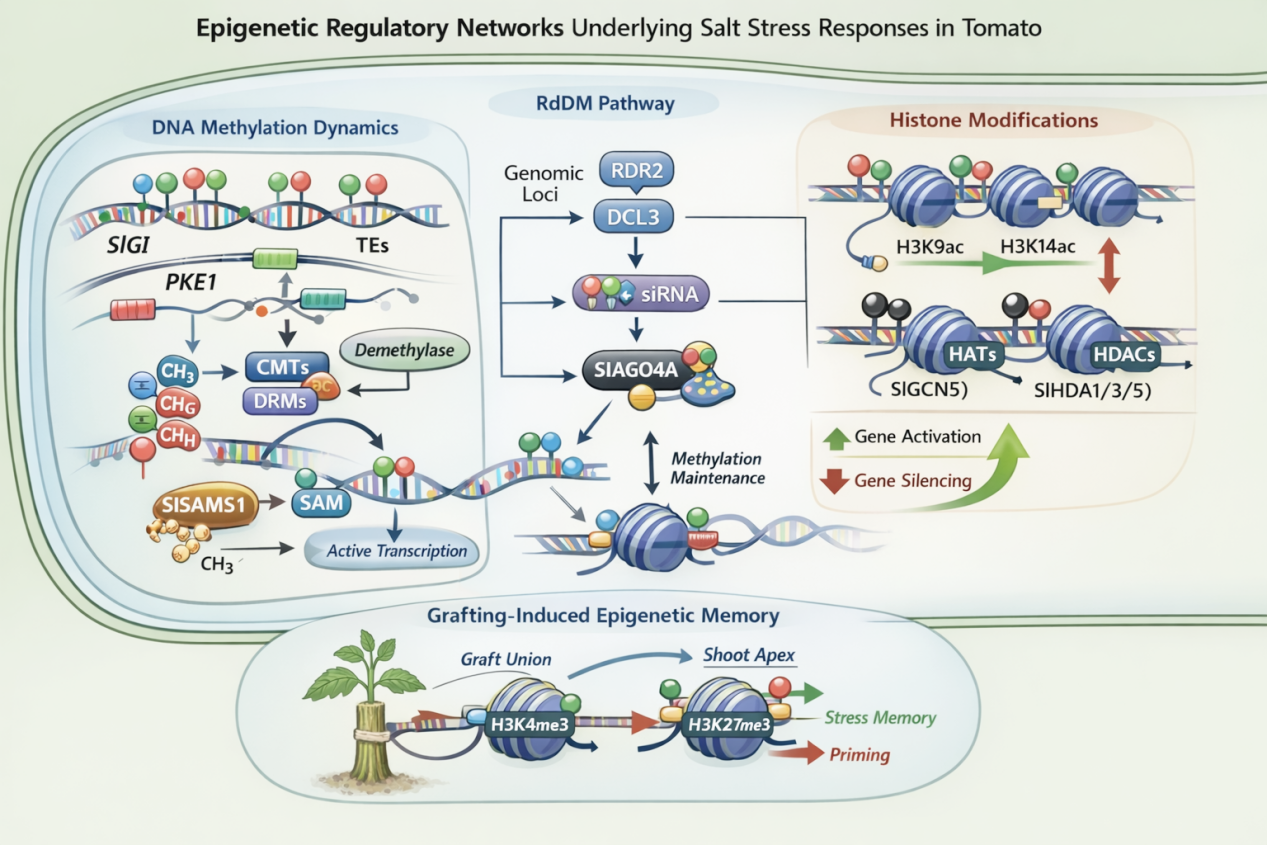
Soil salinization is increasingly threatening global agricultural productivity and food security, currently affecting over 6% of the world’s land and one-third of irrigated areas. Tomato (Solanum lycopersicum L.), a major vegetable crop worldwide, exhibits moderate sensitivity to salinity, which limits both its yield and fruit quality. In recent years, epigenetic regulation has gained attention as a key mechanism enabling flexible and reversible control of gene expression without altering DNA sequences. This review synthesizes current knowledge on the epigenetic control of salt stress responses in tomato, focusing on three interconnected levels: DNA methylation dynamics, RNA-directed DNA methylation (RdDM), and histone modifications. We explore how DNA methyltransferases reshape the methylome under salinity, using examples such as PKE1 and SlGI to illustrate functional gene-body methylation. The RdDM pathway is discussed with emphasis on the unexpected role of SlAGO4A as a negative modulator of stress tolerance and the growing evidence for RdDM-mediated regulation of transcription factors. We also examine the balanced regulation of histone acetylation and deacetylation, highlighting the conserved role of GCN5 in maintaining cell wall integrity and the diverse functions of HDACs (SlHDA1, SlHDA3, SlHDA5) in stress adaptation. Additionally, insights from wild tomato species and grafting-induced epigenetic changes are presented, revealing new dimensions of stress memory. Collectively, these epigenetic mechanisms constitute a complex regulatory framework that integrates stress responses with growth and development, providing potential targets for epigenetic breeding of salt-tolerant tomatoes.