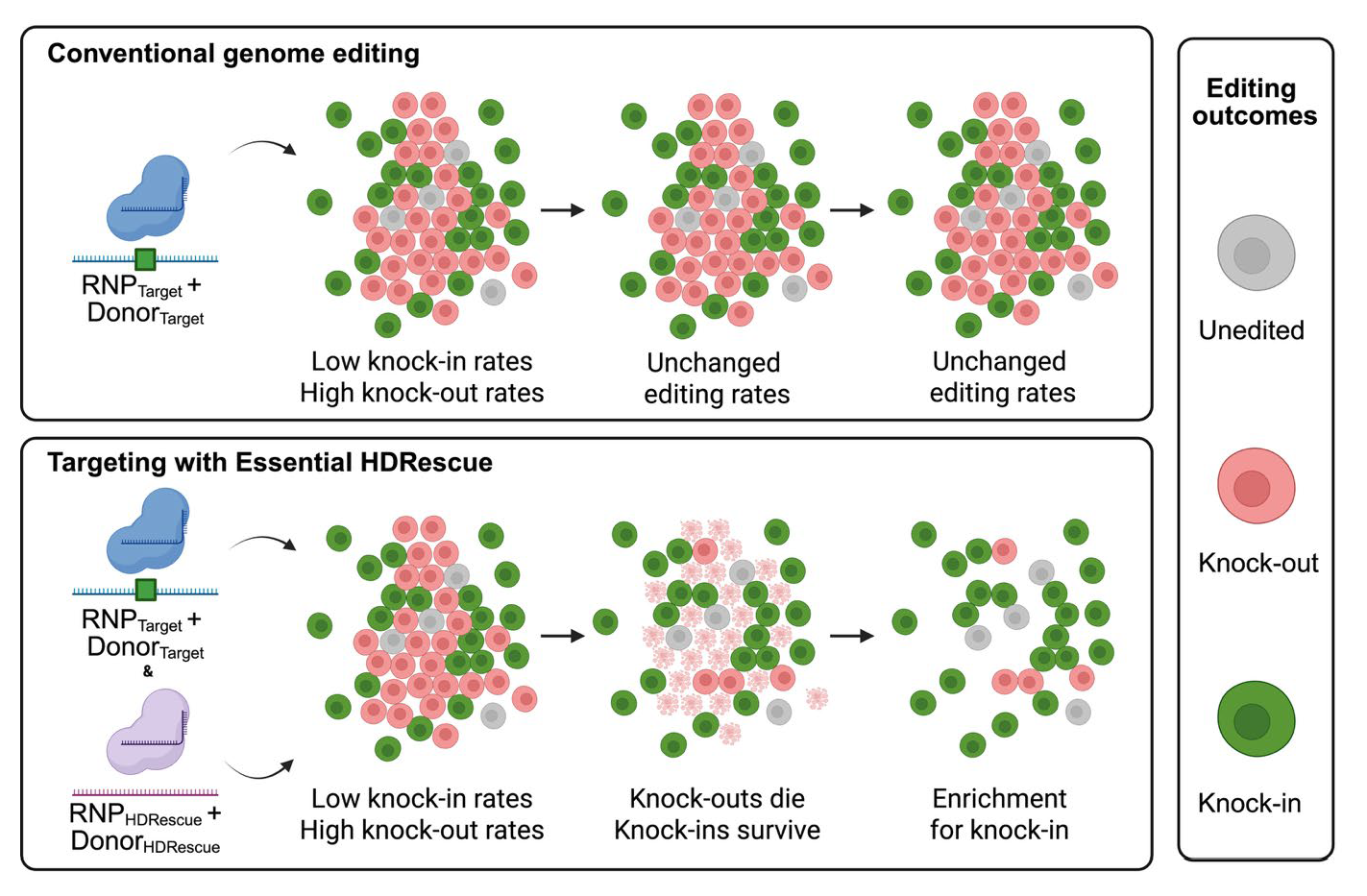
Genome editing is widely used and conceptually simple, yet in practice, it is hindered by laborious workflows and high costs. These challenges stem from the difficulty of identifying and isolating cells that contain the desired user‑defined modifications, a problem compounded by the wide variability in editing efficiencies across cell types. While homology-directed repair (HDR) provides a mechanism for precise genome modification following nuclease-induced double-strand breaks (DSBs), it is frequently outcompeted by the dominant mutagenic non-homologous end-joining (NHEJ) pathway in mammalian cells. Therefore, we developed a novel enrichment method, Essential HDRescue, to increase the frequency of HDR events at a target site by co-targeting an essential genomic locus. Using both intrinsic positive and negative selection at a common essential gene, we enabled enrichment of precise editing events at a second, unlinked target site. We demonstrated that co-targeting essential genes in cancer cell lines and iPSCs increased HDR rates without the need for an exogenous reporter or selective drug. Analysis of resulting clones revealed that Essential HDRescue produced up to a 6‑fold increase in single‑allele edits and almost a four‑fold increase in homozygous edits relative to single‑targeted controls. By harnessing the intrinsic cellular dependencies that arise from DSB repair at essential loci, Essential HDRescue offers a widely applicable method to improve precise genome editing outcomes in mammalian cells, leaving only a minimal, protein-silent scar at the essential gene.