Submitted:
26 March 2026
Posted:
27 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction: Why Biomonitoring 3.0 Now
2. From Taxa Lists to Interaction-Ready Inference: What Changes in 3.0
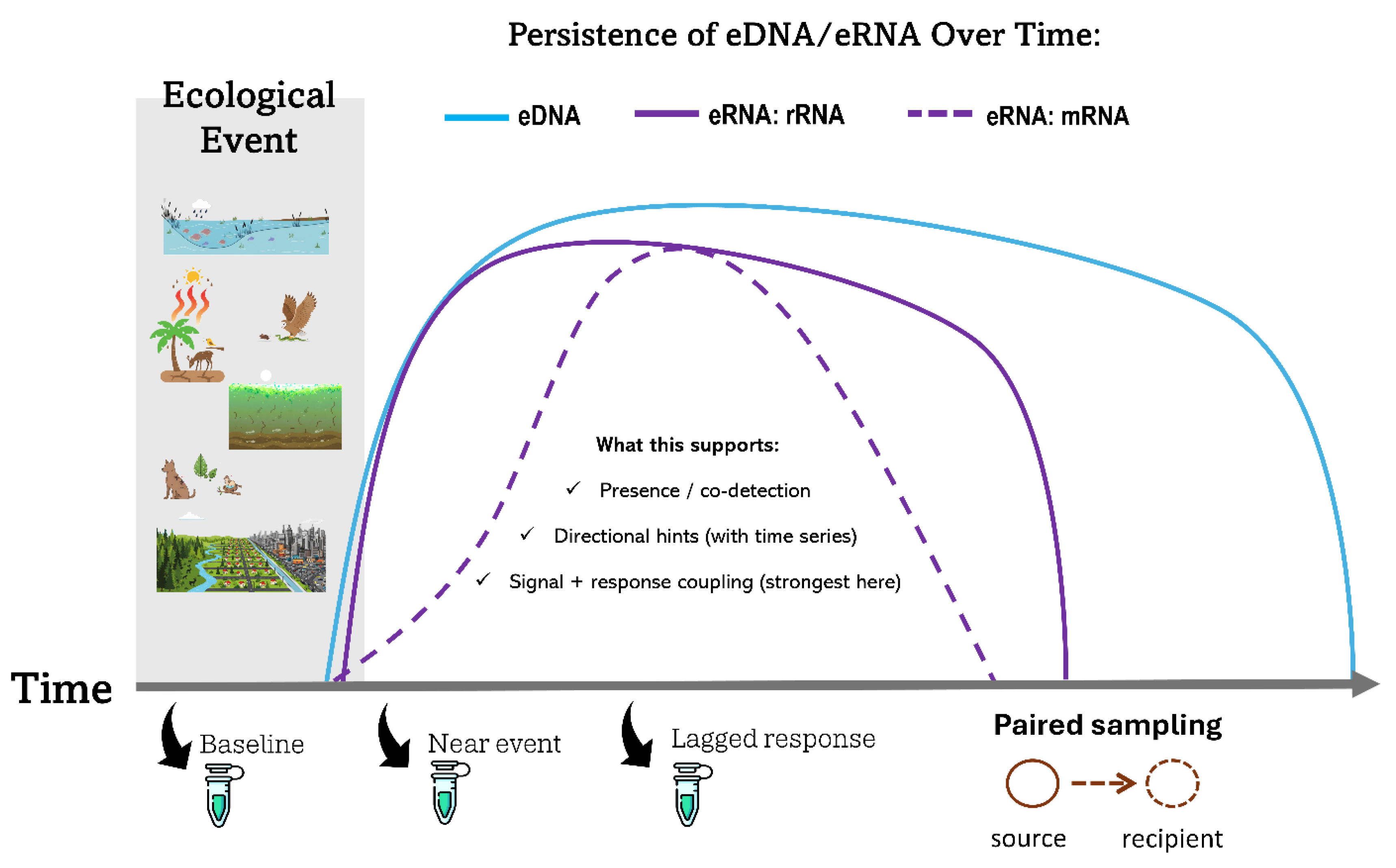
3. Environmental RNA as a Time-Resolved Layer: From Presence to Activity and Response
4. What Counts as an Interaction Signal: An Evidence Ladder for Claims
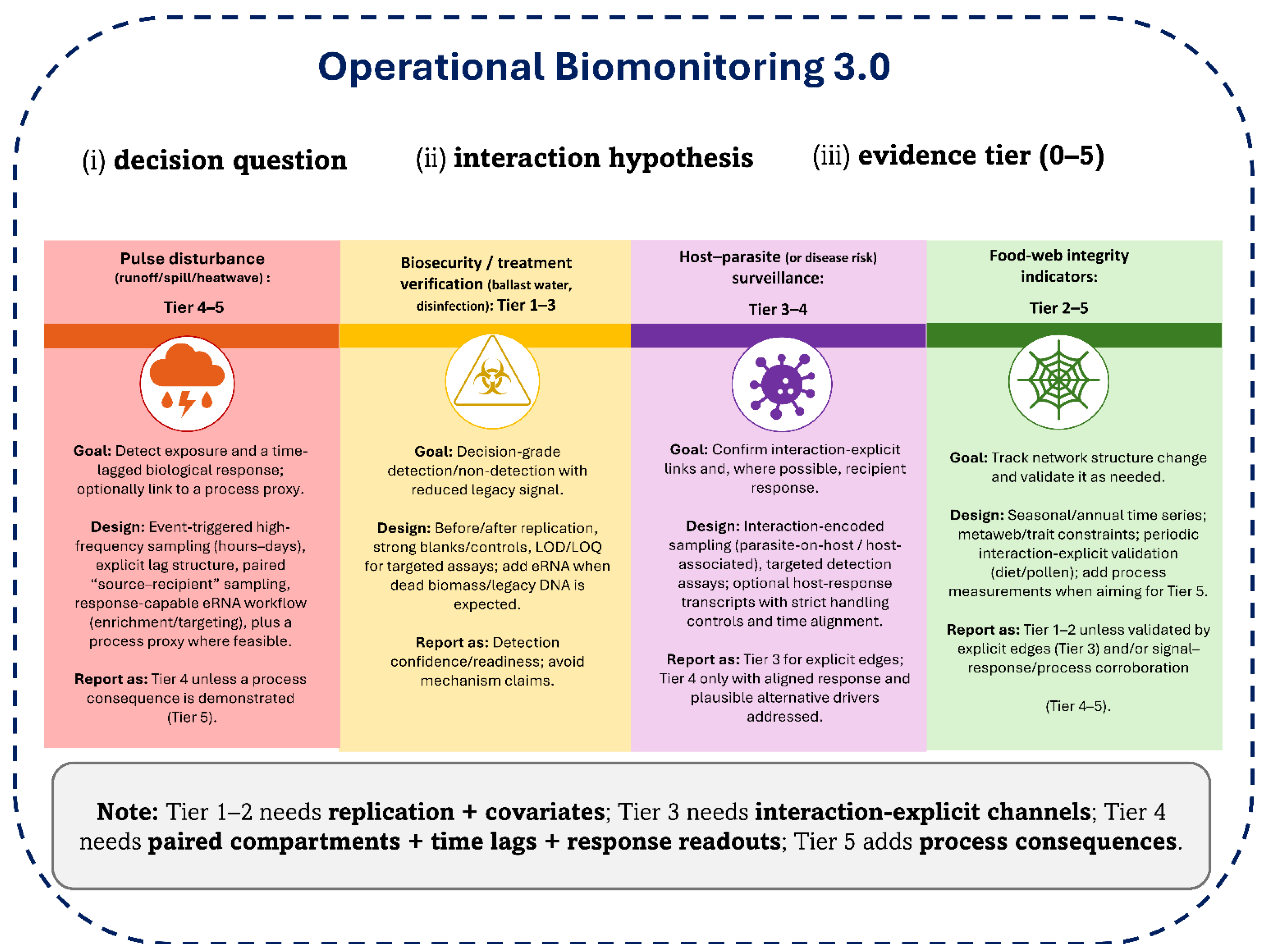
| Box 1 | Biomonitoring 3.0 use-case playbook: design backward from the decision and the evidence tier |
| Biomonitoring 3.0 becomes operational when programs specify (i) the decision question, (ii) the interaction hypothesis, and (iii) the intended evidence tier (0–5), then adopt only the sampling and assay elements needed to meet that tier. Use case A: Pulse disturbance (runoff/spill/heatwave) — Tier 4–5
|
5. Designing Field Programs for Interaction-Ready, Time-Resolved Monitoring
6. Standards, Validation, and Translation to Decisions
7. Conclusion: A Roadmap and Near-Term Priorities for Biomonitoring 3.0
Data availability statement
Acknowledgments
References
- Abdala-Roberts, L., Puentes, A., Finke, D. L., Marquis, R. J., Montserrat, M., Poelman, E. H., … Mooney, K. A. (2025). Connecting the dots: Managing species interaction networks to mitigate the impacts of global change. ELife, 14. [CrossRef]
- Ahi, E. P., & Schenekar, T. (2025). The Promise of Environmental RNA Research Beyond mRNA. Molecular Ecology, 34(12), e17787. [CrossRef]
- Allesina, S., & Tang, S. (2012). Stability criteria for complex ecosystems. Nature 2012 483:7388, 483(7388), 205–208. [CrossRef]
- Baird, D. J., & Hajibabaei, M. (2012). Biomonitoring 2.0: a new paradigm in ecosystem assessment made possible by next-generation DNA sequencing. Molecular Ecology, 21(8), 2039–2044. [CrossRef]
- Barroso-Bergadà, D., Pauvert, C., Vallance, J., Delière, L., Bohan, D. A., Buée, M., & Vacher, C. (2021). Microbial networks inferred from environmental DNA data for biomonitoring ecosystem change: Strengths and pitfalls. Molecular Ecology Resources, 21(3), 762–780. [CrossRef]
- Blackman, R., Couton, M., Keck, F., Kirschner, D., Carraro, L., Cereghetti, E., … Altermatt, F. (2024). Environmental DNA: The next chapter. Molecular Ecology, 33(11), e17355. [CrossRef]
- Blanchet, F. G., Cazelles, K., & Gravel, D. (2020). Co-occurrence is not evidence of ecological interactions. Ecology Letters, 23(7), 1050–1063. [CrossRef]
- Bohan, D. A., Vacher, C., Tamaddoni-Nezhad, A., Raybould, A., Dumbrell, A. J., & Woodward, G. (2017). Next-Generation Global Biomonitoring: Large-scale, Automated Reconstruction of Ecological Networks. Trends in Ecology and Evolution, 32(7), 477–487. [CrossRef]
- Borchardt, M. A., Boehm, A. B., Salit, M., Spencer, S. K., Wigginton, K. R., & Noble, R. T. (2021). The Environmental Microbiology Minimum Information (EMMI) Guidelines: QPCR and dPCR Quality and Reporting for Environmental Microbiology. Environmental Science and Technology, 55(15), 10210–10223. [CrossRef]
- Boyse, E., Robinson, K. P., Carr, I. M., Valsecchi, E., Beger, M., & Goodman, S. J. (2025). Inferring Species Interactions From Co-occurrence Networks With Environmental DNA Metabarcoding Data in a Coastal Marine Food Web. Molecular Ecology, 34(7), e17701. WGROUP:STRING:PUBLICATION. [CrossRef]
- Bustin, S. A., Ruijter, J. M., van den Hoff, M. J. B., Kubista, M., Pfaffl, M. W., Shipley, G. L., … Wittwer, C. T. (2025). MIQE 2.0: Revision of the Minimum Information for Publication of Quantitative Real-Time PCR Experiments Guidelines. Clinical Chemistry, 71(6), 634–651. [CrossRef]
- Chatterjee, A., Zhang, K., & Parker, K. M. (2023). Binding of Dissolved Organic Matter to RNA and Protection from Nuclease-Mediated Degradation. Environmental Science and Technology, 57(42), 16086–16096. [CrossRef]
- Cordier, T., Alonso-Sáez, L., Apothéloz-Perret-Gentil, L., Aylagas, E., Bohan, D. A., Bouchez, A., … Lanzén, A. (2021). Ecosystems monitoring powered by environmental genomics: A review of current strategies with an implementation roadmap. Molecular Ecology, 30(13), 2937–2958. [CrossRef]
- Coyte, K. Z., Schluter, J., & Foster, K. R. (2015). The ecology of the microbiome: Networks, competition, and stability. Science, 350(6261), 663–666. [CrossRef]
- D’Alessandro, S., & Mariani, S. (2021). Sifting environmental DNA metabarcoding data sets for rapid reconstruction of marine food webs. Fish and Fisheries, 22(4), 822–833. [CrossRef]
- Darling, J. A., Jerde, C. L., & Sepulveda, A. J. (2021). What do you mean by false positive? Environmental DNA, 3(5), 879–883. [CrossRef]
- De Brauwer, M., Clarke, L. J., Chariton, A., Cooper, M. K., de Bruyn, M., Furlan, E., … Trujillo-González, A. (2023). Best practice guidelines for environmental DNA biomonitoring in Australia and New Zealand. Environmental DNA, 5(3), 417–423. [CrossRef]
- Deagle, B. E., Thomas, A. C., McInnes, J. C., Clarke, L. J., Vesterinen, E. J., Clare, E. L., … Eveson, J. P. (2019). Counting with DNA in metabarcoding studies: How should we convert sequence reads to dietary data? Molecular Ecology, 28(2), 391–406. [CrossRef]
- Deiner, K., Fronhofer, E. A., Mächler, E., Walser, J. C., & Altermatt, F. (2016). Environmental DNA reveals that rivers are conveyer belts of biodiversity information. Nature Communications 2016 7:1, 7(1), 12544-. [CrossRef]
- Delmas, E., Besson, M., Brice, M. H., Burkle, L. A., Dalla Riva, G. V., Fortin, M. J., … Poisot, T. (2019). Analysing ecological networks of species interactions. Biological Reviews, 94(1), 16–36. ;REQUESTEDJOURNAL:JOURNAL:1469185X;CTYPE:STRING:JOURNAL. [CrossRef]
- Djurhuus, A., Closek, C. J., Kelly, R. P., Pitz, K. J., Michisaki, R. P., Starks, H. A., … Breitbart, M. (2020). Environmental DNA reveals seasonal shifts and potential interactions in a marine community. Nature Communications 2020 11:1, 11(1), 254-. [CrossRef]
- Encinas-Viso, F., Bovill, J., Albrecht, D. E., Florez-Fernandez, J., Lessard, B., Lumbers, J., … Milla, L. (2023). Pollen DNA metabarcoding reveals cryptic diversity and high spatial turnover in alpine plant–pollinator networks. Molecular Ecology, 32(23), 6377–6393. [CrossRef]
- Faust, K., & Raes, J. (2012). Microbial interactions: from networks to models. Nature Reviews Microbiology 2012 10:8, 10(8), 538–550. [CrossRef]
- Ficetola, G. F., Taberlet, P., & Coissac, E. (2016). How to limit false positives in environmental DNA and metabarcoding? Molecular Ecology Resources, 16(3), 604–607. [CrossRef]
- George, S. D., Sepulveda, A. J., Hutchins, P. R., Pilliod, D. S., Klymus, K. E., Thomas, A. C., … Leinonen, E. G. (2024). Field Trials of an Autonomous eDNA Sampler in Lotic Waters. Environmental Science and Technology, 58(47), 20942–20953. [CrossRef]
- Giroux, M. S., Reichman, J. R., Langknecht, T., Burgess, R. M., & Ho, K. T. (2022). Environmental RNA as a Tool for Marine Community Biodiversity Assessments. Scientific Reports 2022 12:1, 12(1), 1–13. [CrossRef]
- Goldberg, C. S., Turner, C. R., Deiner, K., Klymus, K. E., Thomsen, P. F., Murphy, M. A., … Taberlet, P. (2016). Critical considerations for the application of environmental DNA methods to detect aquatic species. Methods in Ecology and Evolution, 7(11), 1299–1307. [CrossRef]
- Haase, P., Pauls, S. U., Schindehütte, K., & Sundermann, A. (2010). First audit of macroinvertebrate samples from an EU Water Framework Directive monitoring program: human error greatly lowers precision of assessment results. 29(4), 1279–1291. [CrossRef]
- Hallam, J., Clare, E. L., Jones, J. I., & Day, J. J. (2023). High frequency environmental DNA metabarcoding provides rapid and effective monitoring of fish community dynamics. Environmental DNA, 5(6), 1623–1640. [CrossRef]
- He, X., Maruki, T., Morgado-Gamero, W. B., Barrett, R. D. H., Fugère, V., Fussmann, G. F., … Cristescu, M. E. (2025). Environmental RNA-Based Metatranscriptomics as a Novel Biomonitoring Tool: A Case Study of Glyphosate-Based Herbicide Effects on Freshwater Eukaryotic Communities. Molecular Ecology, 34(22), e70164; SUBPAGE:STRING:FULL . [CrossRef]
- Hechler, R. M., Yates, M. C., Chain, F. J. J., & Cristescu, M. E. (2023). Environmental transcriptomics under heat stress: Can environmental RNA reveal changes in gene expression of aquatic organisms? Molecular Ecology, 00, 1–15. [CrossRef]
- Hiki, K., & Jo, T. S. (2025). Comprehensive Sequencing of Environmental RNA From Japanese Medaka at Various Size Fractions and Comparison With Skin Swab RNA. Environmental DNA, 7(3), e70137. [CrossRef]
- Hiki, K., Yamagishi, T., & Yamamoto, H. (2023). Environmental RNA as a Noninvasive Tool for Assessing Toxic Effects in Fish: A Proof-of-concept Study Using Japanese Medaka Exposed to Pyrene. Environmental Science and Technology, 57(34), 12654–12662. [CrossRef]
- Jo, T. S. (2023). Utilizing the state of environmental DNA (eDNA) to incorporate time-scale information into eDNA analysis. Proceedings of the Royal Society B: Biological Sciences, 290(1999). [CrossRef]
- Kagzi, K., Hechler, R. M., Fussmann, G. F., & Cristescu, M. E. (2022). Environmental RNA degrades more rapidly than environmental DNA across a broad range of pH conditions. Molecular Ecology Resources, 22(7), 2640–2650. [CrossRef]
- Keck, F., Couton, M., & Altermatt, F. (2023). Navigating the seven challenges of taxonomic reference databases in metabarcoding analyses. Molecular Ecology Resources, 23(4), 742–755. [CrossRef]
- Klymus, K. E., Baker, J. D., Abbott, C. L., Brown, R. J., Craine, J. M., Gold, Z., … Theroux, S. (2024). The MIEM guidelines: Minimum information for reporting of environmental metabarcoding data. Metabarcoding and Metagenomics, 8, 489–518. [CrossRef]
- Klymus, K. E., Merkes, C. M., Allison, M. J., Goldberg, C. S., Helbing, C. C., Hunter, M. E., … Richter, C. A. (2020). Reporting the limits of detection and quantification for environmental DNA assays. Environmental DNA, 2(3), 271–282. [CrossRef]
- Kurtz, Z. D., Müller, C. L., Miraldi, E. R., Littman, D. R., Blaser, M. J., & Bonneau, R. A. (2015). Sparse and Compositionally Robust Inference of Microbial Ecological Networks. PLOS Computational Biology, 11(5), e1004226. [CrossRef]
- Lahoz-Monfort, J. J., Guillera-Arroita, G., & Tingley, R. (2016). Statistical approaches to account for false-positive errors in environmental DNA samples. Molecular Ecology Resources, 16(3), 673–685. [CrossRef]
- Littlefair, J. E., Rennie, M. D., & Cristescu, M. E. (2022). Environmental nucleic acids: A field-based comparison for monitoring freshwater habitats using eDNA and eRNA. Molecular Ecology Resources, 22(8), 2928. [CrossRef]
- Macher, T. H., Arle, J., Beermann, A. J., Frank, L., Hupało, K., Koschorreck, J., … Leese, F. (2024). Is it worth the extra mile? Comparing environmental DNA and RNA metabarcoding for vertebrate and invertebrate biodiversity surveys in a lowland stream. PeerJ, 12(10), e18016. [CrossRef]
- Makiola, A., Compson, Z. G., Baird, D. J., Barnes, M. A., Boerlijst, S. P., Bouchez, A., … Bohan, D. A. (2020). Key Questions for Next-Generation Biomonitoring. Frontiers in Environmental Science, 7, 476977. [CrossRef]
- Marie-Anne Le Guen, C. M., Raoux, A., Tecchio, S., Chauvel, N., Merzi, T., Bellemain, E., … Pezy, J. P. (2025). Environmental DNA as a method to reconstruct food webs and assess ecosystem health. Ecological Indicators, 173, 113399. [CrossRef]
- Marshall, N. T., Vanderploeg, H. A., & Chaganti, S. R. (2021). Environmental (e)RNA advances the reliability of eDNA by predicting its age. Scientific Reports 2021 11:1, 11(1), 1–11. [CrossRef]
- Miyata, K., Inoue, Y., Amano, Y., Nishioka, T., Nagaike, T., Kawaguchi, T., … Honda, H. (2022). Comparative environmental RNA and DNA metabarcoding analysis of river algae and arthropods for ecological surveys and water quality assessment. Scientific Reports 2022 12:1, 12(1), 19828-. [CrossRef]
- Miyata, K., Inoue, Y., Amano, Y., Nishioka, T., Yamane, M., Kawaguchi, T., … Honda, H. (2021). Fish environmental RNA enables precise ecological surveys with high positive predictivity. Ecological Indicators, 128, 107796. [CrossRef]
- Miyata, K., Kusakabe, Y., Inoue, Y., Yamane, M., & Honda, H. (2025). Validation of fish environmental RNA metabarcoding analysis for ecological surveys by additional traditional field surveys in the Naka River. Frontiers in Ecology and Evolution, 13, 1540001. [CrossRef]
- Morgado-Gamero, W. B., Tournayre, O., & Cristescu, M. E. (2025). Comparative Decay Dynamics and Detectability of eDNA and eRNA in Connected and Isolated Freshwater Mesocosms Using Digital PCR. Molecular Ecology Resources, 25(8), e70028. [CrossRef]
- Nicholson, A., McIsaac, D., MacDonald, C., Gec, P., Mason, B. E., Rein, W., … Hanner, R. H. (2020). An analysis of metadata reporting in freshwater environmental DNA research calls for the development of best practice guidelines. Environmental DNA, 2(3), 343–349. [CrossRef]
- Novotny, A., Serandour, B., Kortsch, S., Gauzens, B., Jan, K. M. G., & Winder, M. (2023). DNA metabarcoding highlights cyanobacteria as the main source of primary production in a pelagic food web model. Science Advances, 9(17); SUBPAGE:STRING:FULL . [CrossRef]
- Oña, L., Shreekar, S. K., & Kost, C. (2025). Disentangling microbial interaction networks. Trends in Microbiology, 33(6), 619–634. [CrossRef]
- Parsley, M. B., & Goldberg, C. S. (2024). Environmental RNA can distinguish life stages in amphibian populations. Molecular Ecology Resources, 24(4), e13857. [CrossRef]
- Pawlowski, J., Apothéloz-Perret-Gentil, L., & Altermatt, F. (2020). Environmental DNA: What’s behind the term? Clarifying the terminology and recommendations for its future use in biomonitoring. Molecular Ecology, 29(22), 4258–4264;WGROUP:STRING:PUBLICATION . [CrossRef]
- Pawlowski, J., Bonin, A., Boyer, F., Cordier, T., & Taberlet, P. (2021). Environmental DNA for biomonitoring. Molecular Ecology, 30(13), 2931. [CrossRef]
- Pawlowski, J., Kelly-Quinn, M., Altermatt, F., Apothéloz-Perret-Gentil, L., Beja, P., Boggero, A., … Kahlert, M. (2018). The future of biotic indices in the ecogenomic era: Integrating (e)DNA metabarcoding in biological assessment of aquatic ecosystems. Science of The Total Environment, 637–638, 1295–1310. [CrossRef]
- Pompanon, F., Deagle, B. E., Symondson, W. O. C., Brown, D. S., Jarman, S. N., & Taberlet, P. (2012). Who is eating what: diet assessment using next generation sequencing. Molecular Ecology, 21(8), 1931–1950. [CrossRef]
- Rodriguez, L. K., De Bonis, L., McKee, J., McKenna, J. A., Urvois, T., Barbaccia, E., … Thalinger, B. (2025). Inter-laboratory ring test for environmental DNA extraction protocols: implications for marine megafauna detection using three novel qPCR assays. Metabarcoding & Metagenomics, 9, 1–34. [CrossRef]
- Runge, J., Bathiany, S., Bollt, E., Camps-Valls, G., Coumou, D., Deyle, E., … Zscheischler, J. (2019). Inferring causation from time series in Earth system sciences. Nature Communications 2019 10:1, 10(1), 2553-. [CrossRef]
- Ruppert, K. M., Kline, R. J., & Rahman, M. S. (2019). Past, present, and future perspectives of environmental DNA (eDNA) metabarcoding: A systematic review in methods, monitoring, and applications of global eDNA. Global Ecology and Conservation, 17, e00547. [CrossRef]
- Schrodt, F., Beck, M., Estopinan, J., Bowler, D. E., Fontaine, C., Gaüzère, P., … Thuiller, W. (2025). Advancing causal inference in ecology: Pathways for biodiversity change detection and attribution. Methods in Ecology and Evolution, 16(10), 2276–2304;PAGE:STRING:ARTICLE/CHAPTER. [CrossRef]
- Scriver, M., Zaiko, A., Pochon, X., Stanton, J. A. L., Belonovich, O., Jeunen, G. J., … von Ammon, U. (2025). Biodiversity monitoring in remote marine environments: Advancing environmental DNA/RNA sampling workflows. Marine Environmental Research, 206, 107041. [CrossRef]
- Simaika, J. P., Stribling, J., Lento, J., Bruder, A., Poikane, S., Moretti, M. S., … Macadam, C. R. (2024). Towards harmonized standards for freshwater biodiversity monitoring and biological assessment using benthic macroinvertebrates. Science of The Total Environment, 918, 170360. [CrossRef]
- Stein, E. D., Jerde, C. L., Allan, E. A., Sepulveda, A. J., Abbott, C. L., Baerwald, M. R., … Thielen, P. M. (2024). Critical considerations for communicating environmental DNA science. Environmental DNA, 6(1), e472. [CrossRef]
- Stevens, J. D., & Parsley, M. B. (2023). Environmental RNA applications and their associated gene targets for management and conservation. Environmental DNA, 5(2), 227–239. [CrossRef]
- Sugihara, G., May, R., Ye, H., Hsieh, C. H., Deyle, E., Fogarty, M., & Munch, S. (2012). Detecting causality in complex ecosystems. Science, 338(6106), 496–500. [CrossRef]
- Taberlet, P., Coissac, E., Pompanon, F., Brochmann, C., & Willerslev, E. (2012). Towards next-generation biodiversity assessment using DNA metabarcoding. Molecular Ecology, 21(8), 2045–2050;REQUESTEDJOURNAL:JOURNAL:1365294X;CSUBTYPE:STRING:SPECIAL;PAGE:STRING:ARTICLE/CHAPTER. [CrossRef]
- Takahashi, M., Frøslev, T. G., Paupério, J., Thalinger, B., Klymus, K., Helbing, C. C., … Berry, O. (2025). A Metadata Checklist and Data Formatting Guidelines to Make eDNA FAIR (Findable, Accessible, Interoperable, and Reusable). Environmental DNA, 7(3), e70100. [CrossRef]
- Thalinger, B., Deiner, K., Harper, L. R., Rees, H. C., Blackman, R. C., Sint, D., … Bruce, K. (2021). A validation scale to determine the readiness of environmental DNA assays for routine species monitoring. Environmental DNA, 3(4), 823–836. [CrossRef]
- Thébault, E., & Fontaine, C. (2010). Stability of ecological communities and the architecture of mutualistic and trophic networks. Science, 329(5993), 853–856. [CrossRef]
- Thomas, A. C., Nguyen, P. L., Howard, J., & Goldberg, C. S. (2019). A self-preserving, partially biodegradable eDNA filter. Methods in Ecology and Evolution, 10(8), 1136–1141. [CrossRef]
- Turner, C. R., Barnes, M. A., Xu, C. C. Y., Jones, S. E., Jerde, C. L., & Lodge, D. M. (2014). Particle size distribution and optimal capture of aqueous macrobial eDNA. Methods in Ecology and Evolution, 5(7), 676–684. [CrossRef]
- Tylianakis, J. M., Didham, R. K., Bascompte, J., & Wardle, D. A. (2008). Global change and species interactions in terrestrial ecosystems. Ecology Letters, 11(12), 1351–1363. [CrossRef]
- Valiente-Banuet, A., Aizen, M. A., Alcántara, J. M., Arroyo, J., Cocucci, A., Galetti, M., … Zamora, R. (2015). Beyond species loss: the extinction of ecological interactions in a changing world. Functional Ecology, 29(3), 299–307. [CrossRef]
- Vasselon, V., Rivera, S. F., Ács, É., Almeida, S. B., Andree, K. B., Apothéloz-Perret-Gentil, L., … Paix, B. (2025). Proficiency testing and cross-laboratory method comparison to support standardisation of diatom DNA metabarcoding for freshwater biomonitoring. Metabarcoding and Metagenomics, 3, 35–70. [CrossRef]
- Weiss, S., Van Treuren, W., Lozupone, C., Faust, K., Friedman, J., Deng, Y., … Knight, R. (2016). Correlation detection strategies in microbial data sets vary widely in sensitivity and precision. The ISME Journal, 10(7), 1669–1681. [CrossRef]
- Wieczorek, J., Bloom, D., Guralnick, R., Blum, S., Döring, M., Giovanni, R., … Vieglais, D. (2012). Darwin Core: An Evolving Community-Developed Biodiversity Data Standard. PLOS ONE, 7(1), e29715. [CrossRef]
- Wilkinson, M. D., Dumontier, M., Aalbersberg, Ij. J., Appleton, G., Axton, M., Baak, A., … Mons, B. (2016). The FAIR Guiding Principles for scientific data management and stewardship. Scientific Data 2016 3:1, 3(1), 1–9. [CrossRef]
- Wood, S. A., Biessy, L., Latchford, J. L., Zaiko, A., von Ammon, U., Audrezet, F., … Pochon, X. (2020). Release and degradation of environmental DNA and RNA in a marine system. Science of The Total Environment, 704, 135314. [CrossRef]
- Xu, Z., & Asakawa, S. (2025). Release and degradation of dissolved environmental RNAs from zebrafish cells. RNA Biology, 22(1), 1–12. [CrossRef]
- Xue, Z., Tian, W., Han, Y., Li, S., Guo, J., He, H., … Zhang, W. (2024). Environmental RNA metabarcoding for ballast water microbial diversity: Minimizing false positives. Science of The Total Environment, 955, 176902. [CrossRef]
- Yamahara, K. M., Allan, E. A., Robidart, J., Wilson, W. H., Birch, J. M., Craw, P., … Scholin, C. A. (2025). A State-Of-The-Art Review of Aquatic eDNA Sampling Technologies and Instrumentation: Advancements, Challenges, and Future Prospects. Environmental DNA, 7(4), e70170. [CrossRef]
- Yates, M. C., Derry, A. M., & Cristescu, M. E. (2021). Environmental RNA: A Revolution in Ecological Resolution? Trends in Ecology and Evolution, 36(7), 601–609. [CrossRef]
- Yilmaz, P., Kottmann, R., Field, D., Knight, R., Cole, J. R., Amaral-Zettler, L., … Glöckner, F. O. (2011). Minimum information about a marker gene sequence (MIMARKS) and minimum information about any (x) sequence (MIxS) specifications. Nature Biotechnology 2011 29:5, 29(5), 415–420. [CrossRef]
- Zhang, Y., Qiu, Y., Liu, K., Zhong, W., Yang, J., Altermatt, F., & Zhang, X. (2024). Evaluating eDNA and eRNA metabarcoding for aquatic biodiversity assessment: From bacteria to vertebrates. Environmental Science and Ecotechnology, 21, 100441. [CrossRef]
- Zinger, L., Bonin, A., Alsos, I. G., Bálint, M., Bik, H., Boyer, F., … Taberlet, P. (2019). DNA metabarcoding—Need for robust experimental designs to draw sound ecological conclusions. Molecular Ecology, 28(8), 1857–1862. [CrossRef]
- Zurell, D., Pollock, L. J., & Thuiller, W. (2018). Do joint species distribution models reliably detect interspecific interactions from co-occurrence data in homogenous environments? Ecography, 41(11), 1812–1819. WGROUP:STRING:PUBLICATION. [CrossRef]

| Tier | What you can claim | Minimum data/design requirements | Typical pitfalls / failure modes | Example Biomonitoring 3.0 outputs | Key refs (non-exhaustive) |
|---|---|---|---|---|---|
| 0 | Co-detection / association in space-time | Cross-sectional samples + controls | Habitat filtering, transport, detectability artifacts | Taxa A and B co-occur seasonally | Blanchet et al. 2020; Zinger et al. 2019 |
| 1 | Conditional association network (not mechanism) | Replication + confounder control; uncertainty/stability checks | Compositionality, sparsity, method dependence | Network metrics w/ bootstrap stability | Barroso-Bergadà et al. 2021; Weiss et al. 2016; Kurtz et al. 2015 |
| 2 | Directional influence suggested by time ordering | Time series with adequate frequency, covariates, replication | Unmeasured confounders; aliasing; nonstationarity | Candidate driver → response edges | Runge et al. 2019; Schrodt et al. 2025; Sugihara et al. 2012 |
| 3 | Interaction-explicit links (edge meaning is clear) | Diet/pollen/gut/parasite-on-host designs | Quantitative bias; contamination; ref DB gaps | Trophic or mutualistic networks with explicit edges | Cordier et al. 2021; Pompanon et al. 2012; Deagle et al. 2019; Encinas-Viso et al. 2023 |
| 4 | Coupled signal–response consistent with mechanism | Paired source/recipient + time lag + response readout (often eRNA) | Low target mapping fraction; batch effects; alternative drivers | Stress signature in recipient aligned to exposure | Hechler et al. 2023; Hiki et al. 2023; He et al. 2025; Oña et al. 2025 |
| 5 | Ecological consequence (process-level change) | Process metrics + validation/perturbation + robust linkage | Process multi-causality; scale mismatch | Link shifts associated with productivity/stability proxies | Delmas et al. 2019; Thébault & Fontaine 2010; Allesina & Tang 2012; Abdala-Roberts et al. 2025 |
| Program archetype | Intended tier(s) | Sampling pattern | Matrices & molecules | Lab/seq strategy | Best-fit outputs | Key refs (non-exhaustive) |
|---|---|---|---|---|---|---|
| High-frequency river surveillance | 1–2 (sometimes 4) | Automated + manual calibration; event-based bursts | Water eDNA + optional eRNA | Robust controls; consistent pipeline | Time-resolved change detection; candidate drivers | George et al. 2024; Hallam et al. 2023; Yamahara et al. 2025; Deiner et al. 2016 |
| Paired source–recipient monitoring | 2–4 | Matched sampling upstream/downstream or host/environment; explicit lags | eDNA scaffold + eRNA response | Enrichment if targeting eukaryotic response | Exposure plausibility + recipient response | Hechler et al. 2023; He et al. 2025; Scriver et al. 2025 |
| Biosecurity / disinfection verification | 1–3 | Before/after treatment; replicate controls | eDNA + eRNA to reduce legacy signal | Targeted or 16S/marker panels | Reduced “ghost” detection; compliance evidence | Xue et al. 2024; Darling et al. 2021; Klymus et al. 2020 |
| Food-web indicators via metaweb constraints | 2–5 (depending on validation) | Seasonal surveys + curated trophic info | eDNA inventories + metaweb | Strong taxonomy curation; sensitivity checks | Food-web indicators (connectivity, redundancy) | D’Alessandro & Mariani 2021; Le Guen et al. 2025; Boyse et al. 2025 |
| Interaction-explicit trophic monitoring | 3–4 | Gut/diet sampling + environment | Diet DNA + optional eRNA state | Bias-aware thresholds; controls | Trophic edges with clear meaning | Pompanon et al. 2012; Deagle et al. 2019; Novotny et al. 2023 |
| Reporting element (3.0 add-on) | Why it’s needed for interaction-ready inference | Minimum to report | Applies most to tier(s) | Key refs (non-exhaustive) |
|---|---|---|---|---|
| Tier label (0–5) | Prevents over-interpretation | Tier + rationale | All | Schrodt et al. 2025; Barroso-Bergadà et al. 2021 |
| Time-lag structure | Enables direction/ signal→response tests |
Lag choice, cadence, replication | 2–5 | Runge et al. 2019; Schrodt et al. 2025 |
| Paired compartments | Makes exposure plausible | Source/recipient definition + spatial logic | 2–4 | He et al. 2025; Hechler et al. 2023 |
| Molecule type & transcript class | Time resolution depends on RNA class | DNA vs RNA; rRNA vs mRNA; targets | 1–4 | Yates et al. 2021; Morgado-Gamero et al. 2025 |
| Filtration/size fractions | State affects localization/transport | Pore size(s), fractions, volumes | 1–4 | Jo 2023; Hiki & Jo 2025; Turner et al. 2014 |
| Preservation & time-to-stabilization | RNA is handling-sensitive | Preservative, times, temps, deviations (audit) | 1–4 | Scriver et al. 2025; Thomas et al. 2019; Klymus et al. 2024 |
| Enrichment/library strategy | Determines detectability of response |
rRNA depletion, poly(A), capture, depth | 4 | Hechler et al. 2023; Stevens & Parsley 2023 |
| Control structure (field→bioinfo) | Supports false-positive control | Field blanks, extraction blanks, PCR neg/pos | All | MIEM: Klymus et al. 2024; Ficetola et al. 2016 |
| Provenance + versioning | Avoids “method drift” looking like ecology | Pipeline, parameters, DB build/version | All | Takahashi et al. 2025; Keck et al. 2023 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




