Submitted:
13 April 2026
Posted:
14 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Experimental Animals and Tissue Sample Collection
2.2. Phenotype Data Sources and Measurement Methods
2.3. Quality Control of Genomic DNA
2.4. Whole Genome Resequencing
2.5. Quality Control
2.6. Alignment to Reference Genome
2.7. SNP Detection
2.8. Variant Annotation
2.9. Quality Control
2.10. GWAS
2.11. Candidate Gene and Functional Enrichment Analysis
3. Results
3.1. Descriptive Statistics of Body Size Traits in Yanqi Horses
3.2. Genomic data statistics
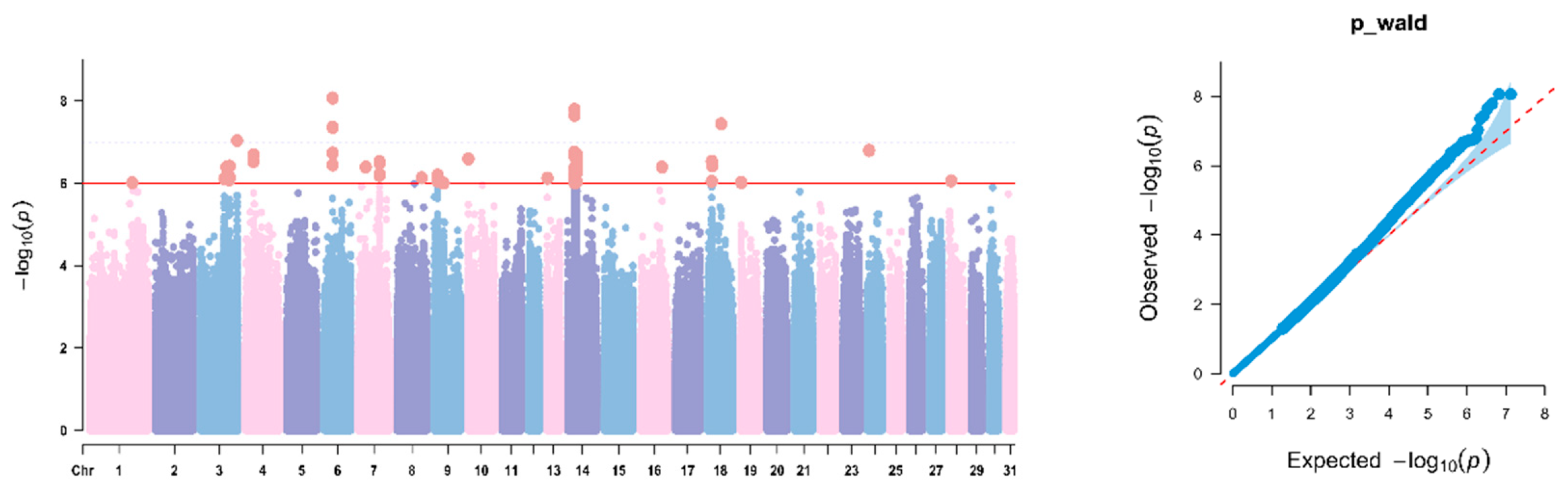
3.3. GWAS of WH Traits in Yanqi Horses
3.4. GWAS of BL Traits in Yanqi Horses
3.5. GWAS of HG Traits in Yanqi Horses
3.6. GWAS of CBC Traits in Yanqi Horses
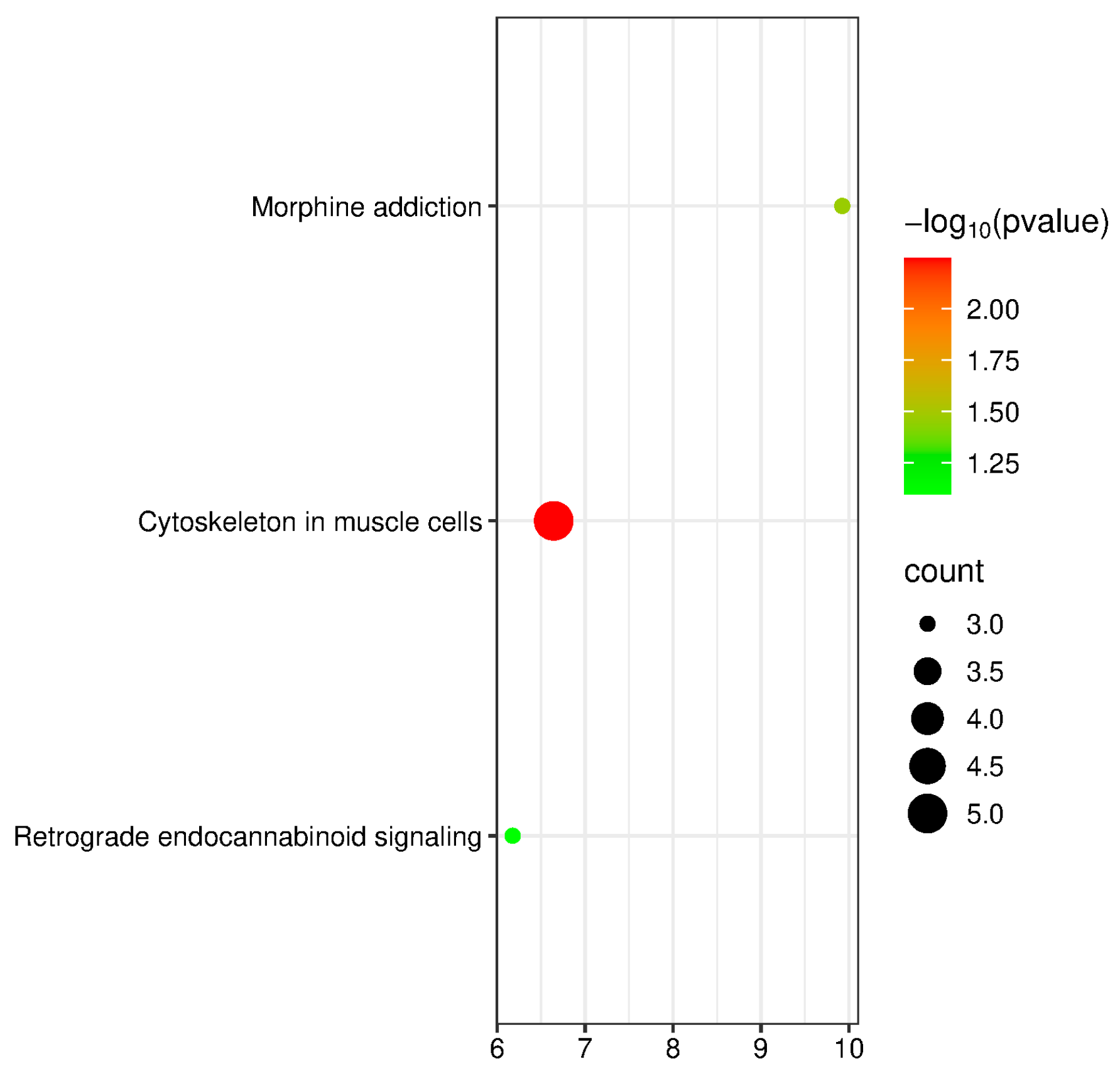
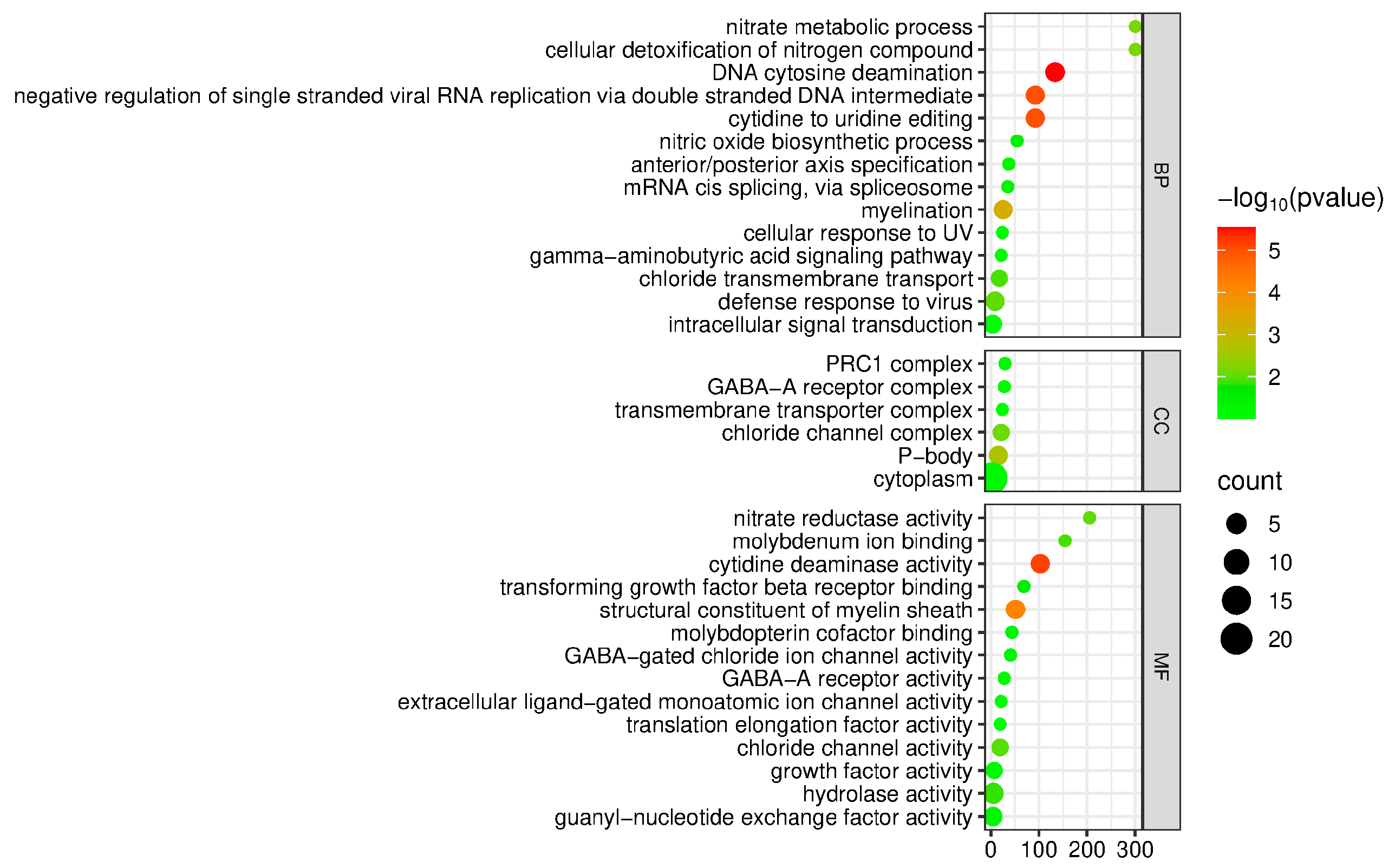
3.7. Functional Annotation and Enrichment Analysis of Candidate Genes for WH Traits in Yanqi Horses


3.8. Functional Annotation and Enrichment Analysis of Candidate Genes for BL Traits in Yanqi Horses
3.9. Functional Annotation and Enrichment Analysis of Candidate Genes for Hg Traits in Yanqi Horses
3.10. Functional Annotation and Enrichment Analysis of Candidate Genes for CBC Traits in Yanqi Horses
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Che, Limuge. Genome-wide Association Study For growth Traits-related Candidate Functional Gene in Chinese Simmental Cattle. Master’s Thesis, Inner Mongolia University, 2023. [Google Scholar]
- Sun, Xueliang. Genetic Diversity and Genome-wide Association Study of Coat Color in Chuanzhong Black Goats. Master’s Thesis, Sichuan Agricultural University, 2023. [Google Scholar]
- Haodong, Li. Study on Whole Genome Selection Technique for Body Weight and other Traits of Sika Deer. Master’s Thesis, Chinese Academy of Agricultural Sciences, 2023. [Google Scholar]
- Hu, Honghong. Genetic Basis and Genomic Selection of Longevity Traits in Holstein Cattle. PhD Thesis, Ningxia University, 2023. [Google Scholar]
- Wu, Ziyi. Genome-wide Association Study and Genomic Selection for Growth-related Traits in Yunong-black. Master’s Thesis, Henan Agricultural University, 2024. [Google Scholar]
- Lu, Xiaotian. Genome-wide Association Analysis of Fleece Traits in Northwest Xizang White Cashmere Goat. Master’s Thesis, Inner Mongolia Agricultural University, 2024. [Google Scholar]
- Wang, Menghan. Genetic Parameter Estimation and Genome-wide Association Study for Growth Traits in East Friesian-Hu hybrid sheep. Master’s Thesis, Northwest A&F University, 2024. [Google Scholar]
- Sun, Yan. Genome-wide Association Study For Numbers of Thoracic and Lumber Vertebrae in Dezhou Donkey and Functional Study of Candidate Genes. PhD Thesis, Shandong Agricultural University, 2023. [Google Scholar]
- Wang, Chuankun. GWAS, ROH and SVs Mining Candidate Genes for Important Traits in Yili Horse. PhD Thesis, Xinjiang Agricultural University, 2024. [Google Scholar]
- Ropka-Molik, K.; Stefaniuk-Szmukier, M.; Musiał, A.D.; Velie, B.D. The Genetics of Racing Performance in Arabian Horses. Int J Genomics 2019, 2019, 9013239. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.C.; Chow, C.C.; Tellier, L.C.; Vattikuti, S.; Purcell, S.M.; Lee, J.J. Second-Generation PLINK: Rising to the Challenge of Larger and Richer Datasets. Gigascience 2015, 4, 7. [Google Scholar] [CrossRef]
- Tang, Y.; Liu, X.; Wang, J.; Li, M.; Wang, Q.; Tian, F.; Su, Z.; Pan, Y.; Liu, D.; Lipka, A.E.; et al. GAPIT Version 2: An Enhanced Integrated Tool for Genomic Association and Prediction. Plant Genome 2016, 9. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Stephens, M. Genome-Wide Efficient Mixed-Model Analysis for Association Studies. Nat Genet 2012, 44, 821–824. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Wang, K. Genomic Variant Annotation and Prioritization with ANNOVAR and wANNOVAR. Nat Protoc 2015, 10, 1556–1566. [Google Scholar] [CrossRef]
- Sherman, B.T.; Hao, M.; Qiu, J.; Jiao, X.; Baseler, M.W.; Lane, H.C.; Imamichi, T.; Chang, W. DAVID: A Web Server for Functional Enrichment Analysis and Functional Annotation of Gene Lists (2021 Update). Nucleic Acids Res 2022, 50, W216–W221. [Google Scholar] [CrossRef]
- Doekes, H.P.; Veerkamp, R.F.; Bijma, P.; Hiemstra, S.J.; Windig, J.J. Trends in Genome-Wide and Region-Specific Genetic Diversity in the Dutch-Flemish Holstein-Friesian Breeding Program from 1986 to 2015. Genet Sel Evol 2018, 50, 15. [Google Scholar] [CrossRef]
- Mebratie, W.; Reyer, H.; Wimmers, K.; Bovenhuis, H.; Jensen, J. Genome Wide Association Study of Body Weight and Feed Efficiency Traits in a Commercial Broiler Chicken Population, a Re-Visitation. Sci Rep 2019, 9, 922. [Google Scholar] [CrossRef]
- Olsen, R.W.; Sieghart, W. GABA A Receptors: Subtypes Provide Diversity of Function and Pharmacology. Neuropharmacology 2009, 56, 141–148. [Google Scholar] [CrossRef]
- Tahir, M.S.; Porto-Neto, L.R.; Gondro, C.; Shittu, O.B.; Wockner, K.; Tan, A.W.L.; Smith, H.R.; Gouveia, G.C.; Kour, J.; Fortes, M.R.S. Meta-Analysis of Heifer Traits Identified Reproductive Pathways in Bos Indicus Cattle. Genes (Basel) 2021, 12, 768. [Google Scholar] [CrossRef]
- Zhao, G.; Liu, Y.; Niu, Q.; Zheng, X.; Zhang, T.; Wang, Z.; Xu, L.; Zhu, B.; Gao, X.; Zhang, L.; et al. Runs of Homozygosity Analysis Reveals Consensus Homozygous Regions Affecting Production Traits in Chinese Simmental Beef Cattle. BMC Genomics 2021, 22, 678. [Google Scholar] [CrossRef] [PubMed]
- Cox, G.A.; Mahaffey, C.L.; Nystuen, A.; Letts, V.A.; Frankel, W.N. The Mouse Fidgetin Gene Defines a New Role for AAA Family Proteins in Mammalian Development. Nat Genet 2000, 26, 198–202. [Google Scholar] [CrossRef]
- Timón-Gómez, A.; Nývltová, E.; Abriata, L.A.; Vila, A.J.; Hosler, J.; Barrientos, A. Mitochondrial Cytochrome c Oxidase Biogenesis: Recent Developments. Semin Cell Dev Biol 2018, 76, 163–178. [Google Scholar] [CrossRef] [PubMed]
- Akter, M.S.; Hada, M.; Shikata, D.; Watanabe, G.; Ogura, A.; Matoba, S. CRISPR/Cas9-Based Genetic Screen of SCNT-Reprogramming Resistant Genes Identifies Critical Genes for Male Germ Cell Development in Mice. Sci Rep 2021, 11, 15438. [Google Scholar] [CrossRef]
- Gimpel, P.; Lee, Y.L.; Sobota, R.M.; Calvi, A.; Koullourou, V.; Patel, R.; Mamchaoui, K.; Nédélec, F.; Shackleton, S.; Schmoranzer, J.; et al. Nesprin-1α-Dependent Microtubule Nucleation from the Nuclear Envelope via Akap450 Is Necessary for Nuclear Positioning in Muscle Cells. Curr Biol 2017, 27, 2999–3009.e9. [Google Scholar] [CrossRef]
- Zeng, J.; Xi, J.; Li, B.; Yan, X.; Dai, Y.; Wu, Y.; Xiao, Y.; Pei, Y.; Zhang, M. Microtubules Play a Crucial Role in Regulating Actin Organization and Cell Initiation in Cotton Fibers. Plant Cell Rep 2022, 41, 1059–1073. [Google Scholar] [CrossRef]
- Pollard, T.D.; Cooper, J.A. Actin, a Central Player in Cell Shape and Movement. Science 2009, 326, 1208–1212. [Google Scholar] [CrossRef]
- Heasman, S.J.; Ridley, A.J. Mammalian Rho GTPases: New Insights into Their Functions from in Vivo Studies. Nat Rev Mol Cell Biol 2008, 9, 690–701. [Google Scholar] [CrossRef] [PubMed]
- Hanson, A.M.; Stodieck, L.S.; Cannon, C.M.A.; Simske, S.J.; Ferguson, V.L. Seven Days of Muscle Re-Loading and Voluntary Wheel Running Following Hindlimb Suspension in Mice Restores Running Performance, Muscle Morphology and Metrics of Fatigue but Not Muscle Strength. J Muscle Res Cell Motil 2010, 31, 141–153. [Google Scholar] [CrossRef] [PubMed]
- Lupu, F.; Terwilliger, J.D.; Lee, K.; Segre, G.V.; Efstratiadis, A. Roles of Growth Hormone and Insulin-like Growth Factor 1 in Mouse Postnatal Growth. Dev Biol 2001, 229, 141–162. [Google Scholar] [CrossRef]
- Bassett, J.H.D.; Williams, G.R. Role of Thyroid Hormones in Skeletal Development and Bone Maintenance. Endocr Rev 2016, 37, 135–187. [Google Scholar] [CrossRef] [PubMed]










| Traits | Number | Mean | SD | Max | Min | CV% |
|---|---|---|---|---|---|---|
| Withers height (cm) | 183 | 132.07 | 0.96 | 152.00 | 98.00 | 0.73 |
| Body length (cm) | 183 | 131.28 | 1.30 | 160.00 | 90.00 | 0.99 |
| Heart girth (cm) | 183 | 150.01 | 1.70 | 180.00 | 90.00 | 1.13 |
| Cannon bone circumference (cm) | 183 | 16.64 | 0.13 | 20.00 | 12.00 | 0.78 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




