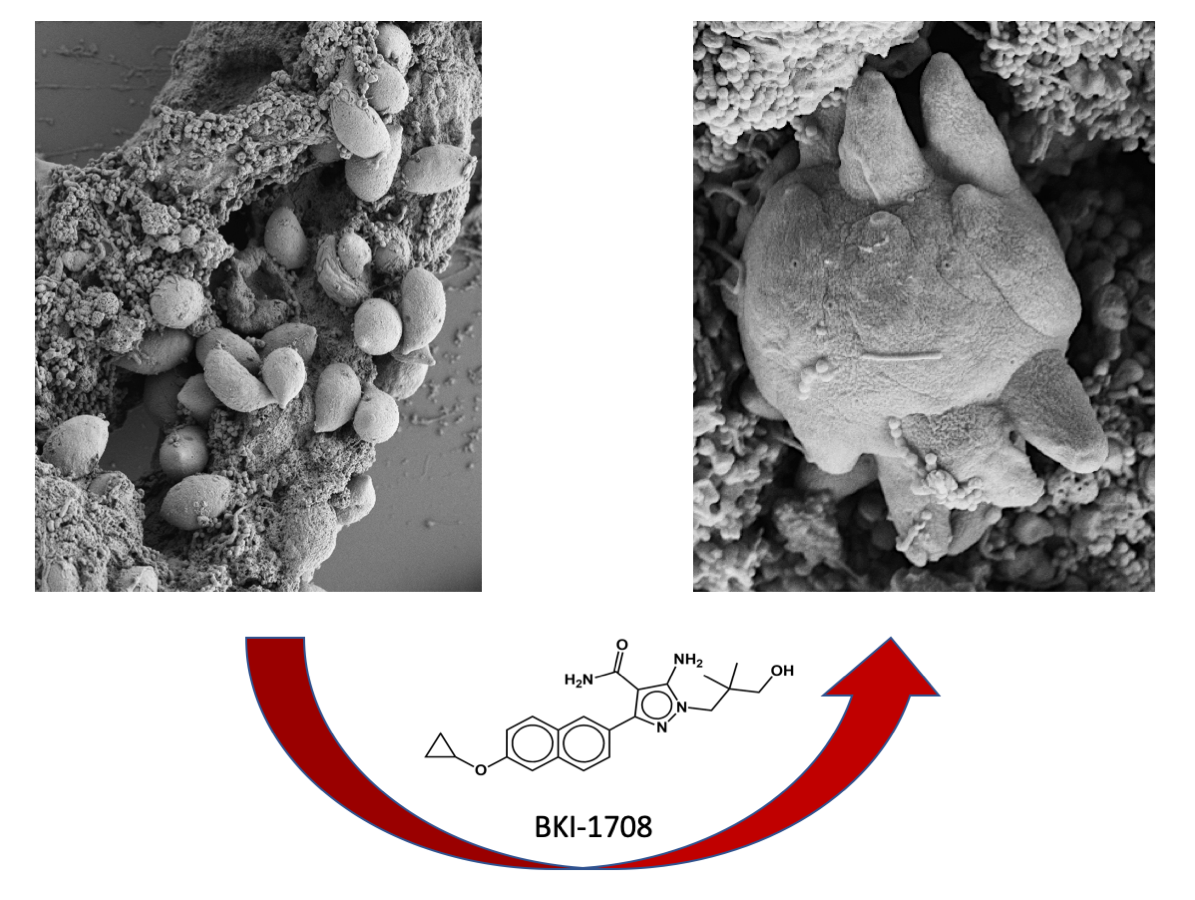
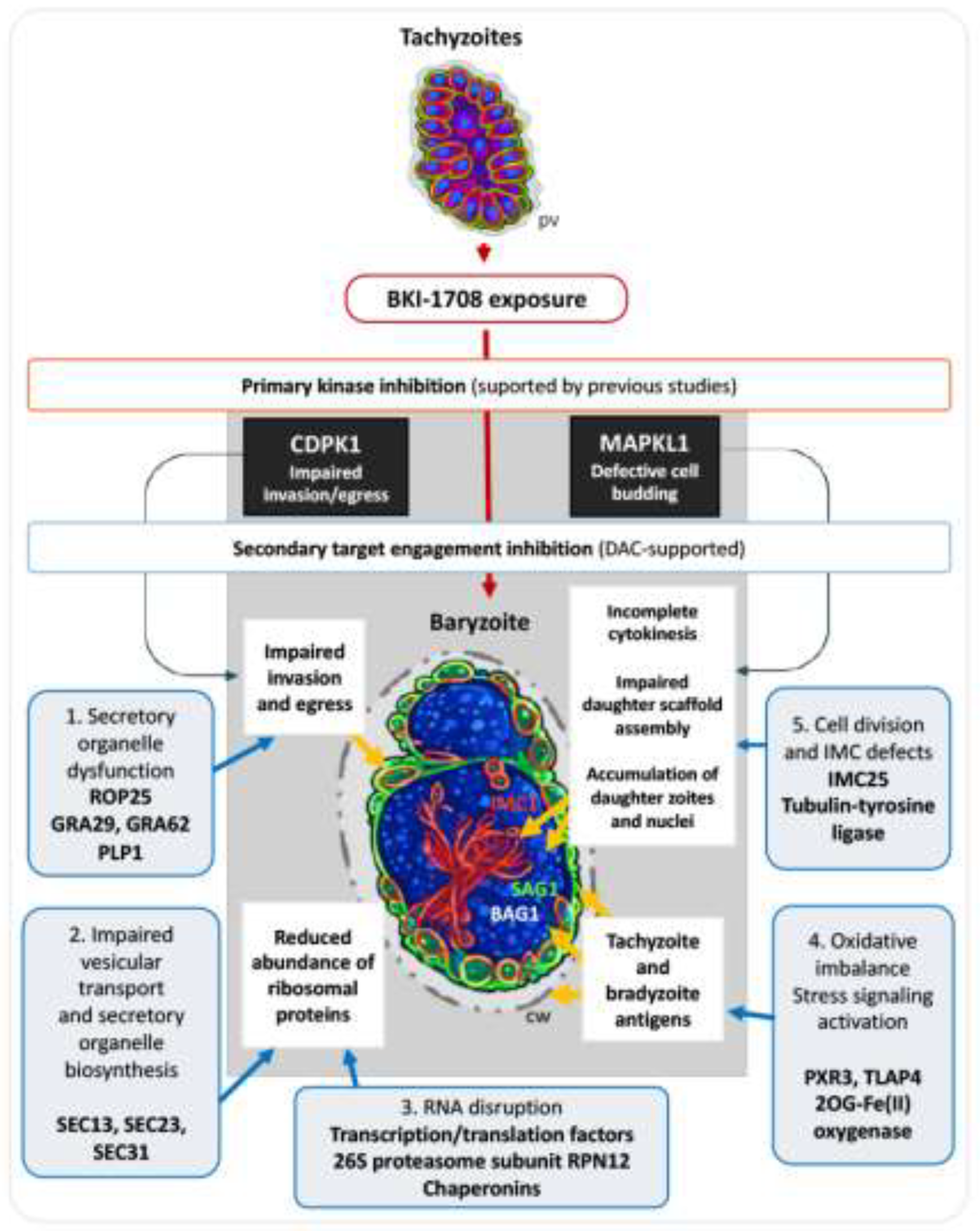
Bumped kinase inhibitor 1708 (BKI-1708), previously demonstrated to target apicomplexan kinases and CDPK1 and MAPKL1, exhibits remarkable activity against Toxoplasma gondii infection both in vitro and in vivo. Notably, BKI-1708 does not affect the viability of mammalian cells. Upon exposure to BKI-1708 exposure, T. gondii tachyzoites form large multinucleated complexes named baryzoites and remain trapped in within host cells. In this study, additional potential molecular targets for BKI-1708 were identified in soluble extracts of T. gondii ME49 tachyzoites and human foreskin fibroblasts (HFF) using differential affinity chromatography coupled to mass spectrometry (DAC-MS). Beyond kinases, secondary interactions in T. gondii involved the binding of proteins associated with cell division, cytoskeleton, vesicular trafficking, secretory organelles, and transcriptional and translational regulators. In non-infected HFFs, BKI-1708 interactors included cytoskeletal regulators along with multiple RNA/DNA-binding proteins. Upon infection, this profile shifted, with cytoskeletal components no longer detected, while nucleic acid–binding proteins remained present, consistent with infection-induced chromatin and transcriptional remodeling. This multi-target interference could contribute to the impaired cytokinesis and the formation of multinucleated baryzoites, aligning with the concept that antiprotozoal drugs exert efficacy through coordinated perturbation of multiple cellular processes rather than a single dominant target.

Keywords:
1. Introduction
2. Materials and Methods
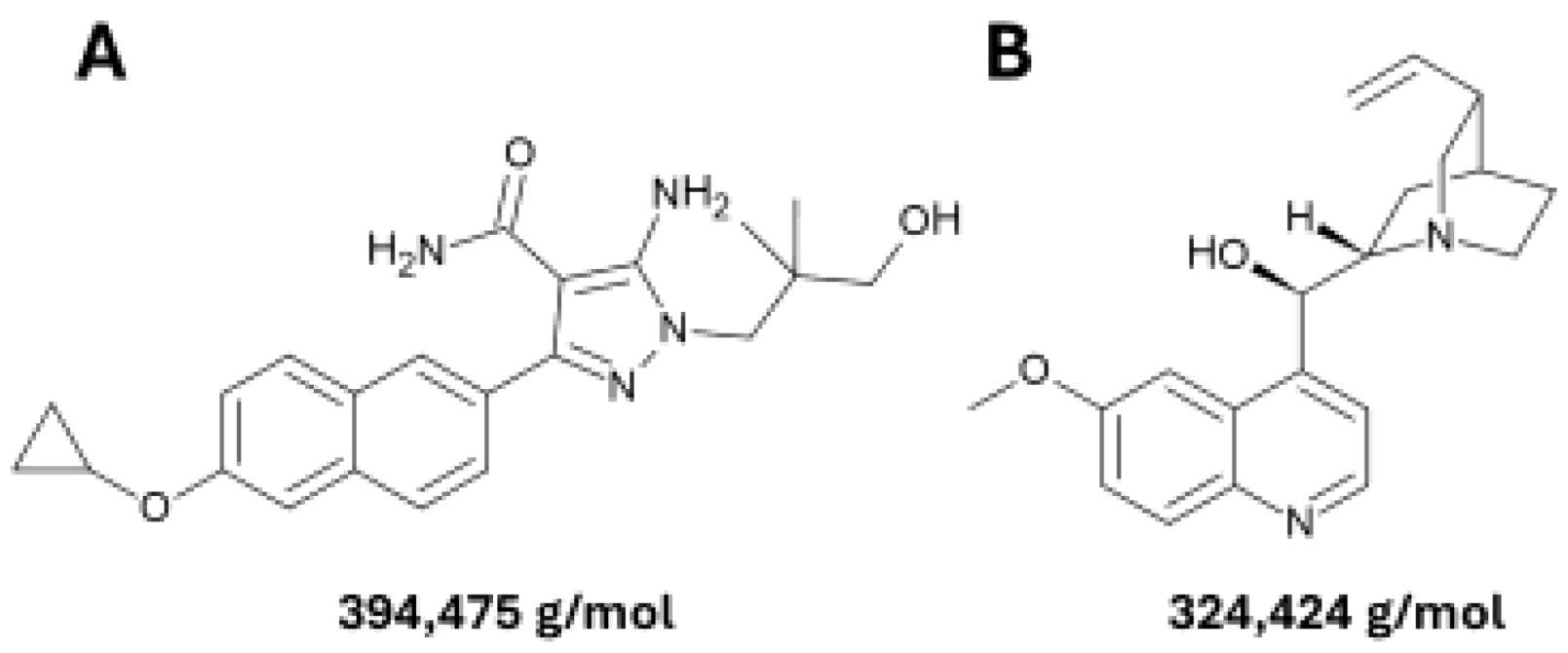
2.1. Parasites, Culture Medium, Biochemicals and Compounds
2.2. SEM and TEM of T. gondii Baryzoites Formed Upon Exposure to BKI-1708.
2.3. Protein Extraction and Differential Affinity Chromatography (DAC)
2.4. Proteomic Analysis of the Eluted Proteins by Mass Spectrometry
3. Results
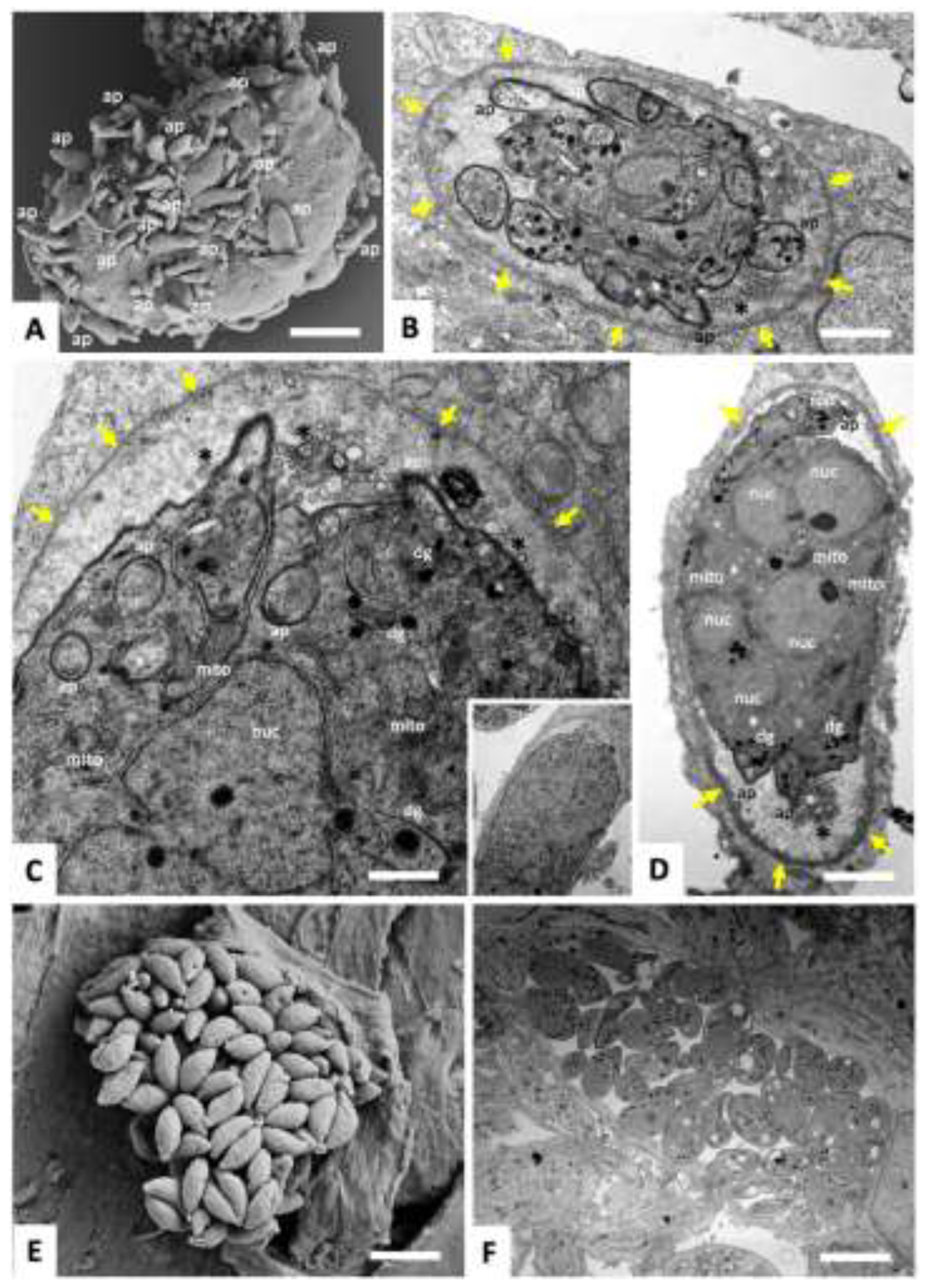
3.1. Exposure of T. gondii Tachyzoites to BKI-1708 Initiates Baryzoite Formation
3.2. DAC-Overview
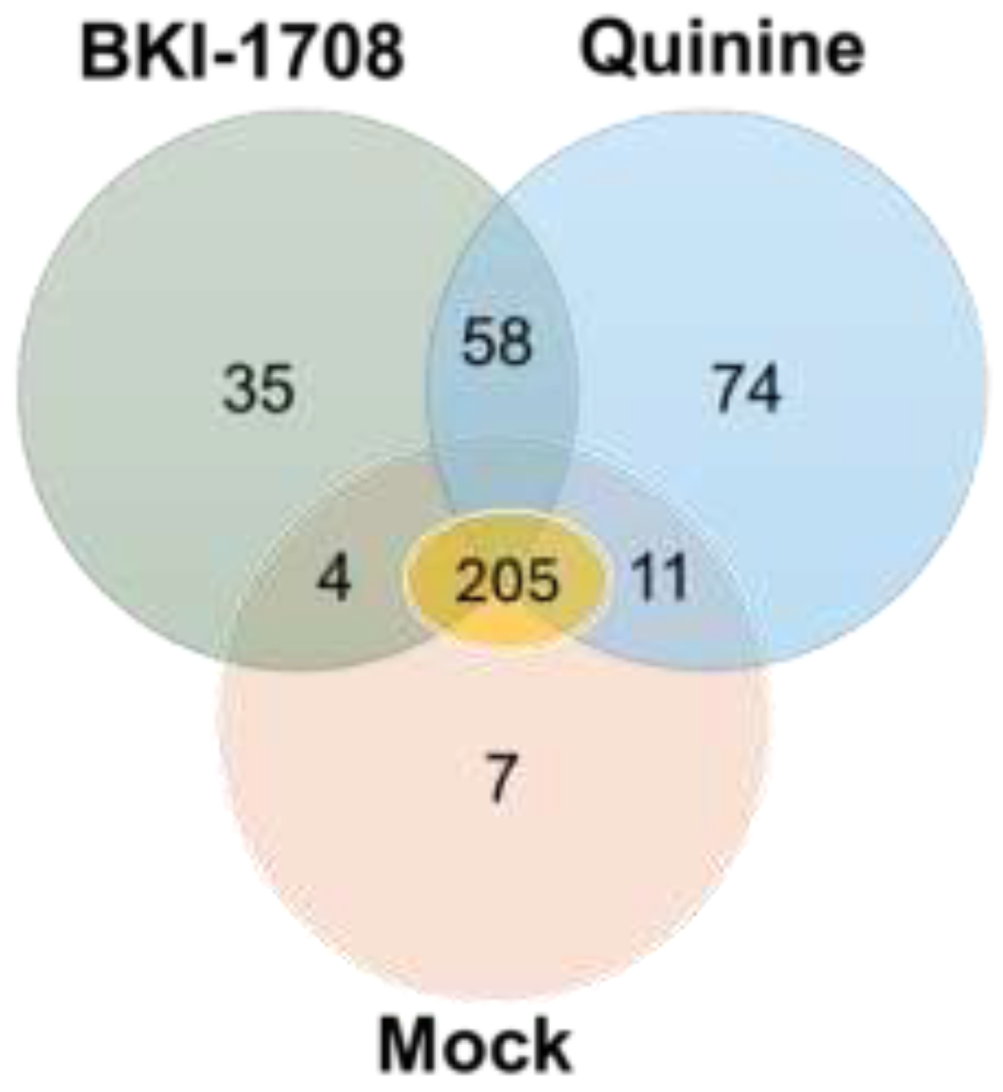
3.3. T. gondii Proteins Binding Identified by BKI-1708 DAC-MS
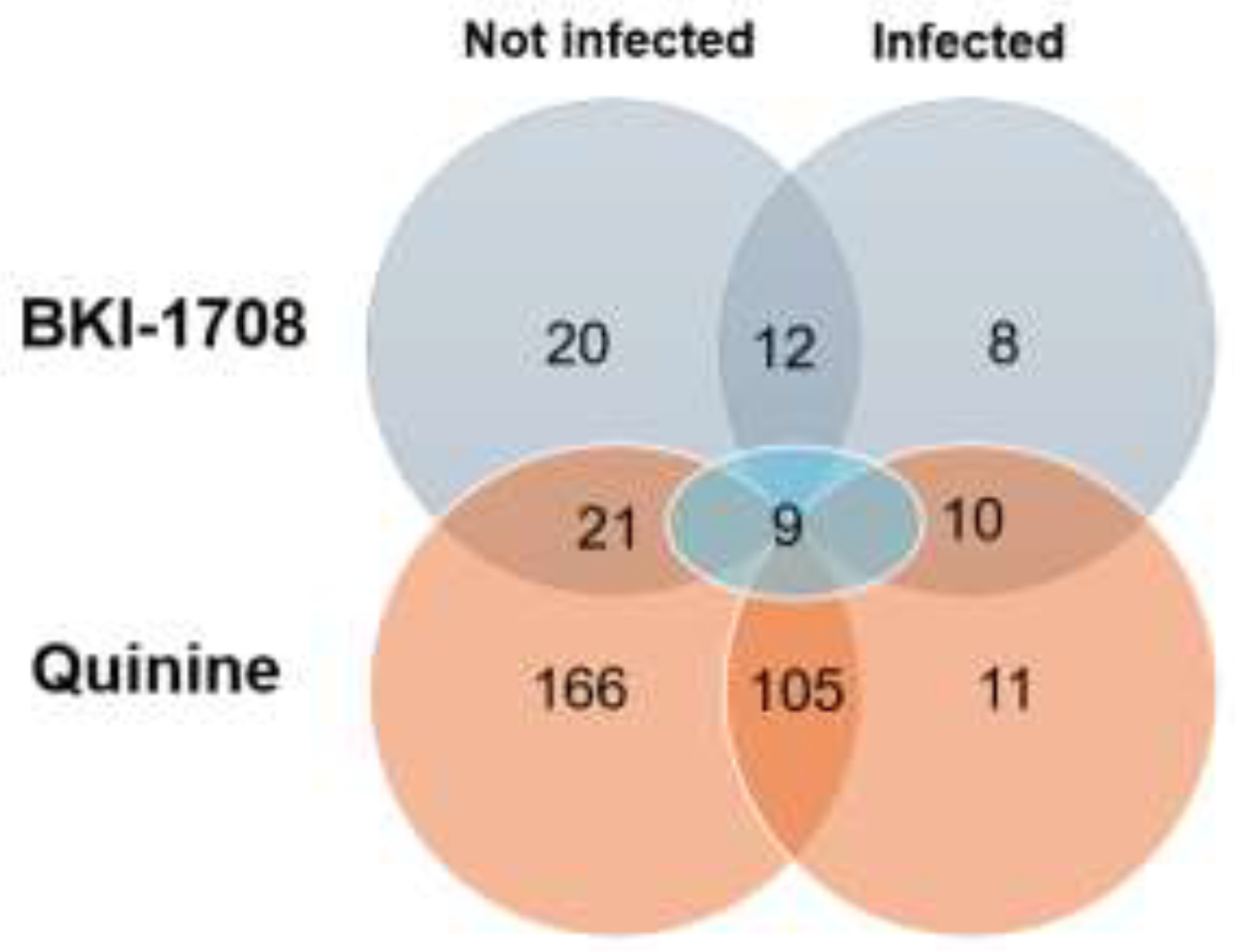
3.4. DAC-MS Revealed Distinct BKI-1708 affino-Proteomes in T. gondii-Infected and Non-Infected HFF
3.4.1. Host Cell Proteins Identified by BKI-1708 DAC-MS in Non-Infected HFF
3.4.2. Proteins Identified by BKI-1708 DAC-MS in T. gondii Infected HFF
3.4.3. Host Cell Proteins Binding Exclusively to BKI-1708 in Both T. gondii-Infected and Non-Infected HFF
3.4.4. Host Cell Proteins Binding to Both BKI-1708 and Quinine in T. gondii-Infected and Non-Infected HFF
3.5. Numbers and Putative Functions of T. gondii Tachzoite and HFF Proteins Specifically Binding to BKI-1708
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- Dubey, J.P. Toxoplasmosis of Animals and Humans; CRC Press, 2021; ISBN 978-1-000-43149-0.
- Martorelli Di Genova, B.; Wilson, S.K.; Dubey, J.P.; Knoll, L.J. Intestinal Delta-6-Desaturase Activity Determines Host Range for Toxoplasma Sexual Reproduction. PLoS Biol 2019, 17, e3000364. [CrossRef]
- Almeria, S.; Dubey, J.P. Foodborne Transmission of Toxoplasma Gondii Infection in the Last Decade. An Overview. Res Vet Sci 2021, 135, 371–385. [CrossRef]
- Holec-Gąsior, L.; Sołowińska, K. Detection of Toxoplasma Gondii Infection in Small Ruminants: Old Problems, and Current Solutions. Animals 2023, 13, undefined-undefined. [CrossRef]
- Sullivan, W.J.; Jeffers, V. Mechanisms of Toxoplasma Gondii Persistence and Latency. FEMS Microbiol Rev 2012, 36, 717–733. [CrossRef]
- Weiss, L.M.; Kim, K. THE DEVELOPMENT AND BIOLOGY OF BRADYZOITES OF TOXOPLASMA GONDII. Front Biosci 2000, 5, D391–D405.
- Montoya, J.; Liesenfeld, O. Toxoplasmosis. The Lancet 2004, 363, 1965–1976. [CrossRef]
- Walana, W.; Odai, S.A.; Tamomh, A.G. Prevalence, Risk Factors, Diagnosis and Outcomes of TOxoplasma Gondii Infection in Pregnancy: A Review. Parasitology International 2026, 110, 103143. [CrossRef]
- Liu, S.; Cai, M.; Liu, Z.; Gao, W.; Li, J.; Li, Y.; Abudouxukuer, X.; Zhang, J. Comprehensive Insights into the Development of Antitoxoplasmosis Drugs: Current Advances, Obstacles, and Future Perspectives. J. Med. Chem. 2024, 67, 20740–20764. [CrossRef]
- Choi, R.; Hulverson, M.A.; Huang, W.; Vidadala, R.S.R.; Whitman, G.R.; Barrett, L.K.; Schaefer, D.A.; Betzer, D.P.; Riggs, M.W.; Doggett, J.S.; et al. Bumped Kinase Inhibitors as Therapy for Apicomplexan Parasitic Diseases: Lessons Learned. International Journal for Parasitology 2020, 50, 413–422.
- Huang, W.; Hulverson, M.A.; Choi, R.; Arnold, S.L.M.; Zhang, Z.; McCloskey, M.C.; Whitman, G.R.; Hackman, R.C.; Rivas, K.L.; Barrett, L.K.; et al. Development of 5-Aminopyrazole-4-Carboxamide-Based Bumped-Kinase Inhibitors for Cryptosporidiosis Therapy. J. Med. Chem. 2019, 62, 3135–3146.
- Imhof, D.; Anghel, N.; Winzer, P.; Balmer, V.; Ramseier, J.; Hänggeli, K.; Choi, R.; Hulverson, M.; Whitman, G.; Arnold, S.; et al. In Vitro Activity, Safety and in Vivo Efficacy of the Novel Bumped Kinase Inhibitor BKI-1748 in Non-Pregnant and Pregnant Mice Experimentally Infected with Neospora Caninum Tachyzoites and Toxoplasma Gondii Oocysts. International Journal for Parasitology: Drugs and Drug Resistance 2021, 16.
- Lourido, S.; Jeschke, G.R.; Turk, B.E.; Sibley, L.D. Exploiting the Unique ATP-Binding Pocket of Toxoplasma Calcium-Dependent Protein Kinase 1 to Identify Its Substrates. ACS Chem Biol 2013, 8, 1155–1162.
- Jacot, D.; Soldati-Favre, D. Does Protein Phosphorylation Govern Host Cell Entry and Egress by the Apicomplexa? Int J Med Microbiol 2012, 302, 195–202.
- Lourido, S.; Shuman, J.; Zhang, C.; Shokat, K.M.; Hui, R.; Sibley, L.D. Calcium-Dependent Protein Kinase 1 Is an Essential Regulator of Exocytosis in Toxoplasma. Nature 2010, 465, 359–362. [CrossRef]
- Wei, F.; Wang, W.; Liu, Q. Protein Kinases of Toxoplasma Gondii: Functions and Drug Targets. Parasitol Res 2013, 112, 2121–2129.
- Montgomery, J.A.; Alday, P.H.; Choi, R.; Khim, M.; Staker, B.L.; Hulverson, M.A.; Ojo, K.K.; Fan, E.; Van Voorhis, W.C.; Doggett, J.S. Bumped Kinase Inhibitors Inhibit Both Toxoplasma Gondii MAPKL1 and CDPK1. ACS Infect. Dis. 2025. [CrossRef]
- Sugi, T.; Kawazu, S.; Horimoto, T.; Kato, K. A Single Mutation in the Gatekeeper Residue in TgMAPKL-1 Restores the Inhibitory Effect of a Bumped Kinase Inhibitor on the Cell Cycle. Int J Parasitol Drugs Drug Resist 2014, 5, 1–8. [CrossRef]
- Winzer, P.; Müller, J.; Aguado-Martínez, A.; Rahman, M.; Balmer, V.; Manser, V.; Ortega-Mora, L.M.; Ojo, K.K.; Fan, E.; Maly, D.J.; et al. In Vitro and In Vivo Effects of the Bumped Kinase Inhibitor 1294 in the Related Cyst-Forming Apicomplexans Toxoplasma Gondii and Neospora Caninum. Antimicrob Agents Chemother 2015, 59, 6361–6374. [CrossRef]
- Winzer, P.; Anghel, N.; Imhof, D.; Balmer, V.; Ortega-Mora, L.-M.; Ojo, K.K.; Van Voorhis, W.C.; Müller, J.; Hemphill, A. Neospora Caninum: Structure and Fate of Multinucleated Complexes Induced by the Bumped Kinase Inhibitor BKI-1294. Pathogens 2020, 9, 382.
- Winzer, P.; Müller, J.; Imhof, D.; Ritler, D.; Uldry, A.-C.; Braga-Lagache, S.; Heller, M.; Ojo, K.K.; Van Voorhis, W.C.; Ortega-Mora, L.-M.; et al. Neospora Caninum: Differential Proteome of Multinucleated Complexes Induced by the Bumped Kinase Inhibitor BKI-1294. Microorganisms 2020, 8, 801. [CrossRef]
- de Sousa, M.C.F.; Imhof, D.; Hänggeli, K.P.A.; Choi, R.; Hulverson, M.A.; Arnold, S.L.M.; Van Voorhis, W.C.; Fan, E.; Roberto, S.-S.; Ortega-Mora, L.M.; et al. Efficacy of the Bumped Kinase Inhibitor BKI-1708 against the Cyst-Forming Apicomplexan Parasites Toxoplasma Gondii and Neospora Caninum in Vitro and in Experimentally Infected Mice. Int J Parasitol Drugs Drug Resist 2024, 25, 100553. [CrossRef]
- Choi, R.; Hulverson, M.A.; Schaefer, D.A.; Betzer, D.P.; Riggs, M.W.; Huang, W.; Sun, V.; Whitman, G.R.; McCloskey, M.C.; Marsh, K.; et al. Anti-Cryptosporidium Efficacy of BKI-1708, an Inhibitor of Cryptosporidium Calcium-Dependent Protein Kinase 1. PLOS Neglected Tropical Diseases 2025, 19, e0013263. [CrossRef]
- de Sousa, M.C.F.; Müller, J.; Hänggeli, K.P.A.; Heller, M.; Uldry, A.-C.; Braga-Lagache, S.; Leitao, A.; Ortega-Mora, L.-M.; Ojo, K.K.; Van Voorhis, W.C.; et al. Bumped Kinase Inhibitor BKI-1708 Interferes in Cytokinesis and Drives Baryzoite Conversion in the Cyst-Forming Apicomplexan Parasites Toxoplasma Gondii, Neospora Caninum and Besnoitia Besnoiti. International Journal of Molecular Sciences 2026, 27, 2914. [CrossRef]
- Müller, J.; Boubaker, G.; Müller, N.; Uldry, A.-C.; Braga-Lagache, S.; Heller, M.; Hemphill, A. Investigating Antiprotozoal Chemotherapies with Novel Proteomic Tools—Chances and Limitations: A Critical Review. International Journal of Molecular Sciences 2024, 25, 6903. [CrossRef]
- Liles, N.W.; Page, E.E.; Liles, A.L.; Vesely, S.K.; Raskob, G.E.; George, J.N. Diversity and Severity of Adverse Reactions to Quinine: A Systematic Review. Am J Hematol 2016, 91, 461–466. [CrossRef]
- Müller, J.; Anghel, N.; Imhof, D.; Hänggeli, K.; Uldry, A.-C.; Braga-Lagache, S.; Heller, M.; Ojo, K.K.; Ortega-Mora, L.-M.; Van Voorhis, W.C.; et al. Common Molecular Targets of a Quinolone Based Bumped Kinase Inhibitor in Neospora Caninum and Danio Rerio. Int J Mol Sci 2022, 23, 2381.
- Ajiboye, J.; Uldry, A.-C.; Heller, M.; Naguleswaran, A.; Fan, E.; Van Voorhis, W.C.; Hemphill, A.; Müller, J. Molecular Targets of the 5-Amido-Carboxamide Bumped Kinase Inhibitor BKI-1748 in Cryptosporidium Parvum and HCT-8 Host Cells. International Journal of Molecular Sciences 2024, 25, 2707. [CrossRef]
- Müller, J.; Schlange, C.; Heller, M.; Uldry, A.-C.; Braga-Lagache, S.; Haynes, R.K.; Hemphill, A. Proteomic Characterization of Toxoplasma Gondii ME49 Derived Strains Resistant to the Artemisinin Derivatives Artemiside and Artemisone Implies Potential Mode of Action Independent of ROS Formation. International Journal for Parasitology: Drugs and Drug Resistance 2023, 21, 1–12.
- Semeraro, M.; Boubaker, G.; Scaccaglia, M.; Müller, J.; Vigneswaran, A.; Hänggeli, K.P.A.; Amdouni, Y.; Kramer, L.H.; Vismarra, A.; Genchi, M.; et al. Transient Adaptation of Toxoplasma Gondii to Exposure by Thiosemicarbazone Drugs That Target Ribosomal Proteins Is Associated with the Upregulated Expression of Tachyzoite Transmembrane Proteins and Transporters. Int J Mol Sci 2024, 25, 9067. [CrossRef]
- Kong, A.T.; Leprevost, F.V.; Avtonomov, D.M.; Mellacheruvu, D.; Nesvizhskii, A.I. MSFragger: Ultrafast and Comprehensive Peptide Identification in Mass Spectrometry–Based Proteomics. Nat Methods 2017, 14, 513–520. [CrossRef]
- Schwanhäusser, B.; Busse, D.; Li, N.; Dittmar, G.; Schuchhardt, J.; Wolf, J.; Chen, W.; Selbach, M. Global Quantification of Mammalian Gene Expression Control. Nature 2011, 473, 337–342. [CrossRef]
- Fereig, R.M.; Nishikawa, Y. Genetic Disruption of Toxoplasma Gondii Peroxiredoxin (TgPrx) 1 and 3 Reveals the Essential Role of TgPrx3 in Protecting Mice from Fatal Consequences of Toxoplasmosis. Int J Mol Sci 2022, 23, 3076. [CrossRef]
- Tang, B.L.; Peter, F.; Krijnse-Locker, J.; Low, S.H.; Griffiths, G.; Hong, W. The Mammalian Homolog of Yeast Sec13p Is Enriched in the Intermediate Compartment and Is Essential for Protein Transport from the Endoplasmic Reticulum to the Golgi Apparatus. Mol Cell Biol 1997, 17, 256–266. [CrossRef]
- Zhang, X.; Zhang, T.; Liu, J.; Li, M.; Fu, Y.; Xu, J.; Liu, Q. Functional Characterization of a Unique Cytochrome P450 in Toxoplasma Gondii. Oncotarget 2017, 8, 115079–115088. [CrossRef]
- Marshall, E.S.; Elshekiha, H.M.; Hakimi, M.-A.; Flynn, R.J. Toxoplasma Gondii Peroxiredoxin Promotes Altered Macrophage Function, Caspase-1-Dependent IL-1β Secretion Enhances Parasite Replication. Vet Res 2011, 42, 80. [CrossRef]
- Bosch, S.S.; Kronenberger, T.; Meissner, K.A.; Zimbres, F.M.; Stegehake, D.; Izui, N.M.; Schettert, I.; Liebau, E.; Wrenger, C. Oxidative Stress Control by Apicomplexan Parasites. BioMed Research International 2015, 2015, 351289. [CrossRef]
- Kafsack, B.F.C.; Pena, J.D.O.; Coppens, I.; Ravindran, S.; Boothroyd, J.C.; Carruthers, V.B. Rapid Membrane Disruption by a Perforin-like Protein Facilitates Parasite Exit from Host Cells. Science 2009, 323, 530–533. [CrossRef]
- Li, M.; Wang, H.; Liu, J.; Hao, P.; Ma, L.; Liu, Q. The Apoptotic Role of Metacaspase in Toxoplasma Gondii. Front Microbiol 2015, 6, 1560. [CrossRef]
- Li, M.; Liu, J.; Wu, Y.; Wu, Y.; Sun, X.; Fu, Y.; Zhang, X.; Liu, Q. Requirement of Toxoplasma Gondii Metacaspases for IMC1 Maturation, Endodyogeny and Virulence in Mice. Parasites & Vectors 2021, 14. [CrossRef]
- Radke, J.B.; Worth, D.; Hong, D.; Huang, S.; Jr, W.J.S.; Wilson, E.H.; White, M.W. Transcriptional Repression by ApiAP2 Factors Is Central to Chronic Toxoplasmosis. PLOS Pathogens 2018, 14, e1007035. [CrossRef]
- Pachano, B.; Farhat, D.C.; Shahinas, M.; Von Velsen, J.; Corrao, C.; Belmudes, L.; De Bock, P.-J.; Mas, C.; Couté, Y.; Bowler, M.W.; et al. An ISWI-Related Chromatin Remodeller Regulates Stage-Specific Gene Expression in Toxoplasma Gondii. Nat Microbiol 2025, 10, 1156–1170. [CrossRef]
- Hurt, J.A.; Obar, R.A.; Zhai, B.; Farny, N.G.; Gygi, S.P.; Silver, P.A. A Conserved CCCH-Type Zinc Finger Protein Regulates mRNA Nuclear Adenylation and Export. J Cell Biol 2009, 185, 265–277. [CrossRef]
- Howell, B.; Larsson, N.; Gullberg, M.; Cassimeris, L. Dissociation of the Tubulin-Sequestering and Microtubule Catastrophe-Promoting Activities of Oncoprotein 18/Stathmin. Mol Biol Cell 1999, 10, 105–118. [CrossRef]
- Sobel, A.; Boutterin, M.C.; Beretta, L.; Chneiweiss, H.; Doye, V.; Peyro-Saint-Paul, H. Intracellular Substrates for Extracellular Signaling. Characterization of a Ubiquitous, Neuron-Enriched Phosphoprotein (Stathmin). J Biol Chem 1989, 264, 3765–3772.
- Kotaka, M.; Kostin, S.; Ngai, S.; Chan, K.; Lau, Y.; Lee, S.M.; Li, H. y; Ng, E.K.; Schaper, J.; Tsui, S.K.; et al. Interaction of hCLIM1, an Enigma Family Protein, with Alpha-Actinin 2. J Cell Biochem 2000, 78, 558–565. [CrossRef]
- Rahajeng, J.; Giridharan, S.S.P.; Naslavsky, N.; Caplan, S. Collapsin Response Mediator Protein-2 (Crmp2) Regulates Trafficking by Linking Endocytic Regulatory Proteins to Dynein Motors. J Biol Chem 2010, 285, 31918–31922. [CrossRef]
- Tang, B.L.; Zhang, T.; Low, D.Y.; Wong, E.T.; Horstmann, H.; Hong, W. Mammalian Homologues of Yeast Sec31p. An Ubiquitously Expressed Form Is Localized to Endoplasmic Reticulum (ER) Exit Sites and Is Essential for ER-Golgi Transport. J Biol Chem 2000, 275, 13597–13604. [CrossRef]
- van der Vaart, B.; Manatschal, C.; Grigoriev, I.; Olieric, V.; Gouveia, S.M.; Bjelic, S.; Demmers, J.; Vorobjev, I.; Hoogenraad, C.C.; Steinmetz, M.O.; et al. SLAIN2 Links Microtubule plus End-Tracking Proteins and Controls Microtubule Growth in Interphase. J Cell Biol 2011, 193, 1083–1099. [CrossRef]
- Gong, C.X.; Wegiel, J.; Lidsky, T.; Zuck, L.; Avila, J.; Wisniewski, H.M.; Grundke-Iqbal, I.; Iqbal, K. Regulation of Phosphorylation of Neuronal Microtubule-Associated Proteins MAP1b and MAP2 by Protein Phosphatase-2A and -2B in Rat Brain. Brain Res 2000, 853, 299–309. [CrossRef]
- Wang, L.; Paudyal, S.C.; Kang, Y.; Owa, M.; Liang, F.-X.; Spektor, A.; Knaut, H.; Sánchez, I.; Dynlacht, B.D. Regulators of Tubulin Polyglutamylation Control Nuclear Shape and Cilium Disassembly by Balancing Microtubule and Actin Assembly. Cell Res 2022, 32, 190–209. [CrossRef]
- Gerke, V.; Moss, S.E. Annexins: From Structure to Function. Physiological Reviews 2002, 82, 331–371. [CrossRef]
- Tartaglia, G.G.; Hollås, H.; Håvik, B.; Vedeler, A.; Pastore, A. The RNA-Binding Properties of Annexins. J Mol Biol 2025, 437, 168933. [CrossRef]
- Benhalevy, D.; Gupta, S.K.; Danan, C.H.; Ghosal, S.; Sun, H.-W.; Kazemier, H.G.; Paeschke, K.; Hafner, M.; Juranek, S.A. The Human CCHC-Type Zinc Finger Nucleic Acid-Binding Protein Binds G-Rich Elements in Target mRNA Coding Sequences and Promotes Translation. Cell Rep 2017, 18, 2979–2990. [CrossRef]
- Pyronnet, S.; Dostie, J.; Sonenberg, N. Suppression of Cap-Dependent Translation in Mitosis. Genes Dev 2001, 15, 2083–2093. [CrossRef]
- Couture, J.-F.; Collazo, E.; Trievel, R.C. Molecular Recognition of Histone H3 by the WD40 Protein WDR5. Nat Struct Mol Biol 2006, 13, 698–703. [CrossRef]
- Beresford, P.J.; Zhang, D.; Oh, D.Y.; Fan, Z.; Greer, E.L.; Russo, M.L.; Jaju, M.; Lieberman, J. Granzyme A Activates an Endoplasmic Reticulum-Associated Caspase-Independent Nuclease to Induce Single-Stranded DNA Nicks. J Biol Chem 2001, 276, 43285–43293. [CrossRef]
- Fan, Z.; Beresford, P.J.; Oh, D.Y.; Zhang, D.; Lieberman, J. Tumor Suppressor NM23-H1 Is a Granzyme A-Activated DNase during CTL-Mediated Apoptosis, and the Nucleosome Assembly Protein SET Is Its Inhibitor. Cell 2003, 112, 659–672. [CrossRef]
- Ridge, R.J.; Sloane, N.H. Partial N-Terminal Amino Acid Sequence of the Anti-Neoplastic Urinary Protein (ANUP) and the Anti-Tumour Effect of the N-Terminal Nonapeptide of the Unique Cytokine Present in Human Granulocytes. Cytokine 1996, 8, 1–5. [CrossRef]
- Iourgenko, V.; Zhang, W.; Mickanin, C.; Daly, I.; Jiang, C.; Hexham, J.M.; Orth, A.P.; Miraglia, L.; Meltzer, J.; Garza, D.; et al. Identification of a Family of cAMP Response Element-Binding Protein Coactivators by Genome-Scale Functional Analysis in Mammalian Cells. Proc Natl Acad Sci U S A 2003, 100, 12147–12152. [CrossRef]
- Screaton, R.A.; Conkright, M.D.; Katoh, Y.; Best, J.L.; Canettieri, G.; Jeffries, S.; Guzman, E.; Niessen, S.; Yates, J.R.; Takemori, H.; et al. The CREB Coactivator TORC2 Functions as a Calcium- and cAMP-Sensitive Coincidence Detector. Cell 2004, 119, 61–74. [CrossRef]
- Takeuchi, A.; Miyamoto, T.; Yamaji, K.; Masuho, Y.; Hayashi, M.; Hayashi, H.; Onozaki, K. A Human Erythrocyte-Derived Growth-Promoting Factor with a Wide Target Cell Spectrum: Identification as Catalase. Cancer Res 1995, 55, 1586–1589.
- Chang, V.; Mahoney, K.E.; Lian, I.; Chen, R.; Chung, N.; Utheim, T.P.; Karlsson, N.G.; Malaker, S.A. In-Depth Analysis of the Tear Fluid Glycoproteome Reveals Diverse Lacritin Glycosylation and Spliceoforms 2025, 2025.06.13.659589.
- Wang, Q.; Moore, M.J.; Adelmant, G.; Marto, J.A.; Silver, P.A. PQBP1, a Factor Linked to Intellectual Disability, Affects Alternative Splicing Associated with Neurite Outgrowth. Genes Dev 2013, 27, 615–626. [CrossRef]
- Waragai, M.; Lammers, C.H.; Takeuchi, S.; Imafuku, I.; Udagawa, Y.; Kanazawa, I.; Kawabata, M.; Mouradian, M.M.; Okazawa, H. PQBP-1, a Novel Polyglutamine Tract-Binding Protein, Inhibits Transcription Activation by Brn-2 and Affects Cell Survival. Hum Mol Genet 1999, 8, 977–987. [CrossRef]
- Chen, Y.-J.; Mahieu, N.G.; Huang, X.; Singh, M.; Crawford, P.A.; Johnson, S.L.; Gross, R.W.; Schaefer, J.; Patti, G.J. Lactate Metabolism Is Associated with Mammalian Mitochondria. Nat Chem Biol 2016, 12, 937–943. [CrossRef]
- Hölzel, M.; Grimm, T.; Rohrmoser, M.; Malamoussi, A.; Harasim, T.; Gruber-Eber, A.; Kremmer, E.; Eick, D. The BRCT Domain of Mammalian Pes1 Is Crucial for Nucleolar Localization and rRNA Processing. Nucleic Acids Res 2007, 35, 789–800. [CrossRef]
- Rohrmoser, M.; Hölzel, M.; Grimm, T.; Malamoussi, A.; Harasim, T.; Orban, M.; Pfisterer, I.; Gruber-Eber, A.; Kremmer, E.; Eick, D. Interdependence of Pes1, Bop1, and WDR12 Controls Nucleolar Localization and Assembly of the PeBoW Complex Required for Maturation of the 60S Ribosomal Subunit. Mol Cell Biol 2007, 27, 3682–3694. [CrossRef]
- Liu, Y.; Mattila, J.; Ventelä, S.; Yadav, L.; Zhang, W.; Lamichane, N.; Sundström, J.; Kauko, O.; Grénman, R.; Varjosalo, M.; et al. PWP1 Mediates Nutrient-Dependent Growth Control through Nucleolar Regulation of Ribosomal Gene Expression. Dev Cell 2017, 43, 240-252.e5. [CrossRef]
- Cuylen, S.; Blaukopf, C.; Politi, A.Z.; Müller-Reichert, T.; Neumann, B.; Poser, I.; Ellenberg, J.; Hyman, A.A.; Gerlich, D.W. Ki-67 Acts as a Biological Surfactant to Disperse Mitotic Chromosomes. Nature 2016, 535, 308–312. [CrossRef]
- Blume, M.; Seeber, F. Metabolic Interactions between Toxoplasma Gondii and Its Host. F1000Res 2018, 7, F1000 Faculty Rev-1719. [CrossRef]
- Sun, H.; Li, J.; Wang, L.; Yin, K.; Xu, C.; Liu, G.; Xiao, T.; Huang, B.; Wei, Q.; Gong, M.; et al. Comparative Proteomics Analysis for Elucidating the Interaction Between Host Cells and Toxoplasma Gondii. Front Cell Infect Microbiol 2021, 11, 643001. [CrossRef]
- Wu, S.-Z.; Wei, H.-X.; Jiang, D.; Li, S.-M.; Zou, W.-H.; Peng, H.-J. Genome-Wide CRISPR Screen Identifies Host Factors Required by Toxoplasma Gondii Infection. Front Cell Infect Microbiol 2020, 9, 460. [CrossRef]
- Arning, S.; Grüter, P.; Bilbe, G.; Krämer, A. Mammalian Splicing Factor SF1 Is Encoded by Variant cDNAs and Binds to RNA. RNA 1996, 2, 794–810.
- Wang, X.; Bruderer, S.; Rafi, Z.; Xue, J.; Milburn, P.J.; Krämer, A.; Robinson, P.J. Phosphorylation of Splicing Factor SF1 on Ser20 by cGMP-Dependent Protein Kinase Regulates Spliceosome Assembly. EMBO J 1999, 18, 4549–4559. [CrossRef]
| Toxo DB ORF | Annotation | rAbu |
|---|---|---|
| TGME49_230410 | peroxiredoxin PRX3 | 604.9 |
| TGME49_311400 | protein transport protein SEC31, putative | 368.6 |
| TGME49_290920 | oxidoreductase, 2OG-Fe(II) oxygenase family protein | 251.6 |
| TGME49_209420 | hypothetical protein | 248.3 |
| TGME49_201700 | protein transport protein SEC13 | 232.4 |
| TGME49_224720 | SPOC domain-containing protein | 216.6 |
| TGME49_205180 | RNA recognition motif-containing protein | 190.9 |
| TGME49_313270 | hypothetical protein | 174.3 |
| TGME49_320600 | cold-shock DNA-binding domain-containing protein | 172.5 |
| TGME49_232370 | CW-type Zinc Finger protein | 160.7 |
| TGME49_273960 | chaperonin GroS protein | 151.5 |
| TGME49_231440 | LsmAD domain-containing protein | 137.2 |
| TGME49_294670 | eukaryotic translation initiation factor 3 subunit G, putative | 127.0 |
| TGME49_263530 | chaperonin, putative | 91.1 |
| TGME49_265250 | RNA recognition motif-containing protein | 84.3 |
| TGME49_202780 | rhoptry kinase family protein ROP25 | 74.6 |
| TGME49_250830 | 26S proteasome regulatory subunit RPN12, putative | 73.9 |
| TGME49_201760 | thioredoxin-like associated protein TLAP4 | 61.1 |
| TGME49_218240 | inner membrane complex protein IMC25 | 54.4 |
| TGME49_278975 | metacaspase MCA2 | 51.5 |
| TGME49_205320 | hypothetical protein | 48.2 |
| TGME49_291680 | protein transport protein SEC23, putative | 44.4 |
| TGME49_309200 | zinc finger (CCCH type) motif-containing protein | 41.6 |
| TGME49_204160 | GYF domain-containing protein | 39.1 |
| TGME49_269690 | dense granule protein GRA29 | 36.0 |
| TGME49_250115 | hypothetical protein | 21.1 |
| TGME49_204130 | perforin-like protein PLP1 | 20.0 |
| TGME49_232280 | hypothetical protein | 18.6 |
| TGME49_215360 | dense granule protein GRA62 | 17.9 |
| TGME49_258240 | chromodomain helicase DNA binding protein CHD1/SWI2/SNF2 | 10.7 |
| TGME49_298610 | GYF domain-containing protein | 9.6 |
| TGME49_233120 | AP2 domain transcription factor AP2VIII-2 | 8.6 |
| TGME49_253750 | PLU-1 family protein | 7.3 |
| TGME49_254940 | MIF4G domain-containing protein | 7.3 |
| TGME49_244500 | Tubulin-tyrosine ligase family protein | 2.8 |
| Toxo DB ORF | Annotation | rAbu BKI-1708 |
rAbu Quinine |
|---|---|---|---|
| TGME49_201860 | hypothetical protein | 1185.1 | 600.9 |
| TGME49_205558 | NAC domain-containing protein | 827.0 | 53.9 |
| TGME49_244110 | nucleosome assembly protein (nap) protein | 300.3 | 34.8 |
| TGME49_314830 | pre-mRNA splicing factor subunit, putative | 202.3 | 160.9 |
| TGME49_232000 | dense granule protein GRA30 | 190.2 | 43.6 |
| TGME49_248250 | translation initiation factor IF-2, putative | 175.8 | 376.6 |
| TGME49_294970 | hypothetical protein | 156.6 | 38.8 |
| TGME49_265530 | RNA recognition motif-containing protein | 130.9 | 8.9 |
| TGME49_257380 | inhibitor of cysteine protease 1 | 122.5 | 181.9 |
| TGME49_263090 | 14-3-3 protein | 115.4 | 50.4 |
| TGME49_500284 | hypothetical protein, conserved | 88.8 | 13.9 |
| TGME49_291330 | RNA recognition motif-containing protein | 88.1 | 105.8 |
| TGME49_260670 | centrin 3 | 86.9 | 61.6 |
| TGME49_213030 | clathrin light chain, putative | 84.2 | 437.1 |
| TGME49_213940 | CHCH domain-containing protein | 78.1 | 35.8 |
| TGME49_280550 | clathrin adaptor complex small chain subfamily protein | 74.7 | 77.3 |
| TGME49_300280 | LSM domain-containing protein | 64.5 | 48.0 |
| TGME49_269180 | MIF4G domain-containing protein | 60.4 | 4.5 |
| TGME49_213050 | hypothetical protein | 60.2 | 41.2 |
| TGME49_313640 | hypothetical protein | 51.6 | 166.6 |
| Protein ID | Annotation | rAbu BKI-1708 |
rAbu Quinine |
|---|---|---|---|
| P47756 | F-actin-capping protein subunit beta | 164.5 | 663.0 |
| P55735 | Protein SEC13 homolog | 161.5 | 351.3 |
| Q9NR12 | PDZ and LIM domain protein 7 | 156.5 | 210.0 |
| P52907 | F-actin-capping protein subunit alpha-1 | 147.5 | 741.8 |
| Q07021 | Complement component 1 Q subcomponent-binding protein, mitochondrial | 135.2 | 151.0 |
| Q8WX93-3 | Isoform 3 of Palladin | 109.7 | 11.8 |
| P51608-2 | Isoform B of Methyl-CpG-binding protein 2 | 92.1 | 45.6 |
| P09493-8 | Isoform 8 of Tropomyosin alpha-1 chain | 71.6 | 463.0 |
| P51659 | Peroxisomal multifunctional enzyme type 2 | 60.5 | 18.0 |
| Q96GY0 | Zinc finger C2HC domain-containing protein 1A | 60.3 | 31.4 |
| Q9C0C2 | 182 kDa tankyrase-1-binding protein | 50.5 | 160.2 |
| Q8ND56-2 | Isoform 2 of Protein LSM14 homolog A | 49.4 | 64.1 |
| Q16531 | DNA damage-binding protein 1 | 35.4 | 18.0 |
| Q13428-2 | Isoform 2 of Treacle protein | 31.0 | 143.0 |
| Q9HAU0-2 | Isoform 2 of Pleckstrin homology domain-containing family A member 5 | 28.7 | 63.5 |
| Q14974 | Importin subunit beta-1 | 18.6 | 449.9 |
| P08123 | Collagen alpha-2(I) chain | 15.2 | 20.9 |
| P50281 | Matrix metalloproteinase-14 | 12.1 | 55.5 |
| P49821-2 | Isoform 2 of NADH dehydrogenase [ubiquinone] flavoprotein 1, mitochondrial | 11.6 | 25.4 |
| O60664-3 | Isoform 3 of Perilipin-3 | 9.3 | 192.6 |
| O14974-5 | Isoform 5 of Protein phosphatase 1 regulatory subunit 12A | 9.2 | 185.0 |
| Protein ID | Annotation | rAbu |
|---|---|---|
| Q9Y2S6 | Translation machinery-associated protein 7 | 5041.5 |
| P16949-2 | Isoform 2 of Stathmin | 1608.4 |
| P55000 | Secreted Ly-6/uPAR-related protein 1 | 324.7 |
| P50995-2 | Isoform 2 of Annexin A11 | 286.3 |
| P82909 | Alpha-ketoglutarate dehydrogenase component 4 | 195.2 |
| Q96FJ2 | Dynein light chain 2, cytoplasmic | 160.2 |
| P01036 | Cystatin-S | 157.5 |
| P62633-2 | Isoform 2 of CCHC-type zinc finger nucleic acid binding protein | 128.7 |
| Q01105 | Protein SET | 77.4 |
| O00151 | PDZ and LIM domain protein 1 | 66.2 |
| P61964 | WD repeat-containing protein 5 | 62.6 |
| P11137-2 | Isoform 2 of Microtubule-associated protein 2 | 61.7 |
| Q6UUV7-3 | Isoform 3 of CREB-regulated transcription coactivator 3 | 57.6 |
| Q16555-2 | Isoform 2 of Dihydropyrimidinase-related protein 2 | 49.7 |
| Q9P270 | SLAIN motif-containing protein 2 | 47.5 |
| O94979-10 | Isoform 10 of Protein transport protein Sec31A | 40.5 |
| P20073-2 | Isoform 2 of Annexin A7 | 30.0 |
| P78344 | Eukaryotic translation initiation factor 4 gamma 2 | 27.7 |
| P04040 | Catalase | 26.7 |
| P27816-5 | Isoform 5 of Microtubule-associated protein 4 | 24.6 |
| Protein ID | Annotation | rAbu |
|---|---|---|
| Q9GZZ8 | Extracellular glycoprotein lacritin | 244.7 |
| O60828-2 | Isoform 2 of Polyglutamine-binding protein 1 | 76.5 |
| P07195 | L-lactate dehydrogenase B chain | 43.1 |
| Q6UWP8-2 | Isoform 2 of Suprabasin | 26.1 |
| P49790-2 | Isoform 2 of Nuclear pore complex protein Nup153 | 23.7 |
| O00541-2 | Isoform 2 of Pescadillo homolog | 23.6 |
| Q9BQG0-2 | Isoform 2 of Myb-binding protein 1A | 22.6 |
| P46013-2 | Isoform Short of Proliferation marker protein Ki-67 | 8.7 |
| Protein ID | Annotation | BKI-1708 rAbu | Quinine rAbu |
|---|---|---|---|
| P11387 | DNA topoisomerase 1 | 97.2 | 27.2 |
| Q9NX24 | H/ACA ribonucleoprotein complex subunit 2 | 85.5 | 98.7 |
| O75400-2 | Isoform 2 of Pre-mRNA-processing factor 40 homolog A | 45.0 | 7.0 |
| Q9NXV6 | CDKN2A-interacting protein | 30.4 | 9.7 |
| P78316 | Nucleolar protein 14 | 21.4 | 5.7 |
| P31943 | Heterogeneous nuclear ribonucleoprotein H | 17.1 | 139.2 |
| Q8NDZ4 | Divergent protein kinase domain 2A | 16.7 | 2.8 |
| P00750-2 | Isoform 2 of Tissue-type plasminogen activator | 12.3 | 17.4 |
| O94776-2 | Isoform 2 of Metastasis-associated protein MTA2 | 11.1 | 15.3 |
| Q8WUM4-2 | Isoform 2 of Programmed cell death 6-interacting protein | 10.3 | 2.4 |
| Protein ID | Annotation | rAbu non-infected |
rAbu infected |
|---|---|---|---|
| Q9Y5V0 | Zinc finger protein 706 | 645.0 | 55.9 |
| Q15637-2 | Isoform 2 of Splicing factor 1 | 456.8 | 584.5 |
| P78406 | mRNA export factor RAE1 | 257.1 | 136.8 |
| Q6E0U4-16 | Isoform 16 of Dermokine | 228.9 | 127.8 |
| O43684-2 | Isoform 2 of Mitotic checkpoint protein BUB3 | 213.9 | 44.1 |
| Q13492-2 | Isoform 2 of Phosphatidylinositol-binding clathrin assembly protein | 194.8 | 66.7 |
| O75223-3 | Isoform 3 of Gamma-glutamylcyclotransferase | 192.0 | 149.3 |
| Q9NPA8-2 | Isoform 2 of Transcription and mRNA export factor ENY2 | 171.9 | 101.7 |
| Q8WWM7-2 | Isoform 2 of Ataxin-2-like protein | 157.0 | 167.9 |
| P0CG12 | Decreased expression in renal and prostate cancer protein | 75.6 | 30.3 |
| P22234-2 | Isoform 2 of Bifunctional phosphoribosylaminoimidazole carboxylase/phosphoribosylaminoimidazole succinocarboxamide synthetase | 15.3 | 51.0 |
| Q96AE4-2 | Isoform 2 of Far upstream element-binding protein 1 | 13.7 | 154.3 |
| Non-infected HFF | Infected HFF | ||||
|---|---|---|---|---|---|
| Protein ID | Annotation | BKI-1708 rAbu | Quinine rAbu | BKI-1708 rAbu | Quinine rAbu |
| P35637-2 | Isoform Short of RNA-binding protein FUS | 571.8 | 257.7 | 391.0 | 69.1 |
| Q14011 | Cold-inducible RNA-binding protein | 506.4 | 299.3 | 203.4 | 63.9 |
| Q9H0D6-2 | Isoform 2 of 5’-3’ exoribonuclease 2 | 16.8 | 41.5 | 65.1 | 18.2 |
| P63167 | Dynein light chain 1, cytoplasmic | 791.7 | 1136.8 | 41.2 | 42.5 |
| Q9UHB6-4 | Isoform 4 of LIM domain and actin-binding protein 1 | 8.7 | 584.4 | 20.7 | 2.4 |
| Q06830 | Peroxiredoxin-1 | 458.2 | 246.8 | 19.5 | 53.7 |
| Q9UN86-2 | Isoform B of Ras GTPase-activating protein-binding protein 2 | 41.5 | 81.4 | 15.9 | 16.1 |
| P12956-2 | Isoform 2 of X-ray repair cross-complementing protein 6 | 19.3 | 61.7 | 11.3 | 56.0 |
| Q8WWI1-3 | Isoform 3 of LIM domain only protein 7 | 20.3 | 137.5 | 6.2 | 1.7 |
| Function | Uninfected HFF proteins | Infected HFF proteins | Infected and uninfected host cell proteins | T. gondii ME49 proteins |
|---|---|---|---|---|
| DNA binding and modification | 2 | 3 | 0 | 5 |
| RNA binding and modification | 4 | 2 | 5 | 6 |
| Protein binding and modification | 3 | 1 | 1 | 5 |
| Cytoskeleton and intracellular transport | 6 | 0 | 1 | 4 |
| Intracellular signalling | 2 | 0 | 0 | 5 |
| Energy and intermediary metabolism | 0 | 1 | 2 | 2 |
| Hypothetical or ambiguous | 3 | 1 | 3 | 6 |
| Total | 20 | 8 | 12 | 35 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




