Submitted:
01 April 2026
Posted:
03 April 2026
You are already at the latest version
Abstract
Keywords:
Introduction
2. Soil Fertility and the Microbiome: The Relationship Between Plant Yield and Soil Type with the Use of Inoculants
2.1. The Influence of Abiotic Factors on the Effectiveness of Microbial Inoculants
2.2. The Combined Influence of Climate and Soil Conditions
2.3. The Role of Soil Type and Fertility Gradients
3. Secondary Antimicrobial Metabolites of Bioinoculants are a Factor in the Formation of the Plant Rhizosphere and Their Impact on Crop Yield and Fertility
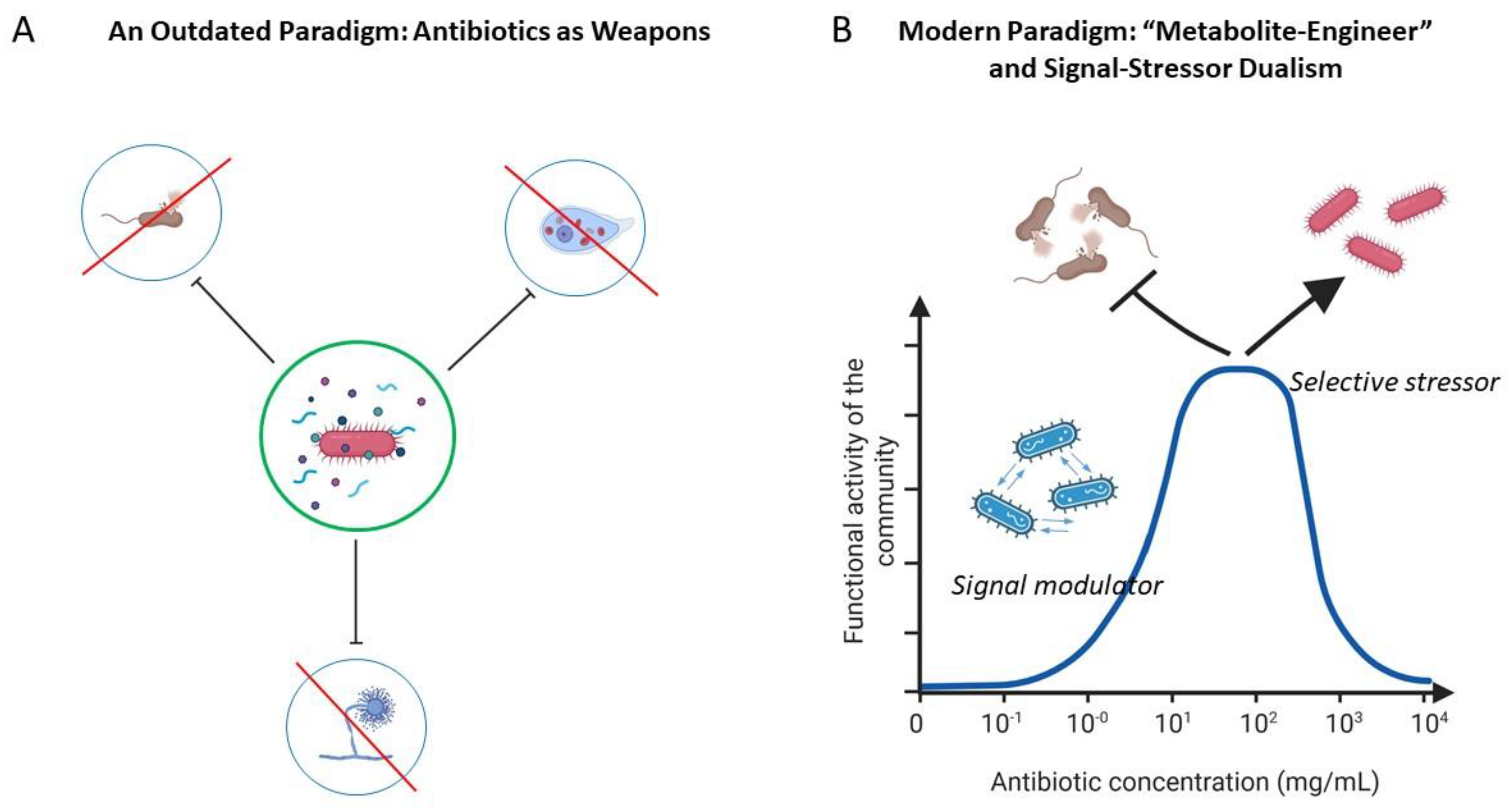
3.1. From Antagonism to Signaling: A Paradigm Shift in Understanding the Role of Antibiotics
3.2. In Situ Evidence Suggests that Bioinoculants Create a Stable "Chemical Environment" in the Rhizosphere
3.3. Mechanisms of Influence: Molecular Interference and Chemical Communication
3.4. Antibiotics as "Ecosystem Engineers": from Cells to Metabolites
3.5. Selective Remodeling of the Rhizobiome by Specific Metabolites
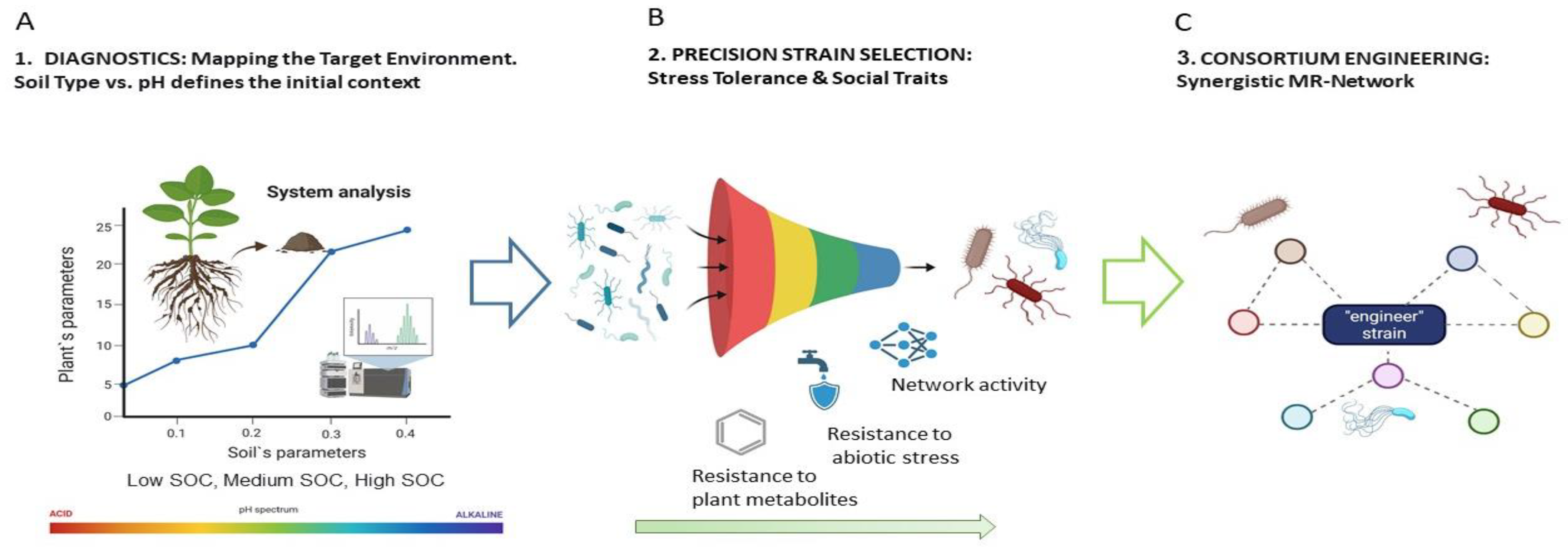
3.6. Towards a Strategy for Targeted Selection of Bioinoculants
4. Practical Recommendations for the Creation of Bioinnoculants
4.2. Consortium Design: the Principle of Synergy Between Moderately Related Communities
4.3. Taking into Account the Soil-Climatic Context: From Universality to Precision
Conclusion
Author Contributions
Funding
Data Availability Statement
Declaration of Competing Interest
References
- Adeleke, BS; Babalola, OO; Glick, BR. Plant growth-promoting root-colonizing bacterial endophytes. Rhizosphere 2021, 20, 100433. [Google Scholar] [CrossRef]
- Onuh, Augustine C.; Manopaek, R; Vangnai, AS; Tiyayon, P; Vinayavekhin, N. Dynamics of Microbial Metabolism in Postharvest Fruit: An Omics Perspective. J Agric Food Chem. 2025, 73, 12525–12536. [Google Scholar] [CrossRef]
- Blagodatskaya, E; Blagodatsky, S; Anderson, T-H; Kuzyakov, Y. Microbial growth and carbon use efficiency in the rhizosphere and root-free soil. PLoS ONE 2014, 9, e93282. [Google Scholar] [CrossRef]
- Böhm, W; Wissemeier, AH. Contribution to the HiStory Series in Plant Nutrition. J Plant Nutr Soil Sci. 2025, 188, 585–592. [Google Scholar] [CrossRef]
- Cao, Y; Pi, H; Chandrangsu, P; et al. Antagonism of two plant-growth promoting Bacillus velezensis isolates against Ralstonia solanacearum and Fusarium oxysporum. Sci Rep. 2018, 8, 4360. [Google Scholar] [CrossRef]
- Chen, J; Zhu, B; Zhang, Y. A meta-analysis on the responses of soil microbial biomass and community structure to antibiotics. Appl Soil Ecol. 2023, 184, 104786. [Google Scholar] [CrossRef]
- Chow, LKM; Ghaly, TM; Gillings, MR. A survey of sub-inhibitory concentrations of antibiotics in the environment. J Environ Sci (China) 2021, 99, 21–27. [Google Scholar] [CrossRef] [PubMed]
- Clifford, JC; Buchanan, A; Vining, O; Kidarsa, TA; Chang, JH; McPhail, KL; Loper, JE. Phloroglucinol functions as an intracellular and intercellular chemical messenger influencing gene expression in Pseudomonas protegens. Environ Microbiol. 2016, 18, 3296–3308. [Google Scholar] [CrossRef] [PubMed]
- Combes-Meynet, E; Pothier, JF; Moënne-Loccoz, Y; Prigent-Combaret, C. The Pseudomonas secondary metabolite 2,4-diacetylphloroglucinol is a signal inducing rhizoplane expression of Azospirillum genes involved in plant-growth promotion. Mol Plant Microbe Interact. 2011, 24, 271–284. [Google Scholar] [CrossRef]
- Correa-Galeote, D; Bedmar, EJ; Arone, GJ. Maize endophytic bacterial diversity as affected by soil cultivation history. Front Microbiol. 2018, 9, 484. [Google Scholar] [CrossRef]
- Davies, J. What are antibiotics? Archaic functions for modern activities. Mol Microbiol. 1990, 4, 1227–1232. [Google Scholar]
- Dawson, M. Nitragin and the nodules of leguminous plants. Philos Trans R Soc Lond B Biol Sci. 1900, 192, 1–28. Available online: http://www.jstor.org/stable/91813 (accessed on 26 November 2025).
- Debois, D; Fernandez, O; Franzil, L; et al. Plant polysaccharides initiate underground crosstalk with bacilli by inducing synthesis of the immunogenic lipopeptide surfactin. Environ Microbiol Rep. 2015, 7, 570–582. [Google Scholar] [CrossRef]
- D'Abramo, F; Neumeyer, S. A historical and political epistemology of microbes. Centaurus 2020, 62, 321–330. [Google Scholar] [CrossRef]
- Fang, J; Blaschkauer, M; Distelfeld, A; et al. Comparison of Rhizosphere Microbiomes Between Domesticated and Wild Wheat in a Typical Agricultural Field: Insights into Microbial Community Structure and Functional Shifts. J Fungi (Basel) 2025, 11, 168. [Google Scholar] [CrossRef]
- Fajardo, A; Martínez, JL. Antibiotics as signals that trigger specific bacterial responses. Curr Opin Microbiol. 2008, 11, 161–167. [Google Scholar] [CrossRef]
- Gfeller, V; Schneider, M; Bodenhausen, N; et al. Crop genotype modulates root rot resistance-associated microbial community composition and abundance of key taxa. Environ Microbiome 2025, 20, 53. [Google Scholar] [CrossRef] [PubMed]
- Glare, T.; Caradus, J; Gelernter, W.; Jackson, T.; Keyhani, N.; Köhl, J.; Marrone, P.; Morin, L.; Stewart, A. Have biopesticides come of age? Trends in Biotechnology 2012, 30(5), 250–258. [Google Scholar] [CrossRef] [PubMed]
- Hamonts, K; Trivedi, P; Garg, A; et al. Field study reveals core plant microbiota and relative importance of their drivers. Environ Microbiol. 2018, 20, 124–140. [Google Scholar] [CrossRef] [PubMed]
- Han, X; Shen, D; Xiong, Q; et al. The plant-beneficial rhizobacterium Bacillus velezensis FZB42 controls the soybean pathogen Phytophthora sojae due to bacilysin production. Appl Environ Microbiol. 2021, 87, e01601–e01621. [Google Scholar] [CrossRef]
- Hernández-Terán, A; Escalante, AE; Rebolleda-Gómez, M. Plant domestication does not reduce diversity in rhizosphere bacterial communities. New Phytol 2025. [Google Scholar] [CrossRef]
- Huang, Y; Tang, S; Liu, R; et al. Soil organic carbon mediates plant immunity-rhizosphere microbiome interactions and controls colonization resistance to microbial inoculants. Cell Host Microbe 2025, 33, 1929–1944.e7. [Google Scholar] [CrossRef]
- Just, BS; Marks, EAN; Roquer-Beni, L; Llenas, L; Ponsà, S; Vilaplana, R. Biofertilization increases soil organic carbon concentrations: results of a meta-analysis. Int J Agric Sustain. 2024, 22, 2361578. [Google Scholar] [CrossRef]
- Kandel, SL; Joubert, PM; Doty, SL. Bacterial Endophyte Colonization and Distribution within Plants. Microorganisms 2017, 5, 77. [Google Scholar] [CrossRef] [PubMed]
- Karasov, TL; Neumann, M; Leventhal, L; Pathodopsis Team. Continental-scale associations of Arabidopsis thaliana phyllosphere members with host genotype and drought. Nat Microbiol. 2024, 9, 2748–2758. [Google Scholar] [CrossRef] [PubMed]
- Kinsella, K; Schulthess, CP; Morris, TF; Stuart, JD. Rapid quantification of Bacillus subtilis antibiotics in the rhizosphere. Soil Biol Biochem. 2009, 41, 374–379. [Google Scholar] [CrossRef]
- Linares, JF; Gustafsson, I; Baquero, F; Martínez, JL. Antibiotics as intermicrobial signaling agents instead of weapons. Proc Natl Acad Sci U S A 2006, 103, 19484–19489. [Google Scholar] [CrossRef]
- Ling, N; Wang, T; Kuzyakov, Y. Rhizosphere bacteriome structure and functions. Nat Commun. 2022, 13, 836. [Google Scholar] [CrossRef]
- Li, T; Li, D; Chen, X; Gérard, F; Li, M; Wu, J; Gao, J. Compartments of roots and mature leaves are key hubs in the connectivity of tea-plant mycobiomes and are influenced by environmental factors and host age. Sci Total Environ. 2023, 893, 164827. [Google Scholar] [CrossRef]
- Li, Y; Sun, M; Raaijmakers, JM; et al. Predicting rhizosphere-competence-related catabolic gene clusters in plant-associated bacteria with rhizoSMASH. Nat Commun. 2025, 16, 8400. [Google Scholar] [CrossRef]
- Liu, Y; Jia, B; Ren, Y; et al. Bacterial social interactions in synthetic Bacillus consortia enhance plant growth. iMeta 2025, 4, e70053. [Google Scholar] [CrossRef]
- Lucas, JM; Sone, BM; Whitmore, D; Strickland, MS. Antibiotics and temperature interact to disrupt soil communities and nutrient cycling. Soil Biol Biochem. 2021, 163, 108437. [Google Scholar] [CrossRef]
- Lundberg, DS; Lebeis, SL; Paredes, SH; et al. Defining the core Arabidopsis thaliana root microbiome. Nature 2012, 488, 86–90. [Google Scholar] [CrossRef]
- Maurhofer, M; Baehler, E; Notz, R; Martinez, V; Keel, C. Cross talk between 2,4-diacetylphloroglucinol-producing biocontrol pseudomonads on wheat roots. Appl Environ Microbiol. 2004, 70, 1990–1998. [Google Scholar] [CrossRef]
- Mayer, T; Teutloff, E; Unger, K; et al. Deterministic colonization arises early during the transition of soil bacteria to the phyllosphere and is shaped by plant-microbe interactions. Microbiome 2025, 13, 102. [Google Scholar] [CrossRef]
- Mittelstrass, J; Sperone, FG; Horton, MW. Using transects to disentangle the environmental drivers of plant-microbiome assembly. Plant Cell Environ. 2021, 44, 3515–3525. [Google Scholar] [CrossRef]
- Mohkam-Singh; Nunes, Márcio R. A 30-Year overview of soil fertility, soil quality, and soil health research in Florida. Ecological Indicators, Volume Volume 176(2025), 113660. [CrossRef]
- Monica, S; Panneerselvam, S; Rajasekaran, R; et al. Recent Advances in Bioinoculant Formulations and Their Shelf-Life: A Review. Curr Microbiol. 2025, 82, 506. [Google Scholar] [CrossRef] [PubMed]
- Nobbe, F; Hiltner, L. Inoculation of the soil for cultivating leguminous plants. US Patent 570,813. 1896. Available online: https://patents.google.com/patent/US570813A/en.
- Okubara, PA; Bonsall, RF. Accumulation of Pseudomonas-derived 2,4-diacetylphloroglucinol on wheat seedling roots is influenced by host cultivar. Biol Control. 2008, 46, 322–331. [Google Scholar] [CrossRef]
- Poshvina, DV; Balkin, AS; Dilbaryan, DS; Vasilchenko, AS. Unravelling the response of the soil microbiome to macrolactin A: A metagenomic study. Chemosphere 2025, 387, 144645. [Google Scholar] [CrossRef] [PubMed]
- Qiao, Y; Wang, Z; Sun, H; et al. Synthetic community derived from grafted watermelon rhizosphere provides protection for ungrafted watermelon against Fusarium oxysporum via microbial synergistic effects. Microbiome 2024, 12, 101. [Google Scholar] [CrossRef]
- Raaijmakers, JM; Mazzola, M. Diversity and natural functions of antibiotics produced by beneficial and plant pathogenic bacteria. Annu Rev Phytopathol. 2012, 50, 403–424. [Google Scholar] [CrossRef]
- Robleto, EA; Borneman, J; Triplett, EW. Effects of bacterial antibiotic production on rhizosphere microbial communities from a culture-independent perspective. Appl Environ Microbiol. 1998, 64, 5020–5022. [Google Scholar] [CrossRef]
- Romero, D; Traxler, MF; López, D; Kolter, R. Antibiotics as signal molecules. Chem Rev. 2011, 111, 5492–5505. [Google Scholar] [CrossRef] [PubMed]
- Sarwar, A; Hassan, MN; Imran, M; et al. Biocontrol activity of surfactin A purified from Bacillus NH-100 and NH-217 against rice bakanae disease. Microbiol Res. 2018, 209, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Schandry, N; Jandrasits, K; Garrido-Oter, R; Becker, C. A practical guide to sequencing and analyzing synthetic bacterial communities. bioRxiv 2021, 2021.01.12.425818. [Google Scholar] [CrossRef]
- Selten, G; de Jonge, R. Bacillus drives functional states in synthetic plant root bacterial communities. Genome Biol. 2025, 26, 270. [Google Scholar] [CrossRef] [PubMed]
- Semenov, MV; Zhelezova, AD; Ksenofontova, NA; Ivanova, EA; Nikitin, DA; Semenov, VM. Microbiological Indicators for Assessing the Effects of Agricultural Practices on Soil Health: A Review. Agronomy 2025, 15, 335. [Google Scholar] [CrossRef]
- Spagnolo, F; Trujillo, M; Dennehy, JJ. Why Do Antibiotics Exist? mBio 2021, 12, e0196621. [Google Scholar] [CrossRef] [PubMed]
- Stringlis, IA; Zhang, H; Pieterse, CMJ; Bolton, MD; de Jonge, R. Microbial small molecules—weapons of plant subversion. Nat Prod Rep. 2018, 35, 410–433. [Google Scholar] [CrossRef]
- Sun, X; Chen, J; Kuzyakov, Y; et al. Meta-analysis shows that planting nitrogen-fixing species increases soil organic carbon stock. Nat Ecol Evol. 2025, 9, 2046–2056. [Google Scholar] [CrossRef]
- Teslya, AV; Gurina, EV; Poshvina, DV; Stepanov, AA; Iashnikov, AV; Vasilchenko, AS. Fungal secondary metabolite gliotoxin enhances enzymatic activity in soils by reshaping their microbiome. Rhizosphere 2024, 32, 100960. [Google Scholar] [CrossRef]
- Teslya, AV; Iashnikov, AV; Poshvina, DV; Stepanov, AA; Vasilchenko, AS. Extracellular enzymes of soils under organic and conventional cropping systems: Predicted functional potential and actual activity. Agronomy 2024, 14, 2634. [Google Scholar] [CrossRef]
- Teslya, AV; Stepanov, AA; Poshvina, DV; Petrushin, IS; Vasilchenko, AS. From lab to field: Context-dependent impacts of Pseudomonas-produced 2,4-diacetylphloroglucinol on soil microbial ecology. Biomolecules 2025, 15, 1578. [Google Scholar] [CrossRef]
- Thoenen, L; Kreuzer, M; Pestalozzi, C; et al. The lactonase BxdA mediates metabolic specialisation of maize root bacteria to benzoxazinoids. Nat Commun. 2024, 15, 6535. [Google Scholar] [CrossRef] [PubMed]
- Tian, B; Zhang, C; Ye, Y; et al. Beneficial traits of bacterial endophytes belonging to the core communities of the tomato root microbiome. Agric Ecosyst Environ. 2017, 247, 149–156. [Google Scholar] [CrossRef]
- Troppens, DM; Chu, M; Holcombe, LJ; Gleeson, O; O’Gara, F; Read, ND; Morrissey, JP. The bacterial secondary metabolite 2,4-diacetylphloroglucinol impairs mitochondrial function and affects calcium homeostasis in Neurospora crassa. Fungal Genet Biol. 2013, 56, 135–146. [Google Scholar] [CrossRef] [PubMed]
- Vasilchenko, AS; Dilbaryan, DS; Poshvina, DV; et al. Bacillus-derived cyclic lipopeptides modulate microbiome structure and enzymatic activity of soil and increase productivity of potato (Solanum tuberosum L.). Rhizosphere 2025, 33, 101033. [Google Scholar] [CrossRef]
- Vasilchenko, AV; Vasilchenko, AS. Plaggic anthrosol in modern research: Genesis, properties and carbon sequestration potential. CATENA 2024, 234, 107626. [Google Scholar] [CrossRef]
- Voeleker, JA. 'Nitragin' or the use of 'pure cultivation' bacteria for leguminous crops. J R Agric Soc Engl. 1896, 7, 253–264. [Google Scholar]
- Waksman, SA. What is an antibiotic or an antibiotic substance? Mycologia 1947, 39, 565–569. [Google Scholar] [CrossRef] [PubMed]
- Wemheuer, F; Kaiser, K; Karlovsky, P; Daniel, R; Vidal, S; Wemheuer, B. Bacterial endophyte communities of three agricultural important grass species differ in their response towards management regimes. Sci Rep. 2017, 7, 40914. [Google Scholar] [CrossRef] [PubMed]
- Xiao, L; Zhao, Q; Deng, J; Cui, L; Zhang, T; Yang, Q; Zhao, S. Comparative analysis of rhizosphere microbiomes in different blueberry cultivars. Horticulturae 2025, 11, 696. [Google Scholar] [CrossRef]
- Yim, G; Wang, HH; Davies, J. Antibiotics as signalling molecules. Philos Trans R Soc Lond B Biol Sci. 2007, 362, 1195–1200. [Google Scholar] [CrossRef]
- Yuan, J; Zhao, M; Li, R; et al. Antibacterial compound macrolactin alters the soil bacterial community and abundance of the gene encoding PKS. Front Microbiol. 2016, 7, 1904. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



