Submitted:
31 March 2026
Posted:
01 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Rearing Conditions of Cockroaches
2.2. Obtaining Germ-Free Cockroaches and Quality Control
2.3. Insect Dissection
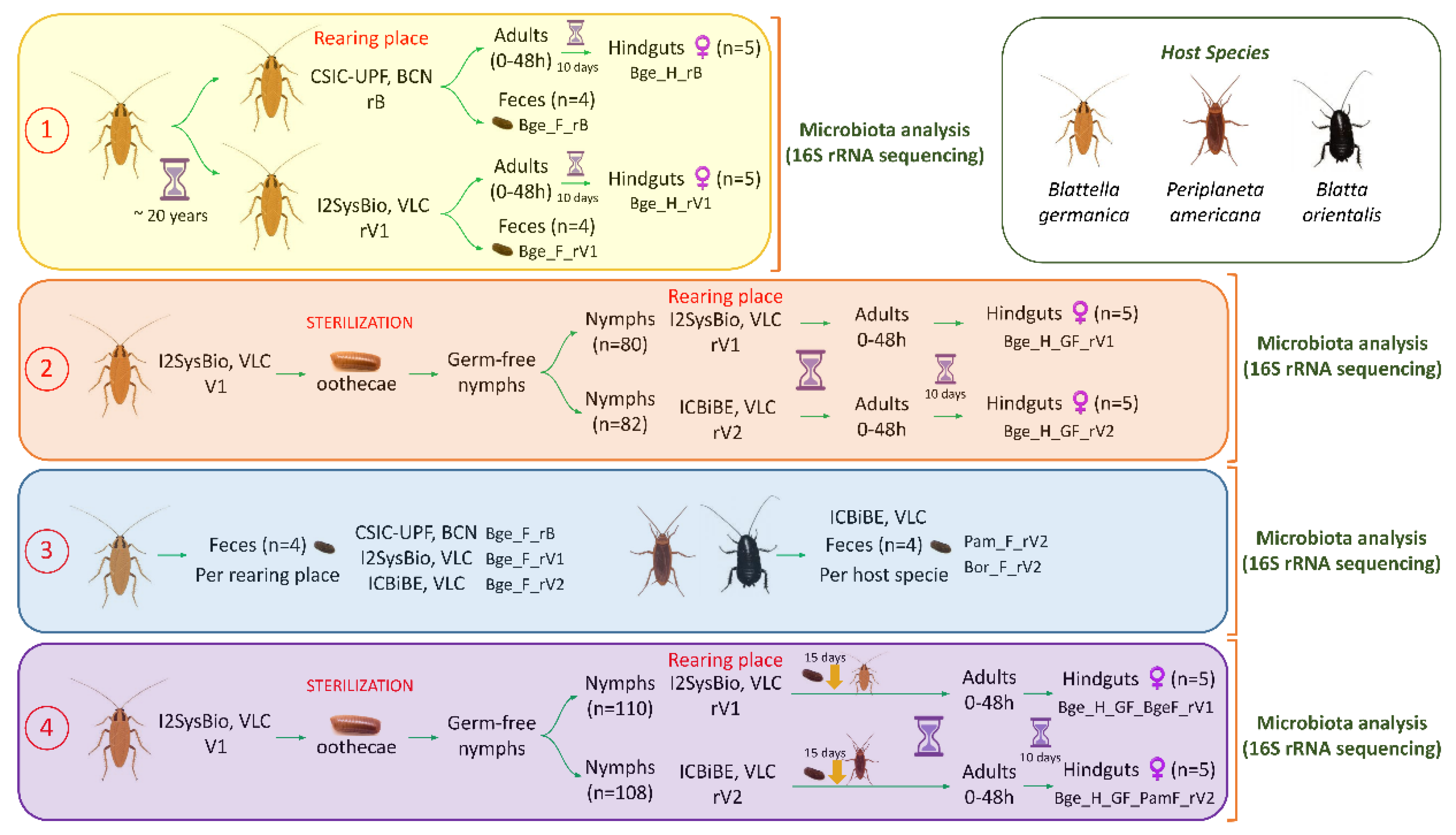
2.4. Experimental Designs
2.5. DNA Extraction and Metabarcoding Sequencing
2.6. Sequence Processing and Taxonomic Assignment
2.7. Microbiome Analysis
3. Results
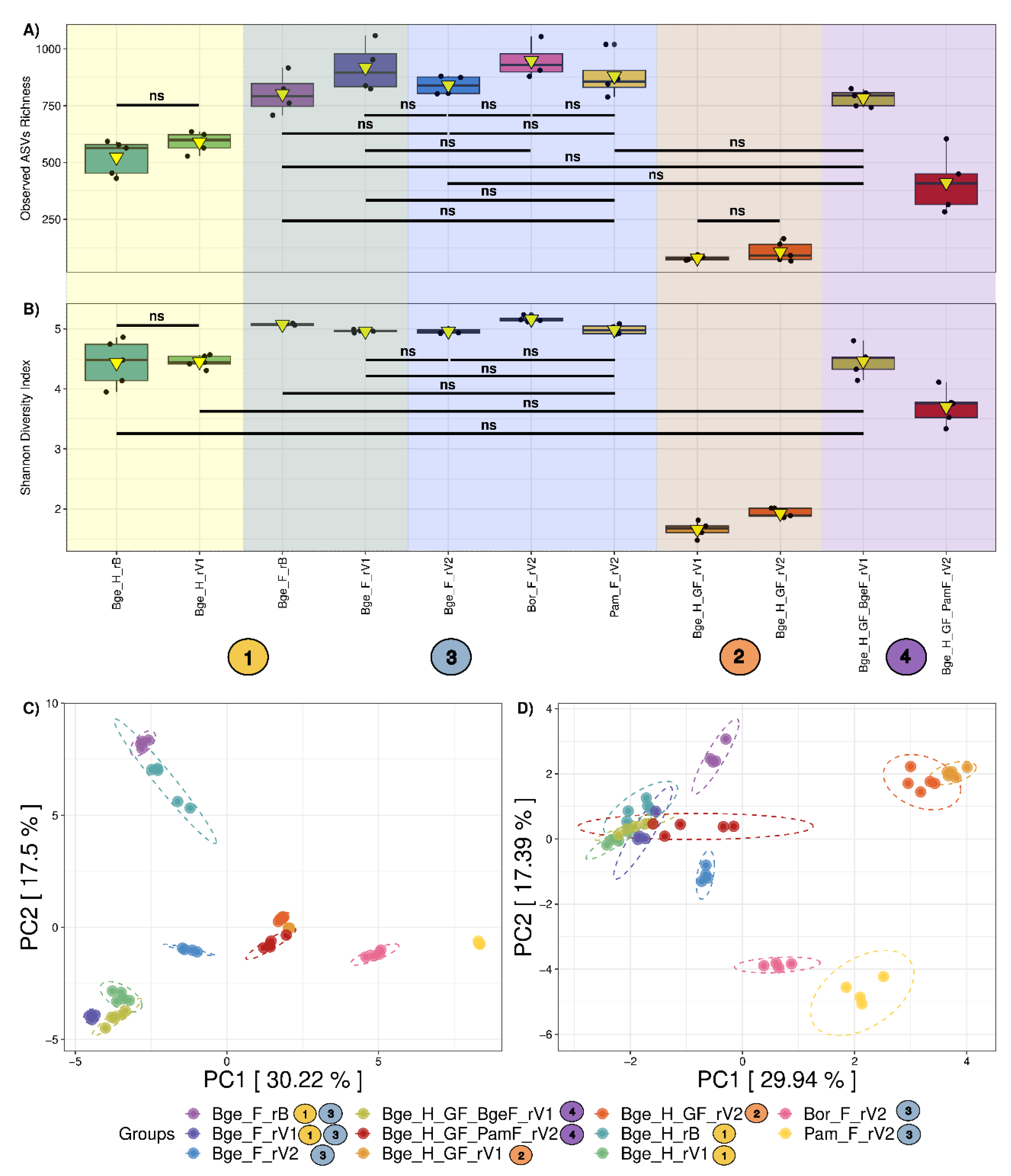
3.1. Global Microbial Diversity
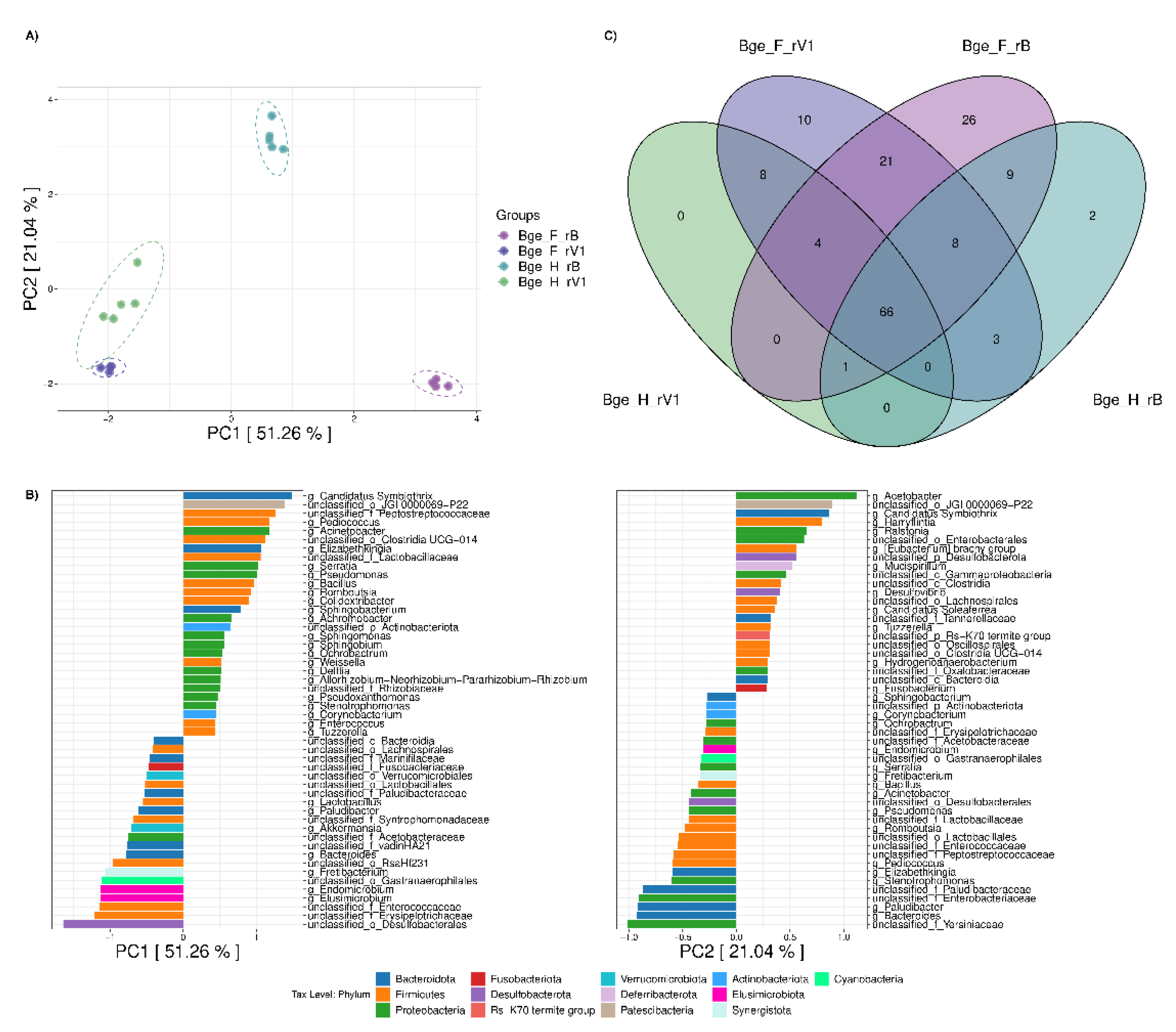
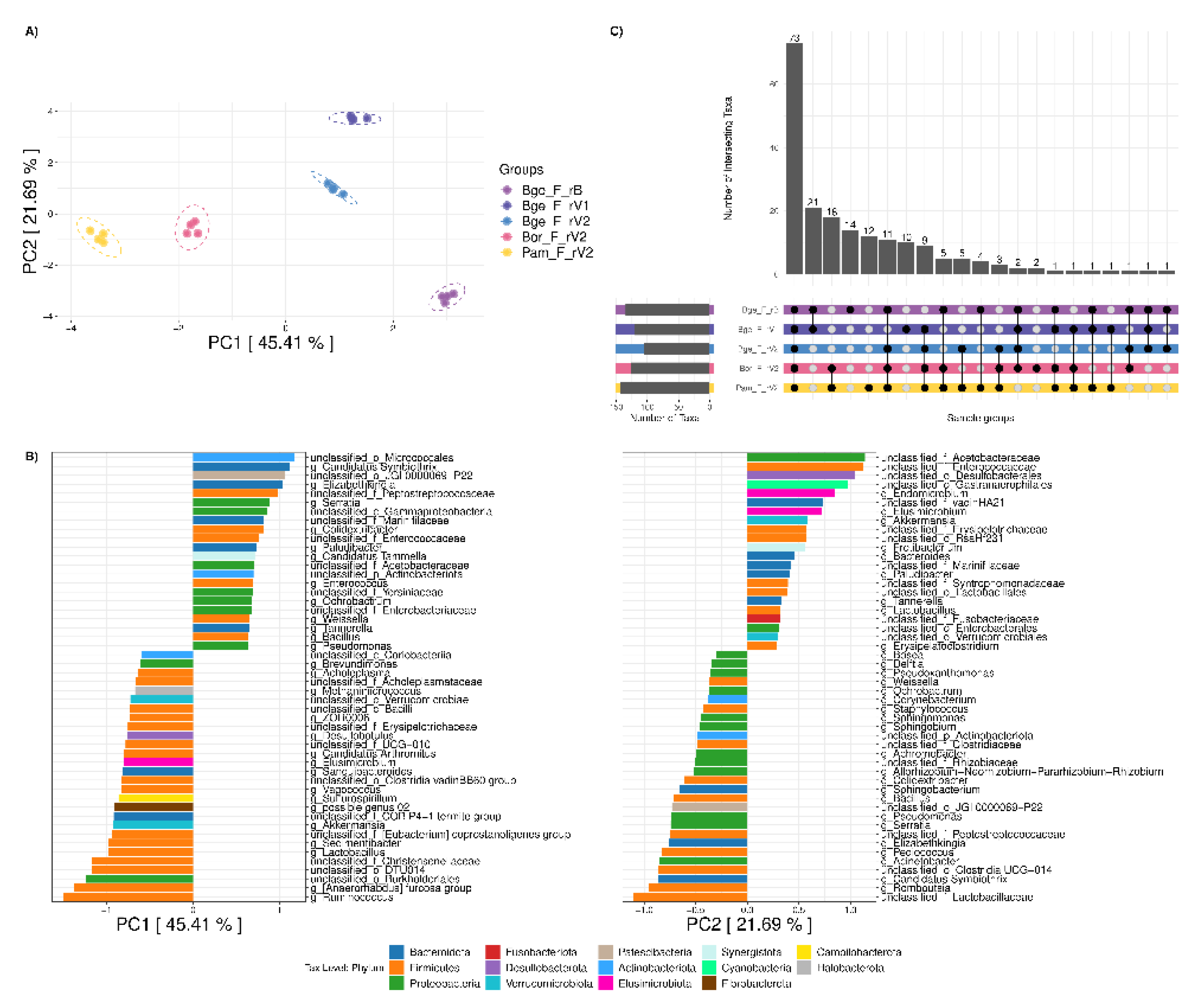
3.2. Comparison of the Gut Microbiota of Twinned Populations of B. germanica
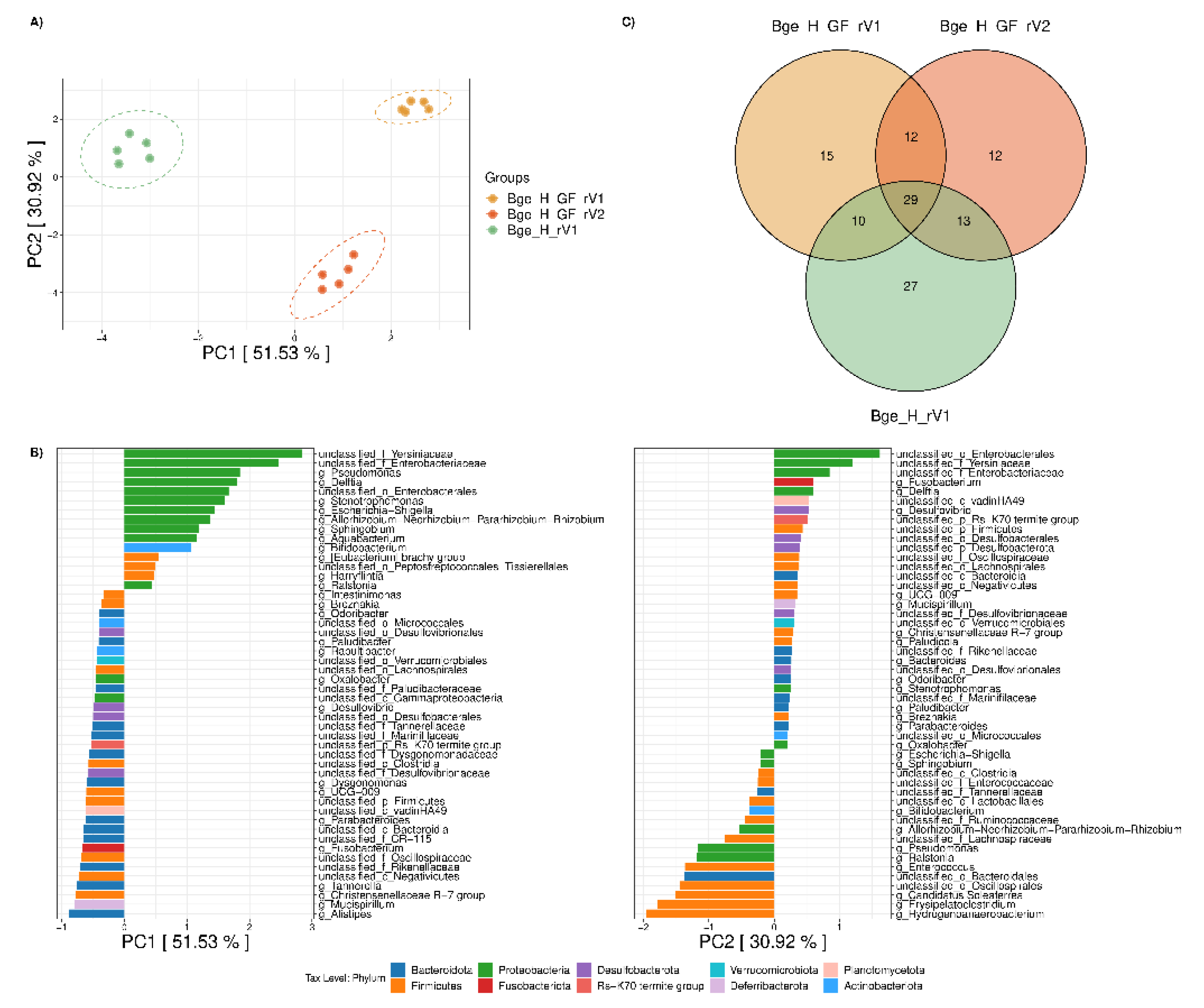
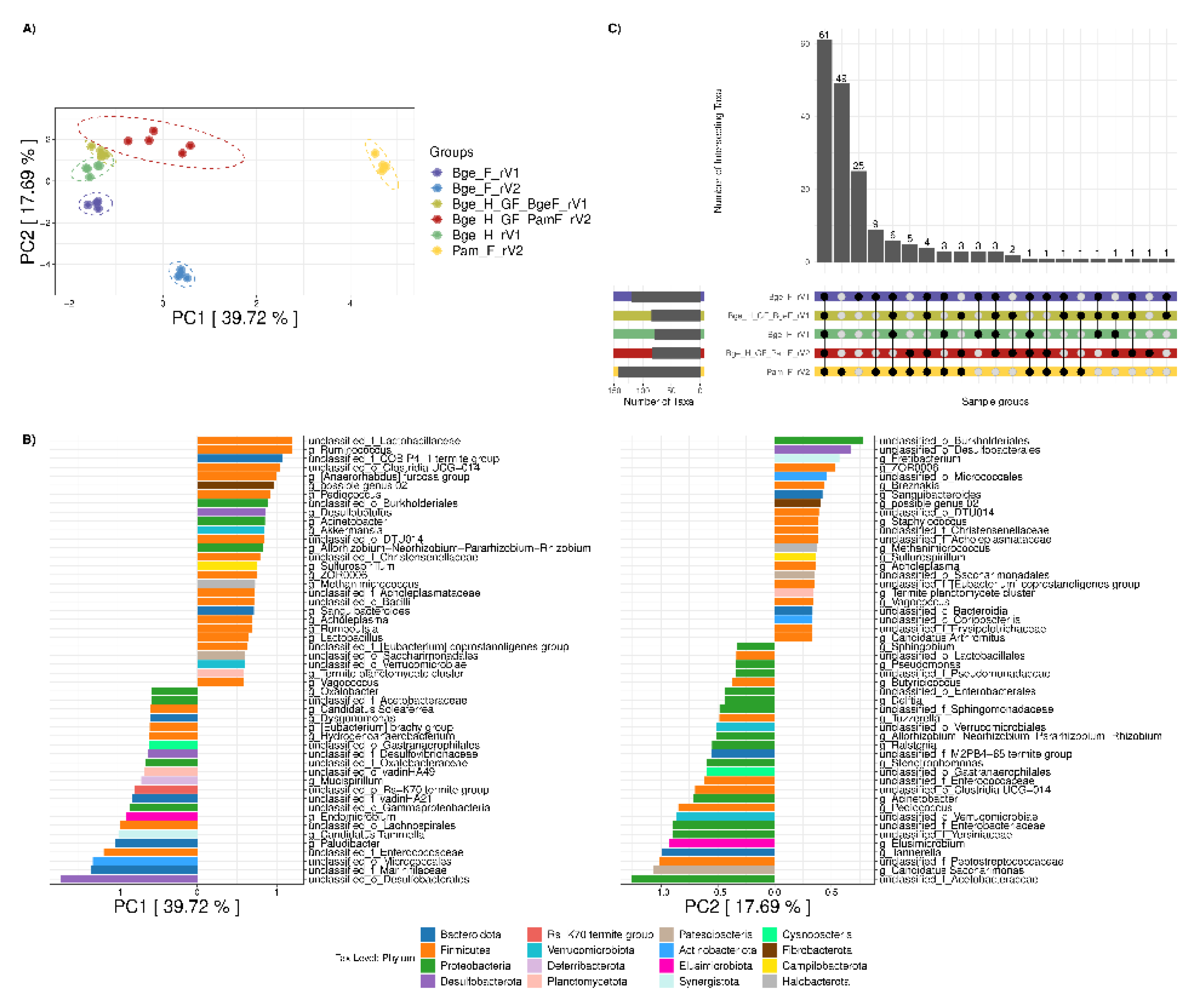
3.3. Exploring the Environmental Effect of Germ-Free Cockroaches
3.4. Comparison of the Fecal Microbiota of Three Cockroach Species
3.5. Inter-Species Cockroaches’ Fecal Transplantation
3.6. Analysis of the Core Microbiome
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| 16S rRNA | 16S ribosomal RNA gene |
| ANOVA | Analysis of Variance |
| ASV | Amplicon Sequence Variant |
| BCN | Barcelona |
| Bge | Blattella germanica |
| BgeF | Fecal input from B. germanica during transplantation |
| BH | Benjamini-Hochberg |
| BHI | Brain Heart Infusion |
| Bor | Blatta orientalis |
| CLR | Centered Log-Ratio transformation |
| CSIC-UPF | Institute of Evolutionary Biology, Spanish National Research Council - University Pompeu Fabra |
| F | Feces |
| GF | Germ-Free |
| GLM | Generalized Linear Model |
| H | Hindgut |
| I2SysBio | Institute for Integrative Systems Biology |
| ICBiBE | Cavanilles Institute of Biodiversity and Evolutionary Biology |
| IMD | Immune Deficiency |
| JAK/STAT | Janus Kinase/Signal Transducers and Activators of Transcription |
| KW | Kruskal Wallace |
| ns | Not Significant |
| Pam | Periplaneta americana |
| PamF | Fecal input from P. americana during transplantation |
| PCA | Principal Component Analysis |
| PERMANOVA | Permutational Multivariate Analysis of Variance |
| rB | rearing place Barcelona (CSIC-UPF) |
| rV1 | rearing place Valencia1 (I2SysBio) |
| rV2 | rearing place Valencia2 (ICBiBE) |
| SDS | Sodium Dodecyl Sulfate |
| USA | United States of America |
| VLC | Valencia |
References
- Sollai, G.; Solari, P. An overview of “Insect Biodiversity”. Diversity 2022, 14, 134. [CrossRef]
- Goryacheva, I.; Andrianov, B. Reproductive parasitism in insects. The interaction of host and bacteria. Biol. Commun. 2021 66, 17–27. [CrossRef]
- Latorre, A.; Moya, A. Role of symbiosis in evolution. In Evolution from the Galapagos: Two Centuries after Darwin; Trueba, G., Montúfar, C., Eds.; Springer: New york, USA, 2013; pp. 63–70. [CrossRef]
- Siddiqui, R.; Elmashak, Y.; Khan, N.A. Cockroaches: A potential source of novel bioactive molecule(s) for the benefit of human health. Appl. Entomol. Zool. 2023, 58, 1–11. [CrossRef]
- Sacchi, L.; Grigolo, A.; Laudani, U.; Ricevuti, G.; Dealessi, F. Behavior of symbionts during oogenesis and early stages of development in the German cockroach, Blattella germanica (Blattodea). J. Invertebr. Pathol. 1985, 46, 139–152. [CrossRef]
- Sacchi, L. Grigolo, A.; Mazzini, M.; Bigliardi, E.; Baccetti, B.; Laudani, U. Symbionts in the oocytes of Blattella germanica (L.) (Dictyoptera: Blattellidae): Their mode of transmission. Int. J. Insect Morphol. Embryol. 1988, 17, 437–446. [CrossRef]
- López-Sánchez, M. J.; Neef, A.; Peretó, J.; Patiño-Navarrete, R.; Pignatelli, M.; Latorre, A.; Moya, A. Evolutionary convergence and nitrogen metabolism in Blattabacterium strain Bge, primary endosymbiont of the cockroach Blattella germanica. PLoS Genet. 2009, 5, e1000721. [CrossRef]
- Carrasco, P.; Pérez-Cobas, A.E.; van de Pol, C.; Baixeras, J.; Moya, A.; Latorre, A. Succession of the gut microbiota in the cockroach Blattella germanica. Int. Microbiol. 2014, 17, 99–109 . [CrossRef]
- Patiño-Navarrete, R.; Piulachs, M.-D.; Belles, X.; Moya, A.; Latorre, A.; Peretó, J. The cockroach Blattella germanica obtains nitrogen from uric acid through a metabolic pathway shared with its bacterial endosymbiont. Biol. Lett. 2014, 10, 20140407. [CrossRef]
- Cazzaniga, M.; Domínguez-Santos, R.; Marín-Miret, J.; Gil, R.; Latorre, A.; García-Ferris, C. Exploring gut microbial dynamics and symbiotic interaction in Blattella germanica using rifampicin. Biology 2023, 12, 955. [CrossRef]
- Schauer, C.; Thompson, C.L.; Brune, A. The bacterial community in the gut of the cockroach Shelfordella lateralis reflects the close evolutionary relatedness of cockroaches and termites. Appl. Environ. Microbiol. 2012, 78, 2758–2767. [CrossRef]
- Schauer, C.; Thompson, C.; Brune, A. Pyrotag sequencing of the gut microbiota of the cockroach Shelfordella lateralis reveals a highly dynamic core but only limited effects of diet on community structure. PLoS ONE 2014, 9, e85861. [CrossRef]
- Pérez-Cobas, A.E.; Maiques, E.; Angelova, A.; Carrasco, P.; Moya, A.; Latorre, A. Diet shapes the gut microbiota of the omnivorous cockroach Blattella germanica. FEMS Microbiol. Ecol. 2015, 91, fiv022. [CrossRef]
- Mikaelyan, A.; Thompson, C.L., Hofer, M.J.; Brune, A. Deterministic assembly of complex bacterial communities in guts of germ-free cockroaches. Appl. Environ. Microbiol. 2016, 82, 1256–1263. [CrossRef]
- Tegtmeier, D.; Thompson, C.L.; Schauer, C.; Brune, A. Oxygen affects gut bacterial colonization and metabolic activities in a gnotobiotic cockroach model. Appl. Environ. Microbiol. 2016, 82, 1080–1089. [CrossRef]
- Kakumanu, M.L.; Maritz, J.M.; Carlton, J.M.; Schal, C. Overlapping community compositions of gut and fecal microbiomes in lab-reared and field-collected German cockroaches. Appl. Environ. Microbiol. 2018, 84, e01037-18. [CrossRef]
- Rosas, T.; García-Ferris, C.; Domínguez-Santos, R.; Llop, P.; Latorre, A.; Moya, A. Rifampicin treatment of Blattella germanica evidences a fecal transmission route of their gut microbiota. FEMS Microbiol. Ecol. 2018, 94, fiy002. [CrossRef]
- Domínguez-Santos, R.; Pérez-Cobas, A.E.; Latorre, A.; Moya, A.; García-Ferris, C. Unraveling assemblage, functions and stability of the gut microbiota of Blattella germanica by antibiotic treatment. Front. Microbiol. 2020, 11, 487. [CrossRef]
- Domínguez-Santos, R.; Pérez-Cobas, A.E.; Latorre, A.; Moya, A.; García-Ferris, C. Interkingdom gut microbiome and resistome of the cockroach Blattella germanica. MSystems 2021, 6, e01213-20. [CrossRef]
- Tinker, K. A.; Ottesen, E.A. Phylosymbiosis across deeply diverging lineages of omnivorous cockroaches (Order Blattodea). Appl. Environ. Microbiol. 2020, 86, e02513-19. [CrossRef]
- Tinker, K.A.; Ottesen, E.A. Differences in gut microbiome composition between sympatric wild and allopatric laboratory populations of omnivorous cockroaches. Front. Microbiol. 2021, 12, 703785. [CrossRef]
- Zhang, F.; Sun, X.X.; Zhang, X.C.; Zhang, S.; Lu, J.; Xia, Y.M.; Huang, Y.H.; Wang, X.J. The interactions between gut microbiota and entomopathogenic fungi: a potential approach for biological control of Blattella germanica (L.). Pest Manag Sci. 2018, 74, 438–447. [CrossRef]
- Motta, E.V.S.; Mak, M.; De Jong, T.K.; Powell, J.E.; O’Donnell, A.; Suhr, K.J.; Riddington, I.M.; Moran, N.A. Oral or topical exposure to glyphosate in herbicide formulation impacts the gut microbiota and survival rates of honey bees. Appl. Environ. Microbiol. 2020, 86, e01150-20. [CrossRef]
- Marín-Miret, J.; González-Serrano, F.; Rosas, T.; Baixeras, J.; Latorre, A.; Pérez-Cobas, A.E.; Moya, A. Temporal variations shape the gut microbiome ecology of the moth Brithys crini. Environ. Microbiol. 2022, 24, 3939–3953. [CrossRef]
- Nakayama, Y.; Suto, C.; Kumada, N. Further studies on the dispersion-inducing substances of the German cockroach, Blattella germanica (LINNE) (Blattaria: Blattellidae). Appl. Entomol. Zool. 1984, 19, 227–236. [CrossRef]
- Lihoreau, M.; Costa, J. T.; Rivault, C. The social biology of domiciliary cockroaches: colony structure, kin recognition and collective decisions. Insect Soc. 2012, 59, 445–452. [CrossRef]
- Wada-Katsumata, A.; Zurek, L.; Nalyanya, G.; Roelofs, W. L.; Zhang, A.; Schal, C. Gut bacteria mediate aggregation in the German cockroach. Proc. Natl. Acad. Sci. U.S.A. 2015, 112, 15678–15683. [CrossRef]
- Ware, J.; Megan, W. Termites, social cockroaches. In Encyclopedia of Biodiversity, (ed. Scheiner, S.M.) 517–541 (Academic Press, 2024). [CrossRef]
- Osawa, R.; Blanshard, W.H.; Ocallaghan, P.G. Microbiological studies of the intestinal microflora of the koala, Phascolarctos cinereus. 2. Pap, a special maternal faeces consumed by juvenile koalas. Aust. J. Zool. 1993, 41, 611–620. [CrossRef]
- Videvall, E., Bensch, H. M., Engelbrecht, A., Cloete, S. & Cornwallis, C. K. Coprophagy rapidly matures juvenile gut microbiota in a precocial bird. Evol. Lett. 2023, 7, 240–251. [CrossRef]
- Domínguez-Santos, R.; Baixeras, J.; Moya, A.; Latorre, A.; Gil, R.; García-Ferris, C. Gut microbiota is not essential for survival and development in Blattella germanica, but affects uric acid storage. Life 2024, 14, 153. [CrossRef]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011, 17, 10–12. [CrossRef]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [CrossRef]
- Murali, A.; Bhargava, A.; Wright, E.S. IDTAXA: a novel approach for accurate taxonomic classification of microbiome sequences. Microbiome 2018, 6, 140. [CrossRef]
- Wright, E.S. Using DECIPHER v2.0 to analyze big biological sequence data in R. R J. 2016, 8, 352–359. https://journal.r-project.org/articles/RJ-2016-025/.
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, D590–D596. [CrossRef]
- Lahti, L.; Shetty, S. Tools for microbiome analysis in R. Bioconductor 2017. [CrossRef]
- McMurdie, P.J.; Holmes, S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 2013, 8, e61217. [CrossRef]
- Oksanen, J.; Simpson, G.L.; Blanchet, F.G.; Kindt, R.; Legendre, P.; Minchin, P.R.; O’Hara, R.B.; Solymos, P.; Stevens, M.H.H.; Szoecs, E.; Wagner, H.; Barbour, M.; Bedward, M.; Bolker, B.; Borcard, D.; Carvalho, G.; Chirico, M.; De Caceres, M.; Durand, S.; Evangelista, H.B.A.; FitzJohn, R.; Friendly, M.; Furneaux, B.; Hannigan, G.; Hill, M.O.; Lahti, L.; McGlinn, D.; Ouellette, M.H.; Ribeiro Cunha, E.; Smith, T.; Stier, A.; Ter Braak, C.J.F.; Weedon, J.H. vegan: Community Ecology Package. R package version 2.6-4. 2022. https://cran.r-project.org/package=vegan.
- Fernandes, A.D.; Reid, J.N.; Macklaim, J.M.; McMurrough, T.A.; Edgell, D.R.; Gloor, G.B. Unifying the analysis of high-throughput sequencing datasets: characterizing RNA-seq, 16S rRNA gene sequencing and selective growth experiments by compositional data analysis. Microbiome 2014, 2, 15. [CrossRef]
- Gu, Z.; Eils, R.; Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 2016, 32, 2847–2849. [CrossRef]
- Krassowski, M.; Arts, M.; Laggery, C. krassowski/complex-upset: v1.3.3. R package version 1.3.3. Zenodo, 2021 . [CrossRef]
- Yan, L. ggvenn: Draw Venn diagram by ‘ggplot2’. R package version 0.1.10 https://cran.r-project.org/package=ggvenn (2023).
- Foster, Z. S., Sharpton, T. J. & Grünwald, N. J. Metacoder: an R package for visualization and manipulation of community taxonomic diversity data. PLoS Comput. Biol. 2017, 13, e1005404. [CrossRef]
- Onchuru, T.O.; Martinez, A.J.; Ingham, C.S.; Kaltenpoth, M. Transmission of mutualistic bacteria in social and gregarious insects. Curr. Opin. Insect Sci. 2018, 28, 50–58. [CrossRef]
- Schapheer, C.; González, L.M.; Villagra, C. Microorganism diversity found in Blatta orientalis L. (Blattodea: Blattidae) cuticle and gut collected in urban environments. Insects 2024, 15, 903. [CrossRef]
- Lee, S.; Kim, J.Y.; Yi, M.H.; Lee, I.Y.; Lee, W.J.; Moon, H.S.; Yong, D.; Yong, T.S. Comparative microbiome analysis of three species of laboratory-reared Periplaneta cockroaches. Korean J. Parasitol. 2020, 58, 537–542. [CrossRef]
- Tinker, K.A.; Ottesen, E.A. The core gut microbiome of the American cockroach, Periplaneta americana, is stable and resilient to dietary shifts. Appl. Environ. Microbiol. 2016, 82, 6603–6610. [CrossRef]
- Leeming, E.R.; Johnson, A.J.; Spector, T.D.; Le Roy, C.I. Effect of diet on the gut microbiota: rethinking intervention duration. Nutrients 2019, 11, 2862. [CrossRef]
- Huang, K.; Wang, J.; Huang, J.; Zhang, S.; Vogler, A.P.; Liu, Q.; Li, Y.; Yang, M.; Li, Y.; Zhou, X. Host phylogeny and diet shape gut microbial communities within bamboo-feeding insects. Front. Microbiol. 2021, 12, 633075. [CrossRef]
- Rinninella, E.; Tohumcu, E.; Raoul, P.; Fiorani, M.; Cintoni, M.; Mele, M.C.; Cammarota, G.; Gasbarrini, A.; Ianiro, G. The role of diet in shaping human gut microbiota. Best Pract. Res. Clin. Gastroenterol. 2023, 62–63, 101828. [CrossRef]
- Jahnes, B.C.; Herrmann, M.; Sabree, Z.L. Conspecific coprophagy stimulates normal development in a germ-free model invertebrate. PeerJ 2019, 7, e6914. [CrossRef]
- Vera-Ponce de León, A.; Jahnes, B.C.; Otero-Bravo, A.; Sabree, Z.L. Microbiota perturbation or elimination can inhibit normal development and elicit a starvation-like response in an omnivorous model invertebrate. MSystems 2021, 6, e00802-21. [CrossRef]
- Qin, M.; Jiang, L.; Qiao, G.; Chen, J. Phylosymbiosis: The eco-evolutionary pattern of insect–symbiont interactions. Int. J. Mol. Sci. 2023, 24, 15836. [CrossRef]
- Li, J.; Wei, X.; Huang, D.; Xiao, J. The phylosymbiosis pattern between the fig wasps of the same genus and their associated microbiota. Front. Microbiol. 2022, 12, 800190. [CrossRef]
- Qin, M.; Jiang, L.; Kholmatov, B.R.; Qiao, G.; Chen, J. Phylosymbiotic structures of the microbiota in Mollitrichosiphum tenuicorpus (Hemiptera: Aphididae: Greenideinae). Microb. Ecol. 2022, 84, 227–239. [CrossRef]
- Lemaitre, B.; Hoffmann, J. The host defense of Drosophila melanogaster. Annu. Rev. Immunol. 2007, 25, 697–743. [CrossRef]
- Kuraishi, T.; Hori, A.; Kurata, S. Host-microbe interactions in the gut of Drosophila melanogaster. Front. Physiol. 2013, 4, 375. [CrossRef]
- Douglas, A.E. The molecular basis of bacterial-insect symbiosis. J. Mol. Biol. 2014, 426, 3830–3837. [CrossRef]
- Brune, A.; Dietrich, C. The gut microbiota of termites: digesting the diversity. Annu. Rev. Microbiol. 2015, 69, 145–166. [CrossRef]
- Zhang, S.; Wang, Z.; Luo, Q.; Zhou, L.; Du, X.; Ren, Y. Effects of microbes on insect host physiology and behavior mediated by the host immune system. Insects 2025, 16, 82. [CrossRef]
- Schmidt, K.; Engel, P. Mechanisms underlying gut microbiota–host interactions in insects. J. Exp. Biol. 2021, 224, jeb207696. [CrossRef]
- Derguini, A.; Basher, N.S. Cockroaches as Vectors of Pathogens and Antimicrobial Resistance: Evidence from Healthcare, Community, and Agricultural Settings. Insects 2026, 17, 310. [CrossRef]
- Renelies-Hamilton, J.; Germer, K.; Sillam-Dussès, D.; Bodawatta, K.H.; Poulsen, M. Disentangling the relative roles of vertical transmission, subsequent colonizations, and diet on cockroach microbiome assembly. MSphere 2021, 6, e01023-20. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



