Submitted:
01 April 2026
Posted:
03 April 2026
You are already at the latest version
Abstract
Keywords:
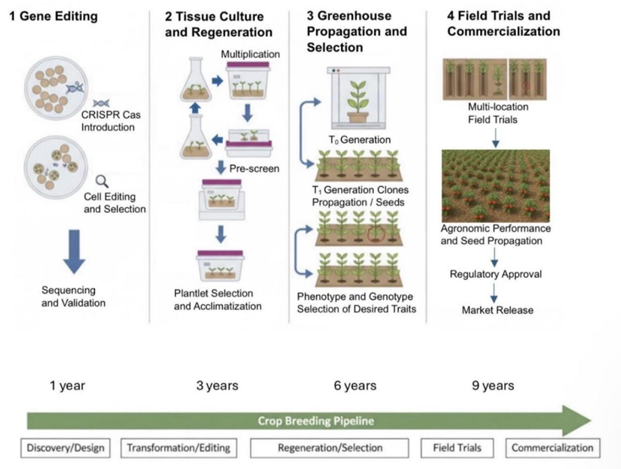
Introduction
Crop Applications
RNA Adaptive Immunity in Plants
CRISPR-Cas Characterization
CRISPR-Cas Delivery
Abiotic Stress Resilience: Single Genes to Network Modulation
Elevating Biotic Resilience: Immunity and Ecological Interactions
Phenotype and Agronomic Performance Improvement
Future Applications of Gene Editing in Plants
Conclusions
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
References
- Ishino, Y.; Shinagawa, H.; Makino, K.; Amemura, M.; Nakata, A. Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli and identification of the gene product. J. Bacteriol. 1987, 169, 5429–5433. [Google Scholar] [CrossRef] [PubMed]
- Jinek, M.; Chylinski, K.; Fonfara, I.; Hauer, M.; Doudna, J.A.; Charpentier, E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 2012, 337, 816–821. [Google Scholar] [CrossRef]
- Klug, A.; Rhodes, D. Zinc fingers: a novel protein fold for nucleic acid recognition. Cold Spring Harb. Symp. Quant. Biol. 1987, 52, 473–482. [Google Scholar] [CrossRef]
- Boch, J. TALEs of genome targeting. Nat. Biotechnol. 2011, 29, 135–136. [Google Scholar] [CrossRef]
- Mojica, F.J.; Montoliu, L. On the origin of CRISPR-Cas technology: from prokaryotes to mammals. Trends Microbiol. 2016, 24, 811–820. [Google Scholar] [CrossRef]
- Mojica, F.J.; Rodriguez-Valera, F. The discovery of CRISPR in archaea and bacteria. FEBS J. 2016, 283, 3162–3169. [Google Scholar] [CrossRef] [PubMed]
- Shah, S.A.; Erdmann, S.; Mojica, F.J.; Garrett, R.A. Protospacer recognition motifs: mixed identities and functional diversity. RNA Biol. 2013, 10, 891–899. [Google Scholar] [CrossRef] [PubMed]
- Mojica, F. J.; Díez-Villaseñor, C.; Soria, E.; Juez, G. Biological significance of a family of regularly spaced repeats in the genomes of archaea, bacteria and mitochondria. Mol. Microbiol. 2000, 36, 244–246. [Google Scholar] [CrossRef]
- Bolotin, A.; Quinquis, B.; Sorokin, A.; Ehrlich, S.D. Clustered regularly interspaced short palindrome repeats (CRISPRs) have spacers of extrachromosomal origin. Microbiol. 2005, 151, 2551–2561. [Google Scholar] [CrossRef]
- Mojica, F. J.; Díez-Villaseñor, C.; García-Martínez, J.; Soria, E. Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J. Mol. Evol. 2005, 60, 174–182. [Google Scholar] [CrossRef]
- Pourcel, C.; Salvignol, G.; Vergnaud, G. CRISPR elements in Yersinia pestis acquire new repeats by preferential uptake of bacteriophage DNA, and provide additional tools for evolutionary studies. Microbiol. 2005, 151, 653–663. [Google Scholar] [CrossRef]
- Barrangou, R.; Fremaux, C.; Deveau, H.; Richards, M.; Boyaval, P.; Moineau, S.; Romero, D. A.; Horvath, P. CRISPR provides acquired resistance against viruses in prokaryotes. Science 2007, 315, 1709–1712. [Google Scholar] [CrossRef]
- Brouns, S. J. J.; Jore, M. M.; Lundgren, M.; Westra, E. R.; Slijkhuis, R. J. H.; Snijders, A. P. L.; Dickman, M. J.; Makarova, K. S.; Koonin, E. V.; van der Oost, J. Small CRISPR RNAs guide antiviral defense in prokaryotes. Science 2008, 321, 960–964. [Google Scholar] [CrossRef]
- Jansen, R.; van Embden, J.D.; Gaastra, W.; Schouls, L.M. Identification of genes that are associated with DNA repeats in prokaryotes. Mol. Microbiol. 2002, 43, 1565–1575. [Google Scholar] [CrossRef]
- Feng, Z.; Zhang, B.; Ding, W.; Liu, X.; Yang, D.-L.; Wei, P.; Cao, F.; Zhu, S.; Zhang, F.; Mao, Y.; Zhu, J.-K. Efficient genome editing in plants using a CRISPR/Cas system. Cell Res. 2013, 23, 1229–1232. [Google Scholar] [CrossRef] [PubMed]
- Shan, Q.; Wang, Y.; Li, J.; Zhang, Y.; Chen, K.; Liang, Z.; Zhang, K.; Liu, J.; Xi, J.J.; Qui, J.L.; Gao, C. Targeted genome modification of crop plants using a CRISPR-Cas system. Nat Biotechnol. 2013, 31, 686–688. [Google Scholar] [CrossRef] [PubMed]
- Li, J.-F.; Norville, J.E.; Aach, J.; McCormack, M.; Zhang, D.; Bush, J.; Church, G.M.; Sheem, J. Multiplex and homologous recombination-mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nat Biotechnol. 2013, 31, 688–691. [Google Scholar] [CrossRef] [PubMed]
- Maher, M.F.; Nasti, R.A.; Vollbrecht, M.; Starker, C.G.; Clark, M.D.; Voytas, D.F. Plant gene editing through de novo induction of meristems. Nat. Biotechnol. 2020, 38, 84–89. [Google Scholar] [CrossRef]
- Komor, A.C.; Kim, Y.B.; Packer, M.S.; Zuris, J.A.; Liu, D.R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 2016, 533, 420–424. [Google Scholar] [CrossRef]
- Abudayyeh, O.O.; Gootenberg, J.S.; Konermann, S.; Joung, J.; Slaymaker, I.M.; Cox, D.B.T.; Shmakov, S.; Makarova, K.S.; Semenova, E.; Minakhin, L.; Severinov, K.; Regev, A.; Lander, E.S.; Koonin, E.V.; Zhang, F. C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 2016, 353, aaf5573. [Google Scholar] [CrossRef]
- Fire, A.; Xu, S.; Montgomery, M.K.; Kostas, S.A.; Driver, S.E.; Mello, C.C. Potent and specific genetic interference by double- stranded RNA in Caenorhabditis elegans. Nature 1998, 391, 806–811. [Google Scholar] [CrossRef]
- Napoli, C.; Lemieux, C.; Jorgensen, R. Introduction of a chimeric chalcone synthase gene into Petunia results in reversible co-suppression of homologous genes in trans. Plant Cell 1990, 2, 279–289. [Google Scholar] [CrossRef]
- Kawchuk, L.M.; Martin, R.R.; McPherson, J. Sense and antisense RNA-mediated resistance to potato leafroll virus in Russet Burbank potato plants. Mol. Plant Microbe Interact. 1991, 4, 247–253. [Google Scholar] [CrossRef]
- Mitter, N.; Worrall, E.A.; Robinson, K.E.; Li, P.; Jain, R.G.; Taochy, C.; Fletcher, S.J.; Carroll, B.J.; Lu, G.Q.M.; Xu, Z.P. Clay nanosheets for topical delivery of RNAi for sustained protection against plant viruses. Nat. Plants 2017, 3, 16207. [Google Scholar] [CrossRef] [PubMed]
- Koeppe, S.; Kawchuk, L.; Kalischuk, M. RNA interference past and future applications in plants. Int. J. Mol. Sci. 2023, 24, 9755. [Google Scholar] [CrossRef] [PubMed]
- Nekrasov, V.; Staskawicz, B.; Weigel, D.; Jones, J.D.G.; Kamoun, S. Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas9 RNA-guided endonuclease. Nat. Biotechnol. 2013, 31, 691–693. [Google Scholar] [CrossRef] [PubMed]
- Ow, D.W.; Wood, K.V.; Deluca, M.; De Wet, J.R.; Helinski, D.R.; Howell, S.H. Transient and stable expression of the firefly luciferase gene in plant cells and transgenic plants. Science 1986, 234, 856–859. [Google Scholar] [CrossRef]
- Shimomura, O.; Johnson, F.H.; Saiga, Y. Extraction, purification and properties of aequorin, a bioluminescent protein from the luminous hydromedusan, Aequorea. J. Cell. Comp. Physiol. 1962, 59, 223–239. [Google Scholar] [CrossRef]
- Qin, G.; Gu, H.; Ma, L.; Peng, Y.; Deng, X.W.; Chen, Z.; Qu, L.-J. Disruption of phytoene desaturase gene results in albino and dwarf phenotypes in Arabidopsis by impairing chlorophyll, carotenoid, and gibberellin biosynthesis. Cell Res. 2007, 17, 471–482. [Google Scholar] [CrossRef]
- Kay, R.; Chan, A.; Daly, M.; McPherson, J. Duplication of CaMV 35S promoter sequences creates a strong enhancer for plant genes. Science 1987, 236, 1299–1302. [Google Scholar] [CrossRef]
- Kawchuk, L.M.; Lynch, D.R.; Martin, R.R.; Kozub, G.C.; Farries, B. Field resistance to the potato leafroll luteovirus in transgenic and somaclone potato plants reduces tuber disease symptoms. Can. J. Plant Path. 1997, 19, 260–266. [Google Scholar] [CrossRef]
- Kawchuk, L.M.; Nielsen, J. Improved bunt inoculation of wheat utilizing adventitious shoots. Can. J. Botany 1987, 65, 1284–1285. [Google Scholar] [CrossRef]
- Weber, E.; Engler, C.; Gruetzner, R.; Werner, S.; Marillonnet, S. A modular cloning system for standardized assembly of multigene constructs. PLoS One 2011, 6, e16765. [Google Scholar] [CrossRef] [PubMed]
- Belhaj, K.; Chaparro-Garcia, A.; Kamoun, S.; Nekrasov, V. Plant genome editing made easy: targeted mutagenesis in model and crop plants using the CRISPR/Cas system. Plant Methods 2013, 9, 39. [Google Scholar] [CrossRef]
- Osakabe, Y.; Watanabe, T.; Sugano, S.S.; Ueta, R.; Ishihara, R.; Shinozaki, K.; Osakabe, K. Optimization of CRISPR/Cas9 genome editing to modify abiotic stress responses in plants. Sci. Rep. 2016, 6, 26685. [Google Scholar] [CrossRef]
- Paixão, J.F.R.; Gillet, F.-X.; Ribeiro, T.P.; Bournaud, C.; Lourenço-Tessutti, I.T.; Noriega, D.D.; Melo, B.P.; de Almeida-Engler, J.; Grossi-de-Sa, M.F. Improved drought stress tolerance in Arabidopsis by CRISPR/dCas9 fusion with a Histone AcetylTransferase. Sci Rep. 2019, 9, 8080. [Google Scholar] [CrossRef]
- Du, Y.-T.; Zhao, M.-J.; Wang, C.-T.; Gao, Y.; Wang, Y.-X.; Liu, Y.-W.; Chen, M.; Chen, J.; Zhou, Y.-B.; Xu, Z.-S.; Ma, Y.-Z. Identification and characterization of GmMYB118 responses to drought and salt stress. BMC Plant Biol. 2018, 18, 320. [Google Scholar] [CrossRef]
- Ogata, T.; Ishizaki, T.; Fujita, M.; Fujita, Y. CRISPR/Cas9-targeted mutagenesis of OsERA1 confers enhanced responses to abscisic acid and drought stress and increased primary root growth under non-stressed conditions in rice. PLoS ONE 2020, 15, e0234156. [Google Scholar] [CrossRef]
- Kim, M.-S.; Ko, S.-R.; Jung, Y.J.; Kang, K.-K.; Lee, Y.-J.; Cho, Y.-G. Knockout mutants of OsPUB7 generated using CRISPR/Cas9 revealed abiotic stress tolerance in rice. Int. J. Mol. Sci. 2023, 24, 5338. [Google Scholar] [CrossRef]
- Zhao, J.; Yao, B.; Peng, Z.; Yang, X.; Li, K.; Zhang, X.; Zhu, H.; Zhou, X.; Wang, M.; Jiang, L.; He, X.; Liang, Y.; Zhan, X.; Wang, X.; Dai, Y.; Yang, Y.; Yang, A.; Dong, M.; Shi, S.; Lu, M.; Du, Y. Splicing defect of StDRO2 intron 1 promotes potato root growth by disturbing auxin transport to adapt to drought stress. Hortic. Plant J. 2025, 11, 706–720. [Google Scholar] [CrossRef]
- Wang, X.; Guo, Y.; Wang, Y.; Peng, Y.; Zhang, H.; Zheng, J. ZmHDT103 negatively regulates drought stress tolerance in maize seedlings. Agronomy 2024, 14, 134. [Google Scholar] [CrossRef]
- Li, P.; Li, X.; Jiang, M. CRISPR/Cas9-mediated mutagenesis of WRKY3 and WRKY4 function decreases salt and Me-JA stress tolerance in Arabidopsis thaliana. Mol. Biol. Rep. 2021, 48, 5821–5832. [Google Scholar] [CrossRef]
- Kim, S.T.; Choi, M.; Bae, S.J.; Kim, J.S. The functional association of Acqos/Victr with salt stress resistance in Arabidopsis thaliana was confirmed by CRISPR-mediated mutagenesis. Int. J. Mol. Sci. 2021, 22, 11389. [Google Scholar] [CrossRef]
- Curtin, S.J.; Xiong, Y.; Michno, J.M.; Campbell, B.W.; Stec, A.O.; Cermak, T.; Starker, C.; Voytas, D.F.; Eamens, A.L.; Stupar, R.M. CRISPR/Cas9 and TALENs generate heritable mutations for genes involved in small RNA processing of Glycine max and Medicago truncatula. Plant Biotechnol. J. 2018, 16, 1125–1137. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Xun, H.; Wang, W.; Ding, X.; Tian, H.; Hussain, S.; Dong, Q.; Li, Y.; Cheng, Y.; Wang, C.; Lin, R.; Li, G.; Qian, X.; Pang, J.; Feng, X.; Dong, Y.; Liu, B.; Wang, S. Mutation of GmAITR genes by CRISPR/Cas9 genome editing results in enhanced salinity stress tolerance in soybean. Front. Plant Sci. 2021, 12, 779598. [Google Scholar] [CrossRef]
- Liu, L.; Wang, J.; Zhang, Q.; Sun, T.; Wang, P. Cloning of the soybean GmNHL1 gene and functional analysis under salt stress. Plants 2023, 12, 3869. [Google Scholar] [CrossRef] [PubMed]
- Yang, R.; Ma, Y.; Yang, Z.; Pu, Y.; Liu, M.; Du, J.; Xu, Z.; Xu, Z.; Zhang, S.; Zhang, H.; Zhang, W.; Yu, D.; Kan, G. Knockdown of β-conglycinin α′ and α subunits alters seed protein composition and improves salt tolerance in soybean. Plant J. 2024, 120, 1488–1507. [Google Scholar] [CrossRef]
- Kloc, Y.; Dmochowska-Boguta, M.; Zebrowska-Rozanska, P.; Laczmanski, L.; Nadolska-Orczyk, A.; Orczyk, W. HvGSK1.1 controls salt tolerance and yield through the brassinosteroid signaling pathway in barley. Int. J. Mol. Sci. 2024, 25, 998. [Google Scholar] [CrossRef] [PubMed]
- Curtin, S.J.; Xiong, Y.; Michno, J.M.; Campbell, B.W.; Stec, A.O.; Čermák, T.; Starker, C.; Voytas, D.F.; Eamens, A.L.; Stupar, R.M. CRISPR/Cas9 and TALENs generate heritable mutations for genes involved in small RNA processing of Glycine max and Medicago truncatula. Plant Biotechnol. J. 2017, 16, 1125–1137. [Google Scholar] [CrossRef]
- Kumar, V.V.S.; Verma, R.K.; Yadav, S.K.; Yadav, P.; Watts, A.; Rao, M.V.; Chinnusamy, V. CRISPR-Cas9 mediated genome editing of drought and salt tolerance (OsDST) gene in indica mega rice cultivar MTU1010. Physiol. Mol. Biol. Plants 2020, 26, 1099–1110. [Google Scholar] [CrossRef]
- Duan, Y.B.; Li, J.; Qin, R.Y.; et al. Identification of a regulatory element responsible for salt induction of rice OsRAV2 through ex situ and in situ promoter analysis. Plant Mol. Biol. 2016, 90, 49–62. [Google Scholar] [CrossRef]
- Zhang, A.; Liu, Y.; Wang, F.; et al. Enhanced rice salinity tolerance via CRISPR/Cas9-targeted mutagenesis of the OsRR22 gene. Mol. Breed. 2019, 39, 47. [Google Scholar] [CrossRef]
- Zhang, X.; Long, Y.; Huang, J.; Xia, J. OsNAC45 is involved in ABA response and salt tolerance in rice. Rice 2020, 13, 79. [Google Scholar] [CrossRef]
- Lan, T.; Zheng, Y.; Su, Z.; Yu, S.; Song, H.; Zheng, X.; Lin, G.; Wu, W. OsSPL10, an SBP-box gene, plays a dual role in salt tolerance and trichome formation in rice (Oryza sativa L.). G3: Genes Genomes Genet. 2019, 9, 4107–4114. [Google Scholar] [CrossRef]
- Ye, N.; Wang, Y.; Yu, H.; Qin, Z.; Zhang, J.; Duan, M.; Liu, L. Abscisic acid enhances trehalose content via OsTPP3 to improve salt tolerance in rice seedlings. Plants 2023, 12, 2665. [Google Scholar] [CrossRef]
- Tran, M.T.; Doan, D.T.H.; Kim, J.; Song, Y.J.; Sung, Y.W.; Das, S.; Kim, E.J.; Son, G.H.; Kim, S.H.; Vu, T.V.; Kim, J.Y. CRISPR/Cas9-based precise excision of SlHyPRP1 domain(s) to obtain salt stress-tolerant tomato. Plant Cell Rep. 2021, 40, 999–1011. [Google Scholar] [CrossRef]
- Xiang, Y.; Jimenez-Gomez, J.M. SlHAK20: a new player in plant salt tolerance. EMBO J. 2020, 39, e104997. [Google Scholar] [CrossRef] [PubMed]
- Makhotenko, A.V.; Khromov, A.V.; Snigir, E.A.; Makarova, S.S.; Makarov, V.V.; Suprunova, T.P.; Kalinina, N.O.; Taliansky, M.E. Functional analysis of coilin in virus resistance and stress tolerance of potato (Solanum tuberosum) using CRISPR-Cas9 editing. Dokl. Biochem. Biophys. 2019, 484, 88–91. [Google Scholar] [CrossRef] [PubMed]
- Zheng, M.; Lin, J.; Liu, X.; Chu, W.; Li, J.; Gao, Y.; An, K.; Song, W.; Xin, M.; Yao, Y.; Peng, H.; Ni, Z.; Sun, Q.; Hu, Z. Histone acetyltransferase TaHAG1 acts as a crucial regulator to strengthen salt tolerance of hexaploid wheat. Plant Physiol. 2021, 186, 1951–1969. [Google Scholar] [CrossRef] [PubMed]
- Cao, Y.; Liang, X.; Yin, P.; Zhang, M.; Jiang, C. A domestication-associated reduction in K⁺-preferring HKT transporter activity underlies maize shoot K⁺ accumulation and salt tolerance. New Phytol. 2018, 222, 301–317. [Google Scholar] [CrossRef]
- Li, R.; Liu, C.; Zhao, R.; Wang, L.; Chen, L.; Yu, W.; Zhang, S.; Sheng, J.; Shen, L. CRISPR/Cas9-mediated SlNPR1 mutagenesis reduces tomato plant drought tolerance. BMC Plant Biol. 2019, 19, 38. [Google Scholar] [CrossRef]
- Bertier, L.D.; Ron, M.; Huo, H.; Bradford, K.J.; Britt, A.B.; Michelmore, R.W. High-resolution analysis of the efficiency, heritability, and editing outcomes of CRISPR/Cas9-induced modifications of NCED4 in lettuce (Lactuca sativa). G3 (Bethesda) 2018, 8, 1513–1521. [Google Scholar] [CrossRef]
- Nandy, S.; Pathak, B.; Zhao, S.; Srivastava, V. Heat-shock-inducible CRISPR/Cas9 system generates heritable mutations in rice. Plant Direct 2019, 3, e00145. [Google Scholar] [CrossRef]
- Malzahn, A.A.; Tang, X.; Lee, K.; Ren, Q.; Sretenovic, S.; Zhang, Y.; Chen, H.; Kang, M.; Bao, Y.; Zheng, X.; Deng, K.; Zhang, T.; Salcedo, V.; Wang, K.; Zhang, Y.; Qi, Y. Application of CRISPR-Cas12a temperature sensitivity for improved genome editing in rice, maize, and Arabidopsis. BMC Biol. 2019, 17, 9. [Google Scholar] [CrossRef]
- Nguyen, T.T.; Pham, D.T.; Nguyen, N.H.; Do, P.T.; To, H.T.M. The germin-like protein gene OsGER4 is involved in heat stress response in rice root development. J. Biosci. 2023, 48, 1201. [Google Scholar] [CrossRef]
- Miao, C.; Xiao, L.; Hua, K.; Zou, C.; Zhao, Y.; Bressan, R.A.; Zhu, J.K. Mutations in a subfamily of abscisic acid receptor genes promote rice growth and productivity. Proc. Natl. Acad. Sci. U.S.A. 2018, 115, 6058–6063. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; He, M.; Li, J.; Chen, L.; Huang, Z.; Zheng, S.; Zhu, L.; Ni, E.; Jiang, D.; Zhao, B.; Zhuang, C. Development of commercial thermo-sensitive genic male sterile rice accelerates hybrid rice breeding using the CRISPR/Cas9-mediated TMS5 editing system. Sci. Rep. 2016, 6, 37395. [Google Scholar] [CrossRef] [PubMed]
- Klap, C.; Yeshayahou, E.; Bolger, A.M.; Arazi, T.; Gupta, S.K.; Shabtai, S.; Usadel, B.; Salts, Y.; Barg, R. Tomato facultative parthenocarpy results from SlAGAMOUS-LIKE 6 loss of function. Plant Biotechnol. J. 2017, 15, 634–647. [Google Scholar] [CrossRef]
- Hu, Z.; Li, J.; Ding, S.; Cheng, F.; Li, X.; Jiang, Y.; Yu, J.; Foyer, C.H.; Shi, K. The protein kinase CPK28 phosphorylates ascorbate peroxidase and enhances thermotolerance in tomato. Plant Physiol. 2021, 186, 1302–1317. [Google Scholar] [CrossRef] [PubMed]
- Yu, W.; Wang, L.; Zhao, R.; Sheng, J.; Zhang, S.; Li, R.; Shen, L. Knockout of SlMAPK3 enhances tolerance to heat stress involving ROS homeostasis in tomato plants. BMC Plant Biol. 2019, 19, 354. [Google Scholar] [CrossRef]
- Wang, H.; Feng, M.; Jiang, Y.; Du, D.; Dong, C.; Zhang, Z.; Wang, W.; Liu, J.; Liu, X.; Li, S. Thermosensitive SUMOylation of TaHsfA1 defines a dynamic ON/OFF molecular switch for the heat stress response in wheat. Plant Cell 2023, 35, 3889–3910. [Google Scholar] [CrossRef]
- Li, J.; Zhang, H.; Si, X.; Tian, Y.; Chen, K.; Liu, J.; Chen, H.; Gao, C. Generation of thermosensitive male-sterile maize by targeted knockout of the ZmTMS5 gene. J. Genet. Genomics 2017, 44, 465–468. [Google Scholar] [CrossRef]
- Li, Z.; Li, Z.; Ji, Y.; Wang, C.; Wang, S.; Shi, Y.; Le, J.; Zhang, M. The heat shock factor 20–HSF4–cellulose synthase A2 module regulates heat stress tolerance in maize. Plant Cell 2024, 36, 2652–2667. [Google Scholar] [CrossRef] [PubMed]
- Nawaz, G.; Han, Y.; Usman, B.; Liu, F.; Qin, B.; Li, R. Knockout of OsPRP1, a gene encoding proline-rich protein, confers enhanced cold sensitivity in rice (Oryza sativa L.) at the seedling stage. 3 Biotech 2019, 9, 254. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Xiang, Z.; Li, J.; Wang, S.; Chen, Y.; Liu, Y.; Mao, D.; Luan, S.; Chen, L. bHLH57 confers chilling tolerance and grain yield improvement in rice. Plant Cell Environ. 2023, 46, 1402–1418. [Google Scholar] [CrossRef]
- Li, R.; Zhang, L.; Wang, L.; Chen, L.; Zhao, R.; Sheng, J.; Shen, L. Reduction of tomato-plant chilling tolerance by CRISPR-Cas9-mediated SlCBF1 mutagenesis. J. Agric. Food Chem. 2018, 66, 9042–9051. [Google Scholar] [CrossRef]
- Yasmeen, A.; Shakoor, S.; Azam, S.; Bakhsh, A.; Shahid, N.; Latif, A.; Shahid, A.A.; Husnain, T.; Rao, A.Q. CRISPR/Cas-mediated knockdown of vacuolar invertase gene expression lowers the cold-induced sweetening in potatoes. Planta 2022, 256, 107. [Google Scholar] [CrossRef]
- Zhang, N.; Wang, S.; Zhao, S.; Chen, D.; Tian, H.; Li, J.; Zhang, L.; Li, S.; Liu, L.; Shi, C.; Yu, X.; Ren, Y.; Chen, F. Global crotonylatome and GWAS revealed a TaSRT1–TaPGK model regulating wheat cold tolerance through mediating pyruvate. Sci. Adv. 2023, 9, eadg1012. [Google Scholar] [CrossRef]
- Li, X.; Cai, Q.; Yu, T.; Li, S.; Li, S.; Li, Y.; Sun, Y.; Ren, H.; Zhang, J.; Zhao, Y.; Zhang, J.; Zuo, Y. ZmG6PDH1 in glucose-6-phosphate dehydrogenase family enhances cold stress tolerance in maize. Front. Plant Sci. 2023, 14, 1116237. [Google Scholar] [CrossRef]
- Baeg, G.J.; Kim, S.H.; Choi, D.M.; Tripathi, S. CRISPR/Cas9-mediated mutation of 5-oxoprolinase gene confers resistance to sulfonamide compounds in Arabidopsis. Plant Biotechnol. Rep. 2021, 15, 803–812. [Google Scholar] [CrossRef]
- Wang, F.Z.; Chen, M.X.; Yu, L.J.; Xie, L.J.; Yuan, L.B.; Qi, H.; Xiao, M.; Guo, W.; Chen, Z.; Yi, K.; Zhang, J.; Qiu, R.; Shu, W.; Xiao, S.; Chen, Q.F. OsARM1, an R2R3 MYB transcription factor, is involved in regulation of the response to arsenic stress in rice. Front. Plant Sci. 2017, 8, 1868. [Google Scholar] [CrossRef] [PubMed]
- Nieves-Cordones, M.; Mohamed, S.; Tanoi, K.; Kobayashi, N.I.; Takagi, K.; Vernet, A.; Guiderdoni, E.; Périn, C.; Sentenac, H.; Véry, A.A. Production of low-Cs⁺ rice plants by inactivation of the K⁺ transporter OsHAK1 with the CRISPR-Cas system. Plant J. 2017, 92, 43–56. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Ye, R.; Liang, Y.; Zhang, S.; Liu, X.; Sun, C.; Li, F.; Yi, J. Generation of low-cadmium rice germplasms via knockout of OsLCD using CRISPR/Cas9. J. Environ. Sci. 2023, 126, 138–152. [Google Scholar] [CrossRef]
- Lu, H.P.; Liu, S.M.; Xu, S.L.; Chen, W.Y.; Zhou, X.; Tan, Y.Y.; Huang, J.Z.; Shu, Q.Y. CRISPR-S: an active interference element for a rapid and inexpensive selection of genome-edited, transgene-free rice plants. Plant Biotechnol. J. 2017, 15, 1371–1373. [Google Scholar] [CrossRef]
- Ding, J.; Ji, C.; Yu, L.; Wang, C.; Ding, G.; Wang, S.; Shi, L.; Xu, F.; Cai, H. OsMYB84, a transcriptional regulator of OsCOPT2 and OsHMA5, modulates copper uptake and transport and yield production in rice. The Crop J. 2024, 12, 456–469. [Google Scholar] [CrossRef]
- Sun, S.-K.; Chen, Y.; Che, J.; Konishi, N.; Tang, Z.; Miller, A.J.; Ma, J.F.; Zhao, F.-J. Decreasing arsenic accumulation in rice by overexpressing OsNIP1;1 and OsNIP3;3 through disrupting arsenite radial transport in roots. New Phytol 2018, 219, 641–653. [Google Scholar] [CrossRef]
- Tang, L.; Mao, B.; Li, Y.; Lv, Q.; Zhang, L.; Chen, C.; He, H.; Wang, W.; Zeng, X.; Shao, Y.; Pan, Y.; Hu, Y.; Peng, Y.; Fu, X.; Li, H.; Xia, S.; Zhao, B. Knockout of OsNramp5 using the CRISPR/Cas9 system produces low Cd-accumulating indica rice without compromising yield. Sci. Rep. 2017, 7, 14438. [Google Scholar] [CrossRef]
- Chu, C.; Huang, R.; Liu, L.; Tang, G.; Xiao, J.; Yoo, H.; Yuan, M. The rice heavy-metal transporter OsNRAMP1 regulates disease resistance by modulating ROS homoeostasis. Plant Cell Environ. 2022, 45, 1109–1126. [Google Scholar] [CrossRef] [PubMed]
- Tan, L.; Qu, M.; Zhu, Y.; Peng, C.; Wang, J.; Gao, D.; Chen, C. ZINC TRANSPORTER5 and ZINC TRANSPORTER9 function synergistically in zinc/cadmium uptake. Plant Physiol. 2020, 183, 1235–1249. [Google Scholar] [CrossRef]
- Mao, X.; Zheng, Y.; Xiao, K.; Wei, Y.; Zhu, Y.; Cai, Q.; Chen, L.; Xie, H.; Zhang, J. OsPRX2 contributes to stomatal closure and improves potassium deficiency tolerance in rice. Biochem. Biophys. Res. Commun. 2018, 495, 461–467. [Google Scholar] [CrossRef]
- Zhang, Y.; Chen, K.; Zhao, F.J.; Sun, C.; Jin, C.; Shi, Y.; Sun, Y.; Li, Y.; Yang, M.; Jing, X.; Luo, J.; Lian, X. OsATX1 interacts with heavy metal P1B-type ATPases and affects copper transport and distribution. Plant Physiol. 2018, 178, 329–344. [Google Scholar] [CrossRef]
- Ibrahim, S.; Saleem, B.; Rehman, N.; Zafar, S.A.; Naeem, M.K.; Khan, M.R. CRISPR/Cas9-mediated disruption of Inositol Pentakisphosphate 2-Kinase 1 (TaIPK1) reduces phytic acid and improves iron and zinc accumulation in wheat grains. J. Adv. Res. 2022, 37, 33–41. [Google Scholar] [CrossRef]
- Wu, J.; Chen, C.; Xian, G.; Liu, D.; Lin, L.; Yin, S.; Sun, Q.; Fang, Y.; Zhang, H.; Wang, Y. Engineering herbicide-resistant oilseed rape by CRISPR/Cas9-mediated cytosine base-editing. Plant Biotechnol. J. 2020, 18, 1857–1859. [Google Scholar] [CrossRef]
- Hummel, A.W.; Chauhan, R.D.; Cermak, T.; Mutka, A.M.; Vijayaraghavan, A.; Boyher, A.; Starker, C.G.; Bart, R.; Voytas, D.F.; Taylor, N.J. Allele exchange at the EPSPS locus confers glyphosate tolerance in cassava. Plant Biotechnol. J. 2018, 16, 1275–1282. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Zhang, X.; Wu, C.; He, Y.; Ma, Y.; Hou, H.; Guo, X.; Du, W.; Zhao, Y.; Xia, L. Engineering herbicide-resistant rice plants through CRISPR/Cas9-mediated homologous recombination of Acetolactate Synthase. Mol. Plant 2016, 9, 628–631. [Google Scholar] [CrossRef] [PubMed]
- Butt, H.; Rao, G.S.; Sedeek, K.; Aman, R.; Kamel, R.; Mahfouz, M. Engineering herbicide resistance via prime editing in rice. Plant Biotechnol. J. 2020, 18, 2370–2372. [Google Scholar] [CrossRef] [PubMed]
- Lyu, Y.S.; Cao, L.M.; Huang, W.Q.; Liu, J.X.; Lu, H.P. Disruption of three polyamine uptake transporter genes in rice by CRISPR/Cas9 gene editing confers tolerance to the herbicide paraquat. aBIOTECH 2022, 3, 140–145. [Google Scholar] [CrossRef]
- Liu, H.; Hu, M.; Wang, Q.; Cheng, L.; Zhang, Z. Role of papain-like cysteine proteases in plant development. Front. Plant Sci. 2018, 9, 1717. [Google Scholar] [CrossRef]
- Hu, S.; Chen, Y.; Qian, C.; Ren, H.; Liang, X.; Tao, W.; Chen, Y.; Wang, J.; Dong, Y.; Han, J.; Ouyang, X.; Huang, X. Nuclear accumulation of rice UV-B photoreceptors is UV-B- and OsCOP1-independent for UV-B responses. Nat. Commun. 2024, 15, 6396. [Google Scholar] [CrossRef]
- Liu, C.; Lin, J.-Z.; Wang, Y.; Tian, Y.; Zheng, H.-P.; Zhou, Z.-K.; Zhou, Y.-B.; Tang, X.-D.; Zhao, X.-H.; Wu, T.; Xu, S.L.; Tang, D.-Y.; Zuo, Z.C.; He, H.; Bai, L.-Y.; Yang, Y.-Z.; Liu, X.-M. The protein phosphatase PC1 dephosphorylates and deactivates CatC to negatively regulate H2O2 homeostasis and salt tolerance in rice. The Plant Cell 2023, 35, 3604–3625. [Google Scholar] [CrossRef] [PubMed]
- Shnaider, Y.; Elad, Y.; Rav-David, D.; Pashkovsky, E.; Leibman, D.; Kravchik, M.; Shtarkman-Cohen, M.; Gal-On, A.; Spiegelman, Z. Development of powdery mildew resistance in cucumber using CRISPR/Cas9-mediated mutagenesis of CsaMLO8. Phytopathology 2023, 113, 786–790. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Naing, A.H.; Kang, H.; Lee, S.Y.; Li, W.; Chung, M.Y.; Kim, C.K. CRISPR/Cas9-mediated editing of PhMLO1 confers powdery mildew resistance in petunia. Plant Biotechnol. Rep. 2023, 17, 767–775. [Google Scholar] [CrossRef]
- Nekrasov, V.; Wang, C.; Win, J.; Lanz, C.; Weigel, D.; Kamoun, S. Rapid generation of a transgene-free powdery mildew resistant tomato by genome deletion. Sci. Rep. 2017, 7, 482. [Google Scholar] [CrossRef]
- Wan, D.Y.; Guo, Y.; Cheng, Y.; Hu, Y.; Xiao, S.; Wang, Y.; Wen, Y.Q. CRISPR/Cas9-mediated mutagenesis of VvMLO3 results in enhanced resistance to powdery mildew in grapevine (Vitis vinifera). Hortic. Res. 2020, 7, 116. [Google Scholar] [CrossRef]
- Santillán Martínez, M.I.; Bracuto, V.; Koseoglou, E.; Appiano, M.; Jacobsen, E.; Visser, R.G.F.; Wolters, A.M.A.; Bai, Y. CRISPR/Cas9-targeted mutagenesis of the tomato susceptibility gene PMR4 for resistance against powdery mildew. BMC Plant Biol. 2020, 20, 284. [Google Scholar] [CrossRef]
- Hong, Y.; Meng, J.; He, X.; Zhang, Y.; Liu, Y.; Zhang, C.; Qi, H.; Luan, Y. Editing miR482b and miR482c simultaneously by CRISPR/Cas9 enhanced tomato resistance to Phytophthora infestans. Phytopathology 2021, 111, 1008–1016. [Google Scholar] [CrossRef]
- Kieu, N.P.; Lenman, M.; Wang, E.S.; Petersen, B.L.; Andreasson, E. Mutations introduced in susceptibility genes through CRISPR/Cas9 genome editing confer increased late blight resistance in potatoes. Sci. Rep. 2021, 11, 4487. [Google Scholar] [CrossRef]
- Hegde, N.; Joshi, S.; Soni, N.; Kushalappa, A.C. The caffeoyl-CoA O-methyltransferase gene SNP replacement in Russet Burbank potato variety enhances late blight resistance through cell wall reinforcement. Plant Cell Rep. 2021, 40, 237–254. [Google Scholar] [CrossRef]
- Lin, X.; Jia, Y.; Heal, R.; Prokchorchik, M.; Sindalovskaya, M.; Olave-Achury, A.; Makechemu, M.; Fairhead, S.; Noureen, A.; Heo, J.; Witek, K.; Smoker, M.; Taylor, J.; Shrestha, R.K.; Lee, Y.; Zhang, C.; Park, S.J.; Sohn, K.H.; Huang, S.; Jones, J.D.G. Solanum americanum genome-assisted discovery of immune receptors that detect potato late blight pathogen effectors. Nat. Genet. 2023, 55, 1579–1588. [Google Scholar] [CrossRef] [PubMed]
- Silva, C.J.; van den Abeele, C.; Ortega-Salazar, I.; Papin, V.; Adaskaveg, J.A.; Wang, D.; Casteel, C.L.; Seymour, G.B.; Blanco-Ulate, B. Host susceptibility factors render ripe tomato fruit vulnerable to fungal disease despite active immune responses. J. Exp. Bot. 2021, 72, 2696–2709. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Blahut-Beatty, L.; Zheng, S.; Clough, S.J.; Simmonds, D.H. The role of a soybean 14-3-3 gene (Glyma05g29080) on white mold resistance and nodulation investigations using CRISPR-Cas9 editing and RNA silencing. Mol. Plant Microbe Interact. 2023, 36, 159–164. [Google Scholar] [CrossRef]
- He, F.; Wang, C.; Sun, H.; Tian, S.; Zhao, G.; Liu, C.; Wan, C.; Guo, J.; Huang, X.; Zhan, G.; et al. Simultaneous Editing of Three Homoeologues of TaCIPK14 Confers Broad-spectrum Resistance to Stripe Rust in Wheat. Plant Biotechnol. J. 2022, 21, 354–368. [Google Scholar] [CrossRef]
- Dai, Z.; Yang, Q.; Chen, D.; Li, B.; Que, J.; Hu, L.; Zhang, B.; Zhang, Z.; Chen, K.; Zhang, S.; et al. ZmAGO18b negatively regulates maize resistance against southern leaf blight. Theor. Appl. Genet. 2023, 136, 158. [Google Scholar] [CrossRef] [PubMed]
- Nawaz, G.; Usman, B.; Peng, H.; Zhao, N.; Yuan, R.; Liu, Y.; Li, R. Knockout of Pi21 by CRISPR/Cas9 and iTRAQ-based proteomic analysis of mutants revealed new insights into M. oryzae resistance in elite rice line. Genes 2020, 11, 735. [Google Scholar] [CrossRef] [PubMed]
- Oliva, R.; Ji, C.; Atienza-Grande, G.; Huguet-Tapia, J.C.; Perez-Quintero, A.; Li, T.; Eom, J.-S.; Li, C.; Nguyen, H.; Liu, B.; Auguy, F.; Sciallano, C.; Luu, V.T.; Dossa, G.S.; Cunnac, S.; Schmidt, S.M.; Slamet-Loedin, I.H.; Cruz, C.V.; Szurek, B.; Frommer, W.B.; White, F.F.; Yang, B. Broad-spectrum resistance to bacterial blight in rice using genome editing. Nat. Biotechnol. 2019, 37, 1344–1350. [Google Scholar] [CrossRef]
- Kim, M.-S.; Le, V.T.; Jung, Y.J.; Kang, K.-K.; Cho, Y.-G. OsPUB9 gene edited by CRISPR/Cas9 enhanced resistance to bacterial leaf blight in rice (Oryza sativa L.). Int. J. Mol. Sci. 2024, 25, 7145. [Google Scholar] [CrossRef]
- Peng, A.; Chen, S.; Lei, T.; Xu, L.; He, Y.; Wu, L.; Yao, L.; Zou, X. Engineering canker-resistant plants through CRISPR/Cas9-targeted editing of the susceptibility gene CsLOB1 promoter in citrus. Plant Biotechnol. J. 2017, 15, 1509–1519. [Google Scholar] [CrossRef] [PubMed]
- Ortigosa, A.; Gimenez-Ibanez, S.; Leonhardt, N.; Solano, R. Design of a bacterial speck resistant tomato by CRISPR/Cas9-mediated editing of SlJAZ2. Plant Biotechnol. J. 2019, 17, 665–673. [Google Scholar] [CrossRef]
- Ortega, A.; Seong, K.; Schultink, A.; de Toledo Thomazella, D.P.; Seo, E.; Zhang, E.; Pham, J.; Cho, M.-J.; Dahlbeck, D.; Warren, J.; Minsavage, G.V.; Jones, J.B.; Sierra-Orozco, E.; Hutton, S.F.; Staskawicz, B. CRISPR/Cas9-Mediated Editing of Bs5 and Bs5L in Tomato Leads to Resistance against Xanthomonas. Plant Biotech. J. 2024, 22, 2785–2787. [Google Scholar] [CrossRef]
- Yang, J.; Fang, Y.; Wu, H.; Zhao, N.; Guo, X.; Mackon, E.; Peng, H.; Huang, S.; He, Y.; Qin, B.; Liu, Y.; Liu, F.; Chen, S.; Li, R. Improvement of resistance to rice blast and bacterial leaf streak by CRISPR/Cas9-mediated mutagenesis of Pi21 and OsSULTR3;6 in rice (Oryza sativa L.). Front. Plant Sci. 2023, 14, 1209384. [Google Scholar] [CrossRef]
- Saikia, B.; S, R.; Debbarma, J.; Maharana, J.; Sastry, G.N.; Chikkaputtaiah, C. CRISPR/Cas9-based genome editing and functional analysis of SlHyPRP1 and SlDEA1 genes of Solanum lycopersicum L. in imparting genetic tolerance to multiple stress factors. Front. Plant Sci. 2024, 15. [Google Scholar] [CrossRef]
- Zhao, A.; Li, Q.; Meng, P.; Liu, P.; Wu, S.; Lang, Z.; Song, Y.; Macho, A.P. Reduced content of gamma-aminobutyric acid enhances resistance to bacterial wilt disease in tomato. Plant Biotech. J. 2025, 23, 792–806. [Google Scholar] [CrossRef] [PubMed]
- Baltes, N.J.; Hummel, A.W.; Konecna, E.; Cegan, R.; Bruns, A.N.; Bisaro, D.M.; Voytas, D.F. conferring resistance to geminiviruses with the CRISPR-Cas prokaryotic immune system. Nat. Plants 2015, 1, 15145. [Google Scholar] [CrossRef]
- Ji, X.; Zhang, H.; Zhang, Y.; Wang, Y.; Gao, C. Establishing a CRISPR–Cas-like immune system conferring dna virus resistance in plants. Nature Plants 2015, 1, 15144. [Google Scholar] [CrossRef]
- Ali, Z.; Abulfaraj, A.; Idris, A.; Ali, S.; Tashkandi, M.; Mahfouz, M.M. CRISPR/Cas9-mediated viral interference in plants. Genome Biol. 2015, 16, 238. [Google Scholar] [CrossRef]
- Tashkandi, M.; Ali, Z.; Aljedaani, F.; Shami, A.; Mahfouz, M.M. Engineering resistance against tomato yellow leaf curl virus via the CRISPR/Cas9 system in tomato. Plant Signal Behav. 2018, 13, e1525996. [Google Scholar] [CrossRef]
- Yin, K.; Han, T.; Xie, K.; Zhao, J.; Song, J.; Liu, Y. Engineer complete resistance to cotton leaf curl multan virus by the CRISPR/Cas9 system in Nicotiana benthamiana. Phytopath. Res. 2019, 1, 9. [Google Scholar] [CrossRef]
- Kis, A.; Hamar, É.; Tholt, G.; Bán, R.; Havelda, Z. Creating highly efficient resistance against wheat dwarf virus in barley by employing CRISPR/Cas9 system. Plant Biotech. J. 2019, 17, 1004–1006. [Google Scholar] [CrossRef] [PubMed]
- Khan, S.; Mahmood, M.S.; Rahman, S.U.; Rizvi, F.; Ahmad, A. Evaluation of the CRISPR/Cas9 system for the development of resistance against cotton leaf curl virus in model plants. Plant Protect. Sci. 2020, 56, 154–162. [Google Scholar] [CrossRef]
- Liu, H.; Soyars, C.L.; Li, J.; Fei, Q.; He, G.; Peterson, B.A.; Meyers, B.C.; Nimchuk, Z.L.; Wang, X. CRISPR/Cas9-mediated resistance to cauliflower mosaic virus. Plant Direct 2018, 2, e00047. [Google Scholar] [CrossRef]
- Tripathi, J.N.; Ntui, V.O.; Ron, M.; Muiruri, S.K.; Britt, A.; Tripathi, L. CRISPR/Cas9 editing of endogenous banana streak virus in the B genome of Musa spp. overcomes a major challenge in banana breeding. Commun. Biol. 2019, 2, 46. [Google Scholar] [CrossRef] [PubMed]
- Roy, A.; Zhai, Y.; Ortiz, J.; Neff, M.; Mandal, B.; Mukherjee, S.K.; Pappu, H.R. multiplexed editing of a begomovirus genome restricts escape mutant formation and disease development. PLoS One 2019, 14, e0223765. [Google Scholar] [CrossRef]
- Pramanik, D.; Shelake, R.M.; Park, J.; Kim, M.J.; Hwang, I.; Park, Y.; Kim, J.-Y. CRISPR/Cas9-mediated generation of pathogen-resistant tomato against tomato yellow leaf curl virus and powdery mildew. Int. J. Mol. Sci. 2021, 22, 1878. [Google Scholar] [CrossRef]
- Zhang, Z.; Ge, X.; Luo, X.; Wang, P.; Fan, Q.; Hu, G.; Xiao, J.; Li, F.; Wu, J. Simultaneous editing of two copies of Gh14-3-3d confers enhanced transgene-clean plant defense against Verticillium dahliae in allotetraploid upland cotton. Front. Plant Sci. 2018, 9, 842. [Google Scholar] [CrossRef]
- Zhan, X.; Zhang, F.; Zhong, Z.; Chen, R.; Wang, Y.; Chang, L.; Bock, R.; Nie, B.; Zhang, J. Generation of virus-resistant potato plants by RNA genome targeting. Plant Biotechnol. J. 2019, 17, 1814–1822. [Google Scholar] [CrossRef]
- Mehta, D.; Stürchler, A.; Anjanappa, R.B.; Zaidi, S.S.-A.; Hirsch-Hoffmann, M.; Gruissem, W.; Vanderschuren, H. Linking CRISPR-cas9 interference in cassava to the evolution of editing-resistant Geminiviruses. Genome Biol. 2019, 20, 80. [Google Scholar] [CrossRef]
- Zhang, Y.; Tian, X.; Xu, H.; Zhan, B.; Zhou, C.; Li, S.; Zhang, Z. Knockout of SlDCL2b attenuates the resistance of tomato to potato spindle tuber viroid infection. Mol. Plant Pathol. 2024, 25, e13441. [Google Scholar] [CrossRef]
- Liu, C.; Kong, M.; Yang, F.; Zhu, J.; Qi, X.; Weng, J.; Di, D.; Xie, C. Targeted generation of null mutants in ZmGDIα confers resistance against maize rough dwarf disease without agronomic penalty. Plant Biotechnol. J. 2022, 20, 803–805. [Google Scholar] [CrossRef]
- Arra, Y.; Auguy, F.; Stiebner, M.; Chéron, S.; Wudick, M.M.; Miras, M.; Schepler-Luu, V.; Köhler, S.; Cunnac, S.; Frommer, W.B.; Albar, L. Rice yellow mottle virus resistance by genome editing of the Oryza sativa L. ssp. japonica nucleoporin gene OsCPR5.1 but not OsCPR5.2. Plant Biotechnol. J. 2024, 22, 1299–1311. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Liu, D.; Gao, D.; Zhao, W.; Du, H.; Qiu, Z.; Huang, J.; Wen, P.; Wang, Y.; Li, Q.; et al. Cytokinin confers brown planthopper resistance by elevating jasmonic acid pathway in rice. Int. J. Mol. Sci. 2022, 23, 5946. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Alariqi, M.; Wang, Y.; Wang, Q.; Xu, Z.; Zafar, M.N.; Yang, G.; Jia, R.; Hussain, A.; Chen, Y.; et al. Construction of host plant insect-resistance mutant library by high-throughput CRISPR/Cas9 system and identification of a broad-spectrum insect resistance gene. Advanced Science 2024, 11, 2306157. [Google Scholar] [CrossRef]
- Zhang, Y.; Guo, W.; Chen, L.; Shen, X.; Yang, H.; Fang, Y.; Ouyang, W.; Mai, S.; Chen, H.; Chen, S.; et al. CRISPR/Cas9-mediated targeted mutagenesis of GmUGT enhanced soybean resistance against leaf-chewing insects through flavonoids biosynthesis. Front. Plant Sci. 2022, 13, 802716. [Google Scholar] [CrossRef] [PubMed]
- Huang, Q.; Lin, B.; Cao, Y.; Zhang, Y.; Song, H.; Huang, C.; Sun, T.; Long, C.; Liao, J.; Zhuo, K. CRISPR/Cas9-mediated mutagenesis of the susceptibility gene OsHPP04 in rice confers enhanced resistance to rice root-knot nematode. Front. Plant Sci. 2023, 14, 1134653. [Google Scholar] [CrossRef]
- Usovsky, M.; Gamage, V.A.; Meinhardt, C.G.; Dietz, N.; Triller, M.; Basnet, P.; Gillman, J.D.; Bilyeu, K.D.; Song, Q.; Dhital, B.; et al. Loss-of-function of an α-SNAP gene confers resistance to soybean cyst nematode. Nat. Commun. 2023, 14, 7629. [Google Scholar] [CrossRef]
- Hao, J.; Yang, Y.; Futrell, S.; Kelly, E.A.; Lorts, C.M.; Nebie, B.; Runo, S.; Yang, J.; Alvarez, S.; Lasky, J.R.; et al. CRISPR/Cas9-mediated mutagenesis of carotenoid cleavage dioxygenase (CCD) genes in sorghum alters strigolactone biosynthesis and plant biotic interactions. Phytobiomes J. 2023, 7, 339–351. [Google Scholar] [CrossRef]
- Vernié, T.; Rich, M.; Pellen, T.; Teyssier, E.; Garrigues, V.; Chauderon, L.; Medioni, L.; van Beveren, F.; Libourel, C.; Keller, J.; et al. Conservation of symbiotic signaling since the most recent common ancestor of land plants. Proc. Natl. Acad. Sci. U.S.A. 2025, 122, e2408539121. [Google Scholar] [CrossRef] [PubMed]
- Yan, D.; Tajima, H.; Cline, L.C.; Fong, R.Y.; Ottaviani, J.I.; Shapiro, H.Y.; Blumwald, E. Genetic modification of flavone biosynthesis in rice enhances biofilm formation of soil diazotrophic bacteria and biological nitrogen fixation. Plant Biotechnol. J. 2022, 20, 2135–2148. [Google Scholar] [CrossRef]
- Kourelis, J.; Marchal, C.; Posbeyikian, A.; Harant, A.; Kamoun, S. NLR immune receptor–nanobody fusions confer plant disease resistance. Science 2023, 379, 934–939. [Google Scholar] [CrossRef]
- Kalischuk, M.; Müller, B.; Fusaro, A.F.; Wijekoon, C.P.; Waterhouse, P.M.; Prüfer, D.; Kawchuk, L. Amplification of cell signaling and disease resistance by an immunity receptor Ve1Ve2 heterocomplex in plants. Commun. Biol. 2022, 5, 497. [Google Scholar] [CrossRef] [PubMed]
- Guayazán-Palacios, N.; Steinbrenner, A.D. Plant cell surface receptors at the forefront of the growth-defense trade-off. Dev. Cell. 2025, 491–492. [Google Scholar] [CrossRef]
- Sashidhar, N.; Harloff, H.J.; Potgieter, L.; Jung, C. Gene editing of three BnITPK genes in tetraploid oilseed rape leads to significant reduction of phytic acid in seeds. Plant Biotechnol. J. 2020, 18, 2241–2250. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, K.; Kobayashi, A.; Endo, M.; Sage-Ono, K.; Toki, S.; Ono, M. CRISPR/Cas9-mediated mutagenesis of the dihydroflavonol-4-reductase-B (DFR-B) locus in the Japanese morning glory ipomoea (Pharbitis) Nil. Sci. Rep. 2017, 7, 10028. [Google Scholar] [CrossRef]
- Cai, L.; Zhang, L.; Fu, Q.; Xu, Z.F. Identification and expression analysis of cytokinin metabolic genes IPTs, CYP735A and CKXs in the biofuel plant Jatropha curcas. Peer J. 2018, 6, e4812. [Google Scholar] [CrossRef]
- Tian, S.; Jiang, L.; Gao, Q.; Zhang, J.; Zong, M.; Zhang, H.; Ren, Y.; Guo, S.; Gong, G.; Liu, F.; et al. Efficient CRISPR/Cas9-based gene knockout in watermelon. Plant Cell Rep. 2017, 36, 399–406. [Google Scholar] [CrossRef] [PubMed]
- Cai, Y.; Wang, L.; Chen, L.; Wu, T.; Liu, L.; Sun, S.; Wu, C.; Yao, W.; Jiang, B.; Yuan, S.; et al. Mutagenesis of GmFT2a and GmFT5a mediated by CRISPR/Cas9 contributes for expanding the regional adaptability of soybean. Plant Biotechnol. J. 2019, 18, 298–309. [Google Scholar] [CrossRef]
- Kis, A.; Polgári, D.; Dalmadi, Á.; Ahmad, I.; Rakszegi, M.; Sági, L.; Csorba, T.; Havelda, Z. Targeted Mutations in the GW2.1 Gene modulate grain traits and induce yield loss in barley. Plant Science 2024, 340, 111968. [Google Scholar] [CrossRef]
- Jiang, Y.; Li, J.; Liu, B.; Cao, D.; Zong, Y.; Chang, Y.; Li, Y. Novel hina alleles created by genome editing increase grain hardness and reduce grain width in barley. Transgenic Res. 2022, 31, 637–645. [Google Scholar] [CrossRef]
- Zeng, Z.; Han, N.; Liu, C.; Buerte, B.; Zhou, C.; Chen, J.; Wang, M.; Zhang, Y.; Tang, Y.; Zhu, M.; et al. Functional dissection of HGGT and HPT in barley vitamin E biosynthesis via CRISPR/Cas9-enabled genome editing. Ann. Bot. 2020, 126, 929–942. [Google Scholar] [CrossRef]
- Li, Z.; Wang, Y.; Lu, X.; Li, R.; Liu, J.; Fu, S.P.; Hu, X.; Jianchun, G.; Yao, Y. Construction and verification of CRISPR/Cas9 gene editing vector for cassava MeSSIII gene. Mol. Plant Breed. 2020, 11, 1–8. [Google Scholar] [CrossRef]
- Juma, B.S.; Mukami, A.; Mweu, C.; Ngugi, M.P.; Mbinda, W. Targeted mutagenesis of the CYP79D1 gene via CRISPR/Cas9-mediated genome editing results in lower levels of cyanide in cassava. Front. Plant Sci. 2022, 13, 1009860. [Google Scholar] [CrossRef] [PubMed]
- Jiang, M.; Hu, H.; Kai, J.; Traw, M.B.; Yang, S.; Zhang, X. Different knockout genotypes of OsIAA23 in rice using CRISPR/Cas9 generating different phenotypes. Plant Mol. Biol. 2019, 100, 467–479. [Google Scholar] [CrossRef]
- Zheng, X.; Zhang, S.; Liang, Y.; Zhang, R.; Liu, L.; Qin, P.; Zhang, Z.; Wang, Y.; Zhou, J.; Tang, X.; et al. Loss-function mutants of OsCKX gene family based on CRISPR-Cas systems revealed their diversified roles in rice. The Plant Genome 2023, 16, e20283. [Google Scholar] [CrossRef]
- Iwamoto, M. In-frame editing of transcription factor gene RDD1 to suppress miR166 recognition influences nutrient uptake, photosynthesis, and grain quality in rice. Sc.i Rep. 2022, 12, 10795. [Google Scholar] [CrossRef]
- Liu, K.; Sakuraba, Y.; Ohtsuki, N.; Yang, M.; Ueda, Y.; Yanagisawa, S. CRISPR/Cas9-mediated elimination of OsHHO3, a transcriptional repressor of three AMMONIUM TRANSPORTER1 genes, improves nitrogen use efficiency in rice. Plant Biotechnol. J. 2023, 21, 2169–2172. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Li, X.; Zhou, Z.; Wu, P.; Fang, M.; Pan, X.; Lin, Q.; Luo, W.; Wu, G.; Li, H. Reassessment of the four yield-related genes Gn1a, DEP1, GS3, and IPA1 in rice using a CRISPR/Cas9 System. Front. Plant Sci. 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Ashokkumar, S.; Jaganathan, D.; Ramanathan, V.; Rahman, H.; Palaniswamy, R.; Kambale, R.; Muthurajan, R. Creation of novel alleles of fragrance gene OsBADH2 in rice through CRISPR/Cas9 mediated gene editing. PLOS ONE 2020, 15, e0237018. [Google Scholar] [CrossRef]
- Miao, C.; Xiao, L.; Hua, K.; Zou, C.; Zhao, Y.; Bressan, R.A.; Zhu, J. Mutations in a subfamily of abscisic acid receptor genes promote rice growth and productivity. Proc. Natl. Acad. Sci. U.S.A. 2018, 115, 6058–6063. [Google Scholar] [CrossRef]
- Hu, X.; Cui, Y.; Dong, G.; Feng, A.; Wang, D.; Zhao, C.; Zhang, Y.; Hu, J.; Zeng, D.; Guo, L.; et al. Using CRISPR-Cas9 to generate semi-dwarf rice lines in elite landraces. Sci. Rep. 2019, 9, 19096. [Google Scholar] [CrossRef]
- Zhu, Y.; Lin, Y.; Chen, S.; Liu, H.; Chen, Z.; Fan, M.; Hu, T.; Mei, F.; Chen, J.; Chen, L.; Wang, F. CRISPR/Cas9-mediated functional recovery of the recessive rc allele to develop red rice. Plant Biotechnol. J. 2019, 17, 2096–2105. [Google Scholar] [CrossRef]
- Xu, J.; Kang, B.-C.; Naing, A.H.; Bae, S.-J.; Kim, J.-S.; Kim, H.; Kim, C.K. CRISPR/Cas9-mediated editing of 1-aminocyclopropane-1-carboxylate oxidase1 enhances Petunia flower longevity. Plant Biotechnol. J. 2019, 18, 287–297. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Yang, X.; Yang, C.; Li, M.; Guo, Y. Exploiting the CRISPR/Cas9 system for targeted genome mutagenesis in Petunia. Sci. Rep. 2016, 6, 20315. [Google Scholar] [CrossRef]
- Wang, D.; Samsulrizal, N.H.; Yan, C.; Allcock, N.S.; Craigon, J.; Blanco-Ulate, B.; Ortega-Salazar, I.; Marcus, S.E.; Bagheri, H.M.; Perez Fons, L.; et al. Characterization of CRISPR mutants targeting genes modulating pectin degradation in ripening tomato. Plant Physiol. 2019, 179, 544–557. [Google Scholar] [PubMed]
- Ueta, R.; Abe, C.; Watanabe, T.; Sugano, S.S.; Ishihara, R.; Ezura, H.; Osakabe, Y.; Osakabe, K. Rapid breeding of parthenocarpic tomato plants using CRISPR/Cas9. Sci. Rep. 2017, 7, 507. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Zha, M.; Huang, J.; Li, L.; Imran, M.; Zhang, C. StMYB44 negatively regulates phosphate transport by suppressing expression of PHOSPHATE1 in potato. J. Exp. Bot. 2017, 68, 1265–1281. [Google Scholar] [CrossRef]
- Andersson, M.; Turesson, H.; Olsson, N.; Fält, A.-S.; Ohlsson, P.; Gonzalez, M.N.; Samuelsson, M.; Hofvander, P. Genome editing in potato via CRISPR-Cas9 ribonucleoprotein delivery. Physiol. Plantarum 2018, 164, 378–384. [Google Scholar] [CrossRef]
- Sevestre, F.; Facon, M.; Wattebled, F.; Szydlowski, N. Facilitating gene editing in potato: A single-nucleotide polymorphism (SNP) map of the Solanum tuberosum L. Cv. Desiree genome. Sci. Rep. 2020, 10, 2045. [Google Scholar] [CrossRef]
- Tuncel, A.; Corbin, K.R.; Ahn-Jarvis, J.; Harris, S.; Hawkins, E.; Smedley, M.A.; Harwood, W.; Warren, F.J.; Patron, N.J.; Smith, A.M. Cas9-mediated mutagenesis of potato starch-branching enzymes generates a range of tuber starch phenotypes. Plant Biotech. J. 2019, 17, 2259–2271. [Google Scholar] [CrossRef]
- Khromov, A.V.; Gushchin, V.A.; Timerbaev, V.I.; Kalinina, N.O.; Taliansky, M.E.; Makarov, V.V. Guide RNA design for CRISPR/Cas9-mediated potato genome editing. Dokl. Biochem. Biophys. 2018, 479, 90–94. [Google Scholar] [CrossRef]
- Nakayasu, M.; Umemoto, N.; Ohyama, K.; Fujimoto, Y.; Lee, H.J.; Watanabe, B.; Muranaka, T.; Saito, K.; Sugimoto, Y.; Mizutani, M. A dioxygenase catalyzes steroid 16α-hydroxylation in steroidal glycoalkaloid biosynthesis. Plant Physiol. 2017, 175, 120–133. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Z.; Ye, G.; Zhou, Y.; Pu, X.; Su, W.; Wang, J. Editing sterol side chain reductase 2 gene (StSSR2) via CRISPR/cas9 reduces the total steroidal glycoalkaloids in potato. All Life 2021, 14, 401–413. [Google Scholar] [CrossRef]
- González, M.N.; Massa, G.A.; Andersson, M.; Turesson, H.; Olsson, N.; Fält, A.-S.; Storani, L.; Décima Oneto, C.A.; Hofvander, P.; Feingold, S.E. Reduced enzymatic browning in potato tubers by specific editing of a polyphenol oxidase gene via ribonucleoprotein complexes delivery of the CRISPR/Cas9 system. Front. Plant Sci. 2019, 10, 1649. [Google Scholar] [CrossRef]
- Pfotenhauer, A.C.; Occhialini, A.; Harbison, S.A.; Li, L.; Piatek, A.A.; Luckett, C.R.; Yang, Y.; Stewart, C.N.; Lenaghan, S.C. Genome-editing of FtsZ1 for alteration of starch granule size in potato tubers. Plants 2023, 12, 1878. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Zhang, H.; Li, S.; Li, J.; Yan, L.; Xia, L. Increasing yield potential through manipulating of an ARE1 ortholog related to nitrogen use efficiency in wheat by CRISPR/Cas9. J. Integr. Plant Biol. 2021, 63, 1649–1663. [Google Scholar] [CrossRef]
- Abdul Rahim, A.; Uzair, M.; Rehman, N.; Fiaz, S.; Attia, K.A.; Abushady, A.M.; Hwan Yang, S.; Ramzan Khan, M. CRISPR/Cas9 mediated TaRPK1 root architecture gene mutagenesis confers enhanced wheat yield. J. King Saud Univ. Sci. 2024, 36, 103063. [Google Scholar] [CrossRef]
- Ibrahim, S.; Saleem, B.; Naeem, M.K.; Arain, S.M.; Khan, M.R. Next-generation technologies for iron and zinc biofortification and bioavailability in cereal grains. Crop Pasture Sci. 2021. [Google Scholar] [CrossRef]
- Wang, W.; Simmonds, J.; Pan, Q.; Davidson, D.; He, F.; Battal, A.; Akhunova, A.; Trick, H.N.; Uauy, C.; Akhunov, E. Gene editing and mutagenesis reveal inter-cultivar differences and additivity in the contribution [i]of TaGW2 homoeologues to grain size and weight in wheat. Theor. Appl. Genet. 2018, 131, 2463–2475. [Google Scholar] [CrossRef]
- Dong, O.X.; Yu, S.; Jain, R.; Zhang, N.; Duong, P.Q.; Butler, C.; Li, Y.; Lipzen, A.; Martin, J.A.; Barry, K.W.; et al. Marker-free carotenoid-enriched rice generated through targeted gene insertion using CRISPR-Cas9. Nat. Commun. 2020, 11, 1178. [Google Scholar] [CrossRef] [PubMed]
- Figueredo, E.F.; da Cruz, T.A.; de Almeida, J.R.; Batista, B.D.; Marcon, J.; de Andrade, P.A.M.; Hayashibara, C.A. de A.; Rosa, M.S.; Azevedo, J.L.; Quecine, M.C. The Key Role of indole-3-acetic acid biosynthesis by Bacillus Thuringiensis RZ2MS9 in promoting maize growth revealed by the IpdC gene knockout mediated by the CRISPR-Cas9 system. Microbiol. Res. 2023, 266, 127. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Wang, S.; Du, W.; Ding, Y.; Li, W.; Chen, Y.; Zheng, Z.; Wang, Y. Sugar transporter ZmSWEET1b is responsible for assimilate allocation and salt stress response in maize. Funct. Integr. Genom. 2023, 23, 137. [Google Scholar] [CrossRef]
- Wang, Y.; Liu, X.; Zheng, X.; Wang, W.; Yin, X.; Liu, H.; Ma, C.; Niu, X.; Zhu, J.-K.; Wang, F. Creation of aromatic maize by CRISPR/Cas. J. Integr. Plant Biol. 2021, 63, 1664–1670. [Google Scholar] [CrossRef]
- Kalischuk, M.L.; Johnson, D.; Kawchuk, L.M. Priming with a double-stranded DNA virus alters Brassica rapa seed architecture and facilitates a defense response. Gene 2015, 557, 130–137. [Google Scholar] [CrossRef]
- Kalischuk, M.L.; Fusaro, A.F.; Waterhouse, P.M.; Pappu, H.R.; Kawchuk, L.M. Complete genomic sequence of a Rubus Yellow Net Virus isolate and detection of genome-wide pararetrovirus-derived small RNAs. Virus Res. 2013, 178, 306–313. [Google Scholar] [CrossRef]
- Kalischuk, M.; Hachey, J.; Kawchuk, L. Complete genome sequence of phytopathogenic Pectobacterium atrosepticum bacteriophage Peat1. Genome Announc. 2015, 3. [Google Scholar] [CrossRef]
- Kalischuk, M.; Hachey, J.; Thomas, D.; Johnson, D.; Kawchuk, L. Complete genome sequence of phytopathogenic Pectobacterium atrosepticum lytic bacteriophage Peat2. Am. J. Potato Res. 2019, 96, 614–616. [Google Scholar] [CrossRef]
- Kathiria, P.; Sidler, C.; Golubov, A.; Kalischuk, M.; Kawchuk, L.M.; Kovalchuk, I. Tobacco mosaic virus infection results in an increase in recombination frequency and resistance to viral, bacterial, and fungal pathogens in the progeny of infected tobacco plants. Plant Physiol. 2010, 153, 1859–1870. [Google Scholar] [CrossRef]
- Kalischuk, M.; Paret, M.L.; Freeman, J.H.; Raj, D.; Da Silva, S.; Eubanks, S.; Wiggins, D.J.; Lollar, M.; Marois, J.J.; Mellinger, H.C.; et al. An improved crop scouting technique incorporating unmanned aerial vehicle-assisted multispectral crop imaging into conventional scouting practice for gummy stem blight in watermelon. Plant Dis. 2019, 103, 1642–1650. [Google Scholar] [CrossRef]
- Kawchuk, L.M.; Lynch, D.R.; Kozub, G.C.; Nelson, G.A.; Kulcsar, F.; Fujimoto, D.K. Multi-year evaluation of Clavibacter michiganensis subsp. sepedonicus disease symptoms in cultivated potato genotypes. Am. J. Potato Res. 1998, 75, 235–243. [Google Scholar] [CrossRef]
- Hill, B.D.; Kalischuk, M.; Waterer, D.R.; Bizimungu, B.; Howard, R.; Kawchuk, L.M. An environmental model predicting bacterial ring rot symptom expression. Am. J. Potato Res. 2011, 88, 294–301. [Google Scholar] [CrossRef]
- Benson, D.A.; Cavanaugh, M.; Clark, K.; Karsch-Mizrachi, I.; Lipman, D.J.; Ostell, J.; Sayers, E.W. GenBank. Nucleic Acids Res. 2013, 41, D36–D42. [Google Scholar] [CrossRef] [PubMed]
- Kawchuk, L.M.; Armstrong, J.D.; Lynch, D.R.; Knowles, N.R. Potatoes having improved quality characteristics and methods for their production. U.S. Patent, 1999, 5,998,701.
- Kawchuk, L.M.; Hachey, J.; Lynch, D.R.; Kulcsar, F.; van Rooijen, G.; Waterer, D.R.; Robertson, A.; Kokko, E.; Byers, R.; Howard, R.J.; Fischer, R.; Prüfer, D. Tomato Ve disease resistance genes encode cell surface-like receptors. Proc. Natl. Acad. Sci. U.S.A. 2001, 98, 6511–6515. [Google Scholar] [CrossRef] [PubMed]
- Kawchuk, L.M.; Lynch, D.R.; Hachey, J.; Kulcsar, F. Tomato nucleic acid sequences that confer resistance to Verticillium and plants transformed therewith. U.S. Patent, 2003, 6,608,245.
- Lynch, D.R.; Kozub, G.C.; Kawchuk, L.M. The relationship between yield, mainstem number, and tuber number in five maincrop and two early-maturing cultivars. Am. J. Potato Res. 2001, 78, 83–90. [Google Scholar] [CrossRef]
- Berman, A.; Su, N.; Li, Z.; Landau, U.; Chakraborty, J.; Gerbi, N.; Liu, J.; Qin, Y.; Yuan, B.; Wei, W.; Yanai, O.; Mayrose, I.; Zhang, Y. Construction of multi-targeted CRISPR libraries in tomato to overcome functional redundancy at genome-scale level. Nat. Commun. 2025, 16, 4111. [Google Scholar] [CrossRef]
- Khadhair, A.H.; Kawchuk, L.M.; Taillon, R.C.; Botar, G. Detection and molecular characterization of an aster yellows phytoplasma in parsley. Can. J. Plant Path. 1998, 20, 55–61. [Google Scholar] [CrossRef]
- Kalischuk, M.L.; Kawchuk, L.M.; Leggett, F. First report of Rubus yellow net virus on Rubus idaeus in Alberta, Canada. Plant Dis. 2008, 92, 974. [Google Scholar] [CrossRef] [PubMed]
- Henrickson, A.; Kalischuk, M.; Lynn, J.; Meers, S.; Johnson, D.; Kawchuk, L. First report of zebra chip on potato in Canada. Plant Dis. 2019, 1016. [Google Scholar] [CrossRef]
- Kalischuk, M.L.; Al-Mughrabi, K.I.; Peters, R.D.; Howard, R.J.; Platt, H.W.; Kawchuk, L.M. Genetic composition of Phytophthora infestans in Canada reveals migration and increased diversity. Plant Dis. 2012, 96, 1729–1735. [Google Scholar] [CrossRef]
- Devendran, R.; Kavalappara, S.R.; Simmons, A.M.; Bag, S. Whitefly-transmitted viruses of cucurbits in the Southern United States. Viruses 2023, 15, 2278. [Google Scholar] [CrossRef]
- Wijekoon, C.P.; Kalischuk, M.L.; Brunelle, P.; Howard, R.J.; Kawchuk, L.M. Characterization of bronze leaf disease in western Canadian aspen and poplar trees. Can. J. Plant Sci. 2021, 102, 11–19. [Google Scholar] [CrossRef]
- Babu, B.; Paret, M.L.; Martini, X.; Knox, G.W.; Riddle, B.; Ritchie, L.; Aldrich, J.; Kalischuk, M.; Da Silva, S. Impact of foliar application of acibenzolar-S-methyl on rose rosette disease and rose plant quality. Plant Dis. 2022, 106, 818–827. [Google Scholar] [CrossRef]
- Kalischuk, M.; Lynn, J.; Kawchuk, L. First report of Potato mop-top virus infecting potatoes in Alberta. Plant Dis. 2016, 100, 2544. [Google Scholar] [CrossRef]
- McNally, J.; Prapagar, K.; Goldenhar, K.; Pate, E.; Shan, S.; Kalischuk, M. First report of an aggressive species of Neopestalotiopsis affecting strawberry in Canada. New Dis. Rep. 2023, 48, e12210. [Google Scholar] [CrossRef]
- Kalischuk, M.; Hendricks, K.; Hochmuth, R.; Freeman, J.; Roberts, P.D.; Paret, M. A portable recombinase polymerase amplification assay for the rapid detection of cucurbit leaf crumple virus in watermelon leaves and fruits. J. Plant Pathol. 2022, 104, 215–224. [Google Scholar] [CrossRef]
- Luckew, A.; Meru, G.; Wang, Y.Y.; Mwatuwa, R.; Paret, M.; Carvalho, R.; Kalischuk, M.; da Silva, A.L.; Candian, J.; Dutta, B.; Srinivasan, R.; Kavalappara, S.R.; RRD, N.C.K.; Bag, S.; McGregor, C. Field evaluation of Cucurbita germplasm for resistance to whiteflies and whitefly-transmitted viruses. HortScience 2022, 57, 337–344. [Google Scholar] [CrossRef]
- Kalischuk, M.L.; Roberts, P.D.; Paret, M.L. A rapid fluorescence-based real-time isothermal assay for the detection of Cucurbit yellow stunting disorder virus in squash and watermelon plants. Mol. Cell. Probes 2020, 53, 101613. [Google Scholar] [CrossRef]
- Kalischuk, M.; Baek, J.; Koeppe, S.; Shan, S.; Pate, E.; Goldenhar, K. Neopestalotiopsis disease of strawberry in Ontario – 2020 to 2022. Can. Plant Dis. Surv. 2025, 105, 229–233. [Google Scholar]
- Bairoch, A.; Boeckmann, B. The SWISS-PROT protein sequence data bank. Nucleic Acids Res. 1992, 20 Suppl., 2019–2022. [Google Scholar] [CrossRef] [PubMed]
- Jumper, J.; Evans, R.; Pritzel, A.; Tim Green, Michael Figurnov, Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; Bridgland, A.; Meyer, C.; Kohl, S.A.A.; Ballard, A.J.; Cowie, A. Highly accurate protein structure prediction with AlphaFold. Nature 596(2021), 583–589. [CrossRef] [PubMed]
- Kawchuk, L.; Jaag, H.M.; Toohey, K.; Martin, R.; Rohde, W.; Prüfer, D. In planta agroinfection by Canadian and German Potato leafroll virus full-length cDNAs. Can. J. Plant Pathol. 210(24), 239–243.
- Ma, W.J.; Wang, C.; Kothandapani, J.; Luzentales-Simpson, M.; Menzies, S.C.; Bescucci, D.M.; Lange, M.E.; Fraser, A.S.C.; Gusse, J.F.; House, K.E.; Moote, P.E.; Xing, X.; Grondin, J.M.; Hui, B.W.; Clarke, S.T.; Shelton, T.G.; Haskey, N.; Gibson, D.L.; Martens, E.C.; Abbott, D.W.; Inglis, G.D.; Sly, L.M.; Brumer, H. Bespoke plant glycoconjugates for gut microbiota-mediated drug targeting. Science 2025, 388, 1410–1416. [Google Scholar] [CrossRef]
- Prüfer, D.; Kawchuk, L.M.; Rohde, W. Polerovirus ORF0 genes induce a host-specific response resembling viral infection. Can. J. Plant Pathol. 2006, 28, 302–309. [Google Scholar] [CrossRef]
- Fusaro, A.F.; Barton, D.A.; Nakasugi, K.; Jackson, C.; Kalischuk, M.L.; Kawchuk, L.M.; Vaslin, M.F.; Correa, R.L.; Waterhouse, P.M. The luteovirus P4 movement protein is a suppressor of systemic RNA silencing. Viruses 2017, 9, 294. [Google Scholar] [CrossRef] [PubMed]
- Anzalone, A.V.; Randolph, P.B.; Davis, J.R.; Sousa, A.A.; Koblan, L.W.; Levy, J.M.; Chen, P.J.; Wilson, C.; Newby, G.A.; Raguram, A.; Liu, D.R. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 2019, 576, 149–157. [Google Scholar] [CrossRef]
- Tuncel, A.; Pan, C.; Clem, J.S.; Lui, D.; Qi, Y. CRISPR-Cas applications in agriculture and plant research. Nat. Rev. Mol. Cell Biol. 2025, 26, 419–441. [Google Scholar] [CrossRef]
- Bai, S.; Cao, X; Hu, L.; Hu, D.; Li, D.; Sun, Y. Engineering an optimized hypercompact CRISPR/Cas12j-8 system for efficient genome editing in plants. Plant Biotechnol. J. 2025, 23, 1153–1164. [Google Scholar] [CrossRef] [PubMed]
- Waltz, E. GABA-enriched tomato is first CRISPR-edited food to enter market. Nat. Biotech. 2022, 40, 9–11. [Google Scholar] [CrossRef] [PubMed]
| Trait | Plant | Locus | Reference |
|---|---|---|---|
| Drought | Arabidopsis | AtOST2 | 35 |
| AtAREB1 | 36 | ||
| Glycine max | GMMYB118 | 37 | |
| Orza sativa | OsERA1 | 38 | |
| OsPUB7 | 39 | ||
| StDRO2 | 40 | ||
| Zea mays | ZmHDT103 | 41 | |
| Salt | Arabidopsis | AtWRKY3, AtWRKY4 | 42 |
| AtACQOS | 43 | ||
| G. max | GmDrb2a, GmDrb2b | 44 | |
| GmAITR | 45 | ||
| GmNHL1 | 46 | ||
| GmCG | 47 | ||
| Hordeum vulgare | HvGSK1.1 | 48 | |
| Medicago truncatula | MtHEN1 | 49 | |
| O. sativa | OsDST | 50 | |
| OsRAV2 | 51 | ||
| OsRP22 | 52 | ||
| OsNAC45 | 53 | ||
| OsSPL10 | 54 | ||
| OsTPP3 | 55 | ||
| Solanum lycopersium | SIHyPRP1 | 56 | |
| SIHAK20 | 57 | ||
| Solanum tuberosum | Stcoilin | 58 | |
| Triticum aestivum | TaHAG1 | 59 | |
| Zea mays | ZmHKT2 | 60 | |
| Heat | Gossypium hirsutum | GhPGF, GhCLA1 | 61 |
| Lactuca sativa | LsNCED4 | 62 | |
| O. sativa | GmHSP17.5Ep | 63 | |
| OsDEP1, OsROC5 | 64 | ||
| OsGER4 | 65 | ||
| OsPYL | 66 | ||
| OsTMS5 | 67 | ||
| S. lycopersicum | SIAGL6 | 68 | |
| SICPK28 | 69 | ||
| SIMAPK3 | 70 | ||
| T. aestivum | TaHsfA1 | 71 | |
| Z. mays | ZmTMS5 | 72 | |
| ZmHSPs | 73 | ||
| Cold | O. sativa | OsPRP1 | 74 |
| OsbHLH57 | 75 | ||
| S. lycopersicum | SICBF1 | 76 | |
| S. tuberosum | StVInv | 77 | |
| T. aestivum | TaPGK | 78 | |
| Z. mays | ZmG6PDH1 | 79 | |
| Metal | Arabidopsis | Atoxp1 | 80 |
| O. sativa | OsARM1 | 81 | |
| OsHAK1 | 82 | ||
| OsLCD | 83 | ||
| OsLCT1 | 84 | ||
| OsMYB84 | 85 | ||
| OsNIP3 | 86 | ||
| OsNramp5 | 87 | ||
| OsNRAMP1 | 88 | ||
| OsZIP5, OsZIP9 | 89 | ||
| OsPRX2 | 90 | ||
| OsATX1 | 91 | ||
| T. aestivum | TalPK1 | 92 | |
| Herbicide | Brassica napus | BnALS | 93 |
| Manihot esculenta | MeEPSPS | 94 | |
| O. sativa | OsALS | 95 | |
| OsTB1 | 96 | ||
| OsPUT1/2/3 | 97 | ||
| OsACC | 98 | ||
| UV Radiation | O, sativa | OsCOP1 | 99 |
| Oxidation | O. sativa | OsCAT2 | 100 |
| Biotic Stress Type | Pathogen/Pest | Plant | Locus | Reference |
|---|---|---|---|---|
| Fungi/ Fungus-like |
Powdery mildew | Cucumis sativus | CsMLO8 | 101 |
| Petunia x hybrida | PhMLO | 102 | ||
| Solanum lycopersicum | SIMLO | 103 | ||
| S. lycopersicum | SIPMR4 | 104 | ||
| Vitis vinifera | VvMOL3 | 105 | ||
| Late blight | S. lycopersicum | miR482b/c | 106 | |
| Solanum tuberosum | S-genes | 107 | ||
| CCoAOMT | 108 | |||
| Solanum americanum | SaNRL1 | 109 | ||
| Gray mold | S. lycopersicum | SIPL | 110 | |
| White mold | Glycine max | Gm5g29080 | 111 | |
| Stripe rust | Triticum aestivum | TaCIPK14 | 112 | |
| Southern late blight | Zea mays | ZmAGO18b | 113 | |
| Rice blast | Oryza sativa | OsPi21 | 114 | |
| Bacteria | Bacterial blight | O. sativa | OsSWEET115 | 115 |
| OsPUB9 | 116 | |||
| Citrus sinensis | CsLOB1 | 117 | ||
| Bacterial speck | S. lycopersicum | SIJAZ2 | 118 | |
| Bacterial spot | S. lycopersicum | SIBs5 | 119 | |
| Bacterial leaf streak | O. sativa | OsSULTR | 120 | |
| Bacterial wilt | S. lycopersicum | SIPRP1/DEA1 | 121 | |
| SIGAD2 | 122 | |||
| Virus | Bean yellow dwarf | Nicotiana benthamiana | LIR | 123 |
| Beet severe curly top | N. benthamiana | IR | 124 | |
| Tomato yellow leaf curl | N. benthamiana | IR | 125 | |
| Tomato yellow leaf curl | S. lycopersicum | CP/Rep | 126 | |
| Cotton leaf curl | N. benthamiana | IR | 127 | |
| Wheat dwarf | Hordeum vulgare | MP/CP/IP | 128 | |
| Cotton leaf curl | N. benthamiana | Rep | 129 | |
| Cauliflower mosaic | A. thaliana | CP | 130 | |
| Banana streak | Musa spp. | ORF1, 2, 3 | 131 | |
| Chili leaf curl | N. benthamiana | C + V | 132 | |
| Tomato yellow leaf curl | S. lycopersicum | SIPelo | 133 | |
| Cucumber mosaic | N. benthamiana | ORF1a, 3a | 134 | |
| Potato virus Y, S, and A | S. tuberosum | P3, CI, CP | 135 | |
| Geminiviruses | Manihot esculenta | IR, ORFs | 136 | |
| Potato spindle viroid | S. lycopersicon | SIDCL2b | 137 | |
| Maize rough dwarf | Z. mays | ZmGDla | 138 | |
| Yellow mottle | O. sativa | OsCPR5.1 | 139 | |
| Pest | Plant hopper | O. sativa | OsCKX | 140 |
| Insects | Gossypium hirsutum | GhMLP423 | 141 | |
| Chewing insects | G. max | GmUGT | 142 | |
| Root-knot nematode | O. sativa | OsHPP04 | 143 | |
| Cyst nematode | G. max | GmSNAP02 | 144 | |
| Parasitic weeds | Sorghum bicolor | CCD | 145 | |
| Beneficial | Mycorrhizal | Marchantia paleacea | CCaMK | 146 |
| Nitrogen fixation | O. sativa | CYP75 | 147 |
| Plant | Locus | Trait | Reference |
|---|---|---|---|
| Brassica napus | BnITPK | Phytic acid | 151 |
| Ipomoea nil | DFR-B | Flower colour | 152 |
| Jatropha curcas | JcCYP735A, JcCKX | Growth | 153 |
| Citrullus lanatus | PDS gene | Colour | 154 |
| Glycine max | GmFT2a | Flowering | 155 |
| Hordeum vulgare | GW2.1 | Seed set | 156 |
| Hina | Grain hardiness | 157 | |
| HvHGGT, HvHPT | Vitamin E | 158 | |
| Manihot esculenta | MeMSIII | Starch synthesis | 159 |
| MeCYP79D1 | Cyanide | 160 | |
| Oryza sativa | OsIAA23 | Development | 161 |
| OsCKX | Growth and quality | 162 | |
| OsRDD1 | Photosynthesis | 163 | |
| OsHHO3 | Nitrogen uptake | 164 | |
| OsGn1a, OsGL3 | Grain number/size | 165 | |
| OsBADH2 | Fragrance | 166 | |
| OsPYL9 | Yield | 167 | |
| OsSD1 | Lodging | 168 | |
| OsRc, OsRd | Red rice | 169 | |
| Petunia hybrid | PhACO | Longevity | 170 |
| PhPDS | Albino | 171 | |
| Solanum lycopersicon | PI, PG2a, TBG4 | Fruit shelf-life | 172 |
| SIIAA9 | Parthenocarpy | 173 | |
| Solanum tuberosum | StMYB44 | Phosphate transport | 174 |
| StGBBS | Starch quality | 175 | |
| StSS6 | Starch quantity | 176 | |
| StSBE | Starch quality | 177 | |
| STPDS | Carotenoids | 178 | |
| St16DOX | Glycoalkaloids | 179 | |
| StSSR2 | Glycoalkaloids | 180 | |
| StPPO2 | Enzymatic browning | 181 | |
| FtsZ1 | Starch granule size | 182 | |
| Triticum aestivum | TaARE1 | Nitrogen use | 183 |
| TaRPK1 | Yield | 184 | |
| Zn and Fe uptake | 185 | ||
| TaGW2 | Seed size and weight | 186 | |
| Zea mays | SSU-crt1, ZmPSY | Carotenoid increase | 187 |
| ipdC | Improved growth | 188 | |
| ZmSWEET1b | Sugar transport | 189 | |
| Zmbadh2 | Aroma | 190 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




