Submitted:
31 March 2026
Posted:
03 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
3. Results
3.1. Correlation Matrix Among Variables
3.2. Variable-by-Variable Basis
3.2.1. Mean Length of the Setae Surrounding the Compound Eye (SM)
3.2.2. Number of Interommatidial Setae (NEIS)
3.2.3. Number of Ommatidia
3.3. Interocular Correlation
3.3.1. Ommatidia
3.3.2. NEIS
3.4. Univariate Discriminative Power of the Variables
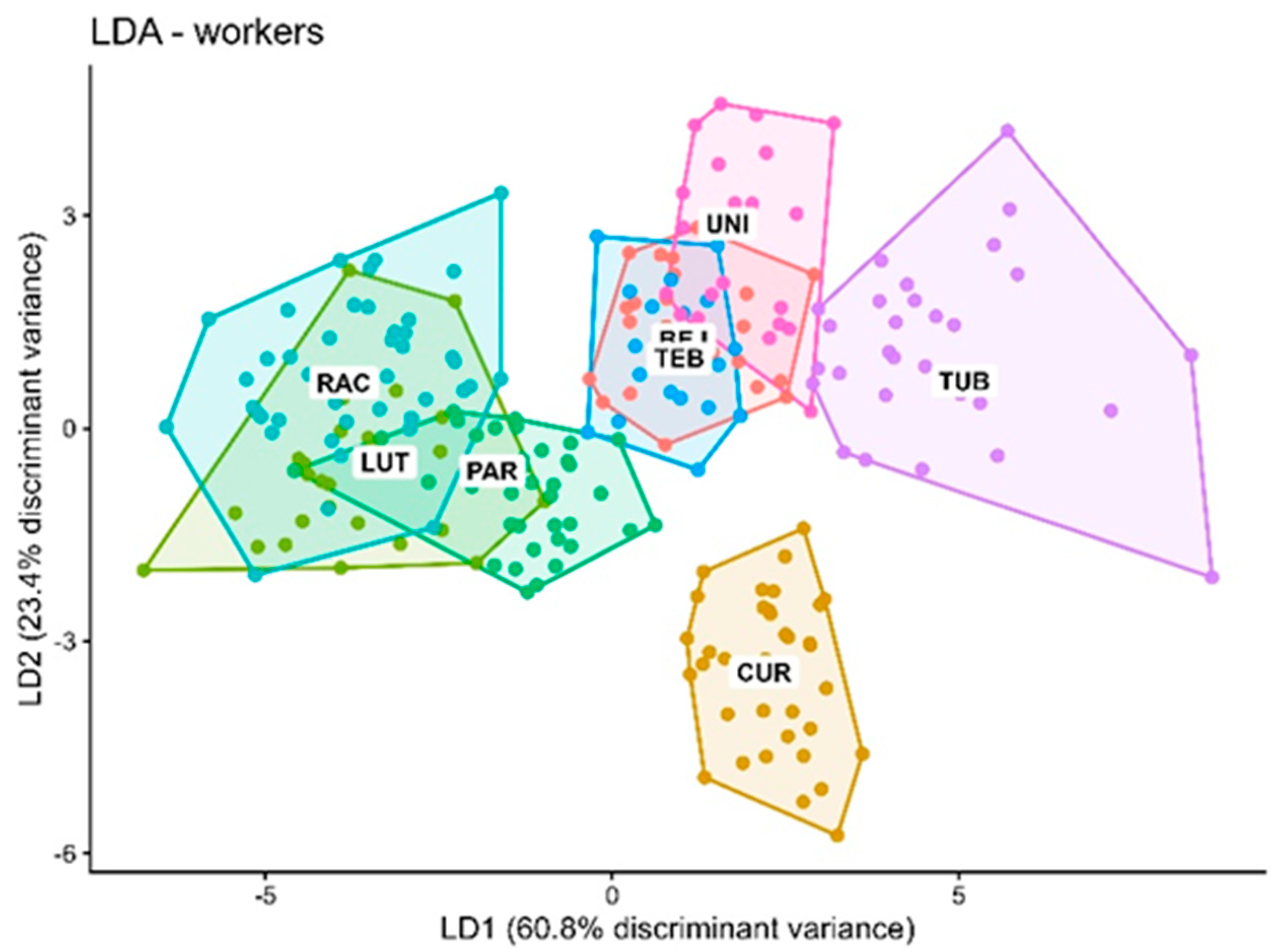
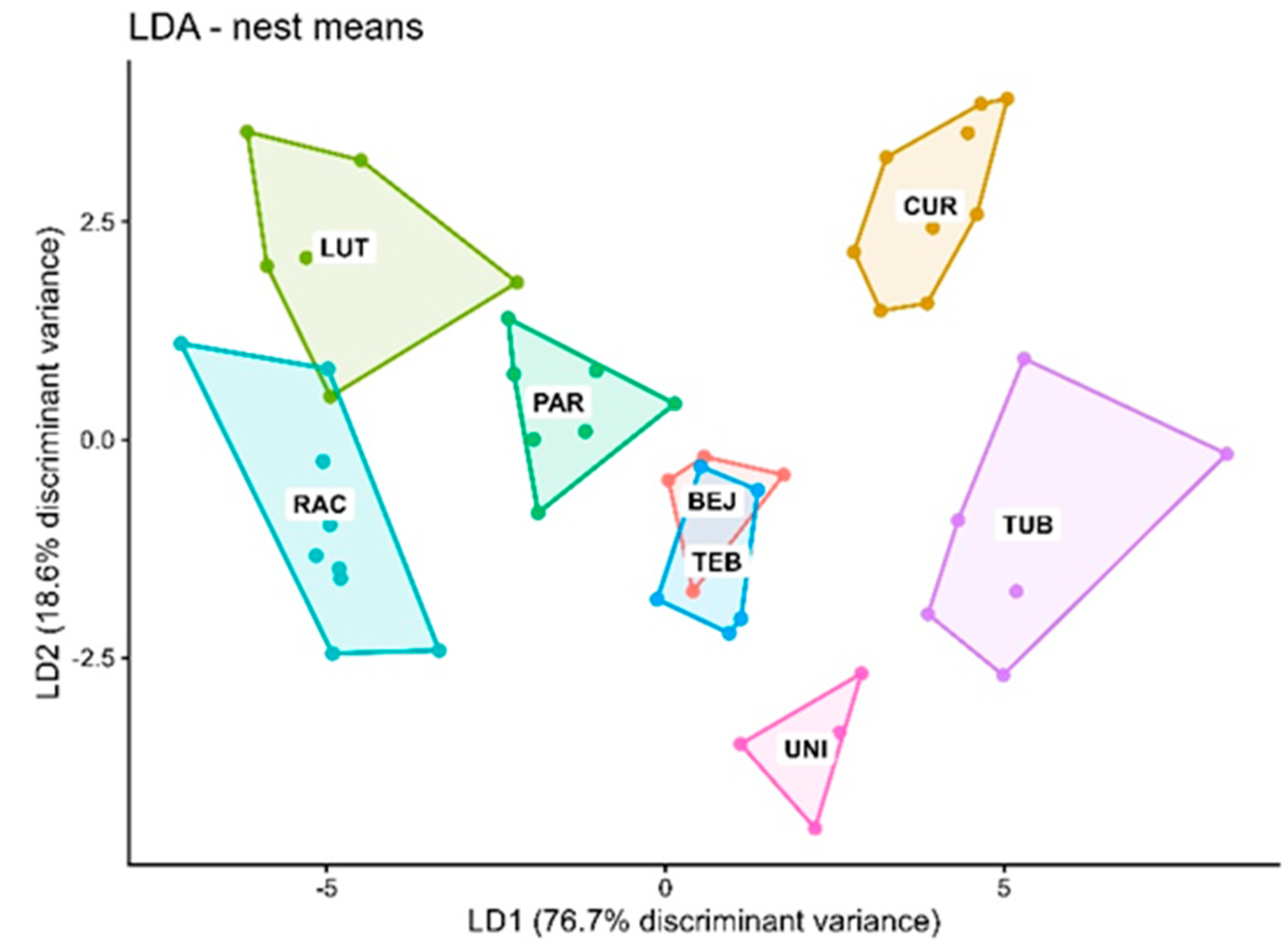
3.5. Comparison of Interspecific Discrimination at the Individual vs. Nest Level
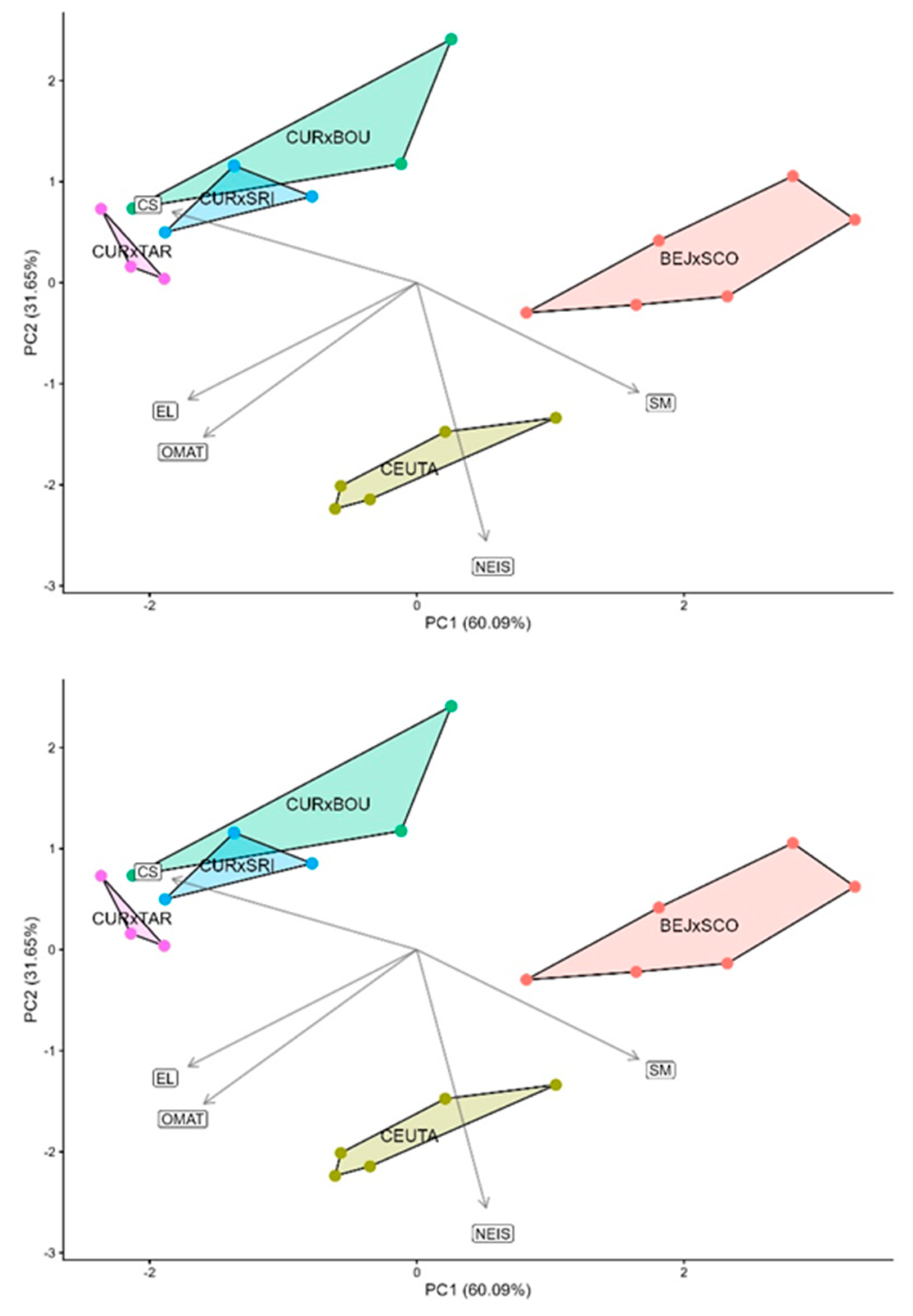
3.6. Application to the Tebesae Complex
4. Discussion
5. Conclusions
Supplementary Materials
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
| species. | zone code | zone | province | latitude | longitude | Alt(m) |
|---|---|---|---|---|---|---|
| Temnothorax bejaraniensis | BEJ | Sierra de Córdoba | Córdoba | 37.918333 | -4.887500 | 410 |
| Temnothorax cf tebessae | TEB-CEU | Ceuta, Spain (N. Africa) | Ceuta | 35.913611 | -5.370834 | 65 |
| Temnothorax cutulus | CUR-EST | Los Alcornocales Natural Park | Cádiz | 36.084667 | -5.533000 | 390 |
| CUR-BOU | Bouhachem National Park (N. Africa) | Chefchaouen | 35.346830 | -5.550360 | 725 | |
| CUR-SSU | Sierras Subbéticas Natural Park | Córdoba | 37.459750 | -4.351432 | 850 | |
| Temnothorax luteus | LUT-CAZ | Sierras de Cazorla, Segura y Las Villas Natural Park | Jaen | 37.813978 | -2.960062 | 1807 |
| LUT-SMA | Sierra Mágina Natural Park | Jaen | 37.715660 | -3.452142 | 1748 | |
| Temnothorax pardoi | PAR-CAZ | Sierras de Cazorla, Segura y Las Villas Natural Park | Jaen | 38.318962 | -2.578504 | 1309 |
| PAR-SGU | PN de la Sierra de Guadarrama Natural Park | Madrid | 40.867149 | -3.763088 | 1508 | |
| Temnothorax racovitzai | RAC-CAR | Sierra de Cardeña y Montoro Natural Park | Córdoba | 38.133328 | -4.276874 | 567 |
| RAC-SCO | Sierra de Córdoba | Córdoba | 37.918333 | -4.887500 | 410 | |
| Temnothorax tuberum | TUB | Sierra de Albarracín | Teruel | 40.484846 | -1.587982 | 1822 |
| Temnothorax unifasciatus | UNI-SGU | Sierra de Guadarrama National Park | Madrid | 40.867149 | -3.763088 | 1508 |
References
- Garcia, F.H.; Fischer, G.; Liu, C.; Audisio, T.L.; Alpert, G.D.; Fisher, B.L.; Economo, E.P. X-Ray microtomography for ant taxonomy: An exploration and case study with two new Terataner (Hymenoptera, Formicidae, Myrmicinae) species from Madagascar. PLOS ONE 2017, 12, e0172641. [Google Scholar] [CrossRef] [PubMed]
- Friedrich, F.; Matsumura, Y.; Pohl, H.; Bai, M.; Hörnschemeyer, T.; Beutel, R.G. Insect morphology in the age of phylogenomics: innovative techniques and its future role in systematics. Èntomol. Sci. 2013, 17, 1–24. [Google Scholar] [CrossRef]
- Mensa, F.S.; Muzzi, M.; Spani, F.; Tromba, G.; Dullin, C.; Di Giulio, A. When the Utility of Micro-Computed Tomography Collides with Insect Sample Preparation: An Entomologist User Guide to Solve Post-Processing Issues and Achieve Optimal 3D Models. Appl. Sci. 2022, 12, 769. [Google Scholar] [CrossRef]
- Roháček, J.; Hammel, J.U.; Baranov, V. Christelenkidae, a new extinct family based on a new taxon from Eocene Baltic amber (Diptera: Acalyptratae), with X-ray synchrotron microtomography imaging of its structures. Arthropod Syst. Phylogeny 2023, 81, 475–498. [Google Scholar] [CrossRef]
- Nelson, L.; Doucet, D.S.; Cook, J.L. Morphology of cuticular structures in the ant genus Strumigenys (Insecta: Hymenoptera: Formicidae). Zoomorphology 2024, 143, 643–652. [Google Scholar] [CrossRef]
- Birkenfeld, V.; Gorb, S.N.; Krings, W. Mandible elemental composition and mechanical properties from distinct castes of the leafcutter ant Atta laevigata (Attini; Formicidae). Interface Focus 2024, 14, 20230048. [Google Scholar] [CrossRef]
- Wang, C.; Chung, F.-Y.; Lin, C.-C.; Gibson, J.C.; McGuire, S.; Suarez, A.V.; Billen, J. The spongiform tissue in Strumigenys ants contains exocrine glands. Arthropod Struct. Dev. 2023, 73, 101246. [Google Scholar] [CrossRef]
- Morris, P. Animal Eyes (Oxford Animal Biology Series) - By Michael F. Land & Dan-Eric Nilsson. Zoöl. J. Linn. Soc. 2012, 166, 912–912. [Google Scholar] [CrossRef]
- Warrant, E.; Dacke, M. Vision and Visual Navigation in Nocturnal Insects. Annu. Rev. Èntomol. 2011, 56, 239–254. [Google Scholar] [CrossRef]
- Agavekar, G.; Garcia, F.H.; Economo, E.P. Taxonomic overview of the hyperdiverse ant genusTetramoriumMayr (Hymenoptera, Formicidae) in India with descriptions and X-ray microtomography of two new species from the Andaman Islands. PeerJ 2017, 5, e3800–e3800. [Google Scholar] [CrossRef]
- Fischer, G.; Garcia, F.H.; Peters, M.K. Taxonomy of the ant genus Pheidole Westwood (Hymenoptera: Formicidae) in the Afrotropical zoogeographic region: definition of species groups and systematic revision of the Pheidole pulchella group. Zootaxa 2012, 3232, 1–43. [Google Scholar] [CrossRef]
- Garcia, F.H.; Fischer, G.; Liu, C.; Audisio, T.L.; Alpert, G.D.; Fisher, B.L.; Economo, E.P. X-Ray microtomography for ant taxonomy: An exploration and case study with two new Terataner (Hymenoptera, Formicidae, Myrmicinae) species from Madagascar. PLOS ONE 2017, 12, e0172641. [Google Scholar] [CrossRef]
- Johnson, R.A.; Rutowski, R.L. Color, activity period, and eye structure in four lineages of ants: Pale, nocturnal species have evolved larger eyes and larger facets than their dark, diurnal congeners. PLOS ONE 2022, 17, e0257779. [Google Scholar] [CrossRef] [PubMed]
- Borowiec, M.; Snelling, R.; Prebus, M. Studies on California ants: a review of the genus Temnothorax (Hymenoptera, Formicidae). ZooKeys 2014, 372, 27–89. [Google Scholar] [CrossRef] [PubMed]
- Prebus, M.M. Taxonomic revision of theTemnothorax salviniclade (Hymenoptera: Formicidae), with a key to the clades of New WorldTemnothorax. PeerJ 2021, 9, e11514. [Google Scholar] [CrossRef] [PubMed]
- Hunt, E.R.; Dornan, C.; Sendova-Franks, A.B.; Franks, N.R. Asymmetric ommatidia count and behavioural lateralization in the ant Temnothorax albipennis. Sci. Rep. 2018, 8, 1–11. [Google Scholar] [CrossRef]
- Cagniant, H.; Espadaler, X. Les Leptothorax, Epimyrma et chalepoxenus du Maroc (Hymenoptera: Formicidae). Clé et catalogue des Espèces. Ann. De La Soc. Èntomol. De Fr. 1997, 33, 259–284. [Google Scholar] [CrossRef]
- Campos, B.B.; Correia, A.O.; Campos, L.A.; Fernandes, J.A.M. Morphology of interommatidial sensilla of Discocephalini (Heteroptera: Pentatomidae: Discocephalinae). Arthropod Struct. Dev. 2022, 72. [Google Scholar] [CrossRef]
- Seifert, B.; Csosz, S. Temnothorax crasecundus sp. n. – a cryptic Eurocaucasian ant species (Hymenoptera, Formicidae) discovered by Nest Centroid Clustering. ZooKeys 2015, 479, 37–64. [Google Scholar] [CrossRef]
- Seifert, B. Cryptic Species in Ants (Hymenoptera: Formicidae) Revisited: We Need a Change in the Alpha-Taxonomic Approach. Myrmecol. News 2009, 12, 149–166. [Google Scholar] [CrossRef]
- Li, X.; Wiens, J.J. Estimating Global Biodiversity: The Role of Cryptic Insect Species. Syst. Biol. 2022, 72, 391–403. [Google Scholar] [CrossRef]
- Ramirez-Esquivel, F.; Leitner, N.E.; Zeil, J.; Narendra, A. The sensory arrays of the ant, Temnothorax rugatulus. Arthropod Struct. Dev. 2017, 46, 552–563. [Google Scholar] [CrossRef]
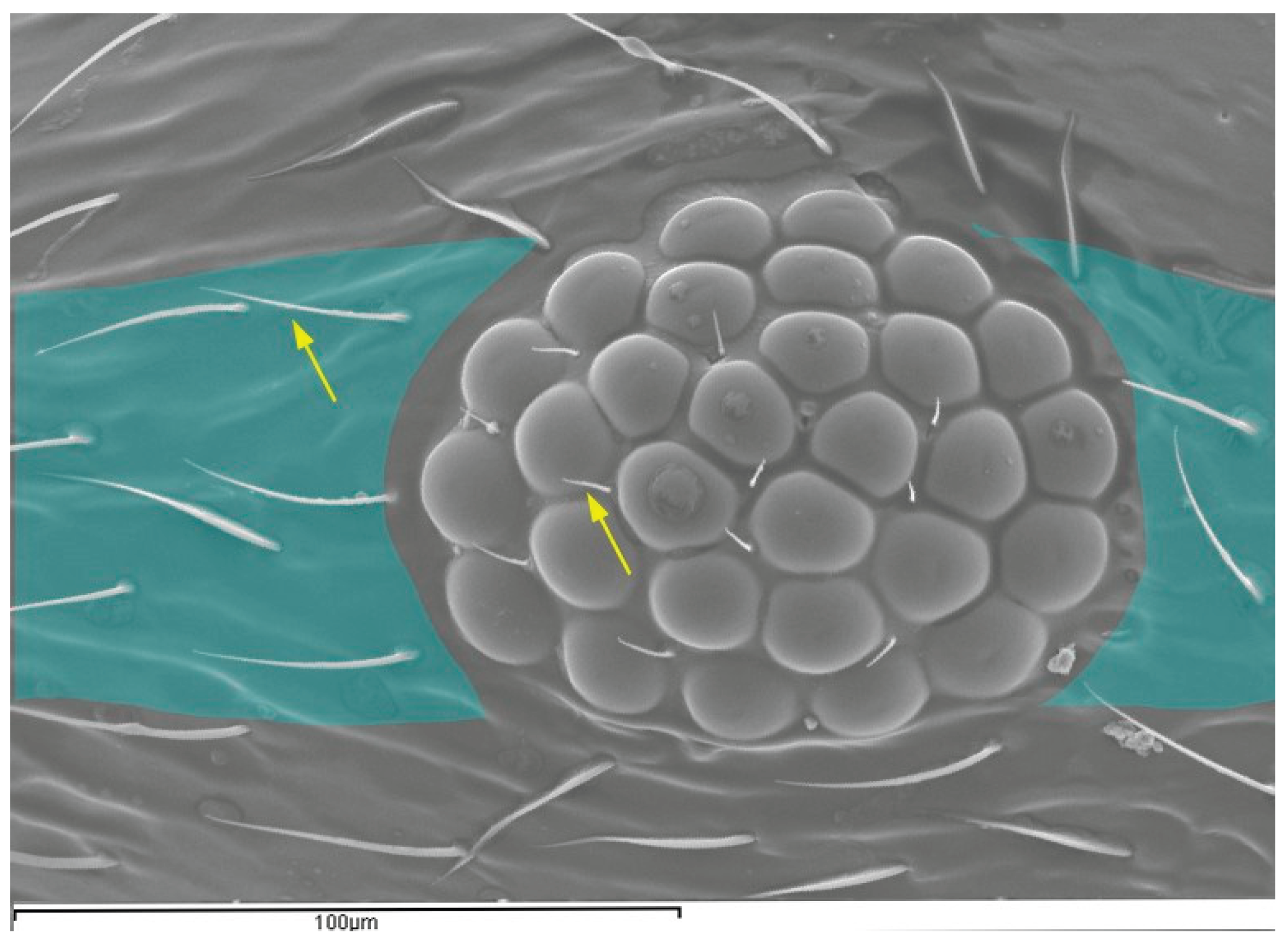
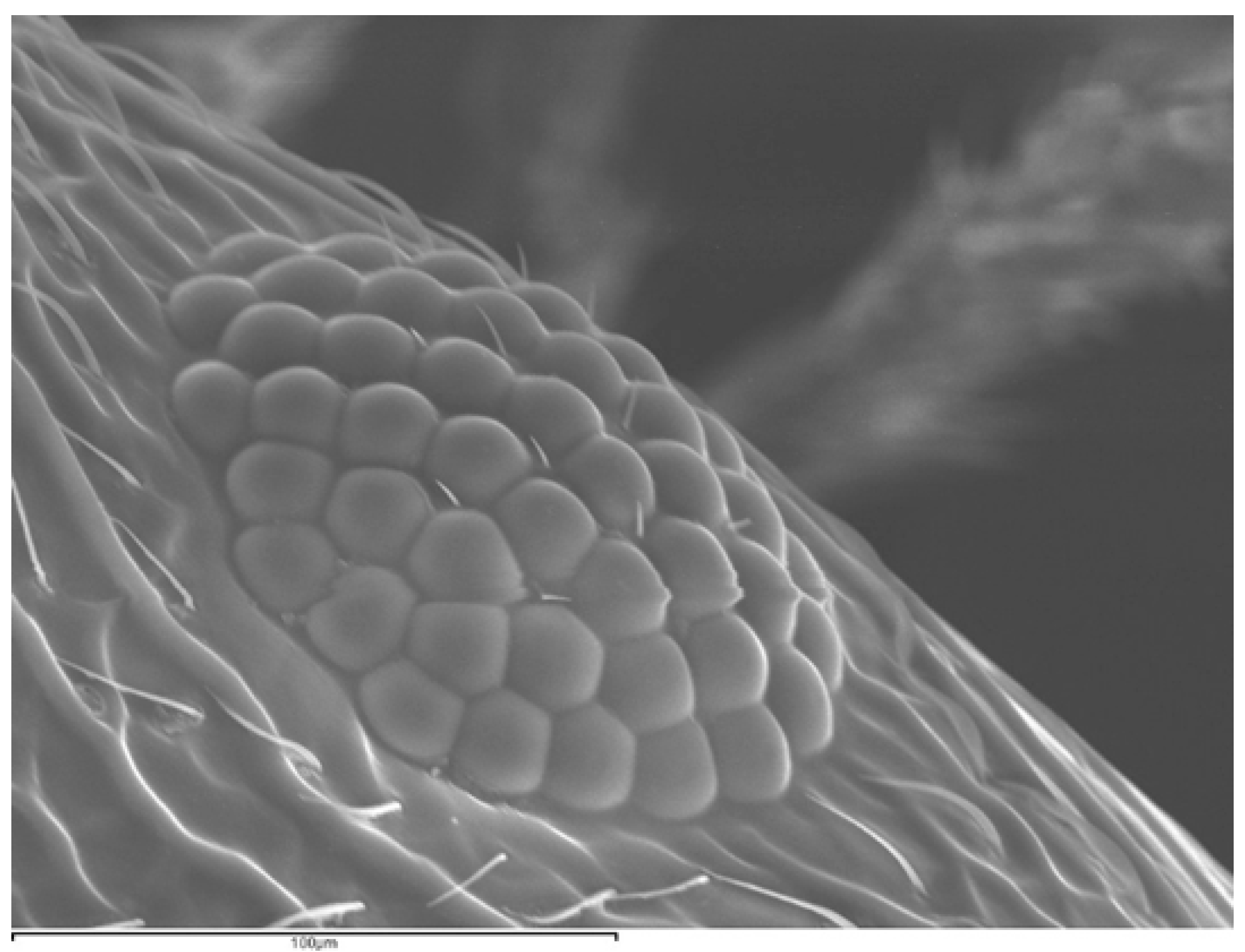
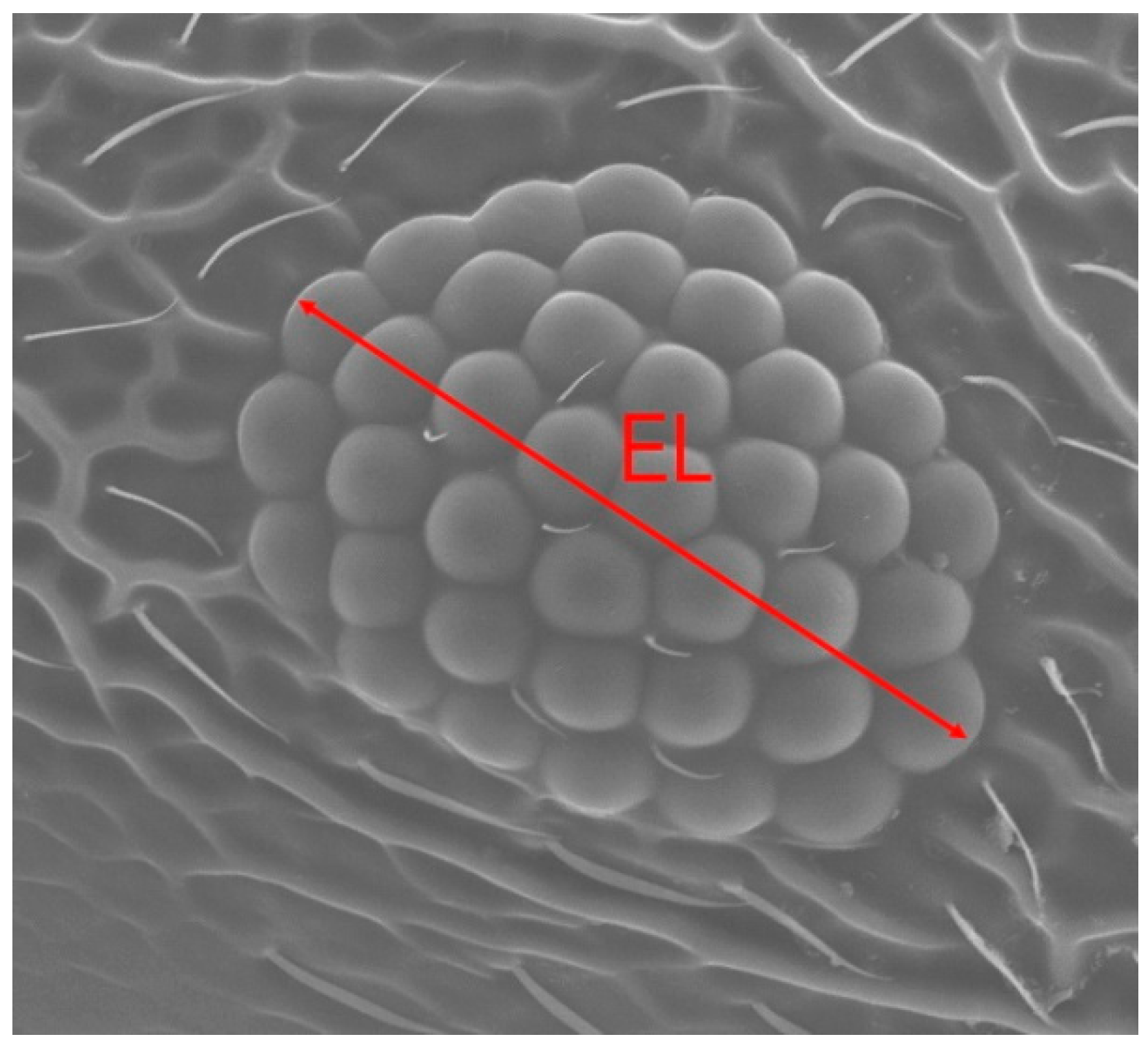
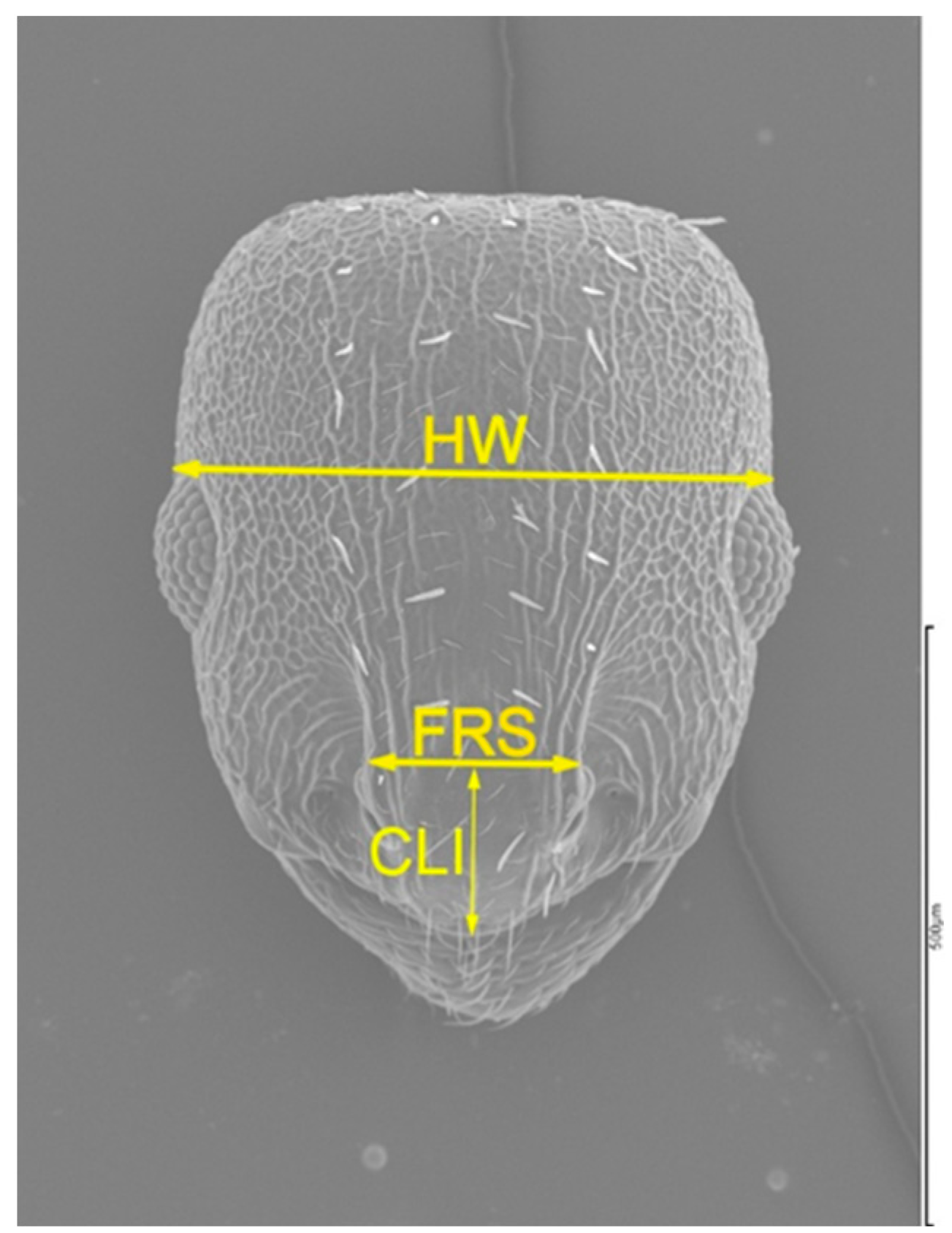
| Variable | SM | OMAT | NEIS | EL |
| CS | -0.165** | 0.854*** | 0.163* | 0.915*** |
| FRS | -0.083ns | 0.858*** | 0.097ns | 0.899*** |
| CLI | -0.331*** | 0.131* | 0.042ns | 0.185** |
| ML | -0.005ns | 0.878*** | 0.180** | 0.895*** |
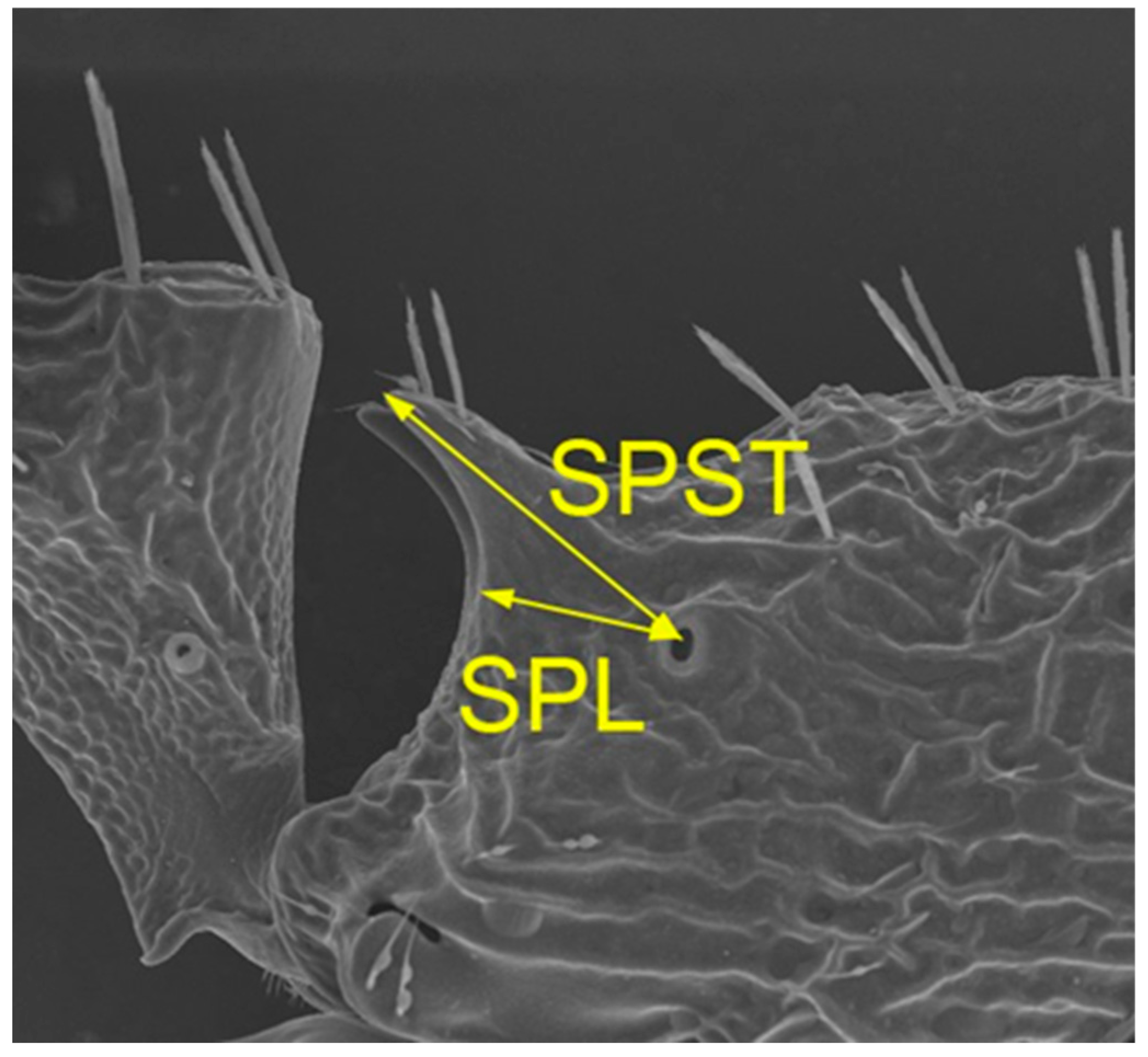
| SPST | -0.158* | 0.473*** | 0.499*** | 0.576*** |
| SPL | 0.008ns | 0.516*** | 0.450*** | 0.594*** |
| SPP | n | SM (µm) | OMAT | NEIS | EL (µm) |
| BEJ | 28 | 29.07 ± 2.73 a | 35.32 ± 3.97 f | 8.18 ± 1.68 c | 114.96 ± 6.70 e |
| CUR | 38 | 21.69 ± 2.32 c | 40.92 ± 5.15 e | 6.76 ± 1.15 d | 122.95 ± 7.42 d |
| LUT | 27 | 25.51 ± 1.77 b | 55.04 ± 5.10 d | 10.19 ± 1.47 b | 141.11 ± 8.08 b |
| PAR | 38 | 22.36 ± 1.18 c | 62.87 ± 4.65 c | 9.68 ± 1.45 b | 145.87 ± 6.89 b |
| RAC | 44 | 24.38 ± 1.76 b | 55.36 ± 8.24 d | 11.75 ± 2.06 a | 138.36 ± 11.43 c |
| TEB | 21 | 26.86 ± 2.64 a | 43.76 ± 4.84 e | 10.62 ± 1.32 a | 124.00 ± 6.16 d |
| TUB | 29 | 27.61 ± 2.60 a | 76.10 ± 8.60 b | 8.90 ± 1.61 b | 155.72 ± 8.73 a |
| UNI | 21 | 25.55 ± 1.41 b | 89.86 ± 9.03 a | 8.90 ± 1.37 b | 160.76 ± 9.44 a |
| Variable | η2 (Welch ANOVA) | LDA accuracy | RF accuracy | Ranking |
| OMAT | 0.860 | 0.505 | 0.415 | 1 |
| FRS | 0.855 | 0.511 | 0.476 | 2 |
| SPST | 0.847 | 0.435 | 0.39 | 3 |
| EL | 0.747 | 0.403 | 0.325 | 4 |
| ML | 0.728 | 0.407 | 0.354 | 5 |
| SM | 0.583 | 0.374 | 0.309 | 6 |
| SPL | 0.542 | 0.309 | 0.284 | 7 |
| NEIS | 0.511 | 0.366 | 0.342 | 8 |
| CLI | 0.191 | 0.195 | 0.325 | 9 |
| Variable (residual) | η2 | LDA accuracy | RF accuracy | Ranking |
| SPST | 0.875 | 0.545 | 0.443 | 1 |
| OMAT | 0.678 | 0.371 | 0.284 | 2 |
| SM | 0.561 | 0.317 | 0.273 | 3 |
| FRS | 0.546 | 0.306 | 0.212 | 4 |
| NEIS | 0.520 | 0.350 | 0.366 | 5 |
| SPL | 0.506 | 0.289 | 0.227 | 6 |
| EL | 0.490 | 0.305 | 0.223 | 7 |
| ML | 0.307 | 0.321 | 0.317 | 8 |
| CLI | 0.191 | 0.118 | 0.318 | 9 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




