Submitted:
12 April 2026
Posted:
14 April 2026
You are already at the latest version
Abstract

Keywords:
1. Introduction
2. Materials and Methods
2.1. Fungal Isolates
2.2. Genes and PCR Primers
2.3. DNA Extraction and PCR
2.4. Phylogenetic Analysis
2.5. Plant Materials
2.6. Preparation of Inoculum
2.7. Plant Inoculation
2.8. Reisolation of Fusarium oxysporum from Infected Plants
2.9. Statistical Analysis
2.10. Genomics Analysis
2.10.1. Genome Assemblies
2.10.2. Structural Annotation of Protein-Coding Genes
2.10.3. Collinearity Analysis
2.10.4. Manual Curation of SIX Gene Annotations
2.10.5. Orthologue Clustering and Evolutionary Analysis
2.10.6. Orthologous Analysis Across the SIX7-10-12 Gene Cluster
3. Results
3.1. Detection of SIX Genes in Fusarium oxysporum Isolates from Ginger
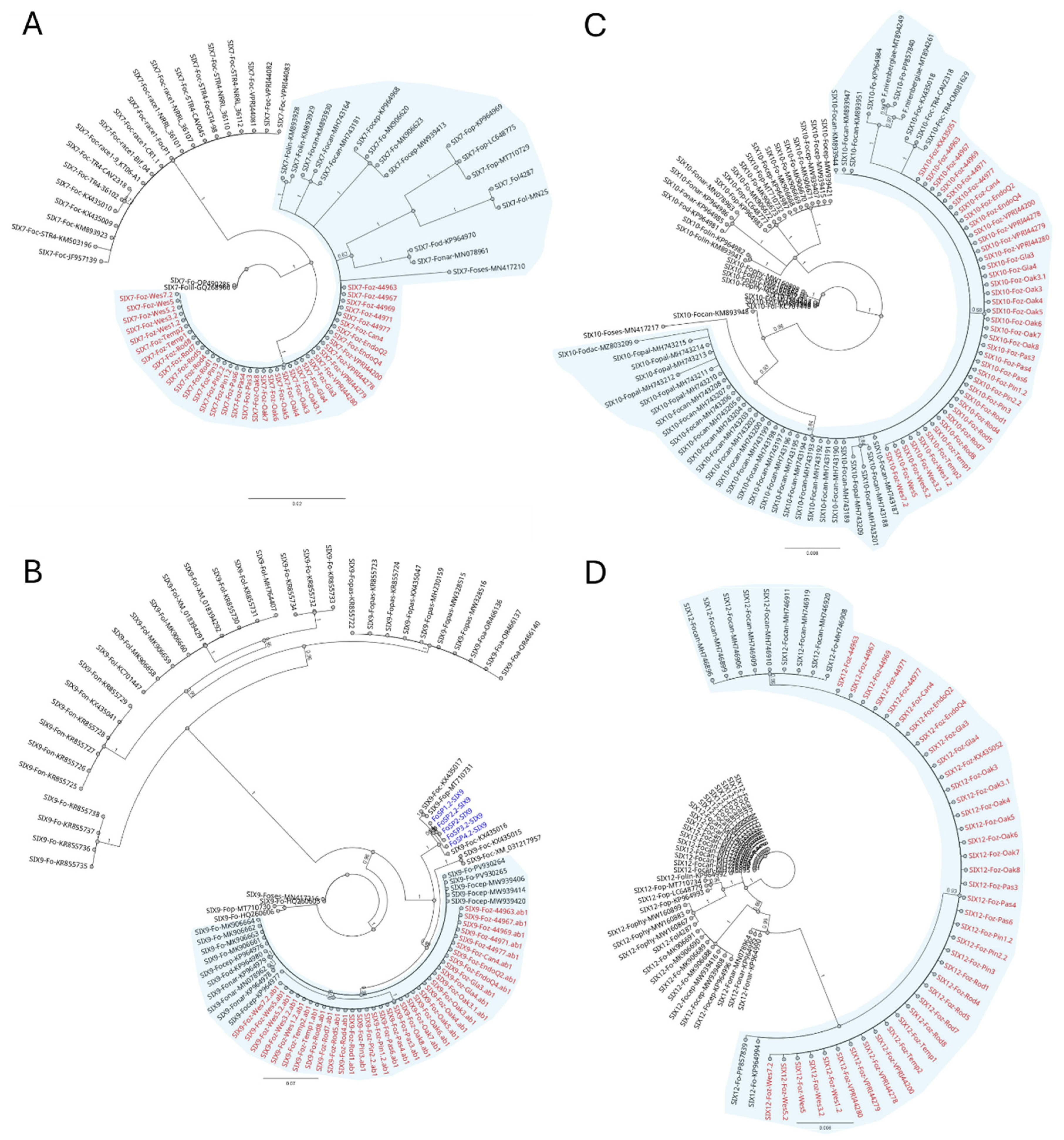
3.2. Phylogenetic Placements of Fusarium oxysporum Isolates Within Fusarium oxysporum Species Complex
3.3. Sequence Analysis of the SIX Genes from the Foz Isolates
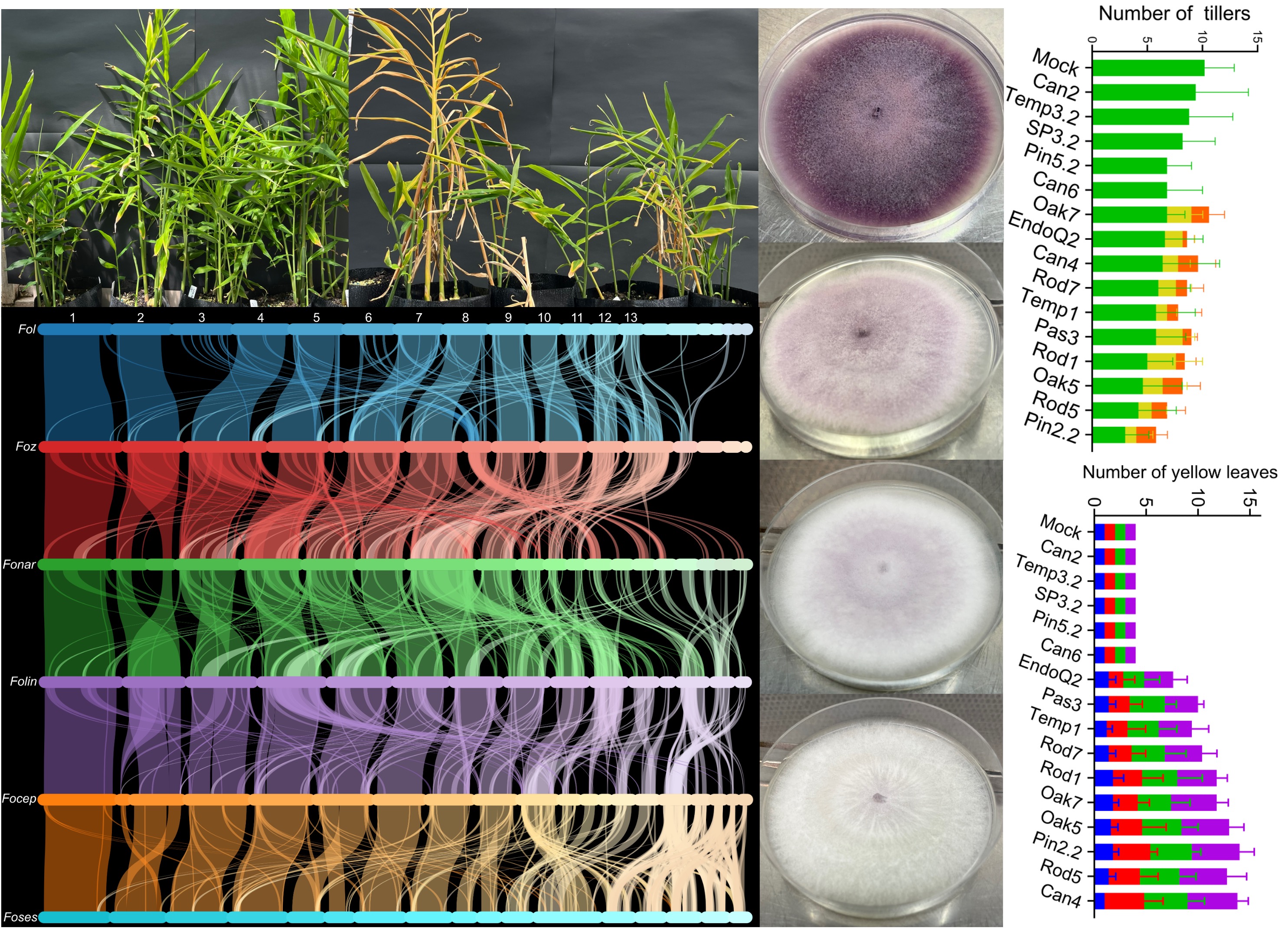
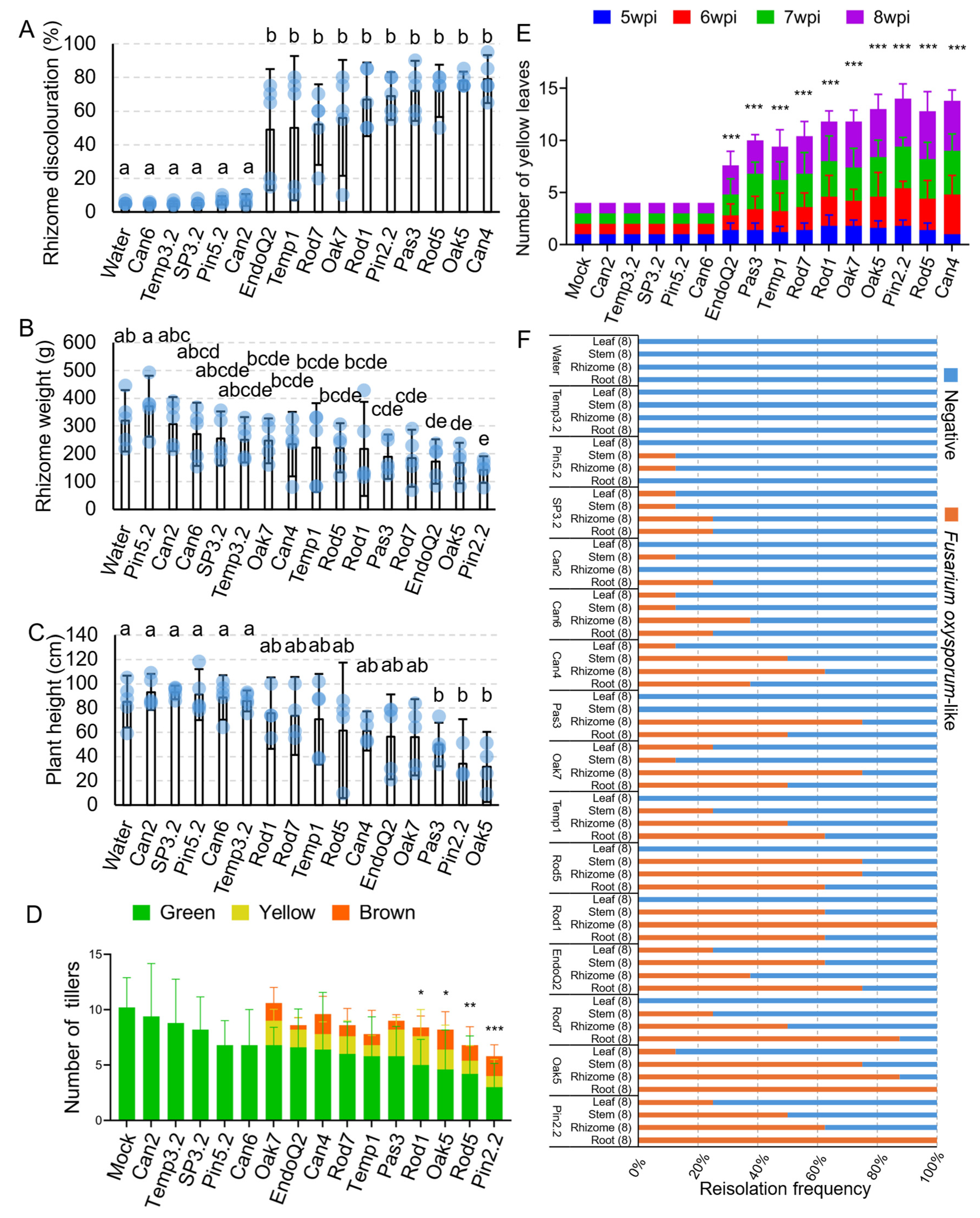
3.4. Greenhouse Pathogenicity Testing Using Pathogenic and Endophytic Fusarium oxysporum Isolates
3.5. Plant Symptomatic Assessment and Comparisons Between Foz and Endophytes
3.6. Reisolation of Fusarium oxysporum from Ginger Tissues at Harvest
3.7. Comparative Genomics
3.7.1. Rational for Choosing Genomes for Comparative Analysis
3.7.2. Genome Structure Analysis
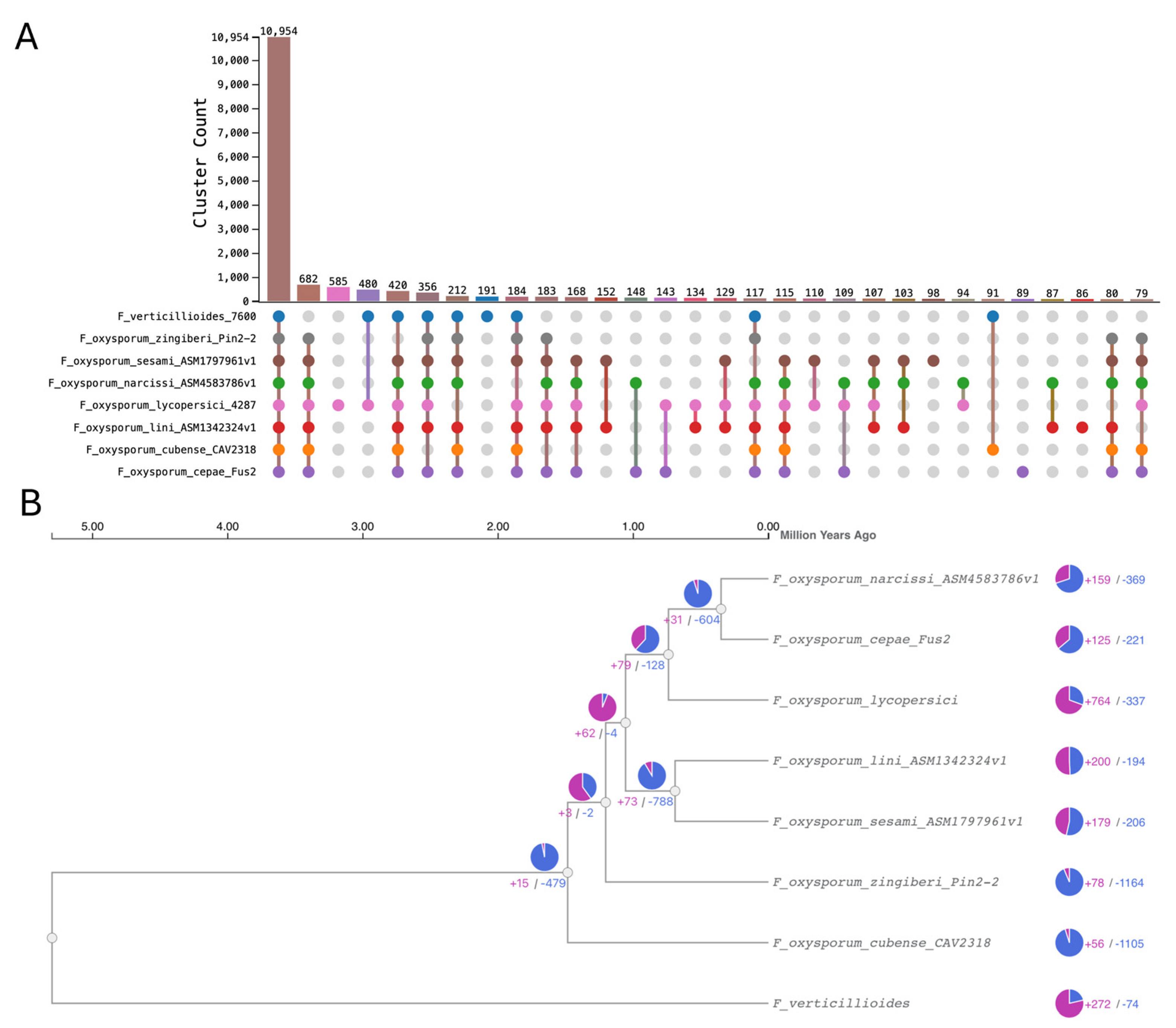
3.7.3. Orthologous Clustering Analysis
3.7.3. Gene Family Expansion and Contraction Analysis
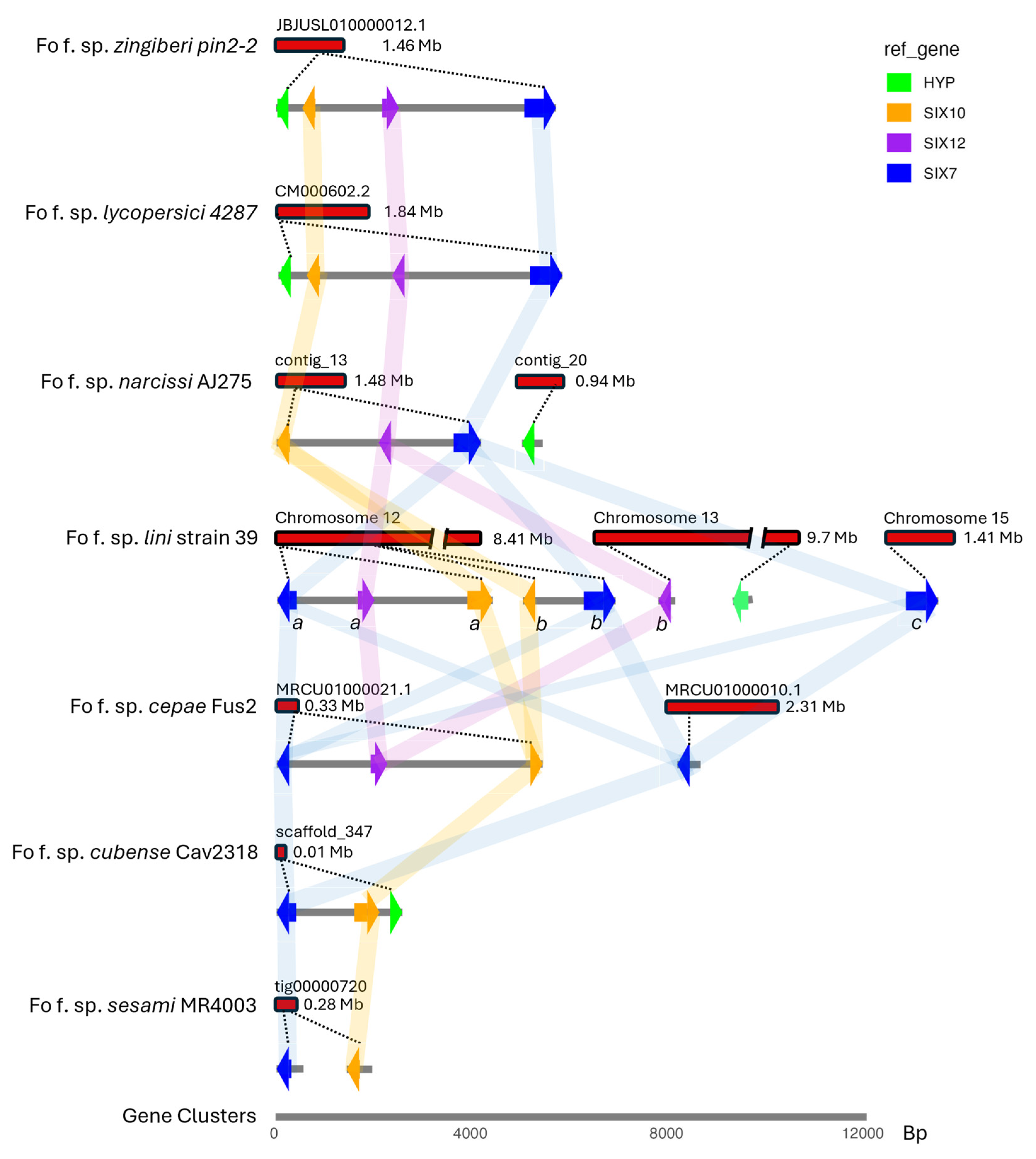
3.7.4. Analysis of the SIX7, SIX10 and SIX12 Virulence Gene Cluster
4. Discussion
4.1. Clonal Nature of the Australian Foz Population
4.2. Endophytic Strains that Colonise Ginger Hosts Asymptomatically
4.3. Role of SIX Gene Effectors in Virulence and Host Specificity
4.4. The Comparative Analysis of the SIX7-SIX10-SIX12 Gene Cluster in Fusarium oxysporum
4.5. Fusarium Foetens Associated with Ginger Hosts
5. Conclusions
6. Patents
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Nair, K.P. The Agronomy and Economy of Ginger. 2013; pp. 225-292.
- Ravindran, P.N.; Babu, K.N.; Shiva, K. Botany and crop improvement of ginger. In Ginger: The Genus Zingiber, Ravindran, P.N., Babu, K.N., Eds.; 2016; pp. 15-85.
- Kizhakkayil, J.; Sasikumar, B. Diversity, characterization and utilization of ginger: a review. Plant genetic resources: characterization and utilization 2011, 9, 464-477. [CrossRef]
- Promdam, N.; Panichayupakaranant, P. [6]-Gingerol: a narrative review of its beneficial effect on human health. Food Chemistry Advances 2022, 1, 100043.
- FAO. FAOSTAT Statistical Database. Available online: https://www.fao.org/faostat/en/ (accessed on 20 October).
- Stirling, A. The causes of poor establishment of ginger (Zingiber officinale) in Queensland, Australia. Australasian Plant Pathology 2004, 33, 203-210.
- Stirling, G.; Turaganivalu, U.; Stirling, A.; Lomavatu, M.; Smith, M. Rhizome rot of ginger (Zingiber officinale) caused by Pythium myriotylum in Fiji and Australia. Australasian Plant Pathology 2009, 38, 453-460.
- Prameela, T.P.; Suseela Bhai, R. Bacterial wilt of ginger (Zingiber officinale Rosc.) incited by Ralstonia pseudosolanacearum-a review based on pathogen diversity, diagnostics and management. Journal of Plant Pathology 2020, 102, 709-719.
- Prasath, D.; Matthews, A.; O'Neill, W.T.; Aitken, E.A.B.; Chen, A. Fusarium Yellows of Ginger (Zingiber officinale Roscoe) Caused by Fusarium oxysporum f. sp. zingiberi Is Associated with Cultivar-Specific Expression of Defense-Responsive Genes. Pathogens 2023, 12. [CrossRef]
- Matthews, A.; Muthukumar, S.P.T.; Hamill, S.; Aitken, E.A.B.; Chen, A. Impact of inoculum density of Fusarium oxysporum f. sp. zingiberi on symptomatic appearances and yield of ginger (Zingiber officinale Roscoe). Access Microbiology 2023, 5. [CrossRef]
- Soesanto, L.; Fakhiroh, Z.; Suharti, W.S. Viability and virulence of Fusarium oxysporum f. sp. zingiberi isolates from Boyolali and temanggung preserved for 17 Years in Sterile Soils. Jurnal Fitopatologi Indonesia 2022, 18, 91-99.
- Chawla, S.; Rafie, R.; Likins, M.; Ren, S.; Ndegwa, E.; Mersha, Z. First Report of Fusarium Yellows and Rhizome Rot Caused by Fusarium oxysporum f. sp. zingiberi on Ginger in the Continental United States. Plant Dis 2021, 105, 3289-3289. [CrossRef]
- Li, Y.; Chi, L.D.; Mao, L.G.; Yan, D.D.; Wu, Z.F.; Ma, T.T.; Guo, M.X.; Wang, Q.X.; Ouyang, C.B.; Cao, A.C. First report of ginger rhizome rot caused by Fusarium oxysporum in China. Plant Dis 2014, 98, 282-282. [CrossRef]
- Waweru, B.; Turoop, L.; Kahangi, E.; Coyne, D.; Dubois, T. Non-pathogenic Fusarium oxysporum endophytes provide field control of nematodes, improving yield of banana (Musa sp.). Biological control 2014, 74, 82-88.
- Constantin, M.E.; De Lamo, F.J.; Vlieger, B.V.; Rep, M.; Takken, F.L. Endophyte-mediated resistance in tomato to Fusarium oxysporum is independent of ET, JA, and SA. Frontiers in plant science 2019, 10, 979.
- Thangavelu, R.; Jayanthi, A. RFLP analysis of rDNA-ITS regions of native non-pathogenic Fusarium oxysporum isolates and their field evaluation for the suppression of Fusarium wilt disease of banana. Australasian Plant Pathology 2009, 38, 13-21.
- Constantin, M.E.; Fokkens, L.; de Sain, M.; Takken, F.L.W.; Rep, M. Number of Candidate Effector Genes in Accessory Genomes Differentiates Pathogenic From Endophytic Fusarium oxysporum Strains. Frontiers in Plant Science 2021, Volume 12 - 2021. [CrossRef]
- Forsyth, L.M.; Smith, L.J.; Aitken, E.A.B. Identification and characterization of non-pathogenic Fusarium oxysporum capable of increasing and decreasing Fusarium wilt severity. Mycological Research 2006, 110, 929-935. [CrossRef]
- Czislowski, E.; Zeil-Rolfe, I.; Aitken, E.A. Effector profiles of endophytic Fusarium associated with asymptomatic banana (Musa sp.) hosts. International Journal of Molecular Sciences 2021, 22, 2508.
- Bubici, G.; Kaushal, M.; Prigigallo, M.I.; Gómez-Lama Cabanás, C.; Mercado-Blanco, J. Biological Control Agents Against Fusarium Wilt of Banana. Frontiers in Microbiology 2019, Volume 10 - 2019. [CrossRef]
- Achari, S.R.; Mann, R.C.; Sharma, M.; Edwards, J. Diagnosis of Fusarium oxysporum f. sp. ciceris causing Fusarium wilt of chickpea using loop-mediated isothermal amplification (LAMP) and conventional end-point PCR. Scientific Reports 2023, 13, 2640. [CrossRef]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. Journal of computational biology 2012, 19, 455-477.
- Ferreira, A.; Glass, V.; Louise, N. PCR from fungal spores after microwave treatment. Fungal Genetics Reports 1996, 43, 25-26.
- O’Donnell, K.; Kistler, H.C.; Cigelnik, E.; Ploetz, R.C. Multiple evolutionary origins of the fungus causing Panama disease of banana: concordant evidence from nuclear and mitochondrial gene genealogies. Proceedings of the National Academy of Sciences 1998, 95, 2044-2049.
- O’Donnell, K.; Rooney, A.P.; Proctor, R.H.; Brown, D.W.; McCormick, S.P.; Ward, T.J.; Frandsen, R.J.; Lysøe, E.; Rehner, S.A.; Aoki, T. Phylogenetic analyses of RPB1 and RPB2 support a middle Cretaceous origin for a clade comprising all agriculturally and medically important fusaria. Fungal Genetics and Biology 2013, 52, 20-31.
- Liu, Y.J.; Whelen, S.; Hall, B.D. Phylogenetic relationships among ascomycetes: evidence from an RNA polymerse II subunit. Molecular biology and evolution 1999, 16, 1799-1808.
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Molecular biology and evolution 2013, 30, 772-780.
- Huelsenbeck, J.P.; Ronquist, F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 2001, 17, 754-755.
- Smith, M.K.; Hamill, S.D. Field evaluation of micropropagated and conventionally propagated ginger in subtropical Queensland. Australian Journal of Experimental Agriculture 1996, 36, 347-354.
- Prasath, D.; Matthews, A.; O’Neill, W.T.; Aitken, E.A.B.; Chen, A. Fusarium Yellows of Ginger (Zingiber officinale Roscoe) Caused by Fusarium oxysporum f. sp. zingiberi Is Associated with Cultivar-Specific Expression of Defense-Responsive Genes. Pathogens 2023, 12, 141.
- Chen, A.; Sun, J.; Matthews, A.; Armas-Egas, L.; Chen, N.; Hamill, S.; Mintoff, S.; Tran-Nguyen, L.T.T.; Batley, J.; Aitken, E.A.B. Assessing Variations in Host Resistance to Fusarium oxysporum f sp. cubense Race 4 in Musa Species, With a Focus on the Subtropical Race 4. Frontiers in Microbiology 2019, Volume 10 - 2019. [CrossRef]
- Manni, M.; Berkeley, M.R.; Seppey, M.; Simão, F.A.; Zdobnov, E.M. BUSCO update: novel and streamlined workflows along with broader and deeper phylogenetic coverage for scoring of eukaryotic, prokaryotic, and viral genomes. Molecular biology and evolution 2021, 38, 4647-4654.
- Stanke, M.; Schöffmann, O.; Morgenstern, B.; Waack, S. Gene prediction in eukaryotes with a generalized hidden Markov model that uses hints from external sources. BMC bioinformatics 2006, 7, 62.
- Flynn, J.M.; Hubley, R.; Goubert, C.; Rosen, J.; Clark, A.G.; Feschotte, C.; Smit, A.F. RepeatModeler2 for automated genomic discovery of transposable element families. Proceedings of the National Academy of Sciences 2020, 117, 9451-9457.
- Chen, N. Using Repeat Masker to identify repetitive elements in genomic sequences. Current protocols in bioinformatics 2004, 5, 4.10. 11-14.10. 14.
- Gabriel, L.; Brůna, T.; Hoff, K.J.; Ebel, M.; Lomsadze, A.; Borodovsky, M.; Stanke, M. BRAKER3: Fully automated genome annotation using RNA-seq and protein evidence with GeneMark-ETP, AUGUSTUS, and TSEBRA. Genome research 2024, 34, 769-777.
- Wang, Y.; Tang, H.; DeBarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.-h.; Jin, H.; Marler, B.; Guo, H. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic acids research 2012, 40, e49-e49.
- Marçais, G.; Delcher, A.L.; Phillippy, A.M.; Coston, R.; Salzberg, S.L.; Zimin, A. MUMmer4: A fast and versatile genome alignment system. PLoS computational biology 2018, 14, e1005944.
- Pertea, G.; Pertea, M. GFF utilities: GffRead and GffCompare. F1000Research 2020, 9, ISCB Comm J-304.
- Sun, J.; Lu, F.; Luo, Y.; Bie, L.; Xu, L.; Wang, Y. OrthoVenn3: an integrated platform for exploring and visualizing orthologous data across genomes. Nucleic acids research 2023, 51, W397-W403.
- Emms, D.M.; Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome biology 2019, 20, 238.
- Kumar, S.; Suleski, M.; Craig, J.M.; Kasprowicz, A.E.; Sanderford, M.; Li, M.; Stecher, G.; Hedges, S.B. TimeTree 5: an expanded resource for species divergence times. Molecular biology and evolution 2022, 39, msac174.
- Mendes, F.K.; Vanderpool, D.; Fulton, B.; Hahn, M.W. CAFE 5 models variation in evolutionary rates among gene families. Bioinformatics 2020, 36, 5516-5518.
- Team, R.C. RA language and environment for statistical computing, R Foundation for Statistical. Computing 2020.
- Van Westerhoven, A.C.; Aguilera-Galvez, C.; Nakasato-Tagami, G.; Shi-Kunne, X.; Martinez de la Parte, E.; Chavarro-Carrero, E.; Meijer, H.J.; Feurtey, A.; Maryani, N.; Ordóñez, N. Segmental duplications drive the evolution of accessory regions in a major crop pathogen. New Phytologist 2024, 242, 610-625.
- Zhou, X.; Everts, K.; Bruton, B. Race 3, a new and highly virulent race of Fusarium oxysporum f. sp. niveum causing Fusarium wilt in watermelon. Plant disease 2010, 94, 92-98.
- Yang, D.; Zhang, X.; Ming, Y.; Liu, C.; Zhang, X.; Liu, S.; Zhu, L. Characterization of the high-quality genome sequence and virulence factors of Fusarium oxysporum f. sp. vasinfectum race 7. Journal of Fungi 2024, 10, 242.
- Le, D.P.; Nguyen, C.P.; Kafle, D.; Scheikowski, L.; Montgomery, J.; Lambeth, E.; Thomas, A.; O’Keeffe, K.; Shakeshaft, B.; Young, A. Surveillance, diversity and vegetative compatibility groups of Fusarium oxysporum f. sp. vasinfectum collected in cotton fields in Australia (2017 to 2022). Pathogens 2022, 11, 1537.
- Dyer, D.R.; Newman, M.; Lawrence, K.S. Diversity and temporal distribution of Fusarium oxysporum f. sp. vasinfectum races and genotypes as influenced by Gossypium cultivar. Frontiers in Fungal Biology 2022, 3, 1022761.
- Ma, L.-J.; van der Does, H.C.; Borkovich, K.A.; Coleman, J.J.; Daboussi, M.-J.; Di Pietro, A.; Dufresne, M.; Freitag, M.; Grabherr, M.; Henrissat, B.; et al. Comparative genomics reveals mobile pathogenicity chromosomes in Fusarium. Nature 2010, 464, 367-373. [CrossRef]
- Gawehns, F.; Ma, L.; Bruning, O.; Houterman, P.M.; Boeren, S.; Cornelissen, B.J.; Rep, M.; Takken, F.L. The effector repertoire of Fusarium oxysporum determines the tomato xylem proteome composition following infection. Front Plant Sci 2015, 6, 967. [CrossRef]
- Gawehns, F.; Houterman, P.M.; Ichou, F.A.; Michielse, C.B.; Hijdra, M.; Cornelissen, B.J.; Rep, M.; Takken, F.L. The Fusarium oxysporum effector Six6 contributes to virulence and suppresses I-2-mediated cell death. Mol Plant Microbe Interact 2014, 27, 336-348. [CrossRef]
- An, B.; Hou, X.; Guo, Y.; Zhao, S.; Luo, H.; He, C.; Wang, Q. The effector SIX8 is required for virulence of Fusarium oxysporum f.sp. cubense tropical race 4 to Cavendish banana. Fungal Biol 2019, 123, 423-430. [CrossRef]
- Houterman, P.M.; Cornelissen, B.J.C.; Rep, M. Suppression of Plant Resistance Gene-Based Immunity by a Fungal Effector. PLOS Pathogens 2008, 4, e1000061. [CrossRef]
- Lievens, B.; Houterman, P.M.; Rep, M. Effector gene screening allows unambiguous identification of Fusarium oxysporum f. sp. lycopersici races and discrimination from other formae speciales. FEMS Microbiol Lett 2009, 300, 201-215. [CrossRef]
- Rep, M.; van der Does, H.C.; Meijer, M.; van Wijk, R.; Houterman, P.M.; Dekker, H.L.; de Koster, C.G.; Cornelissen, B.J. A small, cysteine-rich protein secreted by Fusarium oxysporum during colonization of xylem vessels is required for I-3-mediated resistance in tomato. Mol Microbiol 2004, 53, 1373-1383. [CrossRef]
- Pappalardo, L.; Smith, M.K.; Hamill, S.D.; Stirling, A.M.; McKay, D. DNA amplification fingerprinting analysis of genetic variation within Fusarium oxysporum f.sp. zingiberi. Australasian Plant Pathology 2009, 38, 51-54. [CrossRef]
- Zheng, S.-J.; García-Bastidas, F.A.; Li, X.; Zeng, L.; Bai, T.; Xu, S.; Yin, K.; Li, H.; Fu, G.; Yu, Y.; et al. New Geographical Insights of the Latest Expansion of Fusarium oxysporum f.sp. cubense Tropical Race 4 Into the Greater Mekong Subregion. Frontiers in Plant Science 2018, Volume 9 - 2018. [CrossRef]
- Schulz, B.; Boyle, C. The endophytic continuum. Mycological research 2005, 109, 661-686.
- Rodriguez, R.J.; White Jr, J.; Arnold, A.E.; Redman, a.R.a. Fungal endophytes: diversity and functional roles. New phytologist 2009, 182, 314-330.
- Hardoim, P.R.; Van Overbeek, L.S.; Berg, G.; Pirttilä, A.M.; Compant, S.; Campisano, A.; Döring, M.; Sessitsch, A. The hidden world within plants: ecological and evolutionary considerations for defining functioning of microbial endophytes. Microbiology and molecular biology reviews 2015, 79, 293-320.
- Yates, I.; Bacon, C.; Hinton, D. Effects of endophytic infection by Fusarium moniliforme on corn growth and cellular morphology. Plant disease 1997, 81, 723-728.
- Mousa, W.K.; Raizada, M.N. The diversity of anti-microbial secondary metabolites produced by fungal endophytes: an interdisciplinary perspective. Frontiers in microbiology 2013, 4, 65.
- Houterman, P.M.; Ma, L.; Van Ooijen, G.; De Vroomen, M.J.; Cornelissen, B.J.C.; Takken, F.L.W.; Rep, M. The effector protein Avr2 of the xylem-colonizing fungus Fusarium oxysporum activates the tomato resistance protein I-2 intracellularly. The Plant Journal 2009, 58, 970-978. [CrossRef]
- Simbaqueba, J.; Catanzariti, A.-M.; González, C.; Jones, D.A. Evidence for horizontal gene transfer and separation of effector recognition from effector function revealed by analysis of effector genes shared between cape gooseberry- and tomato-infecting formae speciales of Fusarium oxysporum. Molecular Plant Pathology 2018, 19, 2302-2318. [CrossRef]
- Czislowski, E.; Fraser-Smith, S.; Zander, M.; O'Neill, W.T.; Meldrum, R.A.; Tran-Nguyen, L.T.T.; Batley, J.; Aitken, E.A.B. Investigation of the diversity of effector genes in the banana pathogen, Fusarium oxysporum f. sp. cubense, reveals evidence of horizontal gene transfer. Mol Plant Pathol 2018, 19, 1155-1171. [CrossRef]
- Seong, K.; Krasileva, K.V. Prediction of effector protein structures from fungal phytopathogens enables evolutionary analyses. Nat Microbiol 2023, 8, 174-187. [CrossRef]
- Li, J.; Fokkens, L.; Conneely, L.J.; Rep, M. Partial pathogenicity chromosomes in Fusarium oxysporum are sufficient to cause disease and can be horizontally transferred. Environ Microbiol 2020, 22, 4985-5004. [CrossRef]
- Simbaqueba, J.; Rodríguez, E.A.; Burbano-David, D.; González, C.; Caro-Quintero, A. Putative Novel Effector Genes Revealed by the Genomic Analysis of the Phytopathogenic Fungus Fusarium oxysporum f. sp. physali (Foph) That Infects Cape Gooseberry Plants. Frontiers in Microbiology 2021, Volume 11 - 2020. [CrossRef]
- Yang, W.; Ma, T.; Liang, D.; Zhang, C. Involvement of the SIX10 Gene in the Pathogenicity of Fusarium oxysporum Formae Speciales in Strawberries. International Journal of Molecular Sciences 2025, 26, 1123.
- Sun, J.; Zhang, J.; Gardiner, D.M.; van Dam, P.; Fu, G.; Ferguson, B.J.; Aitken, E.A.; Chen, A. Long-read draft genome sequences of two Fusarium oxysporum f. sp. cubense isolates from banana (Musa spp.). Journal of Fungi 2025, 11, 421.
- van Dam, P.; Fokkens, L.; Schmidt, S.M.; Linmans, J.H.; Kistler, H.C.; Ma, L.J.; Rep, M. Effector profiles distinguish formae speciales of Fusarium oxysporum. Environmental Microbiology 2016, 18, 4087-4102.
- De Sain, M.; Rep, M. The role of pathogen-secreted proteins in fungal vascular wilt diseases. International Journal of Molecular Sciences 2015, 16, 23970-23993.
- van Dam, P.; Rep, M. The Distribution of Miniature Impala Elements and SIX Genes in the Fusarium Genus is Suggestive of Horizontal Gene Transfer. J Mol Evol 2017, 85, 14-25. [CrossRef]
- Croll, D.; McDonald, B.A. The accessory genome as a cradle for adaptive evolution in pathogens. PLoS pathogens 2012, 8, e1002608.
- Dong, S.; Raffaele, S.; Kamoun, S. The two-speed genomes of filamentous pathogens: waltz with plants. Current opinion in genetics & development 2015, 35, 57-65.
- Schroers, H.-J.; Baayen, R.; Meffert, J.; De Gruyter, J.; Hooftman, M.; O’Donnell, K. Fusarium foetens, a new species pathogenic to begonia elatior hybrids (Begonia× hiemalis) and the sister taxon of the Fusarium oxysporum species complex. Mycologia 2004, 96, 393-406.
- Saurat, C.; Fourrier, C.; Wilson, V.; Casset, C.; Ioos, R. First report of begonia elatior wilt disease caused by Fusarium foetens in France. Plant Disease 2013, 97, 144-144.
- Elmer, W.; Vossbrinck, C.; Geiser, D. First report of a wilt disease of Hiemalis begonias caused by Fusarium foetens in the United States. Plant Disease 2004, 88, 1287-1287.
- Tian, X.; Dixon, M.; Zheng, Y. First report of Hiemalis begonias wilt disease caused by Fusarium foetens in Canada. Plant Disease 2010, 94, 1261-1261.
- Lamprecht, S.; Tewoldemedhin, Y. Fusarium species associated with damping-off of rooibos seedlings and the potential of compost as soil amendment for disease suppression. South African Journal of Botany 2017, 110, 110-117.
- Liu, L.; Jin, X.; Lu, X.; Guo, L.; Lu, P.; Yu, H.; Lv, B. Mechanisms of surfactin from Bacillus subtilis SF1 against Fusarium foetens: a novel pathogen inducing potato wilt. Journal of Fungi 2023, 9, 367.
- Yi, B.; Ma, J.; Luo, L.; Ghani, M.I.; Siddique, J.A.; Tang, X.; Cernava, T.; Chen, X. First report of Fusarium foetens causing tobacco root and stem rots in Guizhou, China. Journal of Phytopathology 2025, 173, e70104.
- Wei, X.-J.; Jia, B.-G.; Wang, X.; Li, N.-Y.; Wang, S.-N.; Li, L.; Zhang, A.; Zhang, H.-T.; Wang, L.-P. First report of Fusarium foetens causing root rot on lavender (Lavandula angustifolia) in China. Journal of Plant Pathology 2023, 105, 1173-1174.
- Shang, M.; Zhang, X.; Zhang, X.; Yan, J.; Liu, Q.; Zhai, H.; Gao, S.; Zhao, N.; He, S.; Zhang, H. A chromosome-level genome assembly of Fusarium foetens that causes sweet potato root rot facilitates the identification of a key Fusarium-specific pathogenicity factor. Plant Communications 2025, 6.
- Ono, L.; Silva, J.; Soto, T.; Doná, S.; Iamanaka, B.; Fungaro, M.; Taniwaki, M. Fungal communities in Brazilian cassava tubers and food products. International journal of food microbiology 2023, 384, 109909.
- Summerell, B.A.; Leslie, J.F.; Liew, E.C.; Laurence, M.H.; Bullock, S.; Petrovic, T.; Bentley, A.R.; Howard, C.G.; Peterson, S.A.; Walsh, J.L. Fusarium species associated with plants in Australia. Fungal Diversity 2011, 46, 1-27.
- Laurence, M.H.; Burgess, L.W.; Summerell, B.A.; Liew, E.C. High levels of diversity in Fusarium oxysporum from non-cultivated ecosystems in Australia. Fungal biology 2012, 116, 289-297.



Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.



