Submitted:
04 April 2026
Posted:
07 April 2026
You are already at the latest version
Abstract
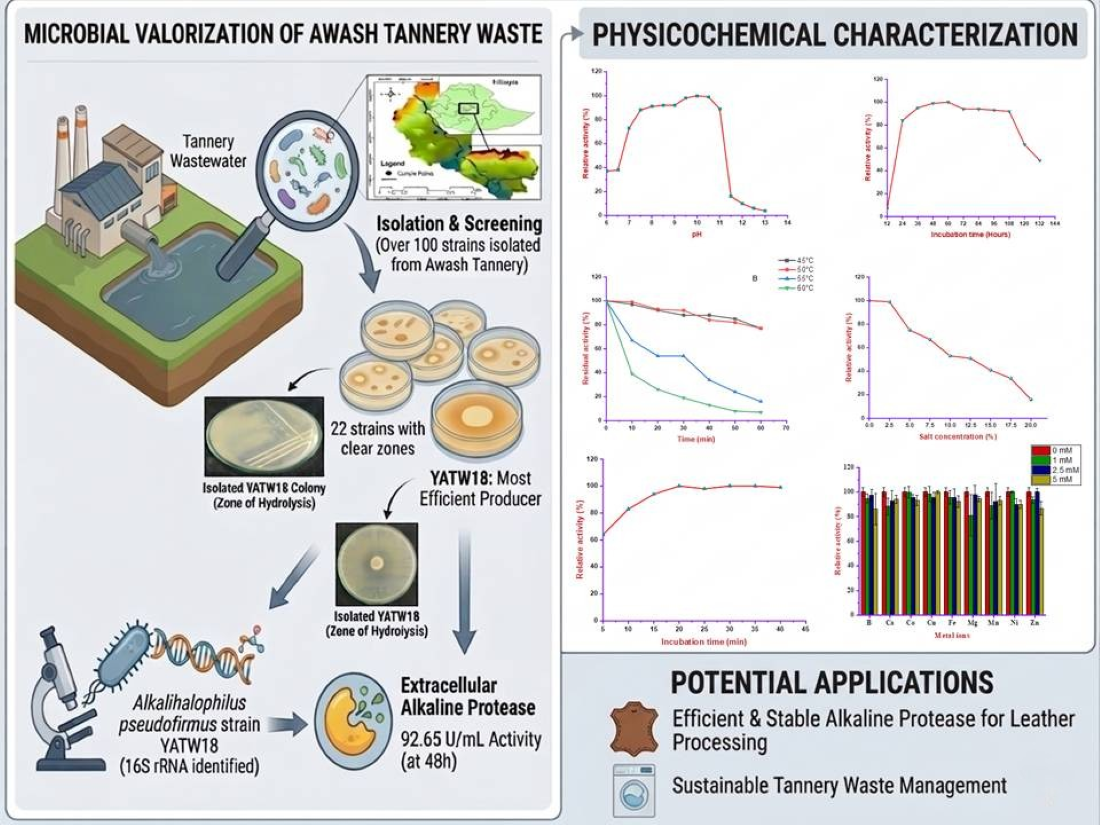
Keywords:
1. Introduction
2. Materials and Methods
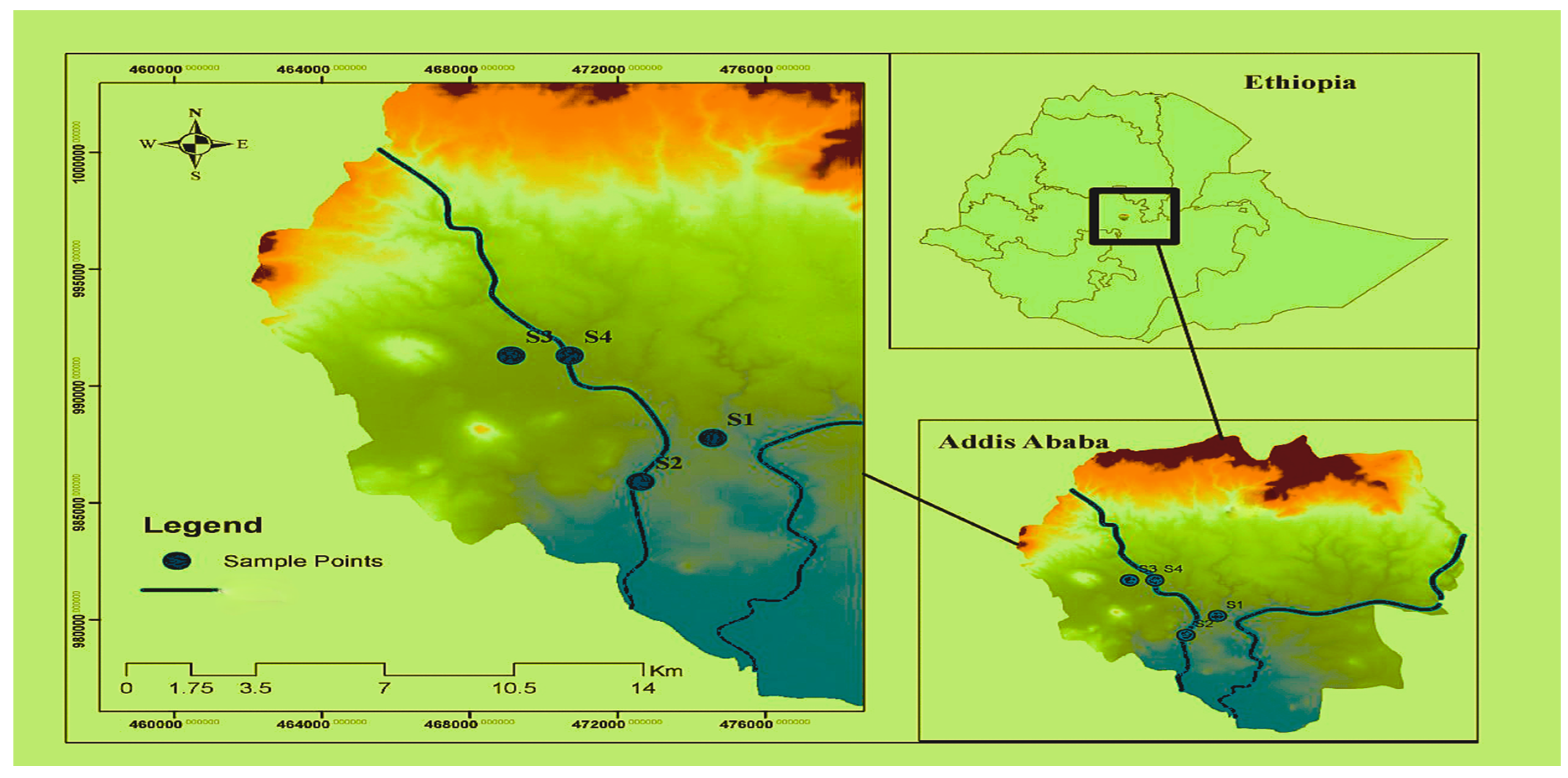
2.1. Sample Area Description
2.2. Sample Collection
2.3. Enrichment of the Samples
2.4. Isolation and Screening of Alkaline Protease Producing Bacteria
2.5. Production of Alkaline Protease
2.6. Molecular Identification
2.6.1. Genomic DNA Extraction, PCR Amplification, Restriction Digestion and Sequencing
2.6.2. Sequence Analysis
2.7. Physicochemical Characterization of Alkaline Protease
2.7.1. Enzyme Activity
2.7.2. Tyrosine Standard Curve Preparation
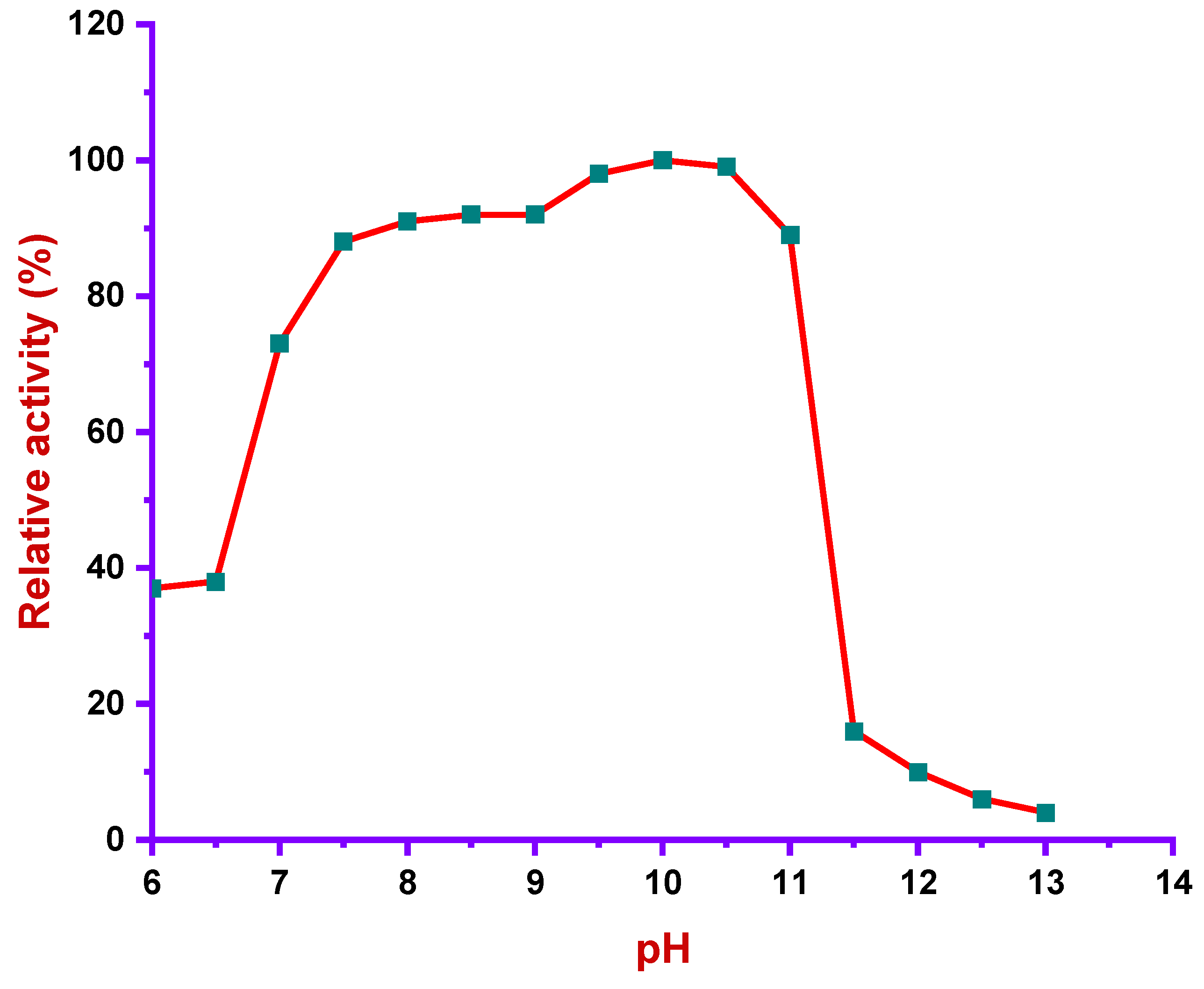
2.7.3. pH Profile
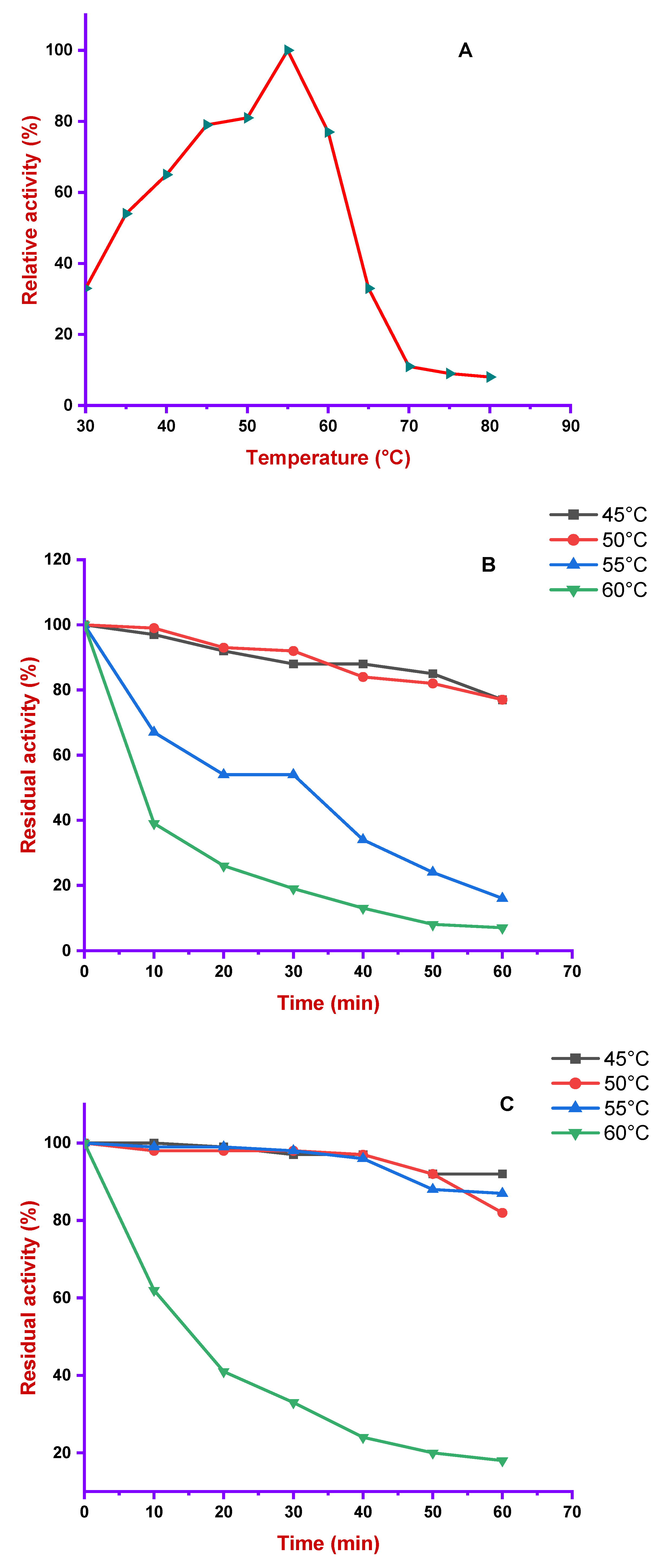
2.7.4. Temperature Profile and Stability
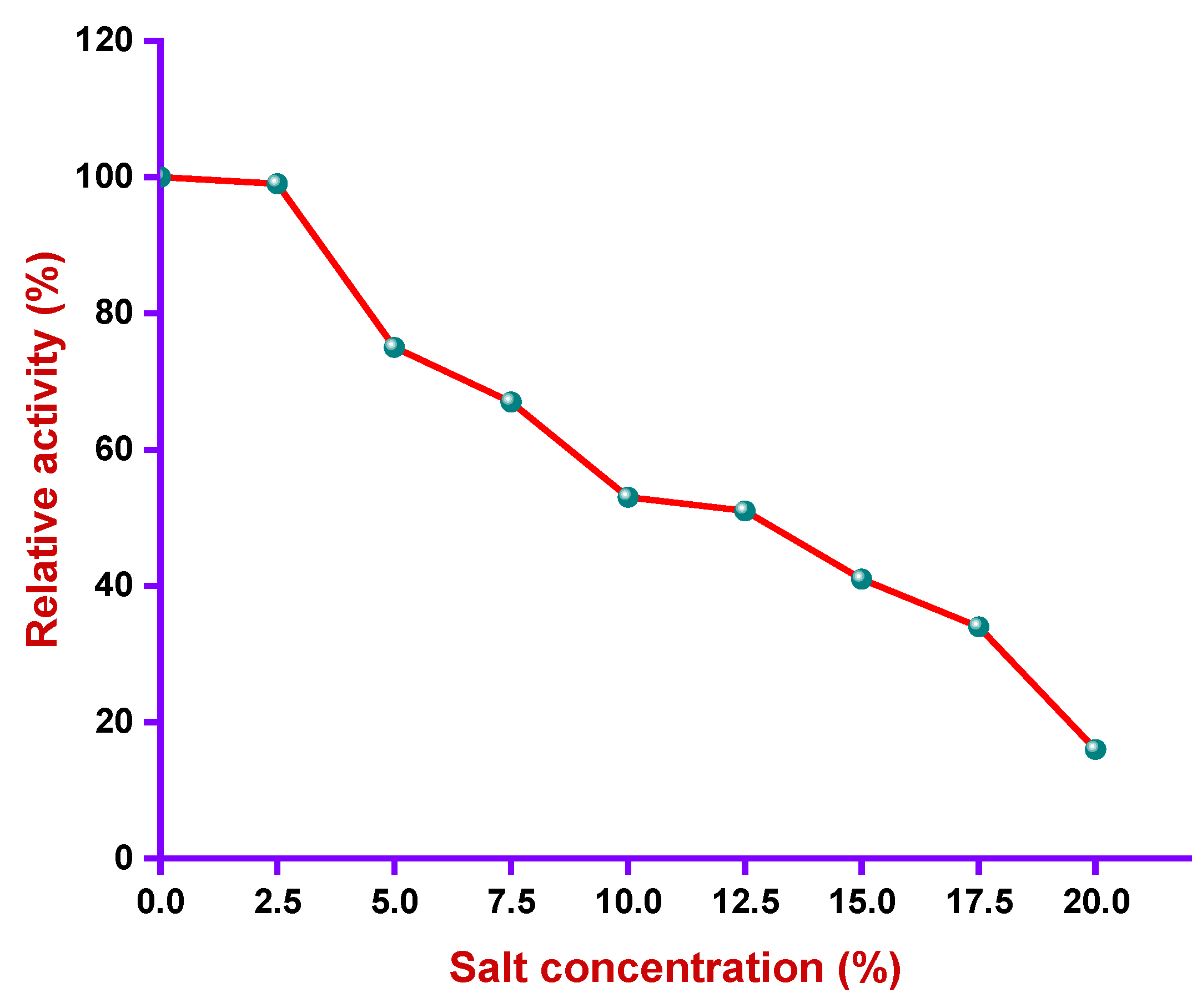
2.7.5. Salt Profile
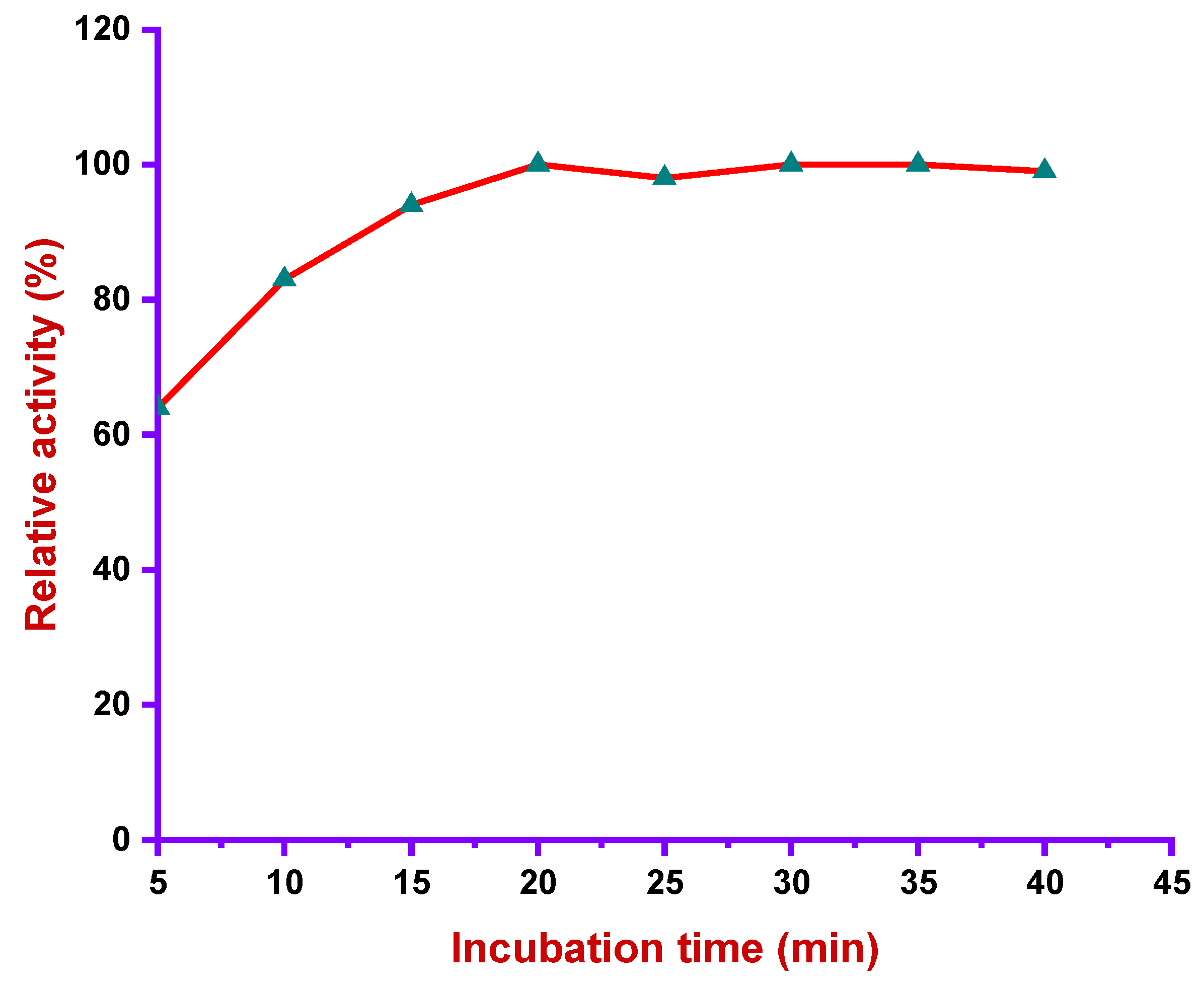
2.7.6. Incubation Time
2.7.7. Metal Ions Profile
3. Results
3.1. Physicochemical Characteristics of the Tannery Waste Water
3.2. Isolation and Screening of Alkaline Protease Producing Bacteria
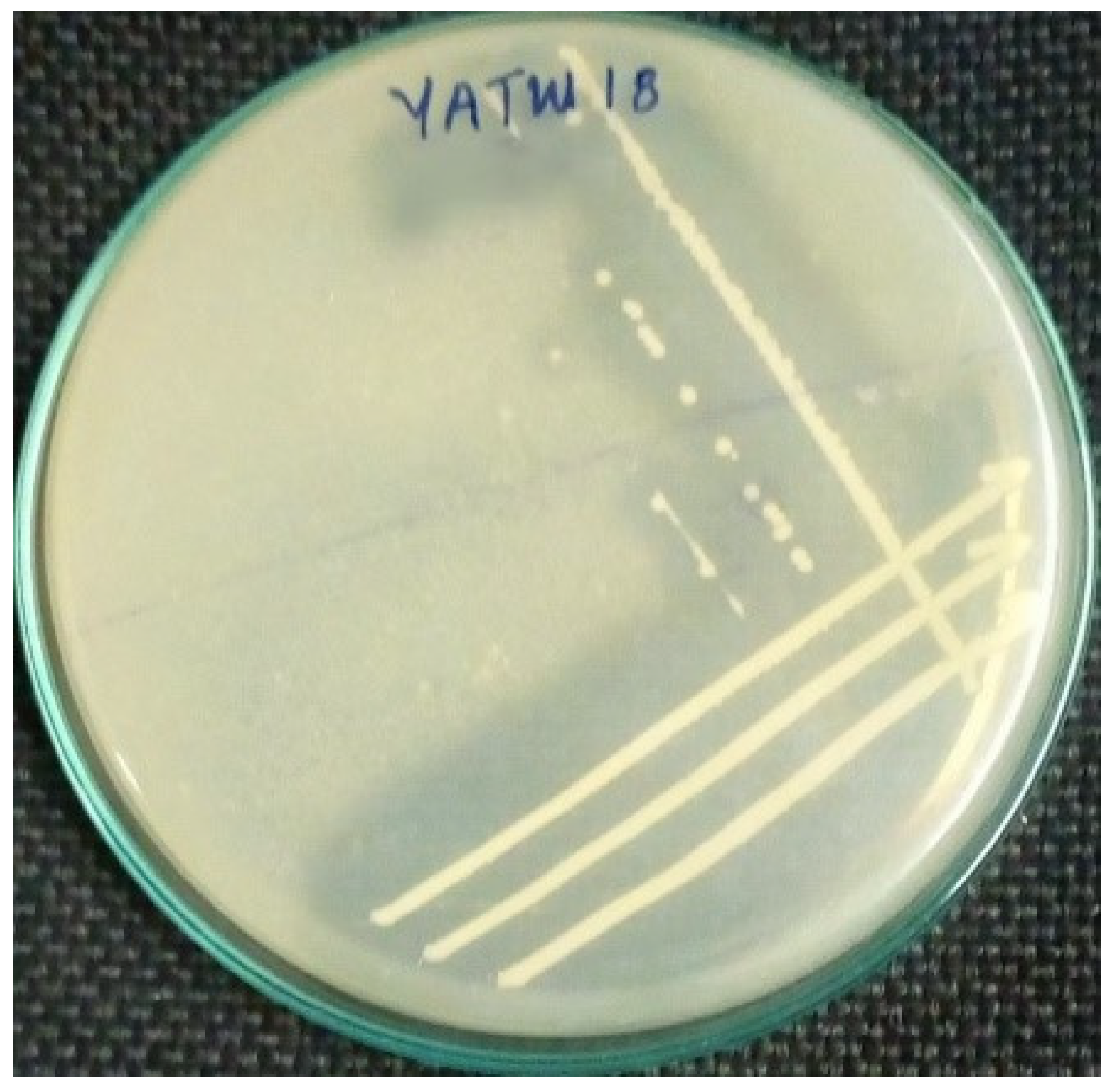
3.2.1. Isolation and Primary Screening
3.2.2. Secondary Screening
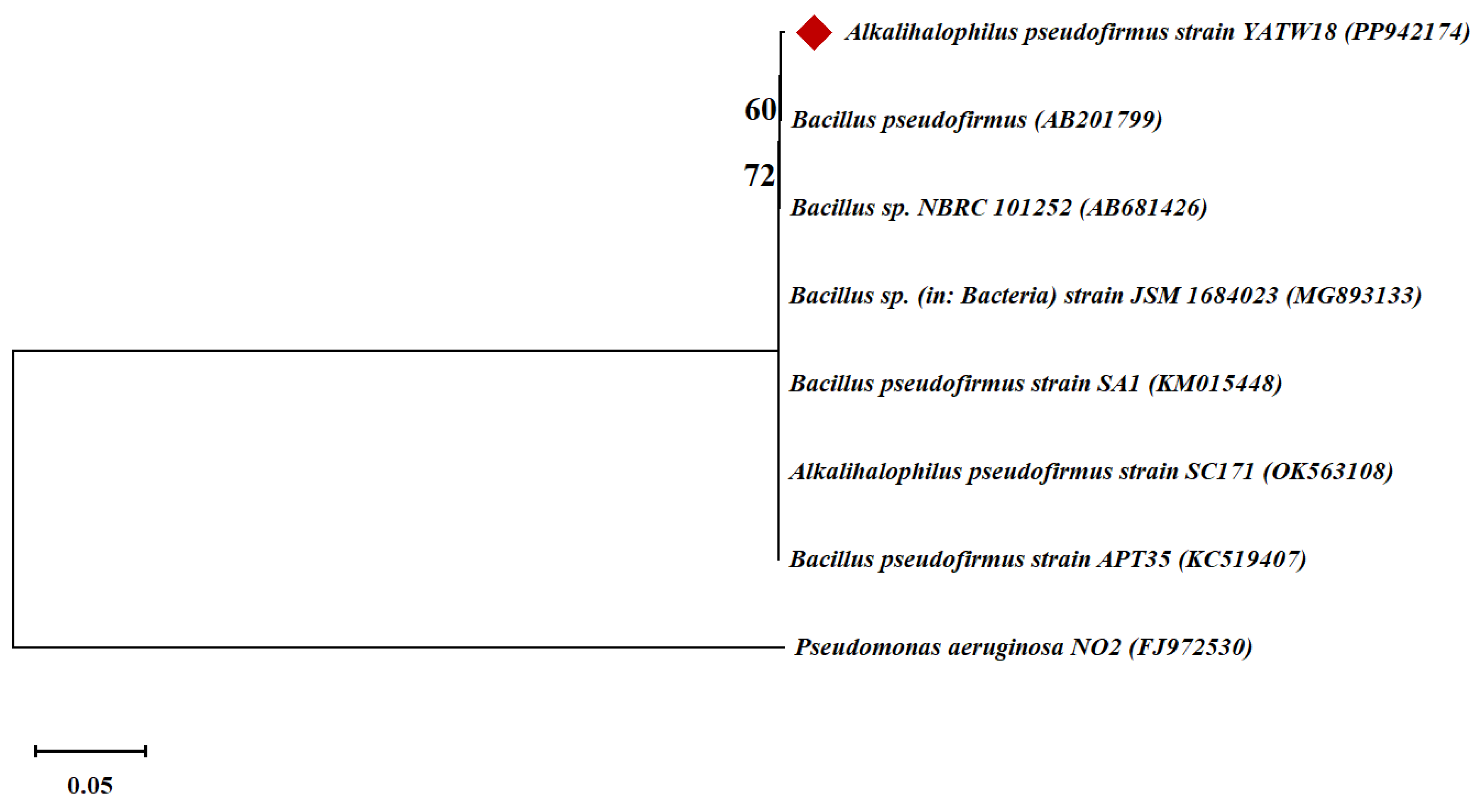
3.3. Molecular Identification
3.4. Physicochemical Characterization of Alkaline Protease
3.4.1. Effect of pH in Alkaliphilic Protease Activity
3.4.2. Effect of Temperature on Alkaliphilic Protease Activity and Stability
3.4.3. Effect of Salt Concentration on Alkaliphilic Protease Activity
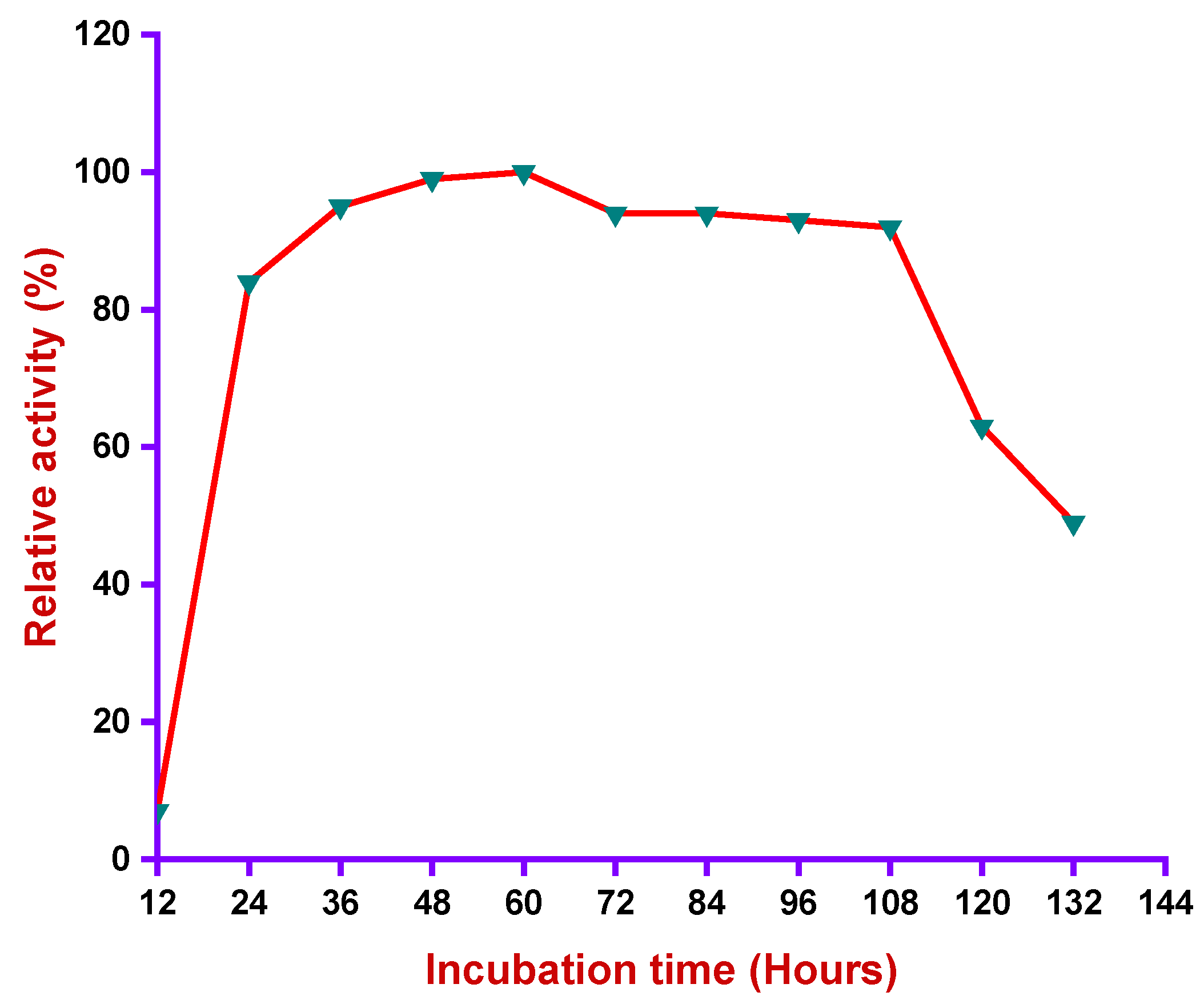
3.4.4. Effect of Incubation Time in Alkaliphilic Protease Activity
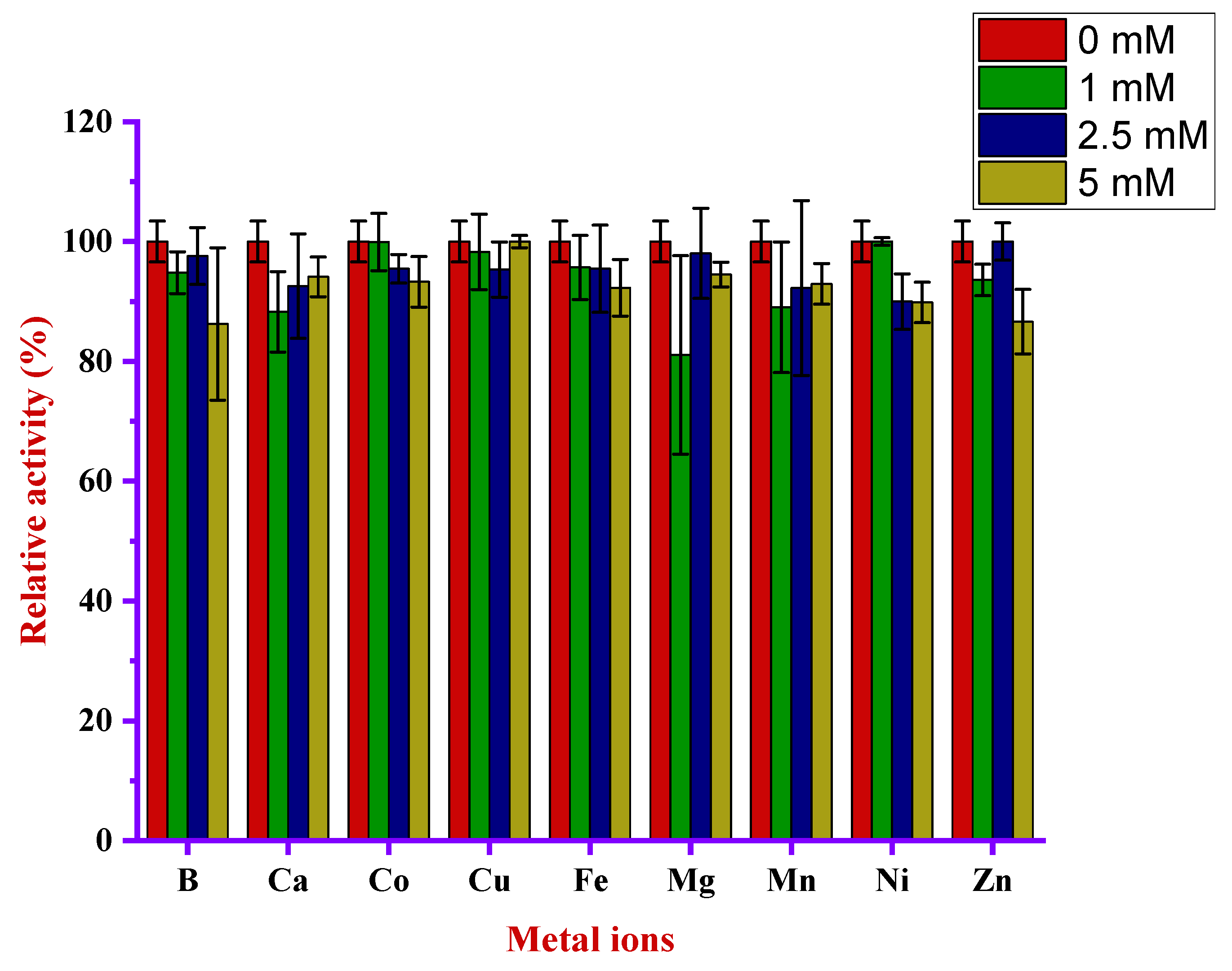
3.4.5. Effect of Metal Ions in Alkaliphilic Protease Activity
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ahsan, S.; Liaqat, A.; Khaliq, A.; Iqbal, R.; Chughtai, M.; Mehmood, T.; Sameed, N. Current Trends and Prospects of Transforming Animal Waste into Food. In Climate Changes Mitigation and Sustainable Bioenergy Harvest through Animal Waste; Springer Nature: Cham, Switzerland, 2023; pp. 469–503.
- Mohamed, A.G.; El Mehalawy, A.A.; Mohamed, S.S. Loofah immobilized with Cladosporium cladosporioides CEL14 is a potential bio-remediating agent for hexavalent chromium in tannery wastewater. Malays. J. Microbiol. 2021, 17, 5.
- Nadeem, M.; Qazi, J.; Syed, Q.; Gulsher, M. Purification and characterization of an alkaline protease from Bacillus licheniformis UV-9 for detergent formulations. Songklanakarin J. Sci. Technol. 2013, 35, 2.
- Khan, Z.; Shafique, M.; Jabeen, N.; Naz, S.; Yasmeen, K.; Ejaz, U.; Sohail, M. Protease from Bacillus subtilis ZMS-2: Evaluation of production dynamics through Response Surface Methodology and application in leather tannery. J. King Saud Univ. Sci. 2023, 35, 102643.
- Amde, A. Environmental impact of Awash leather industry and tannery in Ethiopia. Ethiop. J. Environ. Stud. Manag. 2019, 12, 95–104.
- Gnanavelu, A.; Shanmuganathan, T.; Deepesh, V.; Suresh, S. Validation of a Modified Procedure for the determination of Chemical Oxygen Demand using standard dichromate method in industrial wastewater samples with high calcium chloride content. Indian J. Sci. Technol. 2021, 14, 2391–2399.
- Bennett, V.; MarkManuel, D. Winkler Titration Assessment of Biochemical Oxygen Demand (BOD5) in Oka Creek, Toru-Orua, Bayelsa State. Asian J. Appl. Chem. Res. 2024, 15, 147–156.
- Tiwari, N.; Singh, R.; Gupta, N. Enrichment and isolation of protease-producing alkaliphiles. Biotechnol. J. 2015, 14, 412–417.
- Gessesse, A.; Hatti-Kaul, R.; Gashe, B. Novel alkaline proteases from alkaliphilic bacteria: Purification and characterization. Enzyme Microb. Technol. 2003, 32, 589–594.
- Surzycki, S. Extraction and purification of nucleic acids using CTAB method. In Basic Techniques in Molecular Biology; Springer: Berlin/Heidelberg, Germany, 2012; pp. 163–168.
- Cropper, A. Quantitative assessment of DNA: A spectrophotometric and electrophoretic approach. J. Genet. Res. 2019, 42, 155–162.
- Brukner, I.; Paliouras, M.; Trifiro, M.; Bohbot, M.; Shamir, D.; Kirk, A. Assessing Different PCR Master Mixes for Ultrarapid DNA Amplification: Important Analytical Parameters. Diagnostics 2024, 14, 477.
- Bartlett, J.; Stirling, D. PCR Protocols, 2nd ed.; Humana Press: Totowa, NJ, USA, 2008.
- Altschul, S.; Gish, W.; Miller, W.; Myers, E.; Lipman, D. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410.
- Jukes, T.H.; Cantor, C.R. Evolution of protein molecules. Mamm. Protein Metab. 1969, 3, 132.
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549.
- Felsenstein, J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution 1985, 39, 783–791.
- Rawski, R.; Sanecki, P.; Kalembkiewicz, J. Units and Methods of Proteolytic Activity Determination. Curr. Pharm. Anal. 2020, 16, 661–670.
- Nielsen, P.; Fritze, D.; Priest, F.G. Phenetic diversity of alkaliphilic Bacillus strains: proposal for Alkalihalophilus pseudofirmus sp. nov. Nord. Microbiol. 1995, 45, 120–127.
- Ullah, N.; Mujaddad-ur-Rehman, M.; Sarwar, A.; Nadeem, M.; Nelofer, R.; Irfan, M.; Aziz, T. Effect of bioprocess parameters on alkaline protease production by locally isolated Bacillus cereus AUST-7 using tannery waste in submerged fermentation. Biomass Convers. Biorefin. 2023, 14, 22977–22987.
- Masi, C.; Gemechu, G.; Tafesse, M. Isolation, screening, characterization, and identification of alkaline protease-producing bacteria from leather industry effluent. Ann. Microbiol. 2021, 71, 1–11.
- Singh, K.; Kumari, M.; Prasad, K. Tannery effluents: current practices, environmental consequences, human health risks, and treatment options. CLEAN–Soil Air Water 2023, 51, 2200303.
- Jiang, X.; Xu, X.; Yang, M.; Zhang, M. Emerging applications of solid-state fermentation in bioengineering and biopharmaceuticals. Curr. Opin. Biotechnol. 2021, 70, 54–59.
- Kumar, R.; Tiwari, P.; Jany, K. Comparative production of proteases in Bacillus sp. under submerged fermentation. J. Environ. Biol. 2012, 33, 203–207.
- Gupta, R.; Beg, Q.; Lorenz, P. Bacterial alkaline proteases: Molecular approaches and industrial applications. Appl. Microbiol. Biotechnol. 2002, 59, 15–32.
- Razzaq, A.; Shamsi, S.; Ali, A.; Ali, Q.; Sajjad, M.; Malik, A.; Ashraf, M. Microbial proteases applications. Front. Bioeng. Biotechnol. 2019, 7, 110.
- Wanyonyi, W.W.; Mulaa, F.J. Alkaline proteases: Advances and applications in the leather industry. Enzyme Eng. 2019, 8, 1–10.
- Sharma, R.; Bhatt, M.; Agrawal, A. A comparative study of chemical and enzyme-assisted leather dehairing. Environ. Technol. Innov. 2018, 9, 235–241.
- Taylor, R. Identifying intermolecular atom-atom interactions that are not just bonding but also competitive. CrystEngComm 2020, 22, 7145–7151.
- Govindasamy, V.; Kumar, M. Thermostable alkaline protease production from Bacillus licheniformis under solid-state fermentation: Purification and characterization. J. Genet. Eng. Biotechnol. 2021, 19, 1–10.
- Kieliszek, M.; Pobiega, K.; Piwowarek, K.; Kot, A. Characteristics of the proteolytic enzymes produced by lactic acid bacteria. Molecules 2021, 26, 1858.
- Cortez, D.; Bekke, M.; Liang, Z.; Stamminger, R. The impact of detergent performance on sustainable consumer laundry behavior: a socio-technical challenge. Tenside Surfactants Deterg. 2024, 61, 203–215.
- Stoner, M.R. Mechanistic studies of enzyme degradation in liquid detergent. PhD Thesis, University of Colorado at Boulder, USA, 2004.
- Sivakumar, V. Towards environmental protection and process safety in leather processing–A comprehensive analysis and review. Process Saf. Environ. Prot. 2022, 163, 703–726.
- Galante, Y.; Formantici, C. Enzyme applications in detergency and in manufacturing industries. Curr. Org. Chem. 2003, 7, 1399–1422.
- Samidurai, S.; Khambhaty, Y.; Alagamuthu, T. Bio-preservation of raw hides/skins: A review on greener substitute to conventional salt curing. Environ. Sci. Pollut. Res. 2022, 29, 64513–64535.
- Punekar, N.S. Enzymes: Catalysis, Kinetics and Mechanisms; Springer: Mumbai, India, 2018.
- Romero, E.; Gómez Castellanos, J.; Gadda, G.; Fraaije, M.; Mattevi, A. Same substrate, many reactions: oxygen activation in flavoenzymes. Chem. Rev. 2018, 118, 1742–1769.
- Callender, R.; Dyer, R. The dynamical nature of enzymatic catalysis. Acc. Chem. Res. 2015, 48, 407–413.
- Chapman, J.; Ismail, A.; Dinu, C. Industrial applications of enzymes: Recent advances, techniques, and outlooks. Catalysts 2018, 8, 238.
- Andreini, C.; Bertini, I.; Cavallaro, G.; Holliday, G.; Thornton, J. Metal ions in biological catalysis: From enzyme databases to general principles. J. Biol. Inorg. Chem. 2012, 17, 1091–1098.
- Shahid, M.; Noman, M.; Ahmed, T. Enzyme stabilization: Stabilization of proteases for industrial applications. Crit. Rev. Biotechnol. 2017, 37, 309–322.
- Baldiris, R.; Pulido, P.; Pulido, H.; Montes, J.; Cardozo, A.; Marrugo-Negrete, J. Effects of manganese on oxidative stress in the brain. Neurotox. Res. 2019, 35, 664–670.
- Jakubowski, M.; Kuciel, R.; Chelstowska, A. Heavy metals and enzyme activity: Influence on enzymes and their removal from industrial wastewater. Water Environ. Res. 2011, 83, 278–289.

| No. | Parameters | Measured value | Site of measurement |
| 1. | pH | 8.1 | On-site measurement |
| 2. | Temperature | 28 °C | |
| 3. | Salinity | 1% | |
| 4. | TDS | 9.63 ppt | Off-site measurement |
| 5. | TSS | 1044.7 mg/L | |
| 6. | COD | 1276 mg/L | |
| 7. | BOD | 375.4 mg/L |
| No. | Bacterial isolates | Colony diameter (mm) | Clear zone diameter (mm) | Protolytic index (ratio) |
| 1. | YATW1 | 3.7 | 18.7 | 5.1 |
| 2. | YATW2 | 3 | 19.7 | 6.6 |
| 3. | YATW3 | 3 | 17.3 | 5.8 |
| 4. | YATW4 | 3.3 | 21.3 | 6.5 |
| 5. | YATW5 | 3.7 | 9.3 | 2.5 |
| 6. | YATW6 | 3 | 22 | 7.3 |
| 7. | YATW7 | 2.7 | 20.7 | 7.7 |
| 8. | YATW8 | 3.3 | 20.7 | 6.3 |
| 9. | YATW9 | 2.7 | 21.3 | 7.9 |
| 10. | YATW10 | 2.7 | 17.3 | 6.4 |
| 11. | YATW11 | 3.3 | 22 | 6.7 |
| 12. | YATW12 | 3.7 | 22 | 5.9 |
| 13. | YATW13 | 2.7 | 21.3 | 7.1 |
| 14. | YATW14 | 3 | 8.7 | 2.9 |
| 15. | YATW15 | 3.3 | 21.3 | 6.5 |
| 16. | YATW16 | 2.7 | 8 | 3 |
| 17. | YATW17 | 3 | 9.3 | 3.1 |
| 18. | YATW18 | 3 | 20.7 | 6.9 |
| 19. | YATW19 | 3.7 | 22 | 5.9 |
| 20. | YATW20 | 4 | 20.7 | 5.2 |
| 21. | YATW21 | 2.7 | 20 | 7.4 |
| 22. | YATW22 | 3.3 | 17.3 | 5.2 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




