Submitted:
10 April 2026
Posted:
13 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
1.1. Overview of Celiac Disease
1.2. Current Diagnostic Pathway and Its Limitations
1.3. Immunohistochemistry and In-Situ Hybridization
1.4. Candidate Diagnostic Biomarkers can Be DERIVED from an understanding of CD Pathogenesis
1.5. Aim of the Study
2. Materials and Methods
2.1. Biopsy Selection
2.2. Immunohistochemistry
2.3. Image Scanning and Evaluation
2.4. Staining Analysis
2.5. Statistical Analysis and Data Visualization
3. Results
3.1. Analysis of Biomarker Immunostaining
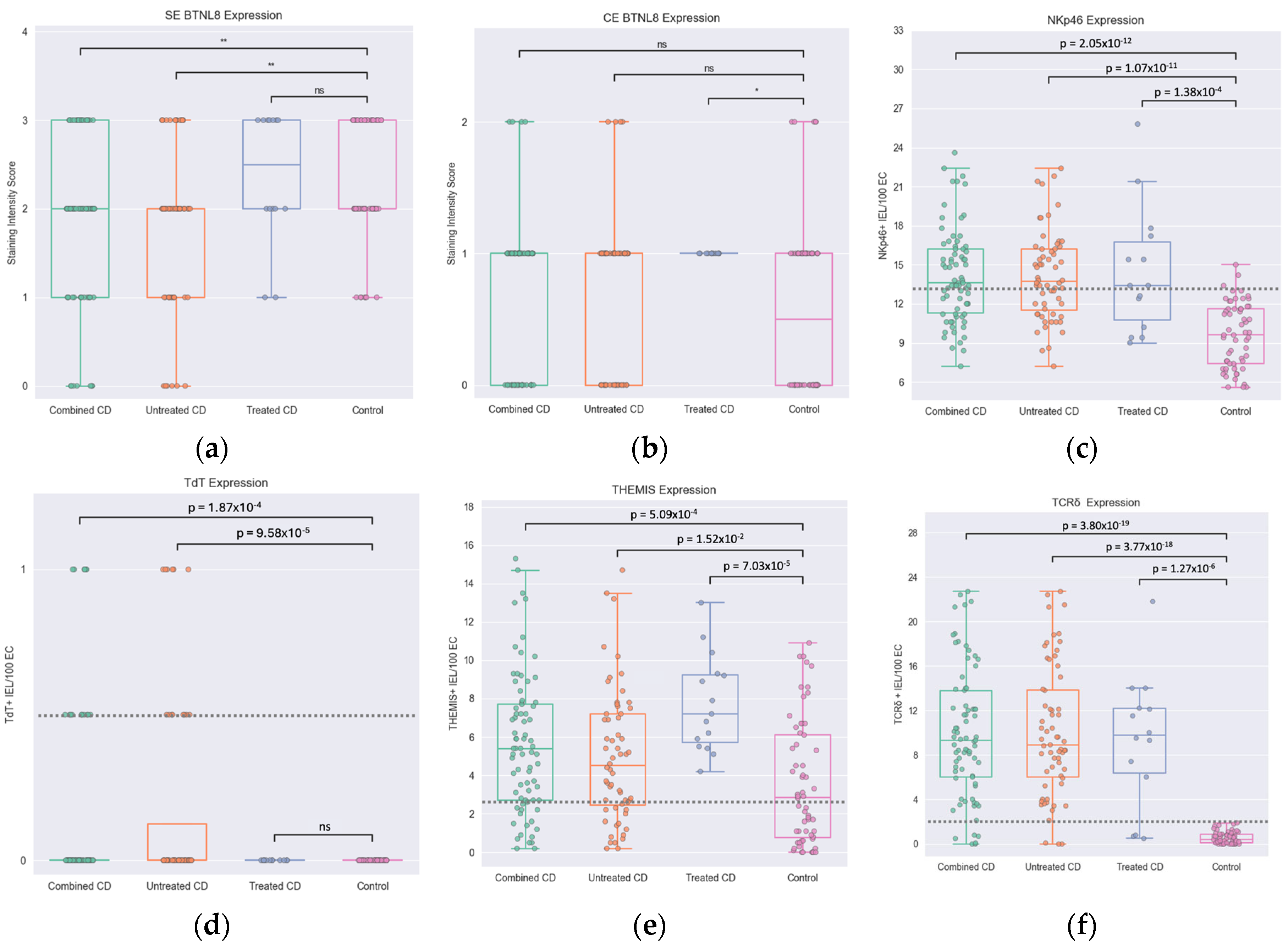
3.2. Statistical Evaluation of Immunostained Markers
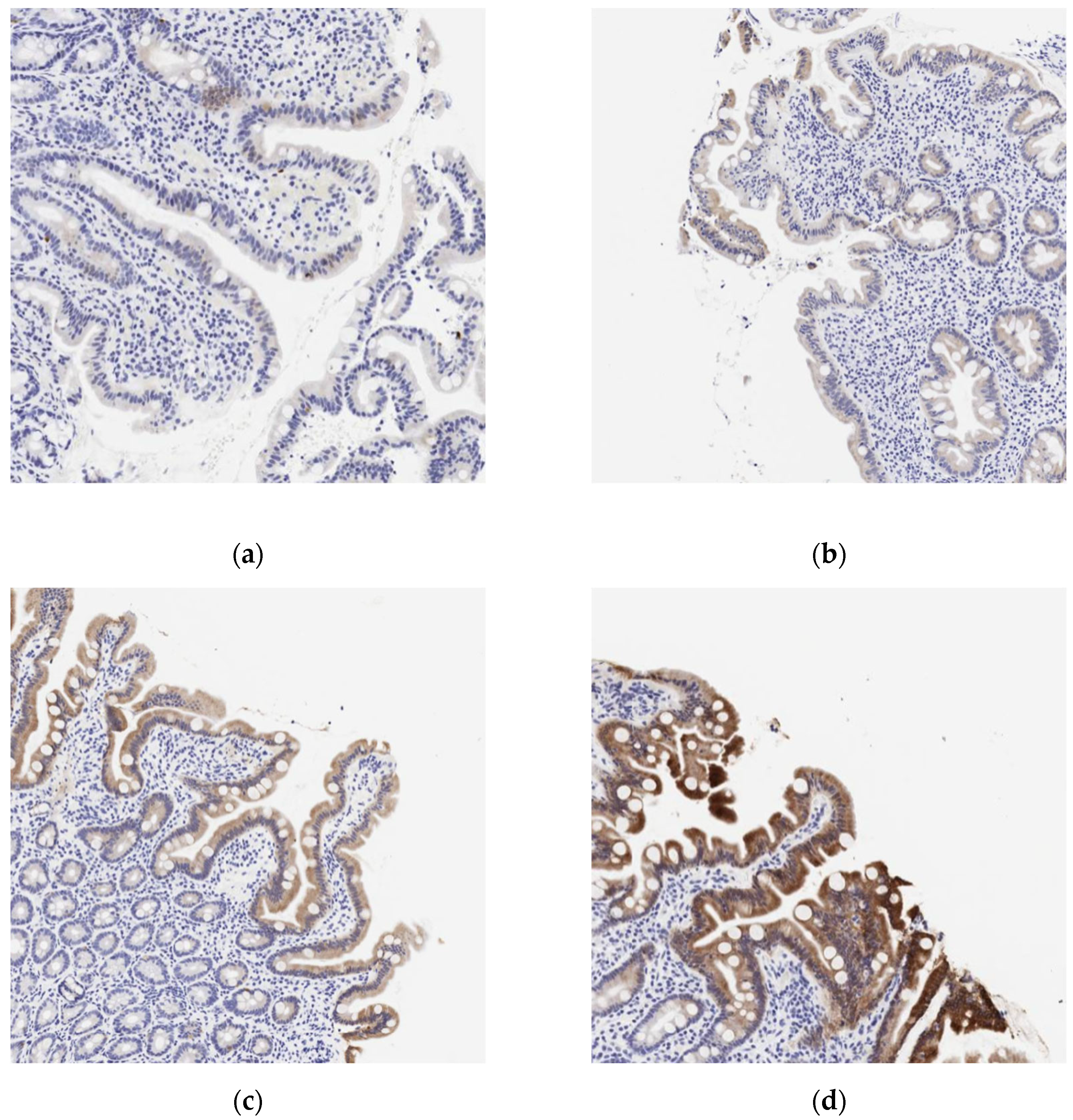
3.2.1. Statistical Evaluation of BTNL8 Immunostaining Results
3.2.2. Statistical Evaluation of NKp46 Immunostaining Results
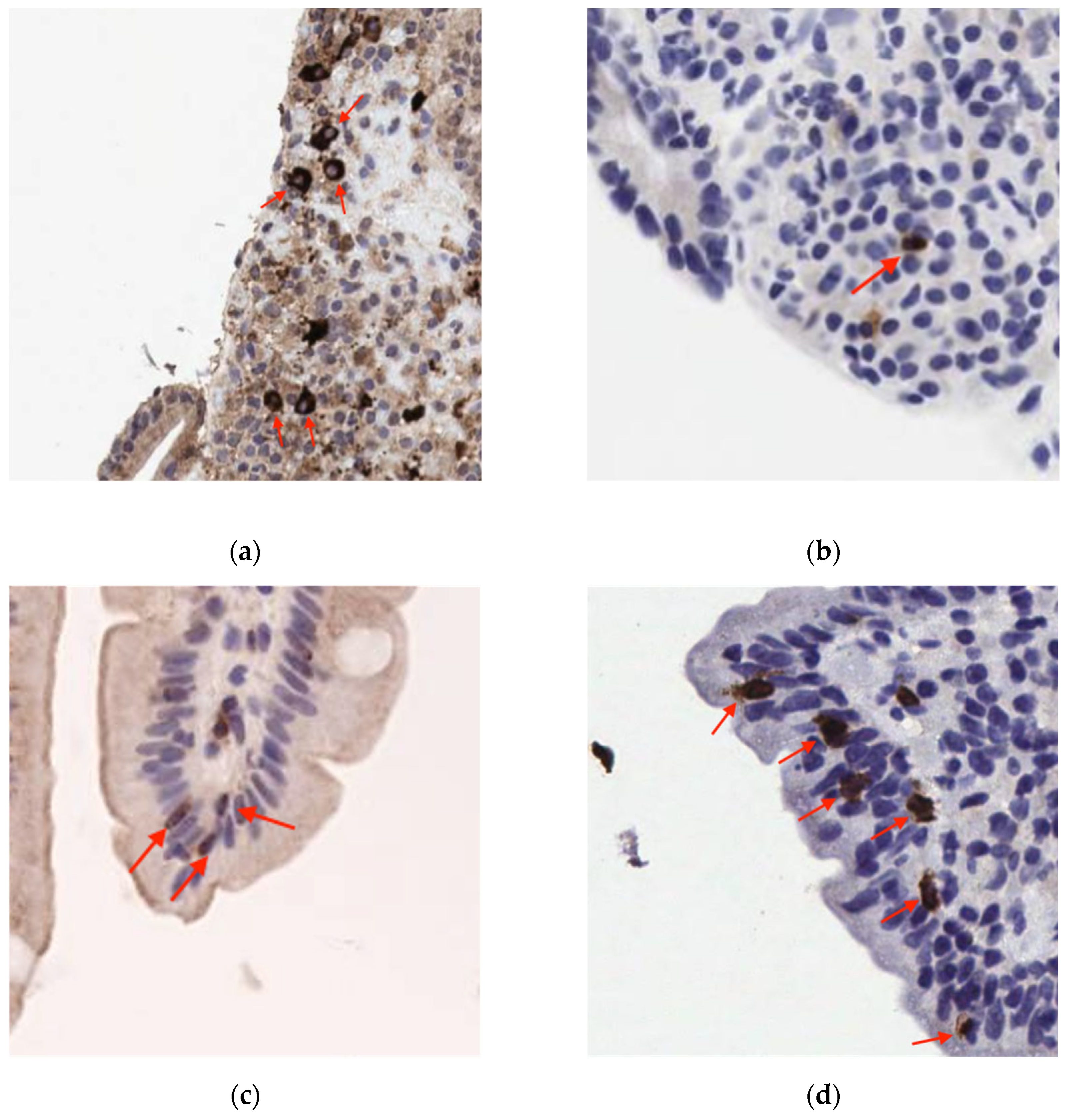
3.2.3. Statistical Evaluation of TdT Immunostaining Results
3.2.4. Statistical Evaluation of THEMIS Immunostaining Results
3.2.5. Statistical Evaluation of TCRδ Immunostaining Results
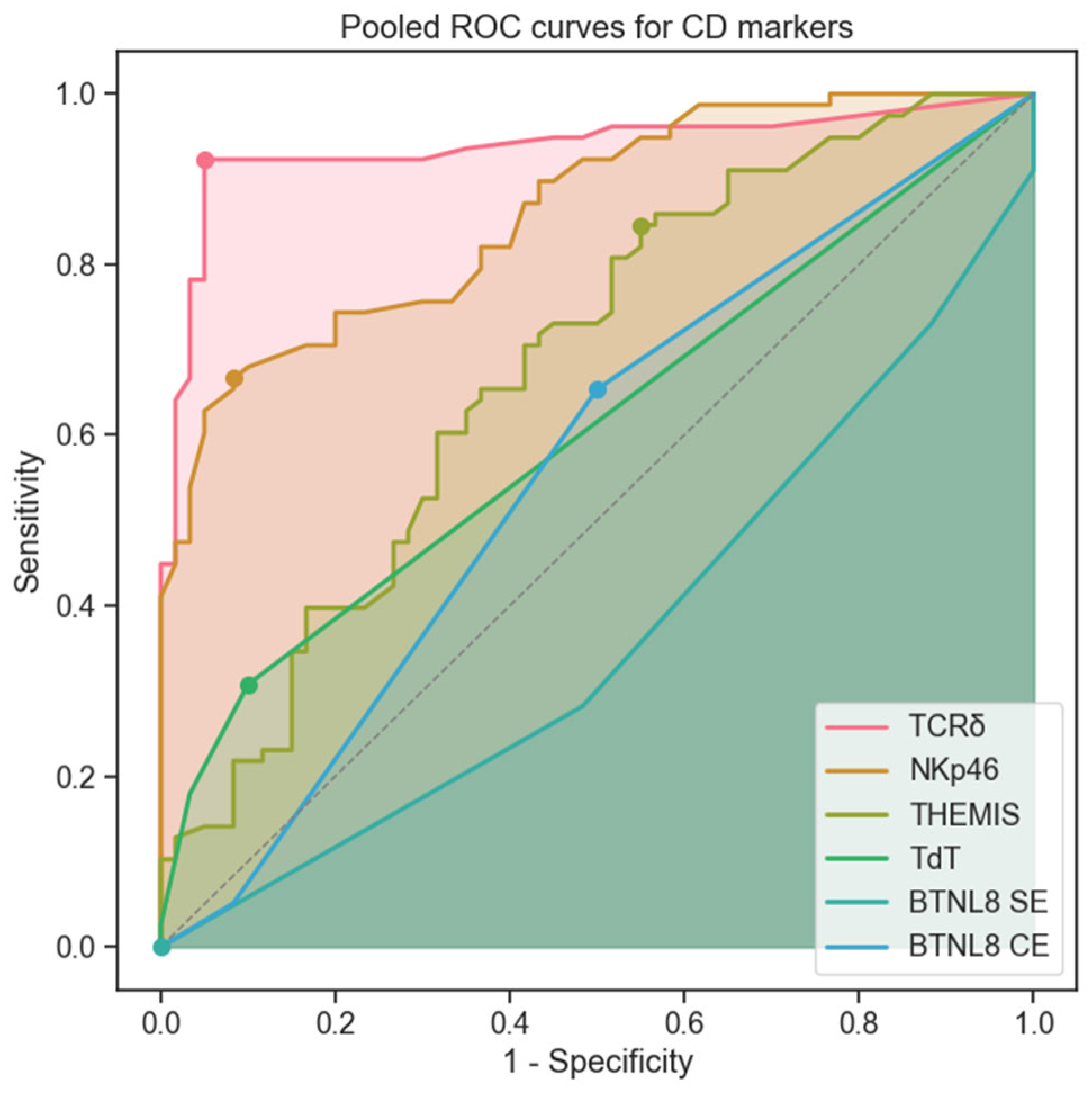
3.3. Sensitivity Analysis and Diagnostic Performance
4. Discussion
4.1. Overall Biomarker Utility
4.2. Utility of TCRδ Immunostaining
4.3. Utility of NKp46 Immunostaining
4.4. Utility of BTNL8 Immunostaining
4.5. Utility of THEMIS Immunostaining
4.6. Utility of TdT Immunostaining
4.7. Study Limitations and Future Work
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AI | Artificial intelligence |
| AUC | Area under the ROC curve |
| BTNL | Butyrophilin-like |
| CD | Celiac disease |
| CE | Crypt epithelium |
| GFD | Gluten-free diet |
| HE | Hematoxylin and eosin |
| ICC | Intraclass coefficient |
| IEL | Intraepithelial lymphocytes |
| IHC | Immunohistochemistry |
| IQR | Interquartile range |
| NCR | Natural cytotoxicity receptors |
| RAG | Recombination-activating genes |
| ROC | Receiver operating characteristic |
| SE | Surface epithelium |
| TCR | T cell receptor |
| TdT | Terminal deoxynucleotidyl transferase |
| THEMIS | Thymocyte-expressed molecule involved in selection |
| tTG | Tissue transglutaminase |
| γδ | Gamma-delta |
Appendix A
| Patient | CD group | Observer 1 | Observer 2 | Observer 3 2 |
|---|---|---|---|---|
| 1 | CD untreated | 2 | 2 | 1 |
| 2 | CD untreated | 2 | 2 | 2 |
| 3 | CD untreated | 1 | 2 | 1 |
| 4 | CD untreated | 2 | 3 | 2 |
| 5 | CD untreated | 1 | 1 | 1 |
| 6 | CD untreated | 2 | 3 | 2 |
| 7 | CD untreated | 3 | 2 | 2 |
| 8 | CD untreated | 2 | 2 | 2 |
| 9 | CD untreated | 3 | 3 | 3 |
| 10 | CD untreated | 2 | 2 | 2 |
| 11 | CD untreated | 0 | 0 | 0 |
| 12 | CD untreated | 3 | 3 | 2 |
| 13 | CD untreated | 2 | 3 | 2 |
| 14 | CD untreated | 2 | 3 | 1 |
| 15 | CD untreated | 2 | 3 | 1 |
| 16 | CD untreated | 1 | 2 | 1 |
| 17 | CD untreated | 1 | 2 | 1 |
| 18 | CD untreated | 2 | 2 | 2 |
| 19 | CD untreated | 3 | 3 | 2 |
| 20 | CD untreated | 3 | 3 | 2 |
| 21 | CD untreated | 0 | 0 | 0 |
| 22 | CD GFD treated | 3 | 3 | 2 |
| 23 | CD GFD treated | 3 | 3 | 2 |
| 24 | CD GFD treated | 0 | 1 | 1 |
| 25 | CD GFD treated | 2 | 3 | 2 |
| 26 | CD GFD treated | 3 | 3 | 2 |
| 27 | Controls | 3 | 3 | 3 |
| 28 | Controls | 3 | 3 | 2 |
| 29 | Controls | 2 | 2 | 1 |
| 30 | Controls | 2 | 3 | 3 |
| 31 | Controls | 2 | 2 | 1 |
| 32 | Controls | 3 | 3 | 3 |
| 33 | Controls | 3 | 3 | 2 |
| 34 | Controls | 3 | 3 | 3 |
| 35 | Controls | 2 | 2 | 1 |
| 36 | Controls | 2 | 2 | 1 |
| 37 | Controls | 2 | 2 | 2 |
| 38 | Controls | 2 | 3 | 2 |
| 39 | Controls | 2 | 2 | 1 |
| 40 | Controls | 3 | 3 | 2 |
| 41 | Controls | 3 | 3 | 3 |
| 42 | Controls | 3 | 3 | 2 |
| 43 | Controls | 3 | 3 | 2 |
| 44 | Controls | 2 | 3 | 1 |
| 45 | Controls | 3 | 3 | 3 |
| 46 | Controls | 2 | 2 | 1 |
| Patient | CD group | Observer 1 | Observer 2 | Observer 3 2 |
|---|---|---|---|---|
| 1 | CD untreated | 1 | 1 | 0 |
| 2 | CD untreated | 0 | 0 | 0 |
| 3 | CD untreated | 0 | 0 | 0 |
| 4 | CD untreated | 1 | 1 | 1 |
| 5 | CD untreated | 1 | 1 | 1 |
| 6 | CD untreated | 2 | 2 | 1 |
| 7 | CD untreated | 0 | 1 | 1 |
| 8 | CD untreated | 0 | 0 | 1 |
| 9 | CD untreated | 1 | 1 | 1 |
| 10 | CD untreated | 0 | 1 | 1 |
| 11 | CD untreated | 0 | 1 | 0 |
| 12 | CD untreated | 1 | 1 | 1 |
| 13 | CD untreated | 0 | 1 | 0 |
| 14 | CD untreated | 0 | 1 | 0 |
| 15 | CD untreated | 1 | 1 | 1 |
| 16 | CD untreated | 0 | 1 | 0 |
| 17 | CD untreated | 1 | 2 | 1 |
| 18 | CD untreated | 1 | 1 | 1 |
| 19 | CD untreated | 0 | 0 | 1 |
| 20 | CD untreated | 1 | 2 | 1 |
| 21 | CD untreated | 0 | 0 | 0 |
| 22 | CD GFD treated | 1 | 0 | 1 |
| 23 | CD GFD treated | 1 | 0 | 1 |
| 24 | CD GFD treated | 1 | 1 | 1 |
| 25 | CD GFD treated | 0 | 1 | 1 |
| 26 | CD GFD treated | 1 | 1 | 1 |
| 27 | Controls | 1 | 1 | 1 |
| 28 | Controls | 1 | 1 | 1 |
| 29 | Controls | 0 | 1 | 0 |
| 30 | Controls | 2 | 2 | 1 |
| 31 | Controls | 0 | 0 | 0 |
| 32 | Controls | 1 | 1 | 0 |
| 33 | Controls | 0 | 1 | 0 |
| 34 | Controls | 2 | 2 | 2 |
| 35 | Controls | 0 | 0 | 0 |
| 36 | Controls | 1 | 0 | 1 |
| 37 | Controls | 0 | 0 | 0 |
| 38 | Controls | 0 | 0 | 0 |
| 39 | Controls | 1 | 1 | 1 |
| 40 | Controls | 0 | 0 | 1 |
| 41 | Controls | 0 | 1 | 0 |
| 42 | Controls | 0 | 1 | 1 |
| 43 | Controls | 1 | 1 | 0 |
| 44 | Controls | 0 | 0 | 0 |
| 45 | Controls | 1 | 1 | 1 |
| 46 | Controls | 0 | 0 | 0 |
| Patient | CD group | Observer 1 | Observer 2 | Observer 3 2 |
|---|---|---|---|---|
| 1 | CD untreated | 21.8 | 16.4 | 19.6 |
| 2 | CD untreated | 11.6 | 11 | 11.2 |
| 3 | CD untreated | 12 | 12.4 | 9.8 |
| 4 | CD untreated | 13.6 | 13.2 | 12 |
| 5 | CD untreated | 16.8 | 15.4 | 15 |
| 6 | CD untreated | 15.6 | 13 | 21.2 |
| 7 | CD untreated | 15.2 | 12.8 | 17.2 |
| 8 | CD untreated | 14.8 | 15.4 | 15 |
| 9 | CD untreated | 23.6 | 15.8 | 18.8 |
| 10 | CD untreated | 16.2 | 13.4 | 13.2 |
| 11 | CD untreated | 15 | 10.6 | 13.4 |
| 12 | CD untreated | 13.4 | 10.6 | 14 |
| 13 | CD untreated | 18.6 | 13.8 | 11 |
| 14 | CD untreated | 22.4 | 9.8 | 13.8 |
| 15 | CD untreated | 36.6 | 35.6 | 16.8 |
| 16 | CD untreated | 16.4 | 12 | 8.4 |
| 17 | CD untreated | 18.6 | 16.2 | 14.8 |
| 18 | CD untreated | 16.6 | 10.6 | 7.2 |
| 19 | CD untreated | 10.6 | 8.6 | 11.2 |
| 20 | CD untreated | 13.6 | 10.2 | 11.2 |
| 21 | CD untreated | 21.4 | 16 | 13 |
| 22 | CD GFD treated | 35.6 | 9.4 | 15.4 |
| 23 | CD GFD treated | 21.4 | 9.4 | 12.4 |
| 24 | CD GFD treated | 12.6 | 9 | 13.4 |
| 25 | CD GFD treated | 25.8 | 17.8 | 13.4 |
| 26 | CD GFD treated | 15.4 | 10.2 | 17.2 |
| 27 | Controls | 6.8 | 5.8 | 13 |
| 28 | Controls | 9.4 | 6.6 | 11.6 |
| 29 | Controls | 14.2 | 12.2 | 12.2 |
| 30 | Controls | 6.2 | 9.2 | 10.6 |
| 31 | Controls | 9.2 | 12.4 | 15 |
| 32 | Controls | 10.8 | 10.8 | 12.4 |
| 33 | Controls | 9.6 | 11.4 | 11.4 |
| 34 | Controls | 7.4 | 13.4 | 11.2 |
| 35 | Controls | 7 | 11.2 | 6.6 |
| 36 | Controls | 12.4 | 8 | 10.4 |
| 37 | Controls | 7 | 7.6 | 13 |
| 38 | Controls | 5.6 | 8.4 | 9.8 |
| 39 | Controls | 7 | 12.6 | 8.6 |
| 40 | Controls | 7.8 | 11.6 | 11.6 |
| 41 | Controls | 8.2 | 9.6 | 7.6 |
| 42 | Controls | 12.4 | 7.6 | 5.6 |
| 43 | Controls | 11.8 | 10 | 11.8 |
| 44 | Controls | 5.6 | 9.8 | 7.4 |
| 45 | Controls | 6.4 | 11.6 | 7 |
| 46 | Controls | 7.6 | 9.4 | 6.6 |
| Patient | CD group | Observer 1 | Observer 2 | Observer 3 2 |
|---|---|---|---|---|
| 1 | CD untreated | 0 | 0 | 0 |
| 2 | CD untreated | 1 | 1 | 0.5 |
| 3 | CD untreated | 0 | 0 | 0.5 |
| 4 | CD untreated | 0 | 0 | 0.5 |
| 5 | CD untreated | 0 | 0 | 1.5 |
| 6 | CD untreated | 1.5 | 1.5 | 1 |
| 7 | CD untreated | 0 | 0 | 0.5 |
| 8 | CD untreated | 0 | 0 | 0 |
| 9 | CD untreated | 0 | 0 | 2.5 |
| 10 | CD untreated | 0 | 0 | 1.5 |
| 11 | CD untreated | 0 | 0 | 0 |
| 12 | CD untreated | 0 | 0 | 0 |
| 13 | CD untreated | 1 | 1 | 1.5 |
| 14 | CD untreated | 0 | 0 | 0.5 |
| 15 | CD untreated | 0 | 0 | 0 |
| 16 | CD untreated | 0 | 0 | 0 |
| 17 | CD untreated | 0 | 0 | 0.5 |
| 18 | CD untreated | 0 | 0 | 1 |
| 19 | CD untreated | 0 | 0 | 0.5 |
| 20 | CD untreated | 0 | 0 | 1.5 |
| 21 | CD untreated | 0 | 0 | 0.5 |
| 22 | CD GFD treated | 0 | 0 | 0 |
| 23 | CD GFD treated | 0 | 0 | 0.5 |
| 24 | CD GFD treated | 0 | 0 | 2.5 |
| 25 | CD GFD treated | 0 | 0 | 0 |
| 26 | CD GFD treated | 0 | 0 | 0.5 |
| 27 | Controls | 0 | 0 | 0 |
| 28 | Controls | 0 | 0 | 0 |
| 29 | Controls | 0 | 0 | 0 |
| 30 | Controls | 0 | 0 | 0 |
| 31 | Controls | 0 | 0 | 0 |
| 32 | Controls | 0 | 0 | 0.5 |
| 33 | Controls | 0 | 0 | 0 |
| 34 | Controls | 0 | 0 | 0 |
| 35 | Controls | 0 | 0 | 0 |
| 36 | Controls | 0 | 0 | 0 |
| 37 | Controls | 0 | 0 | 0 |
| 38 | Controls | 0 | 0 | 0 |
| 39 | Controls | 0 | 0 | 1.5 |
| 40 | Controls | 0.5 | 0.5 | 0 |
| 41 | Controls | 0 | 0 | 0 |
| 42 | Controls | 0 | 0 | 0 |
| 43 | Controls | 0 | 0 | 0 |
| 44 | Controls | 0 | 0 | 1 |
| 45 | Controls | 0 | 0 | 0 |
| 46 | Controls | 0 | 0 | 0.5 |
| Patient | CD group | Observer 1 | Observer 2 | Observer 3 2 |
|---|---|---|---|---|
| 1 | CD untreated | 2.7 | 3.1 | 1.2 |
| 2 | CD untreated | 0.7 | 1.4 | 0.2 |
| 3 | CD untreated | 2.3 | 4.3 | 0.2 |
| 4 | CD untreated | 3.4 | 6.1 | 0.9 |
| 5 | CD untreated | 7 | 13.5 | 4.9 |
| 6 | CD untreated | 20 | 26.5 | 10.7 |
| 7 | CD untreated | 1.5 | 2.7 | 2.5 |
| 8 | CD untreated | 14.7 | 15.3 | 7.8 |
| 9 | CD untreated | 7.8 | 10.2 | 3.6 |
| 10 | CD untreated | 9.3 | 6.9 | 2 |
| 11 | CD untreated | 5.1 | 4.4 | 2.8 |
| 12 | CD untreated | 5.4 | 7.6 | 0.8 |
| 13 | CD untreated | 5.9 | 5.1 | 2.6 |
| 14 | CD untreated | 7.7 | 5.9 | 3.2 |
| 15 | CD untreated | 8.4 | 5.4 | 2.2 |
| 16 | CD untreated | 8.9 | 9.1 | 1.6 |
| 17 | CD untreated | 4.6 | 3.5 | 5.8 |
| 18 | CD untreated | 0.5 | 4.1 | 0.5 |
| 19 | CD untreated | 5.2 | 3.7 | 2.2 |
| 20 | CD untreated | 7.2 | 13.2 | 2.7 |
| 21 | CD untreated | 6.9 | 7.5 | 7.2 |
| 22 | CD GFD treated | 6.8 | 9.2 | 4.2 |
| 23 | CD GFD treated | 5.9 | 5.5 | 5.1 |
| 24 | CD GFD treated | 9.3 | 6.2 | 7.9 |
| 25 | CD GFD treated | 7.2 | 8.9 | 5.4 |
| 26 | CD GFD treated | 11.2 | 13 | 10.4 |
| 27 | Controls | 10.2 | 9.7 | 3 |
| 28 | Controls | 8.6 | 8.6 | 2.8 |
| 29 | Controls | 4.2 | 5.3 | 0.7 |
| 30 | Controls | 2.3 | 1.1 | 0.3 |
| 31 | Controls | 6.7 | 6.7 | 2.9 |
| 32 | Controls | 5.4 | 3.9 | 1.9 |
| 33 | Controls | 6.2 | 6.1 | 3.9 |
| 34 | Controls | 9.9 | 10.2 | 3.1 |
| 35 | Controls | 4.5 | 4.5 | 1.7 |
| 36 | Controls | 4 | 2.9 | 5.6 |
| 37 | Controls | 6.7 | 6.5 | 8.3 |
| 38 | Controls | 0.5 | 0.6 | 1.1 |
| 39 | Controls | 8.1 | 7.1 | 10.9 |
| 40 | Controls | 1.6 | 1.8 | 3.3 |
| 41 | Controls | 0 | 0 | 0 |
| 42 | Controls | 1.1 | 0.8 | 0.9 |
| 43 | Controls | 0.6 | 0.5 | 1.7 |
| 44 | Controls | 0.2 | 0 | 0.1 |
| 45 | Controls | 0 | 0 | 0.2 |
| 46 | Controls | 2.4 | 2.1 | 1.1 |
| Patient | CD group | Observer 1 | Observer 2 | Observer 3 2 |
|---|---|---|---|---|
| 1 | CD untreated | 18.2 | 11 | 17.8 |
| 2 | CD untreated | 6.7 | 5.2 | 3.4 |
| 3 | CD untreated | 12 | 9.2 | 6.1 |
| 4 | CD untreated | 13.8 | 10.1 | 7.7 |
| 5 | CD untreated | 3.7 | 4 | 3.5 |
| 6 | CD untreated | 9.4 | 5.9 | 6.5 |
| 7 | CD untreated | 11.6 | 8.4 | 8.1 |
| 8 | CD untreated | 22.7 | 21.5 | 12.1 |
| 9 | CD untreated | 18.1 | 12.1 | 18.9 |
| 10 | CD untreated | 7.7 | 3.4 | 3.6 |
| 11 | CD untreated | 22.4 | 21.3 | 17.4 |
| 12 | CD untreated | 7.3 | 5.4 | 4 |
| 13 | CD untreated | 18.8 | 8.2 | 9.6 |
| 14 | CD untreated | 0 | 0 | 0.1 |
| 15 | CD untreated | 15 | 8.4 | 10.4 |
| 16 | CD untreated | 16 | 11.6 | 9.6 |
| 17 | CD untreated | 13.9 | 12.4 | 8.2 |
| 18 | CD untreated | 16.7 | 8.5 | 8.9 |
| 19 | CD untreated | 16.6 | 10.4 | 6.9 |
| 20 | CD untreated | 3.8 | 3 | 2.1 |
| 21 | CD untreated | 16.9 | 8.3 | 8.8 |
| 22 | CD GFD treated | 26.1 | 10 | 14 |
| 23 | CD GFD treated | 12.2 | 9.5 | 7.4 |
| 24 | CD GFD treated | 21.8 | 14 | 11.5 |
| 25 | CD GFD treated | 12.1 | 6 | 9.3 |
| 26 | CD GFD treated | 0.7 | 0.5 | 0.8 |
| 27 | Controls | 0 | 0 | 0 |
| 28 | Controls | 1.1 | 1.6 | 1.3 |
| 29 | Controls | 0 | 0 | 0.2 |
| 30 | Controls | 0.3 | 0 | 0.1 |
| 31 | Controls | 0 | 0.1 | 0.2 |
| 32 | Controls | 0.2 | 0 | 0.1 |
| 33 | Controls | 1.1 | 1.1 | 1.8 |
| 34 | Controls | 7.7 | 5.3 | 9.8 |
| 35 | Controls | 0.8 | 0.9 | 0.5 |
| 36 | Controls | 0.7 | 0.7 | 0.7 |
| 37 | Controls | 1.9 | 0.7 | 0.8 |
| 38 | Controls | 1.2 | 0.8 | 1 |
| 39 | Controls | 0.6 | 0 | 0.1 |
| 40 | Controls | 1.7 | 1.3 | 1.4 |
| 41 | Controls | 0.5 | 0.3 | 0.3 |
| 42 | Controls | 0.9 | 0.4 | 0.6 |
| 43 | Controls | 0.7 | 0.7 | 1 |
| 44 | Controls | 0 | 0 | 0.1 |
| 45 | Controls | 0 | 0.1 | 0.2 |
| 46 | Controls | 0.3 | 0.2 | 0.3 |
References
- Ahmadzadeh, A.; Rezaei-Tavirani, M. Pathogenesis and Genetics of Celiac Disease; a Systematic Review. Egyptian Journal of Medical Human Genetics 2025, 26(1). [Google Scholar] [CrossRef]
- Durazzo, M.; Ferro, A.; Brascugli, I.; Mattivi, S.; Fagoonee, S.; Pellicano, R. Extra-Intestinal Manifestations of Celiac Disease: What Should We Know in 2022? Journal of Clinical Medicine 2022, 11(1), 258. [Google Scholar] [CrossRef]
- Lindfors, K.; Ciacci, C.; Kurppa, K.; Lundin, K. E. A.; Makharia, G. K.; Mearin, M. L.; Murray, J. A.; Verdu, E. F.; Kaukinen, K. Coeliac Disease. Nature Reviews Disease Primers 2019, 5(1). [Google Scholar] [CrossRef]
- Kang, J. Y.; Kang, A. H. Y.; Green, A.; Gwee, K. A.; Ho, K. Y. Systematic Review: Worldwide Variation in the Frequency of Coeliac Disease and Changes over Time. Alimentary Pharmacology & Therapeutics 2013, 38(3), 226–245. [Google Scholar] [CrossRef] [PubMed]
- Leonard, M. M.; Sapone, A.; Catassi, C.; Fasano, A. Celiac Disease and Nonceliac Gluten Sensitivity. JAMA 2017, 318(7), 647. [Google Scholar] [CrossRef] [PubMed]
- Leffler, D. A.; Green, P. H. R.; Fasano, A. Extraintestinal Manifestations of Coeliac Disease. Nature Reviews Gastroenterology & Hepatology 2015, 12(10), 561–571. [Google Scholar] [CrossRef] [PubMed]
- Castillo, N. E.; Theethira, T. G.; Leffler, D. A. The Present and the Future in the Diagnosis and Management of Celiac Disease. Gastroenterology Report 2014, 3(1), 3–11. [Google Scholar] [CrossRef]
- Ludvigsson, J. F.; Leffler, D. A.; Bai, J. C.; Biagi, F.; Fasano, A.; Green, P. H. R.; Hadjivassiliou, M.; Kaukinen, K.; Kelly, C. P.; Leonard, J. N.; Lundin, K. E. A.; Murray, J. A.; Sanders, D. S.; Walker, M. M.; Zingone, F.; Ciacci, C. The Oslo Definitions for Coeliac Disease and Related Terms. Gut 2012, 62(1), 43–52. [Google Scholar] [CrossRef]
- Spijkerman, M.; Tan, I. L.; Kolkman, J. J.; Withoff, Sebo; Wijmenga, Cisca; Visschedijk, M. C.; Weersma, R. K. A Large Variety of Clinical Features and Concomitant Disorders in Celiac Disease – a Cohort Study in the Netherlands. Digestive and Liver Disease 2016, 48(5), 499–505. [Google Scholar] [CrossRef]
- Caio, G.; Volta, U.; Sapone, A.; Leffler, D. A.; De Giorgio, R.; Catassi, C.; Fasano, A. Celiac Disease: A Comprehensive Current Review. BMC Medicine 2019, 17(1). [Google Scholar] [CrossRef]
- National Institute for Health and Care Excellence (NICE). Internal Clinical Guidelines Team Final Version Coeliac Disease Recognition, Assessment and Management Clinical Guideline NG20 Methods, Evidence and Recommendations. 2015. Available online: https://www.nice.org.uk/guidance/ng20/evidence/full-guideline-pdf-438530077.
- Walker, M. M.; Murray, J. A. An Update in the Diagnosis of Coeliac Disease. Histopathology 2010, no–no. [Google Scholar] [CrossRef] [PubMed]
- Rubio-Tapia, A.; Kyle, R. A.; Kaplan, E. L.; Johnson, D. R.; Page, W.; Erdtmann, F.; Brantner, T. L.; Kim, W. R.; Phelps, T. K.; Lahr, B. D.; Zinsmeister, A. R.; Melton, L. J.; Murray, J. A. Increased Prevalence and Mortality in Undiagnosed Celiac Disease. Gastroenterology 2009, 137(1), 88–93. [Google Scholar] [CrossRef] [PubMed]
- Logan, R. F.; Rifkind, E. A.; Turner, I. D.; Ferguson, A. Mortality in Celiac Disease. Gastroenterology 1989, 97(2), 265–271. [Google Scholar] [CrossRef] [PubMed]
- Marsh, M. N. Grains of Truth: Evolutionary Changes in Small Intestinal Mucosa in Response to Environmental Antigen Challenge. Gut 1990, 31(1), 111–114. [Google Scholar] [CrossRef]
- Arguelles-Grande, C.; Tennyson, C. A.; Lewis, S. K.; Green, P. H. R.; Bhagat, G. Variability in Small Bowel Histopathology Reporting between Different Pathology Practice Settings: Impact on the Diagnosis of Coeliac Disease. Journal of Clinical Pathology 2011, 65(3), 242–247. [Google Scholar] [CrossRef]
- Picarelli, A.; Borghini, R.; Donato, G.; Di Tola, M.; Boccabella, C.; Isonne, C.; Giordano, M.; Di Cristofano, C.; Romeo, F.; Di Cioccio, G.; Marcheggiano, A.; Villanacci, V.; Tiberti, A. Weaknesses of Histological Analysis in Celiac Disease Diagnosis: New Possible Scenarios. Scandinavian Journal of Gastroenterology 2014, 49(11), 1318–1324. [Google Scholar] [CrossRef]
- Villanacci, V.; Lorenzi, L.; Donato, F.; Auricchio; Dziechciarz, R.; Piotr; Gyimesi, Judit; Koletzko, Sibylle; Mišak, Zrinjka; Laguna, V. M.; Polanco, I.; Ramos, D.; Shamir, R.; Troncone, R.; Vriezinga, S. L.; Mearin, M. Luisa. Histopathological Evaluation of Duodenal Biopsy in the PreventCD Project. An Observational Interobserver Agreement Study. Apmis 2018, 126(3), 208–214. [Google Scholar] [CrossRef]
- Alessio, M. G.; Tonutti, E.; Brusca, I.; Radice, A.; Licini, L.; Sonzogni, A.; Florena, A.; Schiaffino, E.; Marus, W.; Sulfaro, S.; Villalta, D. Correlation between IgA Tissue Transglutaminase Antibody Ratio and Histological Finding in Celiac Disease. Journal of Pediatric Gastroenterology and Nutrition 2012, 55(1), 44–49. [Google Scholar] [CrossRef]
- Montén, C.; Bjelkenkrantz, K.; Gudjonsdottir, A. H.; Browaldh, L.; Arnell, H.; Naluai, Å. T.; Agardh, D. Validity of Histology for the Diagnosis of Paediatric Coeliac Disease: A Swedish Multicentre Study. Scandinavian Journal of Gastroenterology 2015, 51(4), 427–433. [Google Scholar] [CrossRef]
- Corazza, G. R.; Villanacci, V.; Zambelli, C.; Milione, M.; Luinetti, O.; Vindigni, C.; Chioda, C.; Albarello, L.; Bartolini, D.; Donato, F. Comparison of the Interobserver Reproducibility with Different Histologic Criteria Used in Celiac Disease. Clinical Gastroenterology and Hepatology 2007, 5(7), 838–843. [Google Scholar] [CrossRef] [PubMed]
- Denholm, J.; Schreiber, B. A.; Jaeckle, F.; Wicks, M. N.; Benbow, E. W.; Bracey, T. S.; Chan, J. Y. H.; Farkas, L.; Fryer, E.; Gopalakrishnan, K.; Hughes, C. A.; Kirkwood, K. J.; Langman, G.; Mahler-Araujo, B.; McMahon, R. F. T.; Myint, K. L. W.; Natu, S.; Robinson, A.; Sanduka, A.; Sheppard, K. A. CD, or Not CD, That Is the Question: A Digital Interobserver Agreement Study in Coeliac Disease. BMJ Open Gastroenterology 2024, 11(1), e001252. [Google Scholar] [CrossRef] [PubMed]
- Tsung, J. S. H. Institutional Pathology Consultation. The American Journal of Surgical Pathology 2004, 28(3), 399–402. [Google Scholar] [CrossRef] [PubMed]
- Frable, W. J. Surgical Pathology—Second Reviews, Institutional Reviews, Audits, and Correlations: What’s out There? Error or Diagnostic Variation? Archives of Pathology & Laboratory Medicine 2006, 130(5), 620–625. [Google Scholar] [CrossRef]
- Ordi, J.; Castillo, P.; Saco, A.; del Pino, M.; Ordi, O.; Rodríguez-Carunchio, L.; Ramírez, J. Validation of Whole Slide Imaging in the Primary Diagnosis of Gynaecological Pathology in a University Hospital. Journal of Clinical Pathology 2014, 68(1), 33–39. [Google Scholar] [CrossRef]
- Villanacci, V. Celiac Disease: Histology-Differential Diagnosis-Complications. A Practical Approach. Pathologica 2020, 112(3), 186–196. [Google Scholar] [CrossRef]
- Chae, Y. K.; Arya, A.; Chiec, L.; Shah, H.; Rosenberg, A.; Patel, S.; Raparia, K.; Choi, J.; Wainwright, D. A.; Villaflor, V.; Cristofanilli, M.; Giles, F. Challenges and Future of Biomarker Tests in the Era of Precision Oncology: Can We Rely on Immunohistochemistry (IHC) or Fluorescence in Situ Hybridization (FISH) to Select the Optimal Patients for Matched Therapy? Oncotarget 2017, 8(59), 100863–100898. [Google Scholar] [CrossRef]
- Dickson, B. C.; Streutker, C. J.; Chetty, R. Coeliac Disease: An Update for Pathologists. Journal of Clinical Pathology 2006, 59(10), 1008–1016. [Google Scholar] [CrossRef]
- Popp, A.; Taavela, J.; Graziano, P.; Parente, P.; Covelli, C.; Lamacchia, C.; Andriulli, A.; Mäki, M.; Isola, J. A New Intraepithelial γδ T-Lymphocyte Marker for Celiac Disease Classification in Formalin-Fixed Paraffin-Embedded (FFPE) Duodenal Biopsies. Digestive Diseases and Sciences 2020, 66(10), 3352–3358. [Google Scholar] [CrossRef]
- Camarero, C.; De Andrés, A.; García-Hoz, C.; Roldán, B.; Muriel, A.; León, F.; Roy, G. Assessment of Duodenal Intraepithelial Lymphocyte Composition (Lymphogram) for Accurate and Prompt Diagnosis of Celiac Disease in Pediatric Patients. Clinical and translational gastroenterology 2021, 12(11), e00426. [Google Scholar] [CrossRef] [PubMed]
- Dalton, J. E.; Cruickshank, S. M.; Egan, C. E.; Mears, R.; Newton, D. J.; Andrew, E. M.; Lawrence, B.; Howell, G. R.; Else, K. J.; Gubbels, M.-J.; Striepen, B.; Smith, J. A.; White, S.; Carding, S. R. Intraepithelial γδ+ Lymphocytes Maintain the Integrity of Intestinal Epithelial Tight Junctions in Response to Infection. Gastroenterology AGA 2006, 131(3), 818–829. [Google Scholar] [CrossRef]
- Hu, Y.; Hu, Q.; Li, Y.; Lu, L.; Xiang, Z.; Yin, Z.; Kabelitz, D.; Wu, Y. γδ T Cells: Origin and Fate, Subsets, Diseases and Immunotherapy. Signal Transduction and Targeted Therapy 2023, 8(1), 1–38. [Google Scholar] [CrossRef]
- Santillo, D.; Bellos, E.; Sancho-Shimizu, V. Unravelling γδ T-Cell Dysregulation in the Gut and Its Implications for Immune-Mediated Diseases. Disease Models & Mechanisms 2025, 18(9). [Google Scholar] [CrossRef]
- Jandke, A.; Melandri, D.; Monin, L.; Ushakov, D. S.; Laing, A. G.; Vantourout, P.; East, P.; Nitta, T.; Narita, T.; Takayanagi, H.; Feederle, R.; Hayday, A. Butyrophilin-like Proteins Display Combinatorial Diversity in Selecting and Maintaining Signature Intraepithelial γδ T Cell Compartments. Nature Communications 2020, 11(1). [Google Scholar] [CrossRef]
- Mayassi, T.; Ladell, K.; Gudjonson, H.; McLaren, J. E.; Shaw, D. G.; Tran, M. T.; Rokicka, J. J.; Lawrence, I.; Grenier, J.-C.; van Unen, V.; Ciszewski, C.; Dimaano, M.; Sayegh, H. E.; Kumar, V.; Wijmenga, C.; Green, P. H. R.; Gokhale, R.; Jericho, H.; Semrad, C. E.; Guandalini, S. Chronic Inflammation Permanently Reshapes Tissue-Resident Immunity in Celiac Disease. Cell 2019, 176(5), 967–981.e19. [Google Scholar] [CrossRef]
- Coënon, L.; Geindreau, M.; Ghiringhelli, F.; Villalba, M.; Bruchard, Mélanie. Natural Killer Cells at the Frontline in the Fight against Cancer. Cell Death and Disease 2024, 15(8). [Google Scholar] [CrossRef]
- Mikulak, J.; Oriolo, F.; Bruni, E.; Roberto, A.; Colombo, F.; Villa, A.; Bosticardo, M.; Bortolomai; Lo Presti, I.; Elena; Meraviglia, S.; Dieli, F.; Vetrano, S.; Danese; Della Bella, S.; Silvia; Carvello, M.; Sacchi, M.; Cugini, G.; Colombo, G.; Klinger, M.; Scocco, P. NKp46-Expressing Human Gut-Resident Intraepithelial Vδ1 T Cell Subpopulation Exhibits High Antitumor Activity against Colorectal Cancer. JCI insight 2019, 4(24). [Google Scholar] [CrossRef]
- Pazina, T.; Shemesh, A.; Brusilovsky, M.; Porgador, A.; Campbell, K. S. Regulation of the Functions of Natural Cytotoxicity Receptors by Interactions with Diverse Ligands and Alterations in Splice Variant Expression. Frontiers in Immunology 2017, 8. [Google Scholar] [CrossRef]
- Barrow, A. D.; Martin, C. J.; Colonna, M. The Natural Cytotoxicity Receptors in Health and Disease. Frontiers in Immunology 2019, 10. [Google Scholar] [CrossRef]
- Fugmann, S. D. RAG1 and RAG2 in V(D)J Recombination and Transposition. Immunologic Research 2001, 23(1), 23–40. [Google Scholar] [CrossRef]
- Lynch, S.; Kelleher, D.; McManus, R.; O’Farrelly, C. RAG1 and RAG2 Expression in Human Intestinal Epithelium: Evidence of Extrathymic T Cell Differentiation. European Journal of Immunology 1995, 25(5), 1143–1147. [Google Scholar] [CrossRef] [PubMed]
- Bas, A.; Forsberg, G.; Sjoberg, V.; Hammarstrom, S.; Hernell, O.; Hammarstrom, M-L. Aberrant Extrathymic T Cell Receptor Gene Rearrangement in the Small Intestinal Mucosa: A Risk Factor for Coeliac Disease? Gut 2008, 58(2), 189–195. [Google Scholar] [CrossRef] [PubMed]
- Lesourne, R.; Uehara, S.; Lee, J.; Song, K.-D.; Li, L.; Pinkhasov, J.; Zhang, Y.; Weng, N.-P.; Wildt, K. F.; Wang, L.; Bosselut, Remy; Love, P. E. Themis, a T Cell–Specific Protein Important for Late Thymocyte Development. Nature Immunology 2009, 10(8), 840–847. [Google Scholar] [CrossRef] [PubMed]
- Bondar, C.; Plaza-Izurieta, L.; Fernandez-Jimenez, N.; Irastorza, I.; Withoff, S.; Wijmenga, C.; Chirdo, F.; Bilbao, J. R. THEMIS and PTPRK in Celiac Intestinal Mucosa: Coexpression in Disease and after in Vitro Gliadin Challenge. European Journal of Human Genetics 2013, 22(3), 358–362. [Google Scholar] [CrossRef] [PubMed]
- Shrout, P. E.; Fleiss, J. L. Intraclass Correlations: Uses in Assessing Rater Reliability. Psychological Bulletin 1979, 86(2), 420–428. [Google Scholar] [CrossRef]
- Sali, R.; Ehsan, L.; Kowsari, Kamran; Khan, Marium Mateen; Moskaluk, C. A.; Syed, S.; Brown, D. E. CeliacNet: Celiac Disease Severity Diagnosis on Duodenal Histopathological Images Using Deep Residual Networks. 2019 IEEE International Conference on Bioinformatics and Biomedicine (BIBM) 2019. [Google Scholar] [CrossRef]
- Jaeckle, F.; Denholm, J.; Schreiber, B.; Evans, S. C.; Wicks, M. N.; Chan, J. Y. H.; Bateman, A. C.; Natu, S.; Arends, M. J.; Soilleux, E. Machine Learning Achieves Pathologist-Level Celiac Disease Diagnosis. NEJM AI 2025, 2(4). [Google Scholar] [CrossRef]
| Score | Definition |
|---|---|
| 0 | No staining |
| 1 | Mild staining |
| 2 | Moderate staining |
| 3 | Strong staining |
| Diagnostic group/ Observer | Observer 1 median (IQR) | Observer 2 median (IQR) | Observer 3 median (IQR) | Median (IQR) for all 3 observers combined | p-value compared with control |
|---|---|---|---|---|---|
| Median surface BTNL8 staining intensity | - | ||||
| Combined CD | 2 (1.25-3) | 2.5 (2-3) | 2 (1-2) | 2 (1-3) | 0.0039 |
| Untreated CD | 2 (1-2) | 2 (2-3) | 2 (1-2) | 2 (1-2) | 0.0012 |
| Treated CD | 3 (2-3) | 3 (3) | 2 (2) | 2 (2-3) | 1 (ns) |
| Control | 2.5 (2-3) | 3 (2-3) | 2 (1-3) | 2 (2-3) | 1 |
| Median crypt BTNL8 staining intensity | |||||
| Combined CD | 1 | 1 (0.25-1) | 1 (0-1) | 1 (0-1) | 0.16 (ns) |
| Untreated CD | 0 | 1 (1) | 1 (0-1) | 1 (0-1) | 0.29 (ns) |
| Treated CD | 1 | 1 (0-1) | 1 (1) | 1 (1) | 0.011 |
| Control | 0 | 1 (0-1) | 0 (0-1) | 0.5 (0-1) | 1 |
| Median TCRδ+ IEL count per 100 enterocytes | |||||
| Combined CD | 13.9(8.1-17.8) | 8.5(5.5-10.85) | 8.2(4.5-10.2) | 9.4(6.0-13.9) | 3.80 × 10⁻¹⁹ |
| Untreated CD | 13.9(7.7-16.9) | 8.4(5.4-11) | 8.1(4-9.6) | 8.9(6.0-13.9) | 3.77 × 10⁻¹⁸ |
| Treated CD | 12.2(12.1-21.8) | 9.5(6-10) | 9.3(7.4-11.5) | 10.0(6.7-13.1) | 1.27 × 10⁻⁶ |
| Control | 0.7(0.15-1.1) | 0.4(0-0.8) | 0.4(0.2-1) | 0.5(0.1-1.0) | 1 |
| Median NKp46+ IEL count per 100 enterocytes | |||||
| Combined CD | 16.3 (13.9-21.4) | 12.6 (10.3-15.4) | 13.4 (11.4-15.3) | 13.8 (11.7-16.6) | 2.05 × 10⁻¹² |
| Untreated CD | 16.2 (13.6-18.6) | 13 (10.6-15.4) | 13.4 (11.2-15) | 13.8 (11.8-16.4) | 1.07 × 10⁻¹¹ |
| Treated CD | 21.4 (15.4-25.8) | 9.4 (9.4-10.2) | 13.4 (13.4-15.4) | 13.4 (11.3-17.5) | 1.38 × 10⁻⁴ |
| Control | 7.7 (6.95-9.9) | 9.9 (8.3-11.6) | 10.9 (7.55-11.9) | 9.6 (7.4-11.6) | 1 |
| Median THEMIS+ IEL count per 100 enterocytes | |||||
| Combined CD | 6.9(4.6-8.3) | 6.2(4.3-9.2) | 2.75(1.7-5.3) | 5.4(2.7-7.8) | 5.09 × 10⁻⁴ |
| Untreated CD | 5.9(3.4-7.8) | 5.9(4.1-9.1) | 2.5(1.2-3.6) | 4.9(2.6-7.6) | 0.015 |
| Treated CD | 7.2(6.8-9.3) | 8.9(6.2-9.2) | 5.4(5.1-7.9) | 7.2(5.7-9.3) | 7.03 × 10⁻⁵ |
| Control | 4.1(1.0-6.7) | 3.4(0.8-6.6) | 1.8(0.9-3.2) | 2.9(0.8-6.1) | 1 |
| Median TdT+ IEL count per 100 enterocytes | |||||
| Combined CD | 0.0(0.0) | 0.0(0.0) | 0.5(0.0-1.0) | 0.0(0.0-0.5) | 1.87 × 10⁻⁴ |
| Untreated CD | 0.0(0.0) | 0.0(0.0) | 0.5(0.0-1.0) | 0.0(0.0-0.5) | 9.58 × 10⁻⁵ |
| Treated CD | 0.0(0.0) | 0.0(0.0) | 0.5(0.0-0.5) | 0.0(0.0) | 1 (ns) |
| Control | 0.0(0.0) | 0.0(0.0) | 0.0(0.0) | 0.0(0.0) | 1 |
| Marker | ICC(2,1) | Threshold | AUC |
|---|---|---|---|
| TCRδ | 0.82 | >2.1 TCRδ+/100EC | 0.94 |
| NKp46 | 0.82 | >12.8 NKp46+ IEL/100 EC | 0.86 |
| THEMIS | 0.64 | >2.2 THEMIS+ IEL/100EC | 0.68 |
| TdT | 0.22 | >0.5 TdT+ IEL/100EC | 0.61 |
| BTNL8 CE | 0.52 | N/A | 0.56 |
| BTNL8 SE | 0.64 | N/A | 0.37 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




