Submitted:
09 April 2026
Posted:
13 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Nitrogen Uptake and Assimilation
3. Amino Acid Transport in the Plant
4. Plant Amino Acid Transporters
Amino Acid Transporter Families
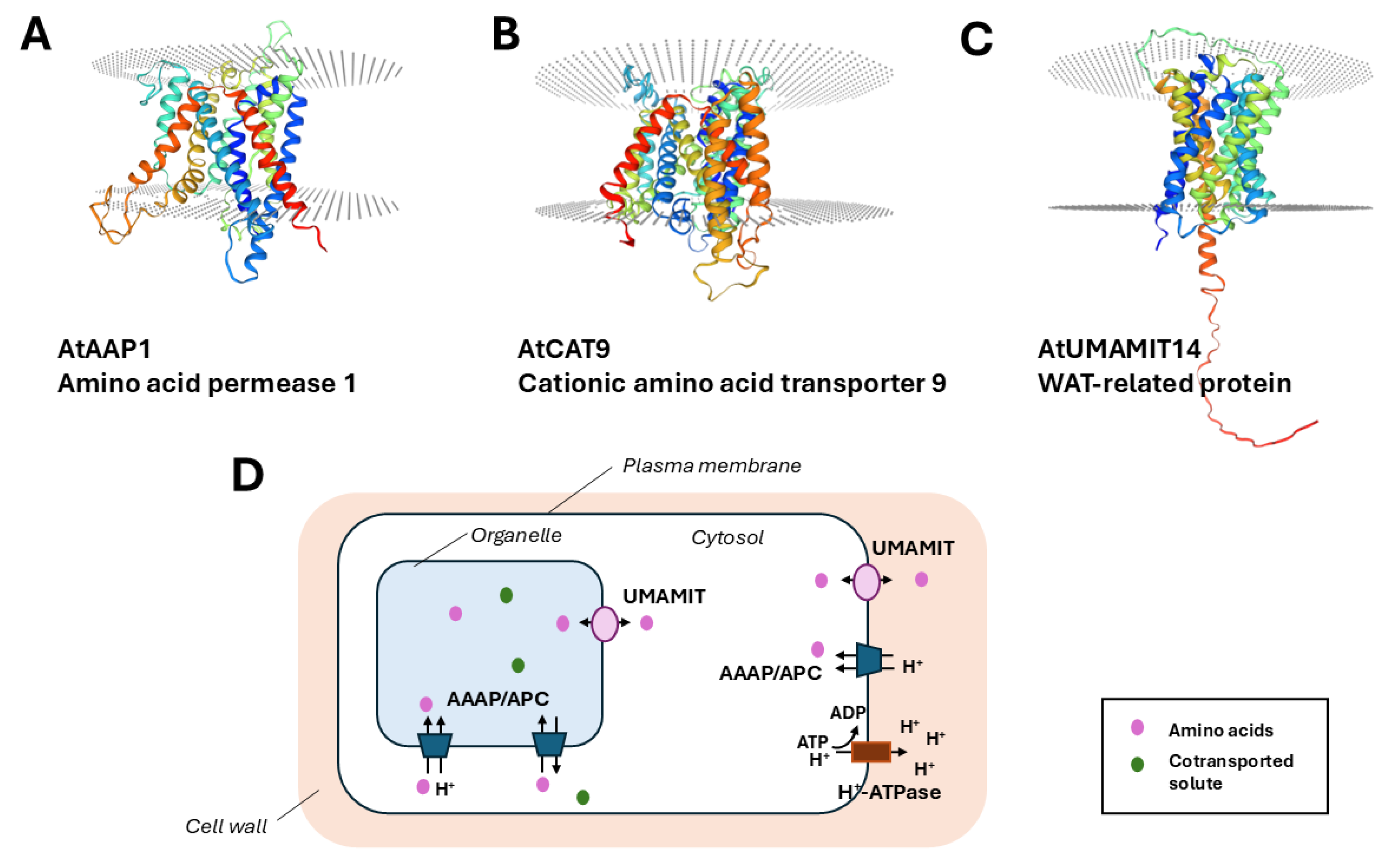
Structure and Transport Characteristics of Amino Acid Permeases
5. Physiological Functions of Amino Acid Permeases
Amino Acid Transport in the Roots
Xylem Transport to the Shoot
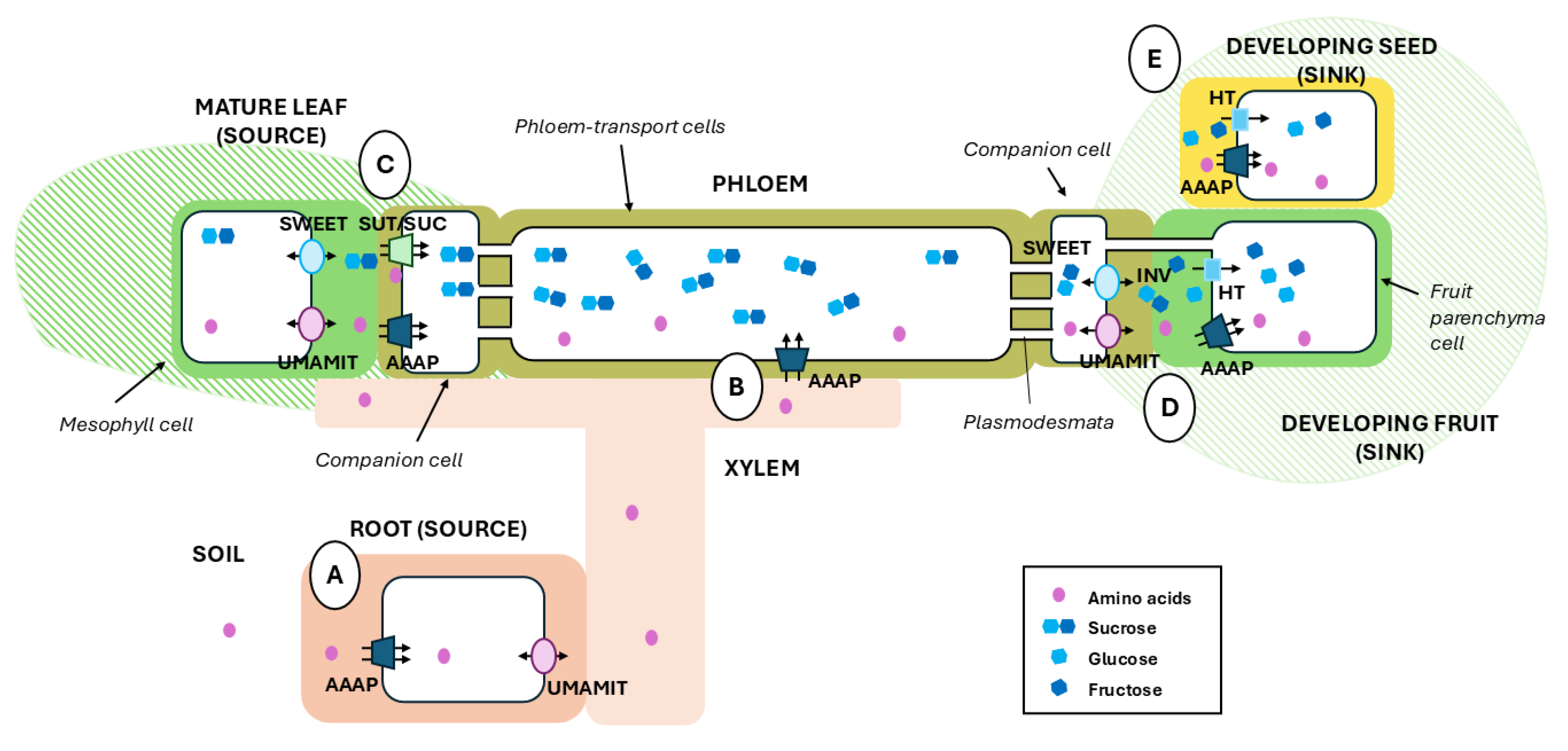
Phloem Partitioning to Sinks
Subcellular Transport
6. Role of Amino Acid Permeases in the Determination of Yield, Nutrient Use Efficiency and Quality of Crops
Role of Amino Acid Permeases in the Determination of Crop Yield
Impact of Amino Acid Transporters on Nutritional Value and Crop Quality
Role of Amino Acid Permeases in the Determination of Nitrogen Use Efficiency (NUE) in Crops
7. Conclusions and Future Perspectives
Funding
Conflicts of Interest
References
- Agyenim-Boateng, KG; Halkier, BA; Xu, D. Transport engineering of glucosinolates for future brassica crops. Trends in Plant Science 2025. [Google Scholar] [CrossRef] [PubMed]
- Andrews, M. The partitioning of nitrate assimilation between root and shoot of higher plants. Plant, Cell and Environment 1986, 9, 511–519. [Google Scholar] [CrossRef]
- Awasthi, A; Rialch, I; Kaur, H; Sharma, S; Harper, AL; Havlickova, L; Bancroft, I; Banga, SS. Underpinning regions associated with nitrogen use efficiency in Brassica juncea L. Czern: Using associative transcriptomics approach. BMC Plant Biology 2025, 25, 1253. [Google Scholar] [CrossRef] [PubMed]
- Bernard, SM; Habash, DZ. The importance of cytosolic glutamine synthetase in nitrogen assimilation and recycling. New Phytologist 2009, 182, 608–620. [Google Scholar] [CrossRef]
- Besnard, J; Zhao, C; Avice, JC; Vitha, S; Hyodo, A; Pilot, G; Okumoto, S. Arabidopsis UMAMIT24 and 25 are amino acid exporters involved in seed loading. Journal of Experimental Botany 2018, 69, 5221–5232. [Google Scholar] [CrossRef]
- Boorer, KJ; Fischer, WN. Specificity and Stoichiometry of the ArabidopsisH+/Amino Acid Transporter AAP5. Journal of Biological Chemistry 1997, 272, 13040–13046. [Google Scholar] [CrossRef]
- Boorer, KJ; Frommer, WB; Bush, DR; Kreman, M; Loo, DD; Wright, EM. Kinetics and specificity of a H+/amino acid transporter from Arabidopsis thaliana. Journal of Biological Chemistry 1996, 271, 2213–2220. [Google Scholar] [CrossRef]
- Bright, SW; Shewry, PR; Kasarda, DD. Improvement of protein quality in cereals. Critical Reviews in Plant Sciences 1983, 1, 49–93. [Google Scholar] [CrossRef]
- Bush, DR. Proton-coupled sugar and amino acid transporters in plants. Annual Review of Plant Biology 1993, 44, 513–542. [Google Scholar] [CrossRef]
- Bush, DR; Chiou, TJ; Chen, L. Molecular analysis of plant sugar and amino acid transporters. Journal of Experimental Botany 1996, 47, 1205–1210. [Google Scholar] [CrossRef]
- Campbell, WH. NITRATE REDUCTASE STRUCTURE, FUNCTION AND REGULATION: Bridging the Gap between Biochemistry and Physiology. Annual Reviews of Plant Biology 1999, 50, 277–303. [Google Scholar] [CrossRef] [PubMed]
- Cao, C; Qiu, X; Yang, Z; Jin, Y. New insights into the evolution and function of the UMAMIT (USUALLY MULTIPLE ACIDS MOVE IN AND OUT TRANSPORTER) gene family. Journal of Plant Research 2025, 138, 3–17. [Google Scholar] [CrossRef] [PubMed]
- Cartea, ME; Velasco, P. Glucosinolates in Brassica foods: Bioavailability in food and significance for human health. Phytochemistry Reviews 2008, 7, 213–229. [Google Scholar] [CrossRef]
- Chen, L; Bush, DR. LHT1, a lysine-and histidine-specific amino acid transporter in Arabidopsis. Plant Physiology 1997, 115, 1127–1134. [Google Scholar] [CrossRef]
- Chen, N; Wang, S; Yin, Y; Ren, G; Zhang, Y; Qu, L; Ling, L. Genome-wide identification of the AAAP gene family and expression analysis under tissue-specific expression in five legumes. BMC Genomics 2025, 26, 173. [Google Scholar] [CrossRef]
- Chen, Z; Zhang, Y; Zhang, J; Fan, B; Zhou, Y; Cui, X. Expression of AtAAP gene family and endosperm-specific expression of AtAAP1 gene promotes amino acid absorption in Arabidopsis thaliana and maize. Agronomy 2021, 11, 1668. [Google Scholar] [CrossRef]
- Cheng, L; Yuan, HY; Ren, R; Zhao, SQ; Han, YP; Zhou, QY; Ke, DX; Wang, YX; Wang, L. Genome-wide identification, classification, and expression analysis of amino acid transporter gene family in Glycine max. Frontiers in Plant Science 2016, 7, 515. [Google Scholar] [CrossRef]
- Choi, J; Eom, S; Shin, K; Lee, RA; Choi, S; Lee, JH; Lee, S; Soh, MS. Identification of Lysine Histidine Transporter 2 as an 1-Aminocyclopropane Carboxylic Acid Transporter in Arabidopsis thaliana by Transgenic Complementation Approach. Frontiers in Plant Science 2019, 10, 1092. [Google Scholar] [CrossRef]
- Crawford, NM; Glass, AD. Molecular and physiological aspects of nitrate uptake in plants. Trends in Plant Science 1998, 3, 389–395. [Google Scholar] [CrossRef]
- Dong, C; Li, F; Yang, T; Feng, L; Zhang, S; Li, F; Li, W; Xu, G; Bao, S; Wan, X; et al. Theanine transporters identified in tea plants (Camellia sinensis L.). The Plant Journal 2020, 101, 57–70. [Google Scholar] [CrossRef]
- Dündar, E; Bush, DR. BAT1, a bidirectional amino acid transporter in Arabidopsis. Planta 2009, 229, 1047–1056. [Google Scholar] [CrossRef]
- Egami, T; Wakayama, M; Aoki, N; Sasaki, H; Kisaka, H; Miwa, T; Ohsugi, R. The effects of introduction of a fungal glutamate dehydrogenase gene (gdhA) on the photosynthetic rates, biomass, carbon and nitrogen contents in transgenic potato. Plant Biotechnology 2012, 29, 57–64. [Google Scholar] [CrossRef]
- ElShamey, E; Yang, X; Yang, J; Xia, L; Zeng, Y. A systems review of grain proteins in rice and barley: Biosynthesis, regulation, and impact on end-use quality. Frontiers in Plant Science 2025, 16, 1658144. [Google Scholar] [CrossRef]
- Fan, X; Tang, Z; Tan, Y; Zhang, Y; Luo, B; Yang, M; Lian, X; Shen, Q; Miller, AJ; Xu, G. Overexpression of a pH-sensitive nitrate transporter in rice increases crop yields. Proceedings of the National Academy of Sciences 2016, 113, 7118–7123. [Google Scholar] [CrossRef] [PubMed]
- Fan, T; Wu, C; Yang, W; Lv, T; Zhou, Y; Tian, C. The LHT Gene Family in Rice: Molecular Characterization, Transport Functions and Expression Analysis. Plants 2023, 12, 817. [Google Scholar] [CrossRef]
- Fang, Z; Bai, G; Huang, W; Wang, Z; Wang, X; Zhang, M. The rice peptide transporter OsNPF7.3 is induced by organic nitrogen, and contributes to nitrogen allocation and grain yield. Frontiers in Plant Science 2017, 8, 1338. [Google Scholar] [CrossRef] [PubMed]
- Fang, Z; Wu, B; Ji, Y. The amino acid transporter OsAAP4 contributes to rice tillering and grain yield by regulating neutral amino acid allocation through two splicing variants. Rice 2021, 14, 2. [Google Scholar] [CrossRef] [PubMed]
- Fischer, WN; Kwart, M; Hummel, S; Frommer, WB. Substrate specificity and expression profile of amino acid transporters (AAPs) in Arabidopsis. Journal of Biological Chemistry 1995, 270, 16315–16320. [Google Scholar] [CrossRef]
- Fischer, WN; Loo, DD; Koch, W; Ludewig, U; Boorer, KJ; Tegeder, M; Rentsch, D; Wright, EM; Frommer, WB. Low and high affinity amino acid H+-cotransporters for cellular import of neutral and charged amino acids. The Plant Journal 2002, 29, 717–731. [Google Scholar] [CrossRef]
- Foyer, CH; Parry, M; Noctor, G. Markers and signals associated with nitrogen assimilation in higher plants. Journal of Experimental Botany 2003, 54, 585–593. [Google Scholar] [CrossRef]
- Frommer, WB; Hummel, S; Riesmeier, JW. Expression cloning in yeast of a cDNA encoding a broad specificity amino acid permease from Arabidopsis thaliana. Proceedings of the National Academy of Sciences 1993, 90, 5944–5948. [Google Scholar] [CrossRef] [PubMed]
- Galili, G; Amir, R; Fernie, AR. The regulation of essential amino acid synthesis and accumulation in plants. Annual Review of Plant Biology 2016, 67, 153–178. [Google Scholar] [CrossRef] [PubMed]
- Gao, Z; Wang, Y; Chen, G; Zhang, A; Yang, S; Shang, L; Wang, D; Ruan, B; Liu, C; Jiang, H; et al. The indica nitrate reductase gene OsNR2 allele enhances rice yield potential and nitrogen use efficiency. Nature Communications 2019, 10, 5207. [Google Scholar] [CrossRef] [PubMed]
- Garneau, MG; Tan, Q; Tegeder, M. Function of pea amino acid permease AAP6 in nodule nitrogen metabolism and export, and plant nutrition. Journal of Experimental Botany 2018, 69, 5205–5219. [Google Scholar] [CrossRef]
- Gil-Villar, D; Arrones, A; Gramazio, P; Vilanova, S; Jiménez-Benavente, E; Plazas, M; Arbona, V; Granell, A; Medina, J; Molina, RV; et al. Variation in responses to N limitation in Solanum lycopersicum var. cerasiforme and S. pimpinellifolium accessions and hybrids reveals genetic potential for improving nitrogen use efficiency (NUE) in tomato breeding. Plant Physiology and Biochemistry 2025, 229, 110428. [Google Scholar] [CrossRef]
- Good, AG; Shrawat, AK; Muench, DG. Can less yield more? Is reducing nutrient input into the environment compatible with maintaining crop production? Trends Plant Science 2004, 9, 597–605. [Google Scholar] [CrossRef]
- Grallath, S; Weimar, T; Meyer, A; Gumy, C; Suter-Grotemeyer, M; Neuhaus, JM; Rentsch, D. The AtProT family. Compatible solute transporters with similar substrate specificity but differential expression patterns. Plant Physiology 2005, 137, 117–126. [Google Scholar] [CrossRef]
- Grant, JE; Ninan, A; Cripps-Guazzone, N; Shaw, M; Song, J; Petřík, I; Novák, O; Tegeder, M; Jameson, PE. Concurrent overexpression of amino acid permease AAP1 (3a) and SUT1 sucrose transporter in pea resulted in increased seed number and changed cytokinin and protein levels. Functional Plant Biology 2021, 48, 889–904. [Google Scholar] [CrossRef]
- Guo, N; Hu, J; Yan, M; Qu, H; Luo, L; Tegeder, M; Xu, G. Oryza sativa Lysine-Histidine-type Transporter 1 functions in root uptake and root-to-shoot allocation of amino acids in rice. The Plant Journal 2020, 103, 395–411. [Google Scholar] [CrossRef]
- Hachiya, T; Ueda, N; Kitagawa, M; Hanke, G; Suzuki, A; Hase, T; Sakakibara, H. Arabidopsis root-type ferredoxin:NADP(H) oxidoreductase 2 is involved in detoxification of nitrite in roots. Plant and Cell Physiology 2016, 57, 2440–2450. [Google Scholar] [CrossRef]
- Hachiya, T; Sakakibara, H. Interactions between nitrate and ammonium in their uptake, allocation, assimilation, and signaling in plants. Journal of Experimental Botany 2017, 68, 2501–2512. [Google Scholar] [CrossRef] [PubMed]
- Haferkamp, I; Schmitz-Esser, S. The plant mitochondrial carrier family: Functional and evolutionary aspects. Frontiers in Plant Science 2012, 3, 2. [Google Scholar] [CrossRef] [PubMed]
- Hammes, UZ; Nielsen, E; Honaas, LA; Taylor, CG; Schachtman, DP. AtCAT6, a sink-tissue-localized transporter for essential amino acids in Arabidopsis. The Plant Journal 2006, 48, 414–426. [Google Scholar] [CrossRef] [PubMed]
- Hammes, UZ; Pedersen, BP. Structure and Function of Auxin Transporters. Annual Review of Plant Biology 2024, 75, 185–209. [Google Scholar] [CrossRef]
- He, C; Amin, B; Liu, X; Wu, B; Yang, X; Kandegama, W; Fang, Z. Amino acid transporter OsAAP8 mediates rice tillering and height by regulating the transport of neutral amino acids. Plant Physiology and Biochemistry 2025, 229, 110452. [Google Scholar] [CrossRef]
- He, C; Yang, X; Zhang, J; Wu, W; Wang, C; Chen, W; Wu, B; Huang, W; Fang, Z. Amino acid transporter OsAAP18 modulates nitrogen allocation to boost yield and grain quality in rice. Rice Science 2026, 1. [Google Scholar] [CrossRef]
- Heinig, U; Gutensohn, M; Dudareva, N; Aharoni, A. The challenges of cellular compartmentalization in plant metabolic engineering. Current Opinion in Biotechnology 2013, 24, 239–246. [Google Scholar] [CrossRef]
- Hirner, B; Fischer, WN; Rentsch, D; Kwart, M; Frommer, WB. Developmental control of H+/amino acid permease gene expression during seed development of Arabidopsis. The Plant Journal 1998, 14, 535–544. [Google Scholar] [CrossRef]
- Hirner, A; Ladwig, F; Stransky, H; Okumoto, S; Keinath, M; Harms, A; Frommer, WB; Koch, W. Arabidopsis LHT1 Is a High-Affinity Transporter for Cellular Amino Acid Uptake in Both Root Epidermis and Leaf Mesophyll. The Plant Cell 2006, 18, 1931–1946. [Google Scholar] [CrossRef]
- Hoyos, ME; Palmieri, L; Wertin, T; Arrigoni, R; Polacco, JC; Palmieri, F. Identification of a mitochondrial transporter for basic amino acids in Arabidopsis thaliana by functional reconstitution into liposomes and complementation in yeast. The Plant Journal 2003, 33, 1027–1035. [Google Scholar] [CrossRef]
- Ji, Y; Huang, W; Wu, B; Fang, Z; Wang, X. The amino acid transporter AAP1 mediates growth and grain yield by regulating neutral amino acid uptake and reallocation in Oryza sativa. Journal of Experimental Botany 2020, 71, 4763–4777. [Google Scholar] [CrossRef]
- Jin, F; Xie, P; Li, Z; Wu, B; Huang, W; Fang, Z. Blocking of amino acid transporter OsAAP7 promoted tillering and yield by determining basic and neutral amino acids accumulation in rice. BMC Plant Biology 2024, 24, 447. [Google Scholar] [CrossRef]
- Kim, JY; Loo, EP; Pang, TY; Lercher, M; Frommer, WB; Wudick, MM. Cellular export of sugars and amino acids: Role in feeding other cells and organisms. Plant Physiology 2021, 187, 1893–1914. [Google Scholar] [CrossRef] [PubMed]
- Koch, W; Kwart, M; Laubner, M; Heineke, D; Stransky, H; Frommer, WB; Tegeder, M. Reduced amino acid content in transgenic potato tubers due to antisense inhibition of the leaf H+/amino acid symporter StAAP1. The Plant Journal 2003, 33, 211–220. [Google Scholar] [CrossRef] [PubMed]
- Kotowska, MM; Wright, IJ; Westoby, M. Parenchyma Abundance in Wood of Evergreen Trees Varies Independently of Nutrients. Frontiers in Plant Science 2020, 11, 86. [Google Scholar] [CrossRef] [PubMed]
- Kuhnert, F; Westhoff, P; Valencia, V; Krüger, S; Vogel, K; Lundquist, PK; Rosar, C; Goss, T; Weber, APM. RETICULATA1 is a plastid-localized basic amino acid transporter. Nat Plants 2025, 11, 1890–1902. [Google Scholar] [CrossRef]
- Ladwig, F; Stahl, M; Ludewig, U; Hirner, AA; Hammes, UZ; Stadler, R; Harter, K; Koch, W. Siliques are Red1 from Arabidopsis acts as a bidirectional amino acid transporter that is crucial for the amino acid homeostasis of siliques. Plant Physiology 2012, 158, 1643–1655. [Google Scholar] [CrossRef]
- Lalonde, S; Tegeder, M; Throne-Holst, M; Frommer, WB; Patrick, JW. Phloem loading and unloading of sugars and amino acids. Plant, Cell and Environment 2003, 26, 37–56. [Google Scholar] [CrossRef]
- Lawlor, DW; Lemaire, G; Gastal, F. Nitrogen, plant growth and crop yield. In Plant nitrogen; Lea, PJ, Morot-Gaudry, JF, Eds.; Springer Berlin Heidelberg, 2001; pp. 343–367. [Google Scholar]
- Lea, PJ; Miflin, BJ. Alternative route for nitrogen assimilation in higher plants. Nature 1974, 251, 614–616. [Google Scholar] [CrossRef]
- Lea, PJ; Miflin, BJ. Nitrogen assimilation and its relevance to crop improvement. Annual Plant Reviews 2011, 42, 1–40. [Google Scholar]
- Lee, YH; Foster, J; Chen, J; Voll, LM; Weber, AP; Tegeder, M. AAP1 transports uncharged amino acids into roots of Arabidopsis. The Plant Journal 2007, 50, 305–319. [Google Scholar] [CrossRef] [PubMed]
- Lehmann, S; Gumy, C; Blatter, E; Boeffel, S; Fricke, W.; Rentsch, D. In planta function of compatible solute transporters of the AtProT family. Journal of Experimental Botany 2011, 62, 787–796. [Google Scholar] [CrossRef] [PubMed]
- Li, MX; Huang, FL; Lin, JH; Qu, JY; Wan, GF; Song, GC; Zhao, J; Peng, Y; Teng, ZN; Liu, L; et al. Regulation of grain protein content by the amino acid permease gene OsAAP4 in rice. Theoretical and Applied Genetics 2026, 139, 50. [Google Scholar] [CrossRef] [PubMed]
- Ling, L; Wang, S; Zhang, H; Yang, F; Qu, L. Genome-wide identification of AAAP gene family and expression analysis under abiotic stresses in Avena sativa. BMC Genomics 2026, 27, 5. [Google Scholar] [CrossRef]
- Liu, Y; von Wirén, N. Ammonium as a signal for physiological and morphological responses in plants. Journal of Experimental Botany 2017, 68, 2581–2592. [Google Scholar] [CrossRef]
- Liu, S; Wang, D; Mei, Y; Xia, T; Xu, W; Zhang, Y; You, X; Zhang, X; Li, L; Wang, NN. Overexpression of GmAAP6a enhances tolerance to low nitrogen and improves seed nitrogen status by optimizing amino acid partitioning in soybean. Plant Biotechnology Journal 2020, 18, 1749–1762. [Google Scholar] [CrossRef]
- Loqué, D; von Wirén, N. Regulatory levels for the transport of ammonium in plant roots. Journal of Experimental Botany 2004, 55, 1293–1305. [Google Scholar] [CrossRef]
- Lu, MZ; Snyder, R; Grant, J; Tegeder, M. Manipulation of sucrose phloem and embryo loading affects pea leaf metabolism, carbon and nitrogen partitioning to sinks as well as seed storage pools. The Plant Journal 2020, 101, 217–236. [Google Scholar] [CrossRef]
- Lu, K; Wu, B; Wang, J; Zhu, W; Nie, H; Qian, J; Huang, W; Fang, Z. Blocking amino acid transporter OsAAP3 improves grain yield by promoting outgrowth buds and increasing tiller number in rice. Plant Biotechnology Journal 2018, 16, 1710–1722. [Google Scholar] [CrossRef]
- Ma, H; Cao, X; Shi, S; Li, S; Gao, J; Ma, Y; Zhao, Q; Chen, Q. Genome-wide survey and expression analysis of the amino acid transporter superfamily in potato (Solanum tuberosum L.). Plant Physiology and Biochemistry 2016, 107, 164–177. [Google Scholar] [CrossRef]
- Maathuis, FJ. Physiological functions of mineral macronutrients. Current Opinion in Plant Biology 2009, 12, 250–258. [Google Scholar] [CrossRef] [PubMed]
- Martin, A; Belastegui-Macadam, X; Quilleré, I; Floriot, M; Valadier, MH; Pommel, B; Andrieu, B; Donnison, I; Hirel, B. Nitrogen management and senescence in two maize hybrids differing in the persistence of leaf greenness: Agronomic, physiological and molecular aspects. New phytologist 2005, 167, 483–492. [Google Scholar] [CrossRef] [PubMed]
- Masclaux-Daubresse, C; Daniel-Vedele, F; Dechorgnat, J; Chardon, F; Gaufichon, L; Suzuki, A. Nitrogen uptake, assimilation and remobilization in plants: Challenges for sustainable and productive agriculture. Annals of botany 2010, 105, 1141–1157. [Google Scholar] [CrossRef] [PubMed]
- Medeiros, DB; Aarabi, F; Rivas, FJM; Fernie, AR. The knowns and unknowns of intracellular partitioning of carbon and nitrogen, with focus on the organic acid-mediated interplay between mitochondrion and chloroplast. Journal of Plant Physiology 2021, 266, 153521. [Google Scholar] [CrossRef]
- Meier, S; Bautzmann, R; Komarova, NY; Ernst, V; Suter-Grotemeyer, M; Schröder, K; Haindrich, AC; Fernández, AV; Robert, CAM; Ward, JM; et al. Stress-regulated Arabidopsis GAT2 is a low affinity γ-aminobutyric acid transporter. Journal of Experimental Botany 2024, 75, 6295–6311. [Google Scholar] [CrossRef]
- Melino, VJ; Tester, MA; Okamoto, M. Strategies for engineering improved nitrogen use efficiency in crop plants via redistribution and recycling of organic nitrogen. Current Opinion in Biotechnology 2022, 73, 263–269. [Google Scholar] [CrossRef]
- Meng, R; Li, Z; Kang, X; Zhang, Y; Wang, Y; Ma, Y; Wu, Y; Dong, S; Li, X; Gao, L; et al. High overexpression of SiAAP9 leads to growth inhibition and protein ectopic localization in transgenic Arabidopsis. International Journal of Molecular Sciences 2024, 25, 5840. [Google Scholar] [CrossRef]
- Meyer, A; Eskandari, S; Grallath, S; Rentsch, D. AtGAT1, a high affinity transporter for γ-aminobutyric acid in Arabidopsis thaliana. Journal of Biological Chemistry 2006, 281, 7197–7204. [Google Scholar] [CrossRef]
- Meyer, C; Stitt, M. Nitrate reductase and signalling. In Plant nitrogen; Lea, PJ, Morot-Gaudry, J-F, Eds.; Springer New York, 2001; pp. 37–59. [Google Scholar]
- Millard, P. The accumulation and storage of nitrogen by herbaceous plants. Plant Cell and Environment 1988, 11, 1–8. [Google Scholar] [CrossRef]
- Moll, RH; Kamprath, EJ; Jackson, WA. Jackson Analysis and interpretation of factors which contribute to efficiency of nitrogen utilization. Agronomy Journal 1982, 74, 562564. [Google Scholar] [CrossRef]
- Moller, IA; Rasmusson, AG; Van Aken, O. Plant mitochondria-past, present and future. Plant J 2021, 108, 912–959. [Google Scholar] [CrossRef]
- Müller, B; Fastner, A; Karmann, J; Mansch, V; Hoffmann, T; Schwab, W; Suter-Grotemeyer, M; Rentsch, D; Truernit, E; Ladwig, F; et al. Amino acid export in developing Arabidopsis seeds depends on UmamiT facilitators. Current Biology 2015, 25, 3126–3131. [Google Scholar] [CrossRef] [PubMed]
- Nie, S; Huang, W; He, C; Wu, B; Duan, H; Ruan, J; Zhao, Q; Fang, Z. Transcription factor OsMYB2 triggers amino acid transporter OsANT1 expression to regulate rice growth and salt tolerance. Plant Physiology 2025, 197, kiae559. [Google Scholar] [CrossRef] [PubMed]
- Okumoto, S; Pilot, G. Amino Acid Export in Plants: A Missing Link in Nitrogen Cycling. Molecular Plant 2011, 4, 453–463. [Google Scholar] [CrossRef] [PubMed]
- Okumoto, S; Schmidt, R; Tegeder, M; Fischer, WN; Rentsch, D; Frommer, WB; Koch, W. High affinity amino acid transporters specifically expressed in xylem parenchyma and developing seeds of Arabidopsis. Journal of Biological Chemistry 2002, 277, 45338–45346. [Google Scholar] [CrossRef]
- Omari Alzahrani, F. Genome wide analysis of amino acid transporter superfamily in Solanum lycopersicum. Plants 2021, 10, 289. [Google Scholar] [CrossRef]
- Pan, R; Liu, J; Wang, S; Hu, J. Peroxisomes: Versatile organelles with diverse roles in plants. New Phytologist 2020, 225, 1410–1427. [Google Scholar] [CrossRef]
- Patrick, JW; Offler, CE. Compartmentation of transport and transfer events in developing seeds. Journal of Experimental Botany 2001, 52, 551–564. [Google Scholar] [CrossRef]
- Peng, B; Kong, H; Li, Y; Wang, L; Zhong, M; Su, L; Gao, G; Zhang, Q; Luo, L; Wang, G; et al. OsAAP6 functions as an important regulator of grain protein content and nutritional quality in rice. Nature Communications 2014, 5, 4847. [Google Scholar] [CrossRef]
- Peng, B; Zhang, Q; Liu, Y; Zhao, Q; Zhao, J; Zhang, Z; Sun, X; Peng, J; Sun, Y; Song, X; et al. OsAAP8 mutation leads to significant improvement in the nutritional quality and appearance of rice grains. Molecular Breeding 2024, 44, 34. [Google Scholar] [CrossRef]
- Perchlik, M; Foster, J; Tegeder, M. Different and overlapping functions of Arabidopsis LHT6 and AAP1 transporters in root amino acid uptake. Journal of Experimental Botany 2014, 65, 5193–5204. [Google Scholar] [CrossRef]
- Perchlik, M; Tegeder, M. Improving Plant Nitrogen Use Efficiency through Alteration of Amino Acid Transport Processes. Plant Physiology 2017, 175, 235–247. [Google Scholar] [CrossRef] [PubMed]
- Perchlik, M; Tegeder, M. Leaf amino acid supply affects photosynthetic and plant nitrogen use efficiency under nitrogen stress. Plant Physiology 2018, 178, 174–188. [Google Scholar] [CrossRef] [PubMed]
- Pohlmeyer, K; Soll, J; Steinkamp, T; Hinnah, S; Wagner, R. Isolation and characterization of an amino acid-selective channel protein present in the chloroplastic outer envelope membrane. Proceedings of the National Academy of Sciences 1997, 94, 9504–9509. [Google Scholar] [CrossRef] [PubMed]
- Pratelli, R; Pilot, G. Regulation of amino acid metabolic enzymes and transporters in plants. Journal of Experimental Botany 2014, 65, 5535–5556. [Google Scholar] [CrossRef]
- Pudelski, B; Kraus, S; Soll, J; Philippar, K. The plant PRAT proteins–preprotein and amino acid transport in mitochondria and chloroplasts. Plant Biology 2010, 12, 42–55. [Google Scholar] [CrossRef]
- Rengel, Z; Cakmak, I; White, PJ. Marschner’s Mineral Nutrition of Plants; Academic Press, 2023. [Google Scholar]
- Renné, P; Dreßen, U; Hebbeker, U; Hille, D; Flügge, UI; Westhoff, P; Weber, AP. The Arabidopsis mutant dct is deficient in the plastidic glutamate/malate translocator DiT2. The Plant Journal 2003, 35, 316–331. [Google Scholar] [CrossRef]
- Rentsch, D; Schmidt, S; Tegeder, M. Transporters for uptake and allocation of organic nitrogen compounds in plants. FEBS letters 2007, 581, 2281–2289. [Google Scholar] [CrossRef]
- Rolletschek, H; Hosein, F; Miranda, M; Heim, U; Götz, KP; Schlereth, A; Borisjuk, L; Saalbach, I; Wobus, U; Weber, H. Ectopic expression of an amino acid transporter (VfAAP1) in seeds of Vicia narbonensis and pea increases storage proteins. Plant Physiology 2005, 137, 1236–1249. [Google Scholar] [CrossRef]
- Sanders, A; Collier, R; Trethewy, A; Gould, G; Sieker, R; Tegeder, M. AAP1 regulates import of amino acids into developing Arabidopsis embryos. The Plant Journal 2009, 59, 540–552. [Google Scholar] [CrossRef]
- Santiago, JP; Tegeder, M. Implications of nitrogen phloem loading for carbon metabolism and transport during Arabidopsis development. Journal of Integrative Plant Biology 2017, 59, 409–421. [Google Scholar] [CrossRef] [PubMed]
- Schjoerring, JK; Husted, S; Mäck, G; Mattsson, M. The regulation of ammonium translocation in plants. Journal of Experimental Botany 2002, 53, 883–890. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, R; Stransky, H; Koch, W. The amino acid permease AAP8 is important for early seed development in Arabidopsis thaliana. Planta 2007, 226, 805–813. [Google Scholar] [CrossRef] [PubMed]
- Shin, K; Lee, S; Song, WY; Lee, RA; Lee, I; Ha, K; Koo, JC; Park, SK; Nam, HG; Lee, Y; et al. Genetic identification of ACC-RESISTANT2 reveals involvement of LYSINE HISTIDINE TRANSPORTER1 in the uptake of 1-aminocyclopropane-1-carboxylic acid in Arabidopsis thaliana. Plant and Cell Physiology 2015, 56, 572–582. [Google Scholar] [CrossRef]
- Snowden, CJ; Thomas, B; Baxter, CJ; Smith, JAC; Sweetlove, LJ. A tonoplast Glu/Asp/GABA exchanger that affects tomato fruit amino acid composition. The Plant Journal 2015, 81, 651–660. [Google Scholar] [CrossRef]
- Su, YH; Frommer, WB; Ludewig, U. Molecular and functional characterization of a family of amino acid transporters from Arabidopsis. Plant Physiology 2004, 136, 3104–3113. [Google Scholar] [CrossRef]
- Suzuki, A; Knaff, DB. Glutamate synthase: Structural, mechanistic and regulatory properties, and role in the amino acid metabolism. Photosynthesis Research 2005, 83, 191–217. [Google Scholar] [CrossRef]
- Svennerstam, H; Ganeteg, U; Näsholm, T. Root uptake of cationic amino acids by Arabidopsis depends on functional expression of amino acid permease 5. New Phytologist 2008, 180, 620–630. [Google Scholar] [CrossRef]
- Svennerstam, H; Jämtgård, S; Ahmad, I; Huss-Danell, K; Näsholm, T; Ganeteg, U. Transporters in Arabidopsis roots mediating uptake of amino acids at naturally occurring concentrations. New Phytologist 2011, 191, 459–467. [Google Scholar] [CrossRef]
- Tan, X; Li, K; Wang, Z; Zhu, K; Tan, X; Cao, J. A review of plant vacuoles: Formation, located proteins, and functions. Plants 2019, 8, 327. [Google Scholar] [CrossRef]
- Tegeder, M. Transporters for amino acids in plant cells: Some functions and many unknowns. Current Opinion in Plant Biology 2012, 15, 315–321. [Google Scholar] [CrossRef] [PubMed]
- Tegeder, M. Transporters involved in source to sink partitioning of amino acids and ureides: Opportunities for crop improvement. Journal of Experimental Botany 2014, 65, 1865–1878. [Google Scholar] [CrossRef] [PubMed]
- Tegeder, M; Hammes, UZ. The way out and in: Phloem loading and unloading of amino acids. Current Opinion in Plant Biology 2018, 43, 16–21. [Google Scholar] [CrossRef] [PubMed]
- Tegeder, M; Masclaux-Daubresse, C. Source and sink mechanisms of nitrogen transport and use. New phytologist 2018, 217, 35–53. [Google Scholar] [CrossRef]
- Tegeder, M; Rentsch, D. Uptake and partitioning of amino acids and peptides. Molecular plant 2010, 3, 997–1011. [Google Scholar] [CrossRef]
- Tegeder, M; Rentsch, D; Patrick, JW. Organic carbon and nitrogen transporters. In The plasma membrane; Murphy, AS, Peer, W, Schulz, B, Eds.; Springer Berlin, 2011; pp. 331–352. [Google Scholar]
- Tegeder, M; Ward, JM. Molecular evolution of plant AAP and LHT amino acid transporters. Frontiers in Plant Science 2012, 3, 21. [Google Scholar] [CrossRef]
- Santiago, JP; Pappenberger, C; Hammes, UZ; Tegeder, M.; The SV. UMAMIT44 is a key player in glutamate export from Arabidopsis chloroplasts. The Plant Cell 2024, 36, 1119–1139. [Google Scholar]
- Snyder, R; Tegeder, M.; The SV. Targeting nitrogen metabolism and transport processes to improve plant nitrogen use efficiency. Frontiers in Plant Science 2021, 11, 628366. [Google Scholar] [CrossRef]
- Tian, R; Yang, Y; Chen, M. Genome-wide survey of the amino acid transporter gene family in wheat (Triticum aestivum L.): Identification, expression analysis and response to abiotic stress. International Journal of Biological Macromolecules 2020, 162, 1372–1387. [Google Scholar] [CrossRef]
- Tilman, D; Cassman, KG; Matson, PA; Naylor, R; Polasky, S. Agricultural sustainability and intensive production practices. Nature 2002, 418, 671–677. [Google Scholar] [CrossRef]
- Vallarino, JG; Kubiszewski-Jakubiak, S; Ruf, S; Rößner, M; Timm, S; Bauwe, H; Carrari, F; Rentsch, D; Bock, R; Sweetlove, LJ; et al. Multi-gene metabolic engineering of tomato plants results in increased fruit yield up to 23%. Scientific Reports 2020, 10, 17219. [Google Scholar] [CrossRef]
- van Bueren, ETL; Struik, PC. Diverse concepts of breeding for nitrogen use efficiency. A review. Agronomy for Sustainable Development 2017, 37, 50. [Google Scholar] [CrossRef]
- Vangelatos, I; Vlachakis, D; Sphianopoulou, V; Diallinas, G. Modelling and mutational evidence identify the substrate binding site and functional elements in APC amino acid transporters. Molecular Membrane Biology 2009, 26, 356–370. [Google Scholar] [CrossRef] [PubMed]
- Vega, A; O’Brien, JA; Gutiérrez, RA. Nitrate and hormonal signaling crosstalk for plant growth and development. Current Opinion in Plant Biology 2019, 52, 155–163. [Google Scholar] [CrossRef] [PubMed]
- von Steimker, J; Wendenburg, R; Klemmer, A; Rosaria, M; Fernie, AR; Alseekh, S; Tripodi, P. Genome-wide association analysis and linkage mapping decipher the genetic control of primary metabolites and quality traits in Capsicum. The Plant Journal 2025, 122, e70300. [Google Scholar] [CrossRef] [PubMed]
- Wan, Y; Wang, Y; Shi, Z; Rentsch, D; Ward, JL; Hassall, K; Sparks, CA; Huttly, AK; Buchner, P; Powers, S; et al. Wheat amino acid transporters highly expressed in grain cells regulate amino acid accumulation in grain. PLoS ONE 2021, 16, e0246763. [Google Scholar] [CrossRef]
- Wang, YY; Hsu, PK; Tsay, YF. Uptake, allocation and signaling of nitrate. Trends in Plant Science 2012, 17, 458–467. [Google Scholar] [CrossRef]
- Wang, S; Jiang, X; Chen, G; Wu, W; Xu, C; Du, M; Xiang, S; Cheng, X; Tian, Y; Tan, J; et al. An Elite Haplotype of Nitrogen-Use-Efficiency Gene LHT5 Enhances Salt Tolerance in Rice. Plant Biotechnology Journal 2026, 0, 1–14. [Google Scholar] [CrossRef]
- Wang, T; Li, Y; Huang, Y; Zhao, X; Dong, Z; Jin, W; Huang, W. Amino acid permease 6 regulates grain protein content in maize. The Crop Journal 2022, 10, 1536–1544. [Google Scholar] [CrossRef]
- Wang, J; Wu, B; Lu, K; Wei, Q; Qian, J; Chen, Y; Fang, Z. The amino acid permease 5 (OsAAP5) regulates tiller number and grain yield in rice. Plant Physiology 2019, 180, 1031–1045. [Google Scholar] [CrossRef]
- Wang, S; Yang, Y; Guo, M; Zhong, C; Yan, C; Sun, S. Targeted mutagenesis of amino acid transporter genes for rice quality improvement using the CRISPR/Cas9 system. The Crop Journal 2020, 8, 457–464. [Google Scholar] [CrossRef]
- Werner, D; Gerlitz, N; Stadler, R. A dual switch in phloem unloading during ovule development in Arabidopsis. Protoplasma 2011, 248, 225–235. [Google Scholar] [CrossRef] [PubMed]
- Widhalm, J; Gutensohn, M; Yoo, H; Adebesin, F; Qian, Y; Guo, L; Jaini, R; Lynch, JH; McCoy, RM; Shreve, JT; et al. Identification of a plastidial phenylalanine exporter that influences flux distribution through the phenylalanine biosynthetic network. Nature communications 2015, 6, 8142. [Google Scholar] [CrossRef] [PubMed]
- Williams, LE; Miller, AJ. Transporters responsible for the uptake and partitioning of nitrogenous solutes. Annual Review of Plant Biology 2001, 52, 659–688. [Google Scholar] [CrossRef]
- Winichayakul, S; Roberts, N. Toward sustainable crops: Integrating vegetative (non-seed) lipid storage, carbon-nitrogen dynamics, and redox regulation. Frontiers in Plant Science 2025, 16, 1589127. [Google Scholar] [CrossRef]
- Winter, H; Robinson, DG; Heldt, HW. Subcellular volumes and metabolite concentrations in barley leaves. Planta 1993, 191, 180–190. [Google Scholar] [CrossRef]
- Witte, CP. Urea metabolism in plants. Plant Science 2011, 180, 431–438. [Google Scholar] [CrossRef]
- Wu, M; Wu, S; Chen, Z; Dong, Q; Yan, H; Xiang, Y. Genome-wide survey and expression analysis of the amino acid transporter gene family in poplar. Tree Genetics & Genomes 2015, 11, 83. [Google Scholar] [CrossRef]
- Xu, G; Fan, X; Miller, AJ. Plant Nitrogen Assimilation and Use Efficiency. Annual Review of Plant Biology 2012, 63, 153–182. [Google Scholar] [CrossRef]
- Xu, D; Sanden, NCH; Hansen, LL; Belew, ZM; Madsen, SR; Meyer, L; Jørgensen, ME; Hunziker, P; Veres, D; Crocoll, C; et al. Export of defensive glucosinolates is key for their accumulation in seeds. Nature 2023, 617, 132–138. [Google Scholar] [CrossRef]
- Yadav, UP; Ayre, BG; Bush, DR. Transgenic approaches to altering carbon and nitrogen partitioning in whole plants: Assessing the potential to improve crop yields and nutritional quality. Frontiers in Plant Science 2015, 6, 275. [Google Scholar] [CrossRef] [PubMed]
- Yanagisawa, S; Akiyama, A; Kisaka, H; Uchimiya, H; Miwa, T. Metabolic engineering with Dof1 transcription factor in plants: Improved nitrogen assimilation and growth under low-nitrogen conditions. Proceedings of the National Academy of Sciences 2004, 101, 7833–7838. [Google Scholar] [CrossRef] [PubMed]
- Yang, H; Krebs, M; Stierhof, YD; Ludewig, U. Characterization of the putative amino acid transporter genes AtCAT2, 3 & 4: The tonoplast localized AtCAT2 regulates soluble leaf amino acids. Journal of Plant Physiology 2014, 171, 594–601. [Google Scholar] [PubMed]
- Yang, G; Wei, Q; Huang, H; Xia, J. Amino acid transporters in plant cells: A brief review. Plants 2020, 9, 967. [Google Scholar] [CrossRef]
- Yang, Y; Zhang, Y; Sun, Z; Shen, Z; Li, Y; Guo, Y; Feng, Y; Sun, S; Guo, M; Hu, Z; et al. Knocking out OsAAP11 to improve rice grain quality using CRISPR/Cas9 system. International Journal of Molecular Sciences 2023, 24, 14360. [Google Scholar] [CrossRef]
- Yao, X; Li, H; Nie, J; Liu, H; Guo, Y; Lv, L; Yang, Z; Sui, X. Disruption of the amino acid transporter CsAAP2 inhibits auxin-mediated root development in cucumber. New Phytologist 2023, 239, 639–659. [Google Scholar] [CrossRef]
- Yu, J; Zhen, X; Li, X; Li, N; Xu, F. Increased autophagy of rice can increase yield and nitrogen use efficiency (NUE). Frontiers in Plant Science 2019, 10, 584. [Google Scholar] [CrossRef]
- Zhang, L; Garneau, MG; Majumdar, R; Grant, J; Tegeder, M. Improvement of pea biomass and seed productivity by simultaneous increase of phloem and embryo loading with amino acids. The Plant Journal 2015, 81, 134–146. [Google Scholar] [CrossRef]
- Zhang, Y; Liu, Y; Wang, D; Li, M; Wang, Y; Li, J; Zhu, Y; Wang, Q; Yan, F. Amino acid transporter GmAAP6-like contributes to seed quality and responds to jasmonic acid pathway under MSX toxicity stress. Plant Physiology and Biochemistry 2025a, 219, 109444. [Google Scholar] [CrossRef]
- Zhang, C; Shi, M; Lin, Y; Chen, Q; Shi, X. Effects of two amino acid transporter-like genes on potato growth. Journal of Plant Physiology 2025b, 304, 154408. [Google Scholar] [CrossRef]
- Zhang, L; Tan, Q; Lee, R; Trethewy, A; Lee, YH; Tegeder, M. Altered xylem-phloem transfer of amino acids affects metabolism and leads to increased seed yield and oil content in Arabidopsis. The Plant Cell 2010, 22, 3603–3620. [Google Scholar] [CrossRef]
- Zhao, H; Ma, H; Yu, L; Wang, X; Zhao, J. Genome-wide survey and expression analysis of amino acid transporter gene family in rice (Oryza sativa L.). PloS one 2012, 7, e49210. [Google Scholar] [CrossRef]
- Zhao, C; Pratelli, R; Yu, S; Shelley, B; Collakova, E; Pilot, G. Detailed characterization of the UMAMIT proteins provides insight into their evolution, amino acid transport properties, and role in the plant. Journal Experimental Botany 2021, 72, 6400–6417. [Google Scholar] [CrossRef]
| Family | Subfamilies | Transport | Functions | Examples |
|---|---|---|---|---|
| AAAP | AAP LHT ProT GAT ANT ATL VAAT |
Import (H+/amino acid symporters) |
Root uptake Xylem loading Xylem unloading Xylem-to-phloem transport Phloem loading Seed loading |
AtAAP1, AtLHT1, AtProT2 CsAAP2 AtLHT1 AtAAP2, AtAAP6, OsAAP3 AtAAP8, PsAAP1 AtAAP1, AtAAP8, AtCAT6 |
| APC | CAT ACT |
Import (H+/amino acid symporters) |
Organelle transport | AtCAT2, AtCAT4, SlCAT9 |
| UMAMIT | Multiple clades | Import/export (Bidirectional facilitators) |
Xylem loading Phloem loading Phloem unloading Organelle transport Seed loading |
AtUMAMIT18 AtUMAMIT11, AtUMAMIT14, AtUMAMIT18 AtUMAMIT11, AtUMAMIT14 AtUMAMIT24, AtUMAMIT44 AtUMAMIT28, AtUMAMIT29 |
| Species | Gene | Function/mechanism | Phenotype | Reference | |
|---|---|---|---|---|---|
| Crop yield | Rice Rice Wheat Pea |
OsAAP1 OsAAP5 TaAAP13 PsAAP1 |
Amino acid transport to grain Amino acid transport to grain Grain sink strength Amino acid partitioning |
Tillering. Grain number and yield Tillering. Grain number and yield Grain size Seed number and yield |
Ji et al. 2020 Wang et al. 2019 Wan et al. 2021 Zhang et al. 2015 |
| NUE | Pea Soybean Rice Rice |
PsAAP1 GmAAP6a OsAAP15 OsLHT1 |
N uptake, assimilation and transport Source to sink N transport Panicle development Amino acid uptake and transport |
NUpE and NUtE NUtE NUpE NUpE and NUtE |
Perchlik and Tegeder 2017 Liu et al. 2020 Yang et al. 2023 Guo et al. 2020 |
| Nutritional quality | Rice Wheat Maize Tomato |
OsAAP6 TaAAP13 ZmAAP6 SlCAT9 |
Amino acid unloading in grain Amino acid unloading in endosperm Amino acid unloading in grain Amino acid storage in vacuole |
Grain protein content Grain amino acid content Grain protein content Fruit amino acid content |
Peng et al. 2014 Wan et al. 2021 Wang et al. 2022 Snowden et al. 2015 |
| NUE: Nitrogen Use Efficiency; NUpE: Nitrogen Uptake Efficiency; NUtE: Nitrogen utilization Efficiency | |||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




