Submitted:
13 March 2026
Posted:
16 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Methods
2.1. Cell Culture and Reagents
2.2. sgRNAs and DNA Donors
2.3. Cell Sorting and Flow Cytometry
2.4. Genomic DNA Isolation and DNA Sequencing
3. Results
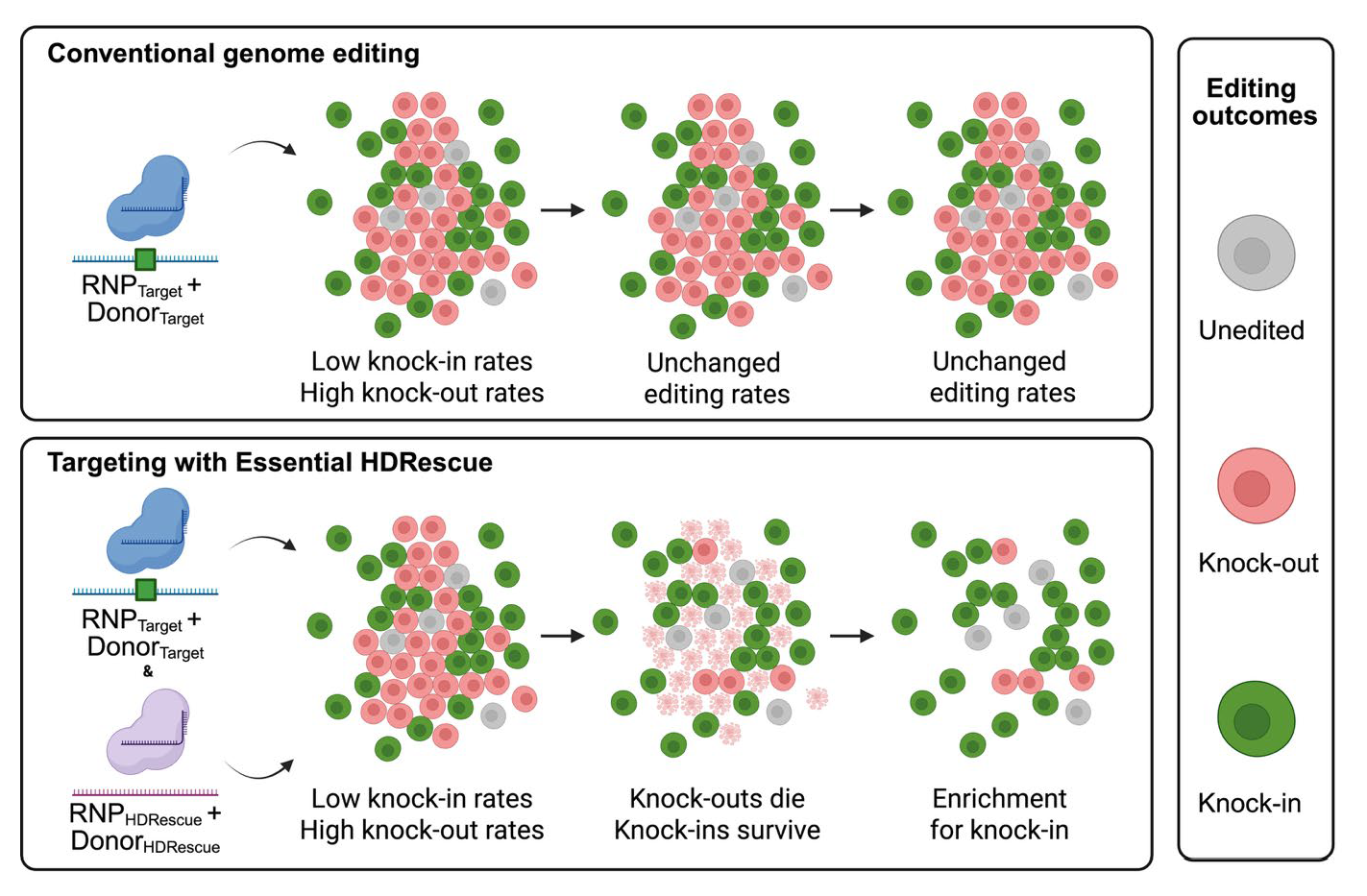
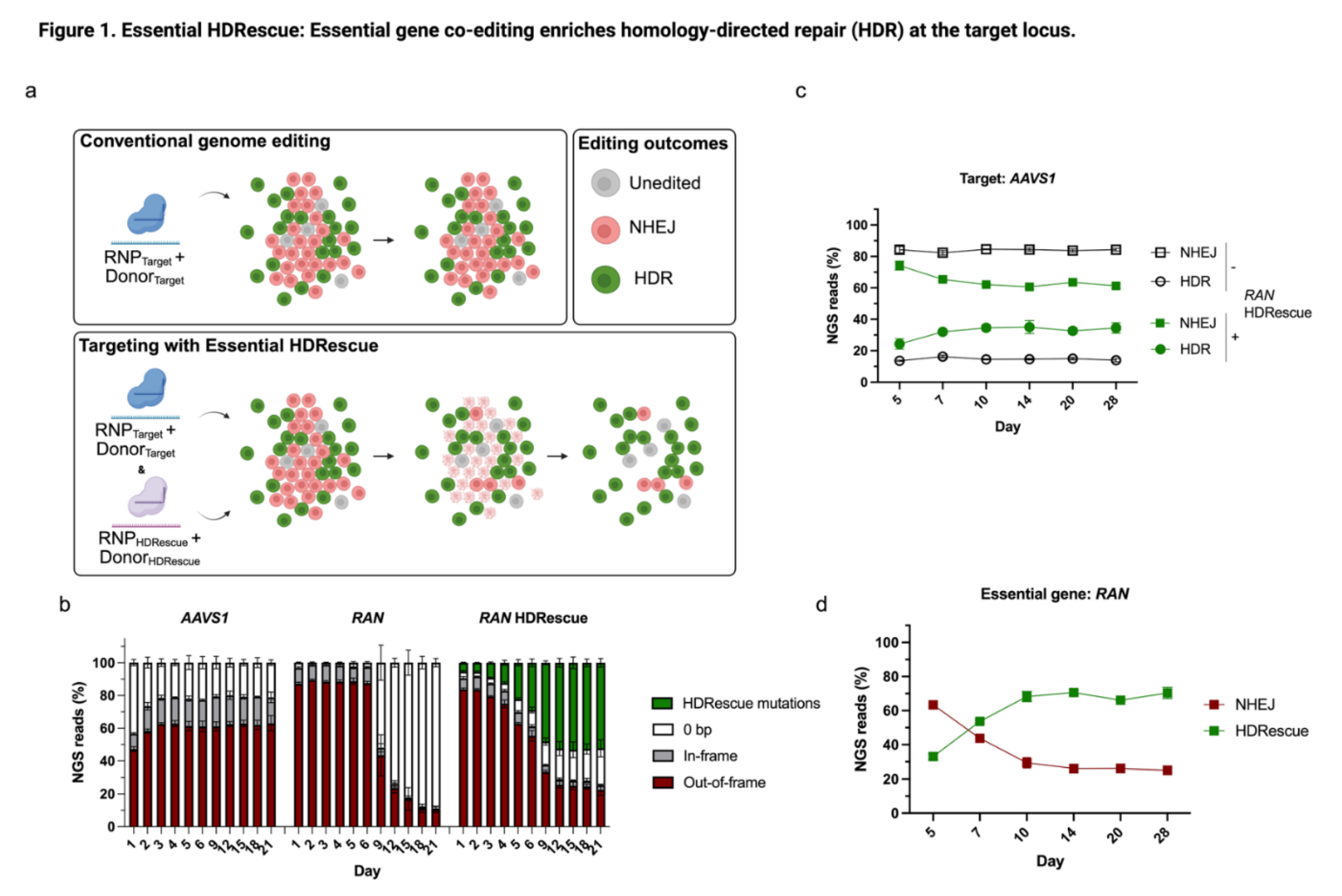
3.1. HDRescue: Essential Gene co-Editing Enriched HDR at the Target Locus
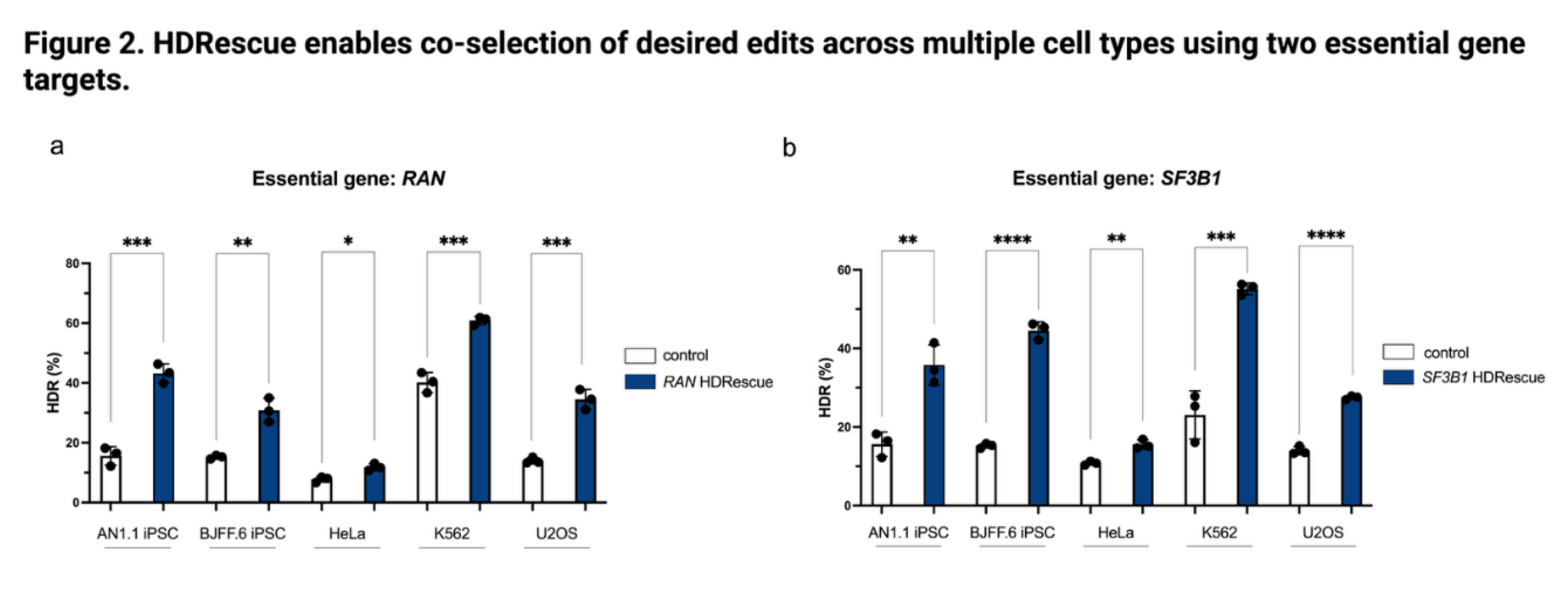
3.2. HDRescue Enabled co-Selection of Desired Edits Across Multiple Cell Types Using Two Essential Gene Targets
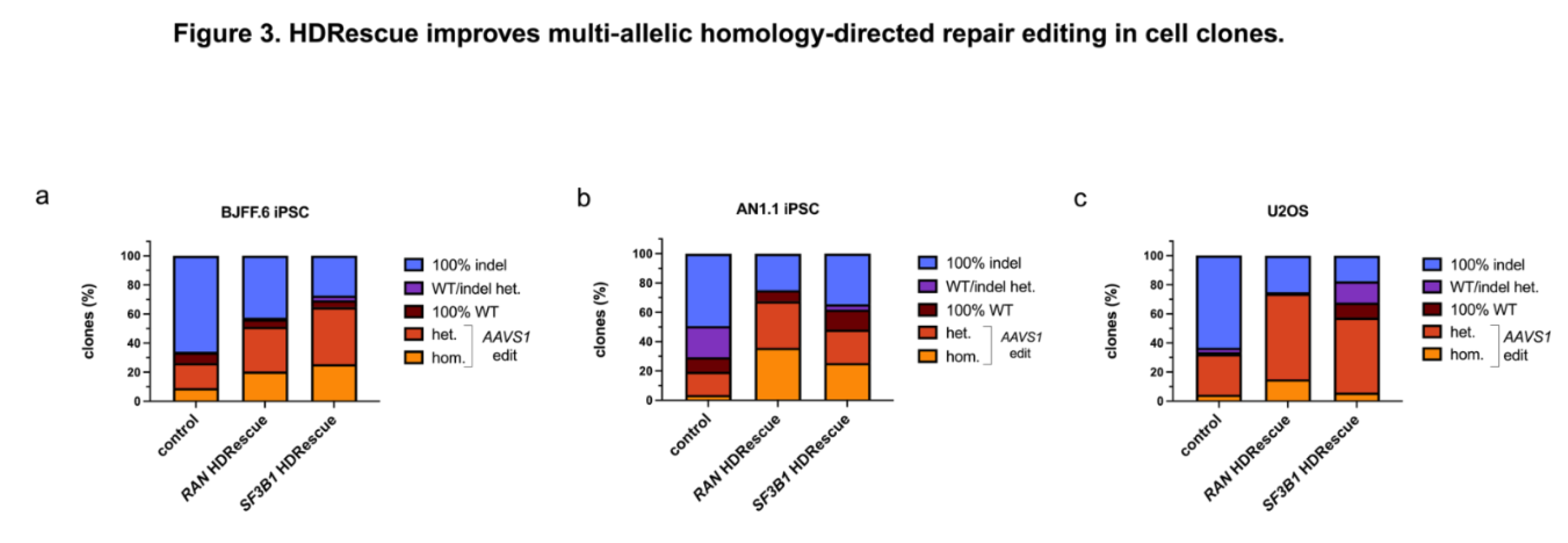
3.3. HDRescue Improved Multi-Allelic HDR Editing in Cell Clones
3.4. Detection and Removal of Aberrant Clones Improves HDRescue Accuracy
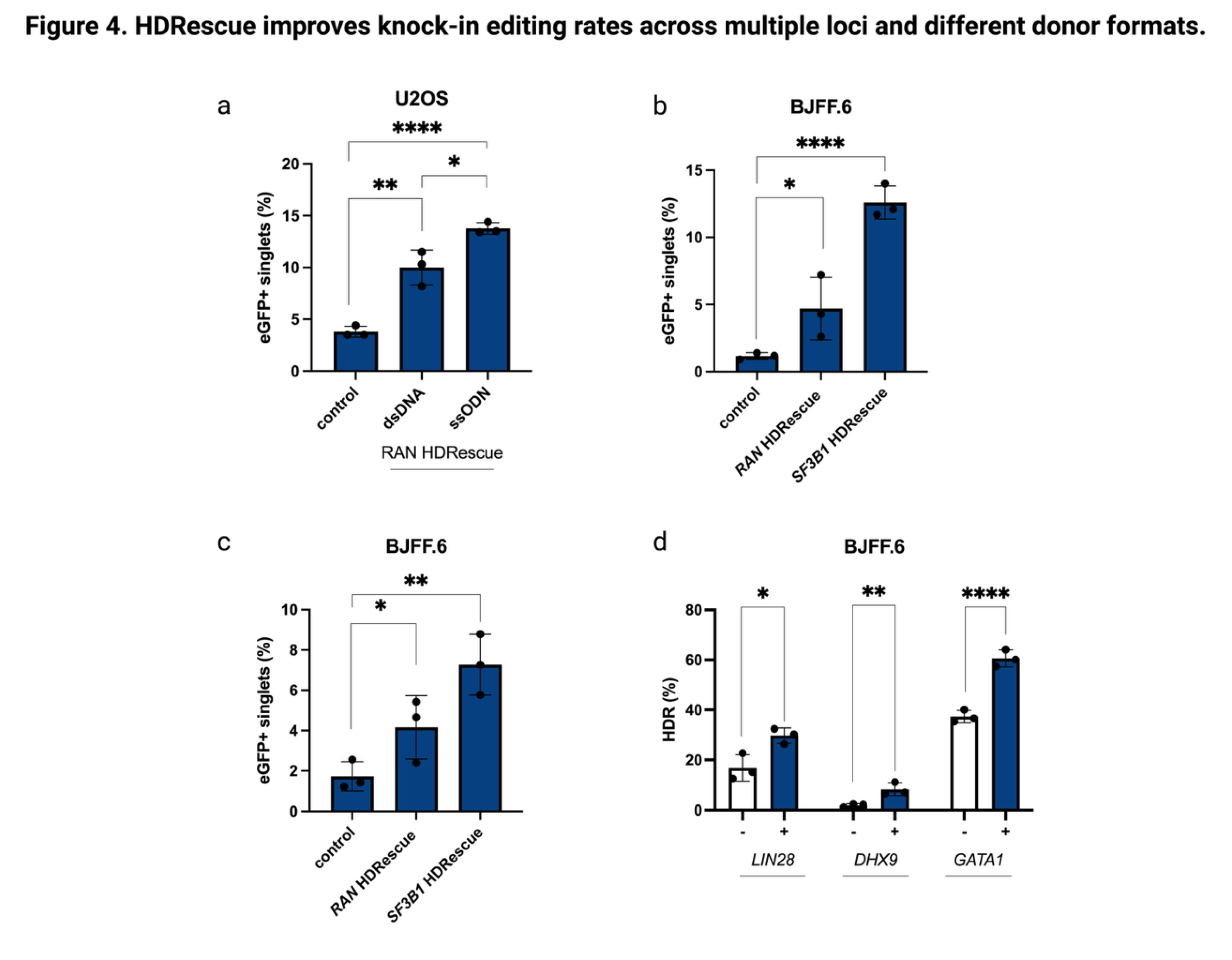
3.5. HDRescue Improved HDR Editing Rates Across Multiple Loci and Different Donor Formats
4. Discussion
Supplementary Materials
Author Contributions
Funding
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Cheng, H.; Jeong, E.; Cho, S.W. Applications of multiplexed CRISPR–Cas for genome engineering. Experimental & Molecular Medicine 2025, 57, 1373–1380. [Google Scholar] [CrossRef]
- Hsu, P.D.; Lander, E.S.; Zhang, F. Development and applications of CRISPR-Cas9 for genome engineering. Cell 2014, 157, 1262–1278. [Google Scholar] [CrossRef] [PubMed]
- Sonoda, E.; Hochegger, H.; Saberi, A.; Taniguchi, Y.; Takeda, S. Differential usage of non-homologous end-joining and homologous recombination in double strand break repair. DNA Repair (Amst) 2006, 5, 1021–1029. [Google Scholar] [CrossRef] [PubMed]
- Jasin, M.; Haber, J.E. The democratization of gene editing: Insights from site-specific cleavage and double-strand break repair. DNA Repair 2016, 44, 6–16. [Google Scholar] [CrossRef] [PubMed]
- Chu, V.T.; Weber, T.; Wefers, B.; Wurst, W.; Sander, S.; Rajewsky, K.; Kühn, R. Increasing the efficiency of homology-directed repair for CRISPR-Cas9-induced precise gene editing in mammalian cells. Nature Biotechnology 2015, 33, 543–548. [Google Scholar] [CrossRef]
- Jasin, M.; Rothstein, R. Repair of strand breaks by homologous recombination. Cold Spring Harb Perspect Biol 2013, 5, a012740. [Google Scholar] [CrossRef]
- Porter, S.N.; Levine, R.M.; Pruett-Miller, S.M. A Practical Guide to Genome Editing Using Targeted Nuclease Technologies. Compr Physiol 2019, 9, 665–714. [Google Scholar] [CrossRef]
- Haider, S.; Mussolino, C. Fine-Tuning Homology-Directed Repair (HDR) for Precision Genome Editing: Current Strategies and Future Directions. Int J Mol Sci 2025, 26. [Google Scholar] [CrossRef]
- Cullot, G.; Aird, E.J.; Schlapansky, M.F.; Yeh, C.D.; van de Venn, L.; Vykhlyantseva, I.; Kreutzer, S.; Mailander, D.; Lewkow, B.; Klermund, J.; et al. Genome editing with the HDR-enhancing DNA-PKcs inhibitor AZD7648 causes large-scale genomic alterations. Nat Biotechnol 2025, 43, 1778–1782. [Google Scholar] [CrossRef]
- Gaudelli, N.M.; Komor, A.C.; Rees, H.A.; Packer, M.S.; Badran, A.H.; Bryson, D.I.; Liu, D.R. Programmable base editing of A*T to G*C in genomic DNA without DNA cleavage. Nature 2017, 551, 464–471. [Google Scholar] [CrossRef]
- Komor, A.C.; Kim, Y.B.; Packer, M.S.; Zuris, J.A.; Liu, D.R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 2016, 533, 420–424. [Google Scholar] [CrossRef] [PubMed]
- Anzalone, A.V.; Randolph, P.B.; Davis, J.R.; Sousa, A.A.; Koblan, L.W.; Levy, J.M.; Chen, P.J.; Wilson, C.; Newby, G.A.; Raguram, A.; et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 2019, 576, 149–157. [Google Scholar] [CrossRef] [PubMed]
- Lavrov, A.V.; Varenikov, G.G.; Skoblov, M.Y. Genome scale analysis of pathogenic variants targetable for single base editing. BMC Med Genomics 2020, 13, 80. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.Y.; Doudna, J.A. CRISPR technology: A decade of genome editing is only the beginning. Science 2023, 379, eadd8643. [Google Scholar] [CrossRef]
- Ren, C.; Xu, K.; Segal, D.J.; Zhang, Z. Strategies for the Enrichment and Selection of Genetically Modified Cells. Trends Biotechnol 2019, 37, 56–71. [Google Scholar] [CrossRef]
- Mikkelsen, N.S.; Bak, R.O. Enrichment strategies to enhance genome editing. J Biomed Sci 2023, 30, 51. [Google Scholar] [CrossRef]
- Certo, M.T.; Ryu, B.Y.; Annis, J.E.; Garibov, M.; Jarjour, J.; Rawlings, D.J.; Scharenberg, A.M. Tracking genome engineering outcome at individual DNA breakpoints. Nat Methods 2011, 8, 671–676. [Google Scholar] [CrossRef]
- Brafman, D.A.; Wang, X.; Nguyen, T.; Schwarz, G.; Brookhouser, N.; Tekel, S.J.; Standage-Beier, K. A transient reporter for editing enrichment (TREE) in human cells. Nucleic Acids Research 2019, 47, e120–e120. [Google Scholar] [CrossRef]
- Reuven, N.; Adler, J.; Myers, N.; Shaul, Y. CRISPR Co-Editing Strategy for Scarless Homology-Directed Genome Editing. Int J Mol Sci 2021, 22. [Google Scholar] [CrossRef]
- Reuven, N.; Shaul, Y. Selecting for CRISPR-Edited Knock-In Cells. Int J Mol Sci 2022, 23. [Google Scholar] [CrossRef]
- Liao, S.; Tammaro, M.; Yan, H. Enriching CRISPR-Cas9 targeted cells by co-targeting the HPRT gene. Nucleic Acids Res 2015, 43, e134. [Google Scholar] [CrossRef]
- Kim, H.; Um, E.; Cho, S.-R.; Jung, C.; Kim, H.; Kim, J.-S. Surrogate reporters for enrichment of cells with nuclease-induced mutations. Nature Methods 2011, 8, 941–943. [Google Scholar] [CrossRef] [PubMed]
- Arribere, J.A.; Bell, R.T.; Fu, B.X.; Artiles, K.L.; Hartman, P.S.; Fire, A.Z. Efficient marker-free recovery of custom genetic modifications with CRISPR/Cas9 in Caenorhabditis elegans. Genetics 2014, 198, 837–846. [Google Scholar] [CrossRef] [PubMed]
- Ward, J.D. Rapid and precise engineering of the Caenorhabditis elegans genome with lethal mutation co-conversion and inactivation of NHEJ repair. Genetics 2015, 199, 363–377. [Google Scholar] [CrossRef]
- Kim, H.; Ishidate, T.; Ghanta, K.S.; Seth, M.; Conte, D., Jr.; Shirayama, M.; Mello, C.C. A co-CRISPR strategy for efficient genome editing in Caenorhabditis elegans. Genetics 2014, 197, 1069–1080. [Google Scholar] [CrossRef]
- Kane, N.S.; Vora, M.; Varre, K.J.; Padgett, R.W. Efficient Screening of CRISPR/Cas9-Induced Events in Drosophila Using a Co-CRISPR Strategy. G3 (Bethesda) 2017, 7, 87–93. [Google Scholar] [CrossRef]
- Ge, D.T.; Tipping, C.; Brodsky, M.H.; Zamore, P.D. Rapid Screening for CRISPR-Directed Editing of the Drosophila Genome Using white Coconversion. G3 (Bethesda) 2016, 6, 3197–3206. [Google Scholar] [CrossRef]
- Yan, N.; Sun, Y.; Fang, Y.; Deng, J.; Mu, L.; Xu, K.; Mymryk, J.S.; Zhang, Z. A Universal Surrogate Reporter for Efficient Enrichment of CRISPR/Cas9-Mediated Homology-Directed Repair in Mammalian Cells. Molecular Therapy - Nucleic Acids 2020, 19, 775–789. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Akrap, N.; Cerboni, S.; Porritt, M.J.; Wimberger, S.; Lundin, A.; Möller, C.; Firth, M.; Gordon, E.; Lazovic, B.; et al. Universal toxin-based selection for precise genome engineering in human cells. Nature Communications 2021, 12, 497. [Google Scholar] [CrossRef]
- Agudelo, D.; Duringer, A.; Bozoyan, L.; Huard, C.C.; Carter, S.; Loehr, J.; Synodinou, D.; Drouin, M.; Salsman, J.; Dellaire, G.; et al. Marker-free coselection for CRISPR-driven genome editing in human cells. Nat Methods 2017, 14, 615–620. [Google Scholar] [CrossRef]
- Mitzelfelt, K.A.; McDermott-Roe, C.; Grzybowski, M.N.; Marquez, M.; Kuo, C.T.; Riedel, M.; Lai, S.; Choi, M.J.; Kolander, K.D.; Helbling, D.; et al. Efficient Precision Genome Editing in iPSCs via Genetic Co-targeting with Selection. Stem Cell Reports 2017, 8, 491–499. [Google Scholar] [CrossRef]
- Moriarity, B.S.; Rahrmann, E.P.; Beckmann, D.A.; Conboy, C.B.; Watson, A.L.; Carlson, D.F.; Olson, E.R.; Hyland, K.A.; Fahrenkrug, S.C.; McIvor, R.S.; et al. Simple and Efficient Methods for Enrichment and Isolation of Endonuclease Modified Cells. PLOS ONE 2014, 9, e96114. [Google Scholar] [CrossRef]
- Shy, B.R.; MacDougall, M.S.; Clarke, R.; Merrill, B.J. Co-incident insertion enables high efficiency genome engineering in mouse embryonic stem cells. Nucleic Acids Res 2016, 44, 7997–8010. [Google Scholar] [CrossRef]
- Li, S.; Akrap, N.; Cerboni, S.; Porritt, M.J.; Wimberger, S.; Lundin, A.; Moller, C.; Firth, M.; Gordon, E.; Lazovic, B.; et al. Universal toxin-based selection for precise genome engineering in human cells. Nat Commun 2021, 12, 497. [Google Scholar] [CrossRef]
- Liu, W.H.; Volse, K.; Senft, D.; Jeremias, I. A reporter system for enriching CRISPR/Cas9 knockout cells in technically challenging settings like patient models. Sci Rep 2021, 11, 12649. [Google Scholar] [CrossRef]
- Allen, A.G.; Khan, S.Q.; Margulies, C.M.; Viswanathan, R.; Lele, S.; Blaha, L.; Scott, S.N.; Izzo, K.M.; Gerew, A.; Pattali, R.; et al. A highly efficient transgene knock-in technology in clinically relevant cell types. Nat Biotechnol 2023. [Google Scholar] [CrossRef]
- Bartha, I.; di Iulio, J.; Venter, J.C.; Telenti, A. Human gene essentiality. Nat Rev Genet 2018, 19, 51–62. [Google Scholar] [CrossRef] [PubMed]
- Huang, N.; Lee, I.; Marcotte, E.M.; Hurles, M.E. Characterising and predicting haploinsufficiency in the human genome. PLoS Genet 2010, 6, e1001154. [Google Scholar] [CrossRef] [PubMed]
- Yilmaz, A.; Peretz, M.; Aharony, A.; Sagi, I.; Benvenisty, N. Defining essential genes for human pluripotent stem cells by CRISPR-Cas9 screening in haploid cells. Nat Cell Biol 2018, 20, 610–619. [Google Scholar] [CrossRef] [PubMed]
- Tsherniak, A.; Vazquez, F.; Montgomery, P.G.; Weir, B.A.; Kryukov, G.; Cowley, G.S.; Gill, S.; Harrington, W.F.; Pantel, S.; Krill-Burger, J.M.; et al. Defining a Cancer Dependency Map. Cell 2017, 170, 564–576 e516. [Google Scholar] [CrossRef]
- Bairoch, A. The Cellosaurus, a Cell-Line Knowledge Resource. J Biomol Tech 2018, 29, 25–38. [Google Scholar] [CrossRef] [PubMed]
- Narina, S.; Connelly, J.P.; Pruett-Miller, S.M. Transgenesis: Methods and Protocols 2023, 155–182.
- Boudhraa, Z.; Carmona, E.; Provencher, D.; Mes-Masson, A.-M. Ran GTPase: A Key Player in Tumor Progression and Metastasis. Frontiers in Cell and Developmental Biology 2020, 8. [Google Scholar] [CrossRef]
- Institute, B. DepMap: The Cancer Dependency Map Project at Broad Institute. Available online: https://depmap.org/portal/.
- Loughran, A.J.; Narina, S.; Klein, J.; Siwak, J.F.; Connelly, J.P.; Pruett-Miller, S.M. Rapid and robust validation of pooled CRISPR knockout screens using CelFi. Sci Rep 2025, 15, 13358. [Google Scholar] [CrossRef]
- Guo, C.; Ma, X.; Gao, F.; Guo, Y. Off-target effects in CRISPR/Cas9 gene editing. Front Bioeng Biotechnol 2023, 11, 1143157. [Google Scholar] [CrossRef]
- Regan, S.B.; Medhi, D.; Xu, Y.; White, T.B.; Jiang, Y.Z.; Kim, J.E.; Wang, S.C.; Deng, Q.; Jia, S.; Baasan, D.; et al. Megabase-scale loss of heterozygosity provoked by CRISPR-Cas9 DNA double-strand breaks. Mol Cell 2025, 85, 4119–4137 e4110. [Google Scholar] [CrossRef]
- Kosicki, M.; Tomberg, K.; Bradley, A. Repair of double-strand breaks induced by CRISPR-Cas9 leads to large deletions and complex rearrangements. Nat Biotechnol 2018, 36, 765–771. [Google Scholar] [CrossRef]
- Liao, H.; Wu, J.; VanDusen, N.J.; Li, Y.; Zheng, Y. CRISPR-Cas9-mediated homology-directed repair for precise gene editing. Mol Ther Nucleic Acids 2024, 35, 102344. [Google Scholar] [CrossRef]
- Li, G.; Yang, X.; Luo, X.; Wu, Z.; Yang, H. Modulation of cell cycle increases CRISPR-mediated homology-directed DNA repair. Cell Biosci 2023, 13, 215. [Google Scholar] [CrossRef] [PubMed]
- Kostyrko, K.; Bosshard, S.; Urban, Z.; Mermod, N. A role for homologous recombination proteins in cell cycle regulation. Cell Cycle 2015, 14, 2853–2861. [Google Scholar] [CrossRef] [PubMed]
- Aussel, C.; Cathomen, T.; Fuster-Garcia, C. The hidden risks of CRISPR/Cas: structural variations and genome integrity. Nat Commun 2025, 16, 7208. [Google Scholar] [CrossRef] [PubMed]
- Dibitetto, D.; Marshall, S.; Sanchi, A.; Liptay, M.; Badar, J.; Lopes, M.; Rottenberg, S.; Smolka, M.B. DNA-PKcs promotes fork reversal and chemoresistance. Mol Cell 2022, 82, 3932–3942 e3936. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



