Submitted:
01 April 2026
Posted:
01 April 2026
You are already at the latest version
Abstract
Introduction: The use of medicinal plants to cure human diseases is going on since the development of human civilization. Also, the discovery of antibiotics had profound impact to reduced death rates against various pathogens but due to rise of resistance against these antibiotics are serious threats for human health. Henceforth, the world is looking for alternative approach and the use of active plant metabolites are one of them. Multidrug resistant Shiga toxin producing Escherichia coli (MDR STEC) are life threatening microorganisms worldwide and their ability to produce aggregated biofilm makes them tolerant to many antibiotics used for treating STEC infections. In this study effect of plant metabolites were studied against MDR STEC samples. Methods: The different parts of 10 medicinal plants reported from central India were used in this study. Extract preparation and active fractions were used to test antimicrobial activity against MDR-STEC through measuring zone of inhibition. The biofilm structure was observed using electron microscopy. Results: Total 20 MDR STECs were identified out of 100 STEC samples. The intimin (eae) gene responsible for drug resistance was present in 18 (90%) MDR STEC samples. STEC were producing more aggregated biofilm layer as compare to sensitive E. coli. The plant extracts isolated from Acacia auriculiformis, Albizia lebbeck and Gliricidia sepium showed significantly high antimicrobial activity against MDR STEC as compared to various antibiotics. Conclusions: The study will be helpful to develop new or alternate antimicrobial agents and therapy against MDR STEC by using metabolites from medicinal plants.

Keywords:
1. Introduction
2. Materials & Methods
2.1. Sample Collection & Isolation of STEC
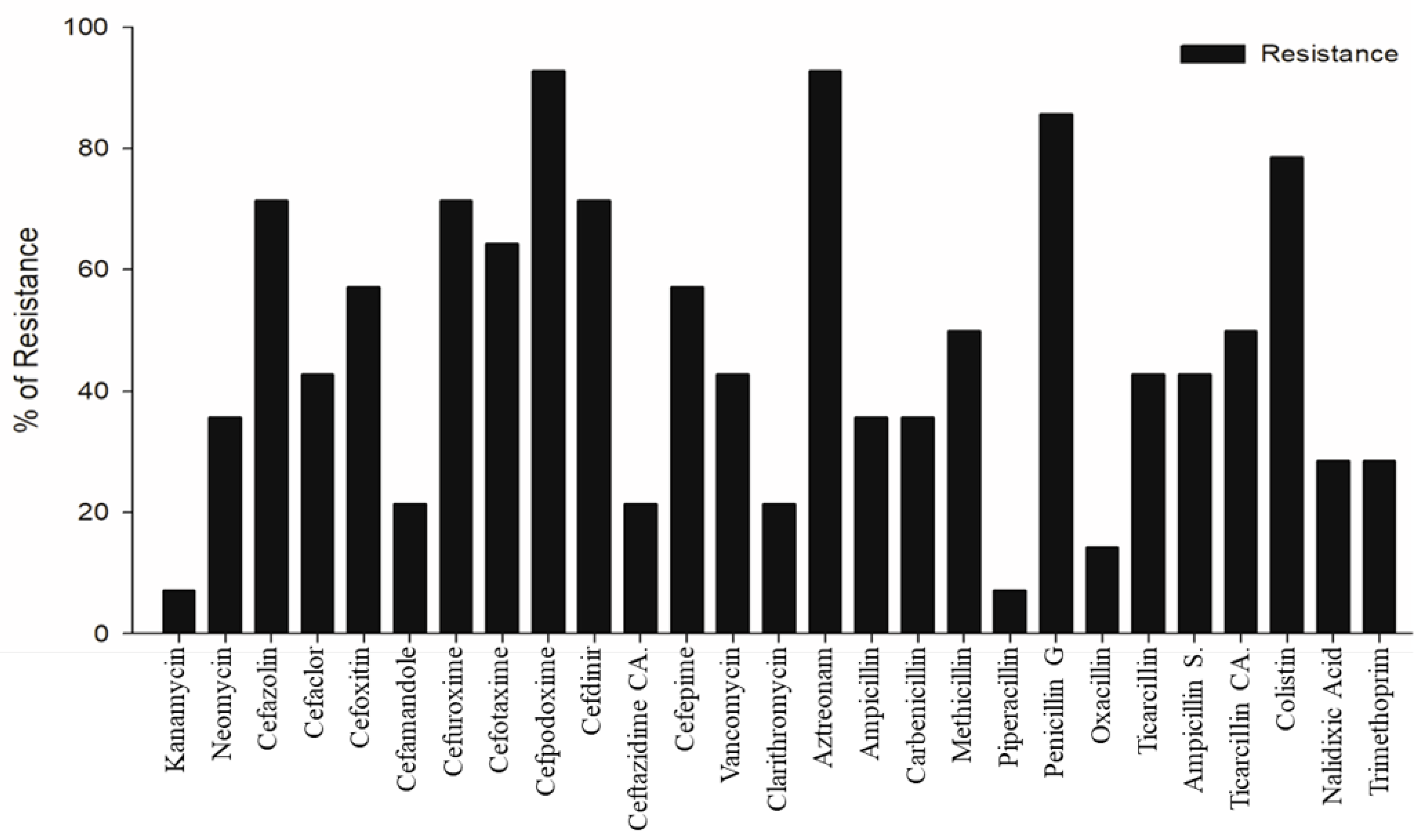
2.2. Antibiotic Sensitivity Test
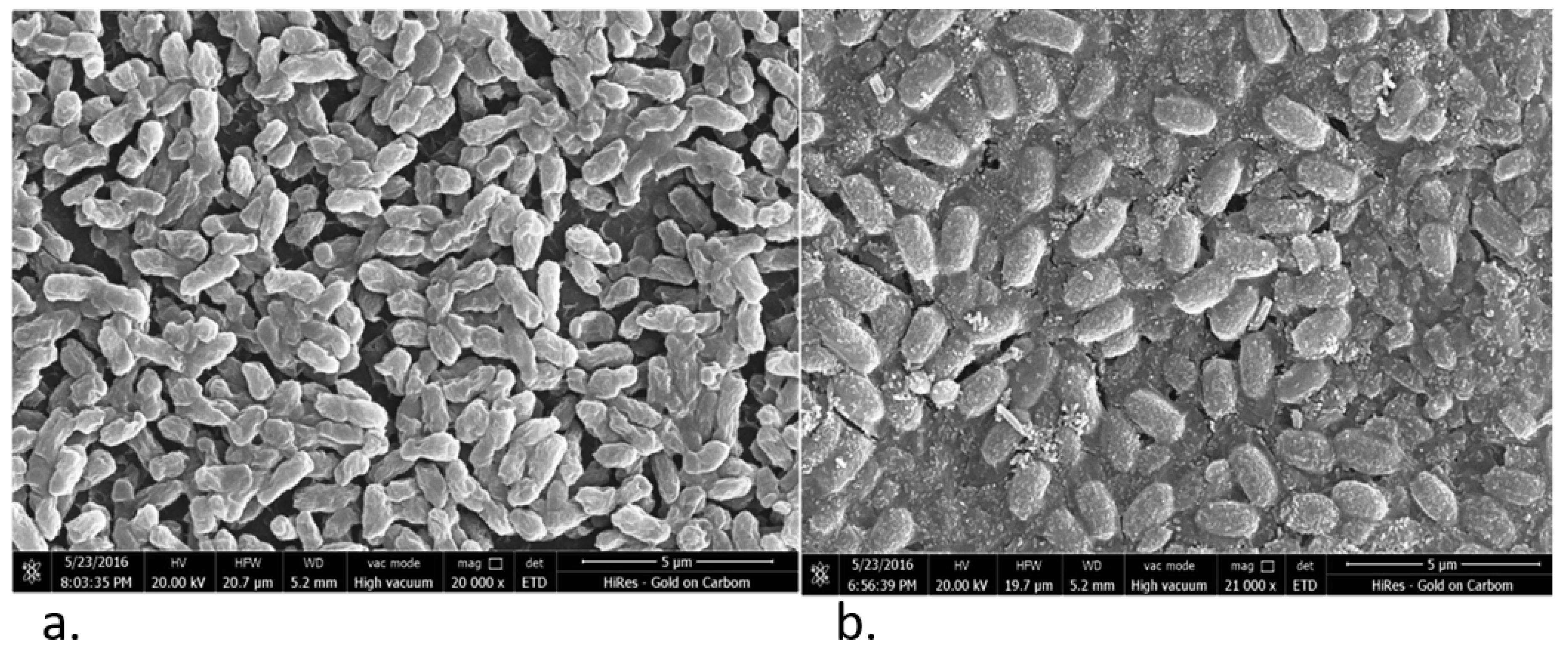
2.3. Biofilm Assay and Scanning Electron Microscopy
2.4. Extraction of Plant Materials
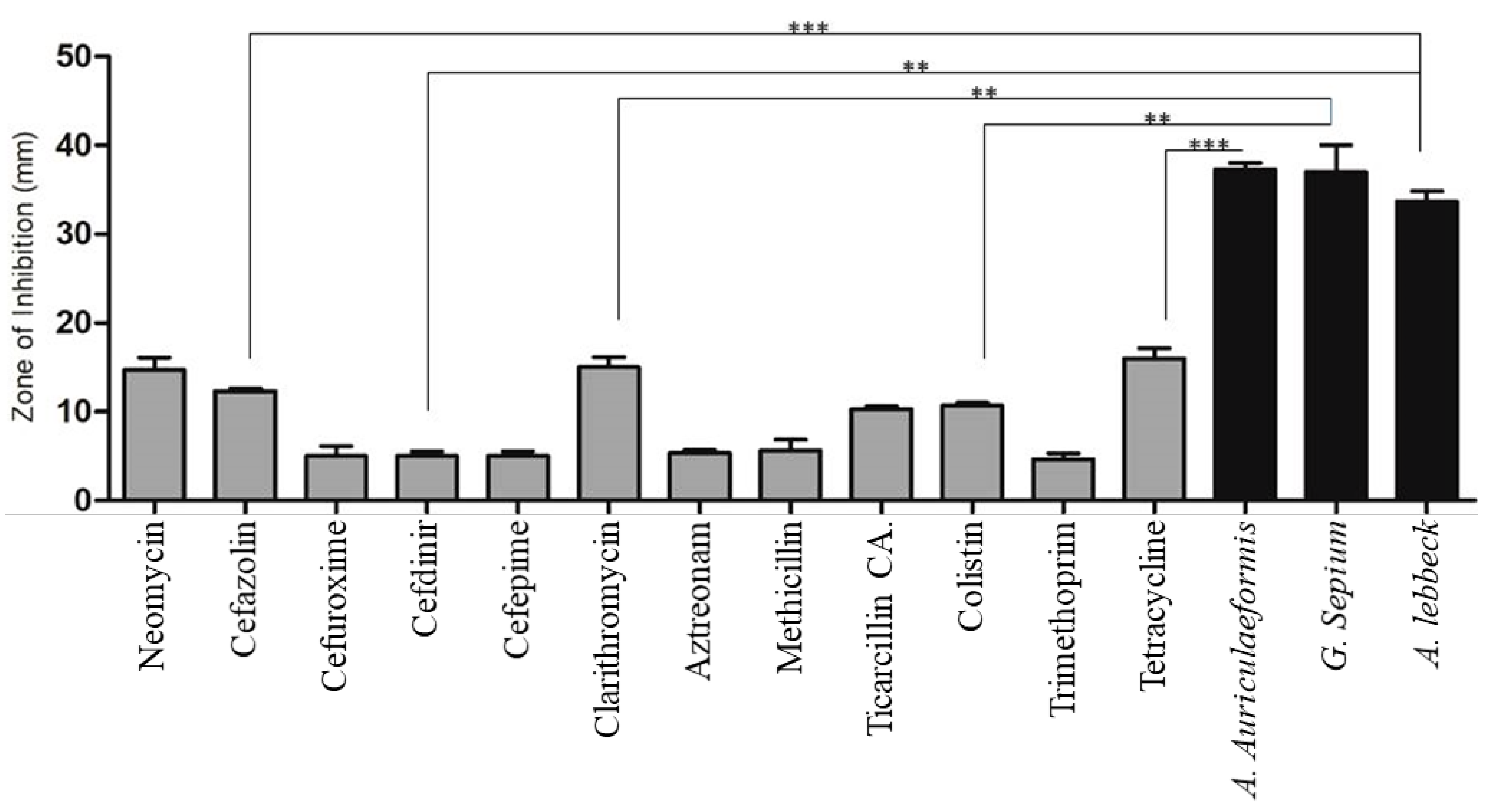
2.5. Susceptibility Test of Plant Extracts Against MDR STEC Isolates
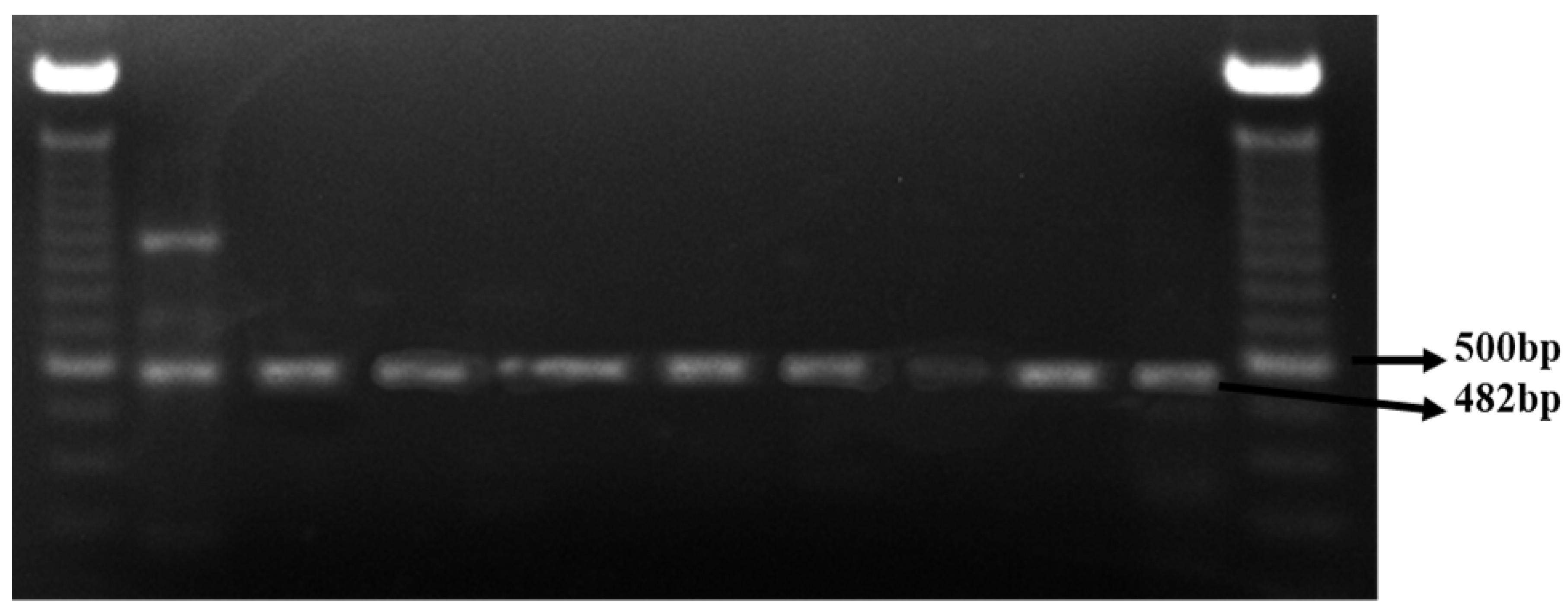
2.6. Intimin (eae) Genotype Detection
2.7. Statics
3. Results and Discussion
3.1. Characterization of MDR STEC
3.2. Antimicrobial Activity of Plant Extracts
3.3. Intimin (eae) Genotype Detection
4. Conclusions
Author Contributions
Financial support
Declaration of Competing Interests
Acknowledgments
References
- WHO Monographs on Selected Medicinal Plants- Volume 2 (1999), WHO, Genava.
- R.C. Agoha, Medicinal plants of Nigeria, Offsetdikker Jifaculteit waskunden, Natnurwenten schopp, pen, Netherlands, 1981, pp. 22-158.
- H.M. Burkill, The useful plants of West Tropical Africa vol 1, Families A-D, Royal Botanical Garden, Kew, 1984, pp. 441-415.
- H.M. Burkill, The useful plants of West Tropical Africa vol 3, Families J-L, Royal Botanical Garden, Kew, 1984, pp. 522.
- A. Sofowara, Medicinal plants and Traditional medicine in Africa. Spectrum Books Ltd, Ibadan, Nigeria, 1993, pp. 289.
- R. Taylor-Smith, Investigations on plants of West Africa, 111 Phytochemical Studies of some plants of Sierra Leone, Bull. Insst. France Afn noire A. 28 (1966) 528-541.
- H.O. Edeoga and A. Gomina, Nutritional values of some noncomventional leafy vegetables of Nigeria, J. Econ. Taxon. Bot. 24 (2000) 7-13.
- A.F. Hill, Economic Botany, A textbook of useful plants and plant products, 2nd ed., McGarw-Hill Book Company Inc, New York, 1952.
- D. Corogeanu, R. Willmes, M. Wolke, G. Plum, O. Utermohlen, M. Kronke, Therapeutic concentrations of antibiotics inhibit Shiga toxin release from enterohemorrhagic E. coli O104:H4 from the 2011 German outbreak, BMC Microbiol. 12 (2012) 160.
- Ahamad et al, Screening of some Indian medicinal plants for their antimi-crobial properties, Journal of Ethno pharmacology. 62 (1998) 183-193.
- J. Davis, Inactivation of the antibiotics and the dissemination of resistance genes, Science. 264 (1994) 375-382.
- S. Monroe and R. Polk, Antimicrobial use and bacterial resistance, Current Opinion Microbiology, 3 (2000) 496-501.
- G.E. Thomson, Further consideration of Asian Medicinal plants in treating common chronic disease in West, Journal of Medicinal Plants Research. 4 (2010) 125.
- S. Ethelberg, K.E. Olsen, F. Scheutz, C. Jensen, P, Schiellerup, J, Engberg et al, Virulence Factors for Hemolytic Uremic Syndrome, Denmark, Emerging Infectious Diseases. 10 (2004) 842-847. [CrossRef]
- M. Blanco, S. Schumacher, T. Tasara, C. Zweifel, J.E. Blanco, G. Dahbi, J. Blanco, R. Stephan, Serotypes, intimin variants and other virulence factors of eae positive Escherichia coli strains isolated from healthy cattle in Switzerland. Identification of a new intimin variant gene (eae-eta2), BMC Microbiology. 5 (2005) 23. [CrossRef]
- R. Wang, N. Kalchayanand, J.L. Bono, J.W. Schmidt, J.M. Bosilevac, Dual-serotype biofilm formation by shiga toxin-producing Escherichia coli O157:H7 and O26:H11 strains, Applied and environmental microbiology. 78 (2012) 6341–6344. [CrossRef]
- B.O. Obdoni, P.O.Ochuko, Phytochemical studies and comparative efficacy of the crude extracts of some Homostatic plants in Edo and Delta States of Nigeria, Global J. Pure Appl. Sci. 8b (2001) 203-208.
- T. Rahila, N. Rukhsandra, A.A. Zaidi, R. Shamishilia, Phytochemical Screening of medicinal plants belonging to Euphorbiaceae Pak, Vet. J. 14 (1994) 160-162.
- A.W. Bauer, D.M. Perry, W.M.M Kirby, Single-Disk Antibiotic-Sensitivity Testing of Staphylococci: An Analysis of Technique and Results, AMA Arch Intern Med. 104 (1959) 208–216. [CrossRef]
- D.S. Binnington, Improved soxhlet extraction apparatus, Fourth ed., Ind Eng Chem Anal Ed., 1932, pp. 125-126.
- Clinical Laboratory Standards Institute, Performance standards for antimicrobial disk susceptibility tests; approved standard—9th Ed. CLSI document M2-A9. 26:1. Clinical Laboratory Standards Institute, Wayne, PA, 2016.
- M.M. Cowan, Plant products as antimicrobial agents, Clin Microbiol Rev. 12 (1999) 564-82.
- J.Y. Lin, C.Y. Tang, Determination of total phenolic and flavonoid conents in selected fruits and vegetables, as well as their stimulatory effects on mouse splenocyte proliferation, Food Chem. 101 (2007) 140–147.

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



