Submitted:
09 April 2026
Posted:
10 April 2026
You are already at the latest version
Abstract
Keywords:
Introduction
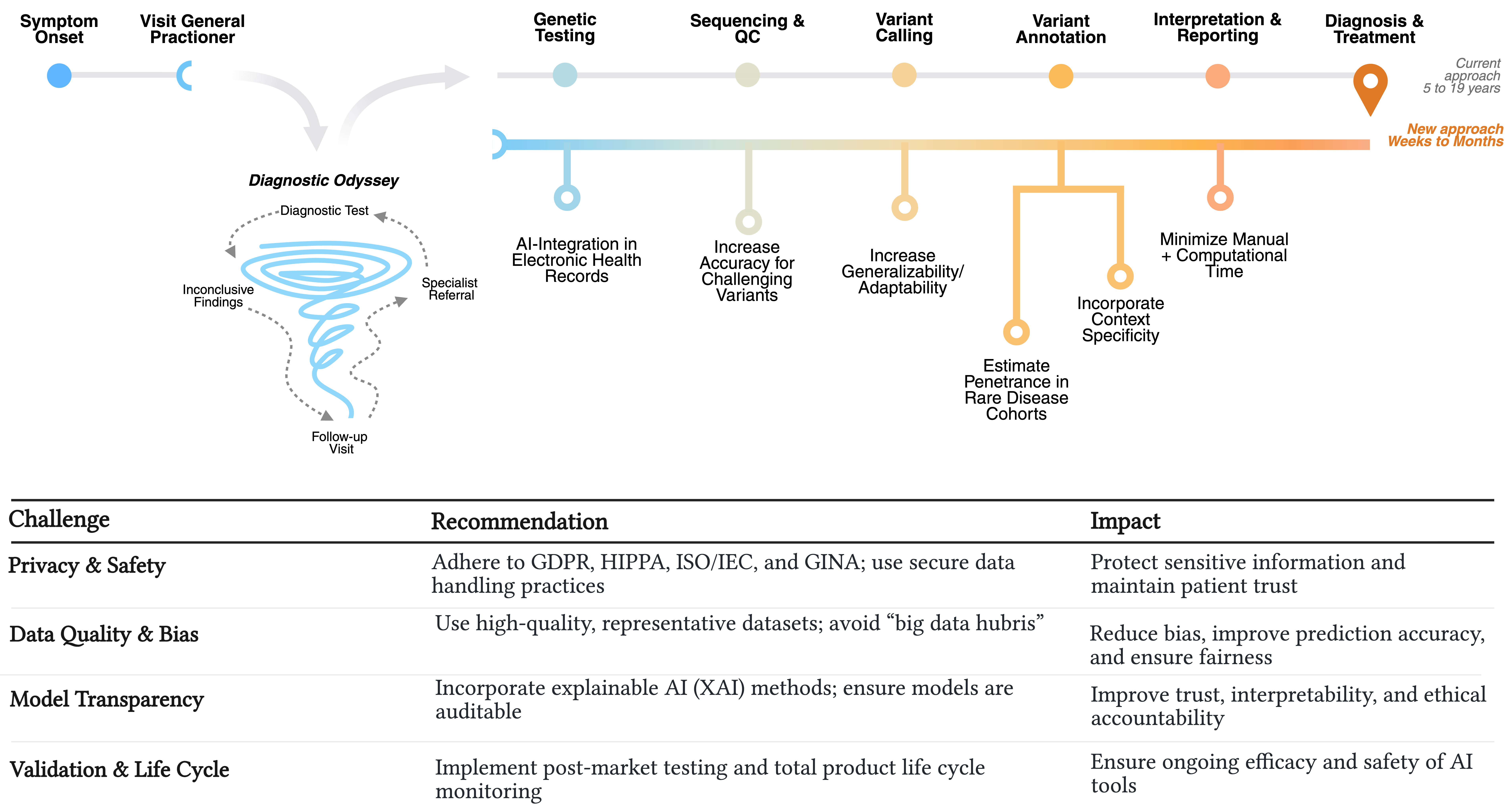
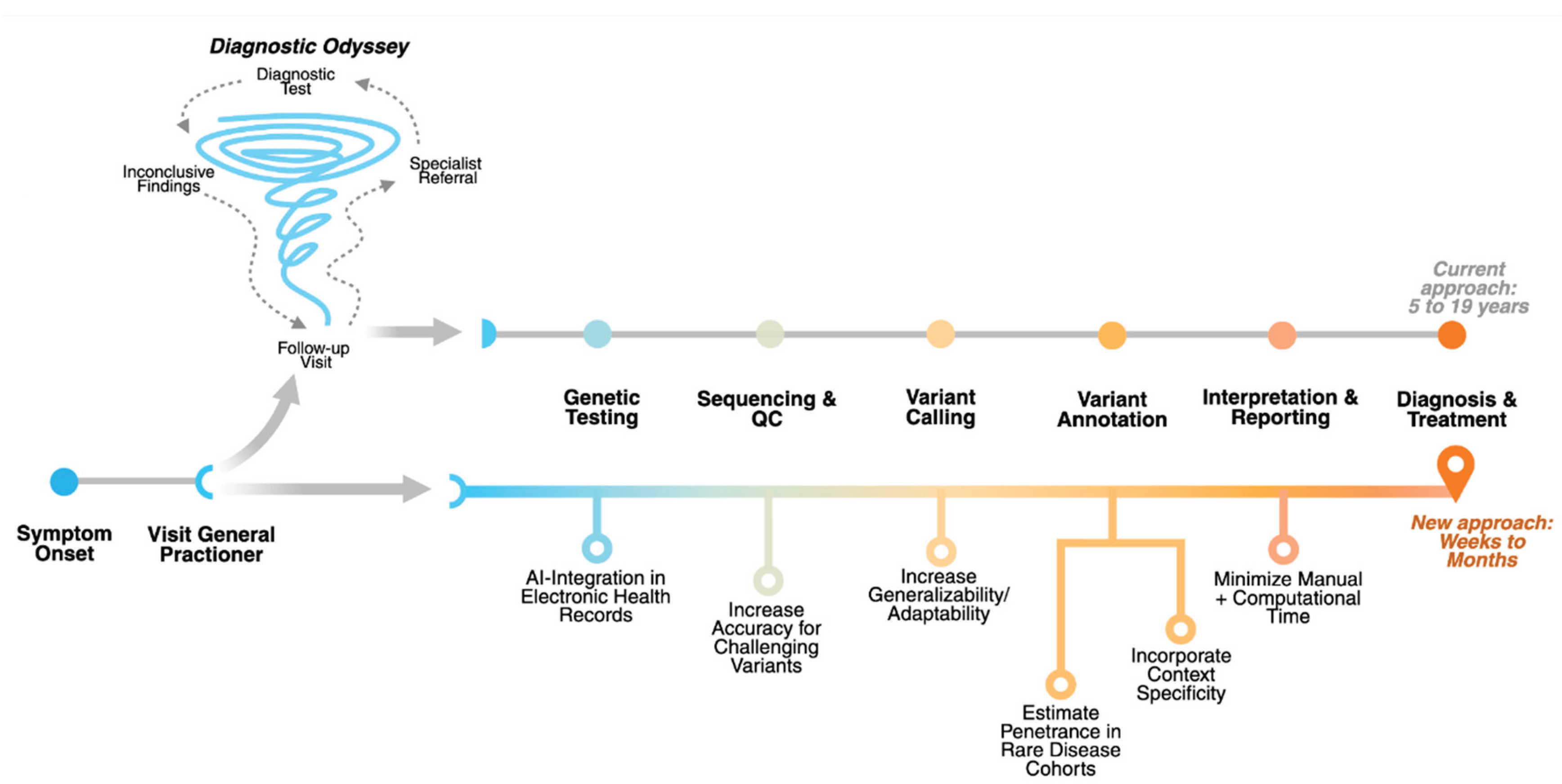
Approach to Variant Analysis
Challenges and Recommendations
Conclusions
Competing Interests
Author Contributions
Funding
Acknowledgments
References
- Abdelwahab, O; Belzile, F; Torkamaneh, D. Performance analysis of conventional and AI-based variant callers using short and long reads. BMC Bioinformatics 2023, 24, 472. [Google Scholar] [CrossRef] [PubMed]
- Abdelwahab, O; Torkamaneh, D. Artificial intelligence in variant calling: a review. Front Bioinform 2025, 5, 1574359. [Google Scholar] [CrossRef] [PubMed]
- Agaoglu, NB; Unal, B; Akgun Dogan, O. Consistency of variant interpretations among bioinformaticians and clinical geneticists in hereditary cancer panels. Eur J Hum Genet 2022, 30, 378–383. [Google Scholar] [CrossRef] [PubMed]
- Allot, A; Wei, C-H; Phan, L. Tracking genetic variants in the biomedical literature using LitVar 2.0. Nat Genet 2023, 55, 901–903. [Google Scholar] [CrossRef]
- Avsec, Ž; Agarwal, V; Visentin, D. Effective gene expression prediction from sequence by integrating long-range interactions. Nat Methods 2021, 18, 1196–1203. [Google Scholar] [CrossRef]
- Balmaña, J; Digiovanni, L; Gaddam, P. Conflicting interpretation of genetic variants and cancer risk by commercial laboratories as assessed by the Prospective Registry of Multiplex Testing. J Clin Oncol 2016, 34, 4071–4078. [Google Scholar] [CrossRef]
- Beutel, G; Geerits, E; Kielstein, JT. Artificial hallucination: GPT on LSD? Crit Care 2023, 27, 148. [Google Scholar] [CrossRef]
- Brand, F; Guski, J; Krawitz, P. Extending DeepTrio for sensitive detection of complex de novo mutation patterns. NAR Genom Bioinform 2024, 6, lqae013. [Google Scholar] [CrossRef]
- Brixi, G; Durrant, MG; Ku, J. Genome modelling and design across all domains of life with Evo 2; Nature, 2026; pp. 1–13. [Google Scholar] [CrossRef]
- Campeau, PM. An all-encompassing variant classification system proposed. Eur J Hum Genet 2022, 30, 139. [Google Scholar] [CrossRef]
- Center for Devices; Radiological Health. Good Machine Learning Practice for Medical Device Development: Guiding Principles. U.S. Food and Drug Administration, 2025a. Available online: https://www.fda.gov/medical-devices/software-medical-device-samd/good-machine-learning-practice-medical-device-development-guiding-principles (accessed on 7 Oct 2025).
- Center for Devices; Radiological Health. Predetermined Change Control Plans for Machine Learning-Enabled Medical Devices: Guiding Principles. U.S. Food and Drug Administration, 2025b. Available online: https://www.fda.gov/medical-devices/software-medical-device-samd/predetermined-change-control-plans-machine-learning-enabled-medical-devices-guiding-principles (accessed on 7 Oct 2025).
- Chen, X; Wang, L; You, M. Evaluating and enhancing large language models’ performance in domain-specific medicine: Development and usability study with DocOA. J Med Internet Res 2024, 26, e58158. [Google Scholar] [CrossRef]
- Daneshjou, R; Vodrahalli, K; Novoa, RA. Disparities in dermatology AI performance on a diverse, curated clinical image set. Sci Adv 2022, 8, eabq6147. [Google Scholar] [CrossRef]
- Dastin, J. Insight - Amazon scraps secret AI recruiting tool that showed bias against women. Reuters 2018. [Google Scholar]
- Delgado, J; de Manuel, A; Parra, I. Bias in algorithms of AI systems developed for COVID-19: A scoping review. J Bioeth Inq 2022, 19, 407–419. [Google Scholar] [CrossRef]
- Diaz, M; Johnson, I; Lazar, A. Addressing age-related bias in sentiment analysis. In Proceedings of the 2018 CHI Conference on Human Factors in Computing Systems, New York, NY, USA; ACM, 2018. [Google Scholar]
- Erikson, SL. Cell phones ≠ self and other problems with big data detection and containment during epidemics. Med Anthropol Q 2018, 32, 315–339. [Google Scholar] [CrossRef]
- European Union (2016) Regulation (EU) 2016/679 General Data Protection Regulation (GDPR). Official Journal of the European Union.
- Faye, F; Crocione, C; Anido de Peña, R. Time to diagnosis and determinants of diagnostic delays of people living with a rare disease: results of a Rare Barometer retrospective patient survey. Eur J Hum Genet 2024, 32, 1116–1126. [Google Scholar] [CrossRef] [PubMed]
- Fieldhouse, R. Too much social media gives AI chatbots “brain rot.”; Nature, 2025. [Google Scholar] [CrossRef]
- Fiorini, MR; Dilliott, AA; Farhan, SMK. Evaluating the utility of REVEL and CADD for interpreting variants in amyotrophic lateral sclerosis genes. Hum Mutat 2023, 2023, 8620557. [Google Scholar] [CrossRef] [PubMed]
- Forrest, IS; Vy, HMT; Rocheleau, G. Machine learning-based penetrance of genetic variants. Science 2025, 389, eadm7066. [Google Scholar] [CrossRef]
- Fowler, DM; Rehm, HL. Will variants of uncertain significance still exist in 2030? Am J Hum Genet 2024, 111, 5–10. [Google Scholar] [CrossRef]
- Freed, D; Pan, R; Chen, H. DNAscope: High accuracy small variant calling using machine learning. bioRxiv 2022. [Google Scholar] [CrossRef]
- Gao, H; Hamp, T; Ede, J. The landscape of tolerated genetic variation in humans and primates. Science 2023, 380, eabn8153. [Google Scholar] [CrossRef] [PubMed]
- Gosiewska, A; Kozak, A; Biecek, P. Simpler is better: Lifting interpretability-performance trade-off via automated feature engineering. Decis Support Syst 2021, 150, 113556. [Google Scholar] [CrossRef]
- Gurovich, Y; Hanani, Y; Bar, O. Identifying facial phenotypes of genetic disorders using deep learning. Nat Med 2019, 25, 60–64. [Google Scholar] [CrossRef] [PubMed]
- Hajek, C; Hutchinson, AM; Galbraith, LN. Improved provider preparedness through an 8-part genetics and genomic education program. Genet Med 2022, 24, 214–224. [Google Scholar] [CrossRef]
- Houge, G; Bratland, E; Aukrust, I. Comparison of the ABC and ACMG systems for variant classification. Eur J Hum Genet 2024, 32, 858–863. [Google Scholar] [CrossRef]
- Houge, G; Laner, A; Cirak, S. Stepwise ABC system for classification of any type of genetic variant. Eur J Hum Genet 2022, 30, 150–159. [Google Scholar] [CrossRef]
- Hu, J; Freed, D; Feng, H. Accelerated, Accurate, Hybrid Short and Long Reads Alignment and Variant Calling. Bioinformatics 2025. [Google Scholar]
- Hu, X; Feng, C; Zhou, Y. DeepTrio: a ternary prediction system for protein-protein interaction using mask multiple parallel convolutional neural networks. Bioinformatics 2022, 38, 694–702. [Google Scholar] [CrossRef]
- International Organization for Standardization; International Electrotechnical Commission. ISO/IEC 27001:2022 Information security, cybersecurity and privacy protection — Information security management systems — Requirements. ISO: Geneva, 2022.
- Ioannidis, NM; Rothstein, JH; Pejaver, V. REVEL: An ensemble method for predicting the pathogenicity of rare missense variants. Am J Hum Genet 2016, 99, 877–885. [Google Scholar] [CrossRef]
- Jaganathan, K; Kyriazopoulou Panagiotopoulou, S; McRae, JF. Predicting splicing from primary sequence with deep learning. Cell 2019, 176, 535–548.e24. [Google Scholar] [CrossRef]
- Janiesch, C; Zschech, P; Heinrich, K. Machine learning and deep learning. Electron Mark 2021, 31, 685–695. [Google Scholar] [CrossRef]
- Jin, Q; Wang, Z; Floudas, CS. Matching patients to clinical trials with large language models. Nat Commun 2024, 15, 9074. [Google Scholar] [CrossRef] [PubMed]
- Kennedy, B; Tyson, A. Americans’ Trust in Scientists, Positive Views of Science Continue to Decline. Pew Research Center. 2023. Available online: https://www.pewresearch.org/science/2023/11/14/confidence-in-scientists-medical-scientists-and-other-groups-and-institutions-in-society/ (accessed on 6 Nov 2025).
- Kessler, MD; Yerges-Armstrong, L; Taub, MA. Challenges and disparities in the application of personalized genomic medicine to populations with African ancestry. Nat Commun 2016, 7, 12521. [Google Scholar] [CrossRef] [PubMed]
- Kim, YE; Ki, CS; Jang, MA. Challenges and considerations in sequence variant interpretation for Mendelian disorders. Ann Lab Med 2019, 39, 421–429. [Google Scholar] [CrossRef]
- Kircher, M; Witten, DM; Jain, P. A general framework for estimating the relative pathogenicity of human genetic variants. Nat Genet 2014, 46, 310–315. [Google Scholar] [CrossRef]
- Kneifati-Hayek, JZ; Zachariah, T; Ahn, W. Bridging the gap in genomic implementation: Identifying user needs for precision nephrology. Kidney Int Rep 2024, 9, 2420–2431. [Google Scholar] [CrossRef]
- Koteluk, O; Wartecki, A; Mazurek, S. How do machines learn? Artificial intelligence as a New Era in medicine. J Pers Med 2021, 11, 32. [Google Scholar] [CrossRef]
- Wetterstrand, Kris A., MS. DNA Sequencing Costs: Data. Genome.gov. 2019. Available online: https://www.genome.gov/about-genomics/fact-sheets/DNA-Sequencing-Costs-Data (accessed on 5 Nov 2025).
- Larson, J; Angwin, J; Kirchner, L; Mattu, S. How We Analyzed the COMPAS Recidivism Algorithm. #creator. 2016. Available online: https://www.propublica.org/article/how-we-analyzed-the-compas-recidivism-algorithm (accessed on 7 Oct 2025).
- Lazer, D; Kennedy, R; King, G; Vespignani, A. Big data. The parable of Google Flu: traps in big data analysis. Science 2014, 343, 1203–1205. [Google Scholar] [CrossRef]
- Lee, J; Cha, H; Hwangbo, Y; Cheon, W. Enhancing large language model reliability: Minimizing hallucinations with dual retrieval-augmented generation based on the latest diabetes guidelines. J Pers Med 2024, 14, 1131. [Google Scholar] [CrossRef]
- Leiser, F; Guse, R; Sunyaev, A. Large language model architectures in health care: Scoping review of research perspectives. J Med Internet Res 2025, 27, e70315. [Google Scholar] [CrossRef]
- Lewin Group T. The cost of delayed diagnosis in rare disease: a health economic study. In EveryLife Foundation for Rare Diseases; 2023. [Google Scholar]
- Lin, L; Pan, H; Qi, Y. Reasons and Resolutions for Inconsistent Variant Interpretation. Human Mutation 2023, 2023, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Lundberg, S; Lee, S-I. A unified approach to interpreting model predictions. arXiv [cs.AI 2017. [Google Scholar]
- Mao, D; Liu, C; Wang, L. AI-MARRVEL - A knowledge-driven AI system for diagnosing Mendelian disorders. NEJM AI 2024, 1. [Google Scholar] [CrossRef] [PubMed]
- May, W; Berghoff, C; Böddinghaus, J. May 2022 Towards Auditable AI Systems From Principles to Practice. 2022. Available online: https://www.semanticscholar.org/paper/Whitepaper-%7C-May-2022-Towards-Auditable-AI-Systems-May-Berghoff/2af03e944a0c0e905ccf1b24e795f53cc780ea36#related-papers (accessed on 22 Oct 2025).
- McCoy, LG; Bihorac, A; Celi, LA. Building health systems capable of leveraging AI: applying Paul Farmer’s 5S framework for equitable global health. BMC Glob Public Health 2025, 3, 39. [Google Scholar] [CrossRef]
- McNeill, A. A new system for variant classification? Eur J Hum Genet 2022, 30, 137–138. [Google Scholar] [CrossRef]
- Meng, L; Attali, R; Talmy, T. Evaluation of an automated genome interpretation model for rare disease routinely used in a clinical genetic laboratory. Genet Med 2023, 25, 100830. [Google Scholar] [CrossRef]
- Mercurio, SA; Chunn, LM; Khursigara, G. ENPP1 deficiency: A clinical update on the relevance of individual variants using a locus-specific patient database. Hum Mutat 2022, 43, 1673–1705. [Google Scholar] [CrossRef]
- Nagy, D; Pennetta, V; Rodger, G. Nanopore long-read-only genome assembly of clinical Enterobacterales isolates is complete and accurate. Microb Genom 2026, 12, 001631. [Google Scholar] [CrossRef]
- Nakayama, LF; Kras, A; Ribeiro, LZ. Global disparity bias in ophthalmology artificial intelligence applications. BMJ Health Care Inform 2022, 29, e100470. [Google Scholar] [CrossRef]
- Nichols, JA; Herbert Chan, HW; Baker, MAB. Machine learning: applications of artificial intelligence to imaging and diagnosis. Biophys Rev 2019, 11, 111–118. [Google Scholar] [CrossRef]
- Obermeyer, Z; Powers, B; Vogeli, C; Mullainathan, S. Dissecting racial bias in an algorithm used to manage the health of populations. Science 2019, 366, 447–453. [Google Scholar] [CrossRef] [PubMed]
- Office for Civil Rights (OCR) Summary of the HIPAA Privacy Rule. Available online: https://www.hhs.gov/hipaa/for-professionals/privacy/laws-regulations/index.html (accessed on 16 Mar 2022).
- Pasquier, L; Minguet, G; Moisdon-Chataigner, S. How do non-geneticist physicians deal with genetic tests? A qualitative analysis. Eur J Hum Genet 2022, 30, 320–331. [Google Scholar] [CrossRef] [PubMed]
- Peabody, J; DeMaria, L; Tamandong-LaChica, D. Low rates of genetic testing in children with developmental delays, intellectual disability, and autism spectrum disorders. Glob Pediatr Health 2015, 2, 2333794X15623717. [Google Scholar] [CrossRef] [PubMed]
- Peng, M; Guo, X; Chen, X. LC-LLM: Explainable lane-change intention and trajectory predictions with Large Language Models. Communications in Transportation Research 2025, 5, 100170. [Google Scholar] [CrossRef]
- Phillips, C; Parkinson, A; Namsrai, T. Time to diagnosis for a rare disease: managing medical uncertainty. A qualitative study. Orphanet J Rare Dis 2024, 19, 297. [Google Scholar] [CrossRef]
- Poplin, R; Chang, P-C; Alexander, D. A universal SNP and small-indel variant caller using deep neural networks. Nat Biotechnol 2018, 36, 983–987. [Google Scholar] [CrossRef]
- Poushter, J; Fagan, M; Corichi, M. (2025) How People Around the World View AI. Pew Research Center. Available online: https://www.pewresearch.org/global/2025/10/15/how-people-around-the-world-view-ai/ (accessed on 6 Nov 2025).
- Ramachandran, A; Lumetta, SS; Klee, EW; Chen, D. HELLO: improved neural network architectures and methodologies for small variant calling. BMC Bioinformatics 2021, 22, 404. [Google Scholar] [CrossRef]
- Rasouly, HM; Balderes, O; Marasa, M. The effect of genetic education on the referral of patients to genetic evaluation: Findings from a national survey of nephrologists. Genet Med 2023, 25, 100814. [Google Scholar] [CrossRef]
- Richards, S; Aziz, N; Bale, S. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 2015, 17, 405–424. [Google Scholar] [CrossRef]
- Ross, C; Swetlitz, I. IBM’s Watson supercomputer recommended “unsafe and incorrect” cancer treatments, internal documents show. STAT. 2018. Available online: https://www.statnews.com/2018/07/25/ibm-watson-recommended-unsafe-incorrect-treatments/ (accessed on 7 Oct 2025).
- Ruiz, J. Machine learning and the right to explanation in GDPR. Open Rights Group. Available online: https://www.openrightsgroup.org/blog/machine-learning-and-the-right-to-explanation-in-gdpr/ (accessed on 7 Oct 2025).
- Russell, S; Norvig, P. Artificial intelligence: A modern approach, global edition, 4th edn; Pearson Education: London, England, 2021. [Google Scholar]
- Sina Gräupner, O; Pawlaszczyk, D; Hummert, C. Basics of Auditable AI Systems. In 2023 Congress in Computer Science, Computer Engineering, & Applied Computing (CSCE); IEEE, 2023; pp. pp 2355–2362. [Google Scholar]
- Sokhansanj, BA; Rosen, GL. Regulating genome language models: navigating policy challenges at the intersection of AI and genetics. Hum Genet 2025, 144, 949–970. [Google Scholar] [CrossRef]
- Su, J; Zheng, Z; Ahmed, SS. Clair3-trio: high-performance Nanopore long-read variant calling in family trios with trio-to-trio deep neural networks. Brief Bioinform 2022, 23, bbac301. [Google Scholar] [CrossRef] [PubMed]
- Tagliafico, E; Bernardis, I; Grasso, M. Workload measurement for molecular genetics laboratory: A survey study. PLoS One 2018, 13, e0206855. [Google Scholar] [CrossRef] [PubMed]
- Tomašev, N; Cornebise, J; Hutter, F. AI for social good: unlocking the opportunity for positive impact. Nat Commun 2020, 11, 2468. [Google Scholar] [CrossRef] [PubMed]
- Tordai, H; Torres, O; Csepi, M. Analysis of AlphaMissense data in different protein groups and structural context. Sci Data 2024, 11, 495. [Google Scholar] [CrossRef]
- Wan, EL; Elkaim, Y; Gao, W; Yoon, R. Zebras among us: Advocating for the 30 million Americans living with rare disease. Med Sci Educ 2023, 33, 1239–1242. [Google Scholar] [CrossRef]
- Xing, S; Hong, J; Wang, Y. LLMs can get “Brain Rot”! arXiv [cs.CL] 2025. [Google Scholar]
- Yang, X; Chen, A; PourNejatian, N. A large language model for electronic health records. NPJ Digit Med 2022, 5, 194. [Google Scholar] [CrossRef]
- Ye, J; Woods, D; Jordan, N; Starren, J. The role of artificial intelligence for the application of integrating electronic health records and patient-generated data in clinical decision support. AMIA Summits Transl Sci Proc 2024, 2024, 459–467. [Google Scholar]
- Zawar, A; Manoj, G; Nair, PP. Variants of uncertain significance: At the crux of diagnostic odyssey. Gene 2025, 962, 149587. [Google Scholar] [CrossRef]
- Zhan, H; Zhang, Z. DYNA: Disease-specific language model for variant pathogenicity. arXiv [q-bio.GN 2024. [Google Scholar]
- Zhao, H; Chen, H; Yang, F. Explainability for large language models: A survey. ACM Trans Intell Syst Technol 2024, 15, 1–38. [Google Scholar] [CrossRef]
- Zukin, E; Culver, JO; Liu, Y. Clinical implications of conflicting variant interpretations in the cancer genetics clinic. Genet Med 2023, 25, 100837. [Google Scholar] [CrossRef]
- Machine Learning vs Deep Learning vs LLMs vs GenAI: Explained and How are they Different from Each Other? Cloud4C. 2024. Available online: https://www.cloud4c.com/blogs/genai-vs-machine-learning-vs-deep-learning-vs-llms (accessed on 14 Nov 2025).
- Franklin - Bioinformatics Software. Bioinformatics Software QIAGEN Digital Insights. 2025a. Available online: https://digitalinsights.qiagen.com/franklin/ (accessed on 14 Oct 2025).
- Enhancing Rare Disease Diagnostics: Updated Performance Evaluation of the AION AI-Driven Variant Interpretation Platform. 2025b. Available online: https://www.nostos-genomics.com. (accessed on 30 Mar 2026).
- Genetic Information Nondiscrimination Act of 2008 (GINA). 2008.
- Annual Report 2023. Partners in Health 2023.
- (2025c) AI & Variant Interpretation: From Data Tsunami to Diagnostic Clarity. Available online: https://www.nostos-genomics.com. (accessed on 7 Oct 2025).
| Challenge | Recommendation | Impact |
|---|---|---|
| Privacy & Safety | Adhere to *GDPR, *HIPPA, *ISO/IEC, and *GINA; use secure data handling practices | Protect sensitive information and maintain patient trust |
| Data Quality & Bias | Use high-quality, representative datasets; avoid “big data hubris” | Reduce bias, improve prediction accuracy, and ensure fairness |
| Model Transparency | Incorporate explainable AI (XAI) methods; ensure models are auditable | Improve trust, interpretability, and ethical accountability |
| Validation & Life Cycle | Implement post-market testing and total product life cycle monitoring | Ensure ongoing efficacy and safety of AI tools |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




