Submitted:
07 March 2026
Posted:
09 March 2026
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
1.1. Contributions of This Paper
- 1.
- Black cotton as a new design target. We propose, for the first time, a dual-pathway genetic strategy to produce near-black cotton fiber by combining (a) eumelanin biosynthesis via heterologous expression of tyrosinase (TYR) from Streptomyces antibioticus and polyphenol oxidase (PPO) from Vitis vinifera, with (b) hyperaccumulation and laccase-catalyzed oxidative polymerization of proanthocyanidins in the fiber lumen. We specify the complete gene list, donor organisms, construct architecture (six transcription units in a Level 2 Golden Gate binary vector), and fiber-specific promoter assignments for this new target (Section 3.5, Table 2, Figure 3).
- 2.
- Unified six-color design framework. We present the first single paper that systematically covers the genetic design procedures for all six cotton fiber colors — white (reference), brown, green, pink, blue, and black — within one integrated biochemical pathway map (Figure 2), enabling direct comparison of gene requirements, donor organisms, and engineering complexity across the full color spectrum.
- 3.
- Reproducible bench-level laboratory protocols. Unlike previous reviews that describe strategies at a conceptual level, we provide step-by-step, experimentally reproducible protocols with specific reagents, concentrations, media compositions, instrument settings, and quantitative decision gates (Protocols 1.1–5.1). These cover: RNA-seq-based pathway profiling, codon-optimized gene synthesis, Golden Gate multigene assembly (Level 0 → Level 1 → Level 2), Agrobacterium-mediated transformation of cotton cv. Coker 312, tiered molecular screening (PCR, Southern blot, RT-qPCR, LC-MS/MS with diagnostic m/z values for each pigment class), and HVI fiber quality evaluation against defined threshold and target values.
- 4.
- Complete 15-step lab-to-field translational pipeline. We map the entire journey from T0 transgenic plant to commercial farmer deployment through 15 explicit stages spanning five phases — Laboratory, Greenhouse, Contained Field Trials, Regulatory, and Commercial — including protocols for confined field trial design (RCBD, biosafety monitoring), multi-environment G×E analysis (AMMI/GGE biplot), regulatory dossier content (molecular characterization, substantial equivalence, toxicity, allergenicity, environmental risk), seed multiplication chain (breeder → foundation → certified seed), and farmer extension with insect resistance management (Figure 4, Protocols 6.1–8.1). This operational-level translational detail has not been previously consolidated in a single colored-cotton publication.
2. Bt-Cotton: Mechanism, Generations, and Global Impact
2.1. Mechanism of Action
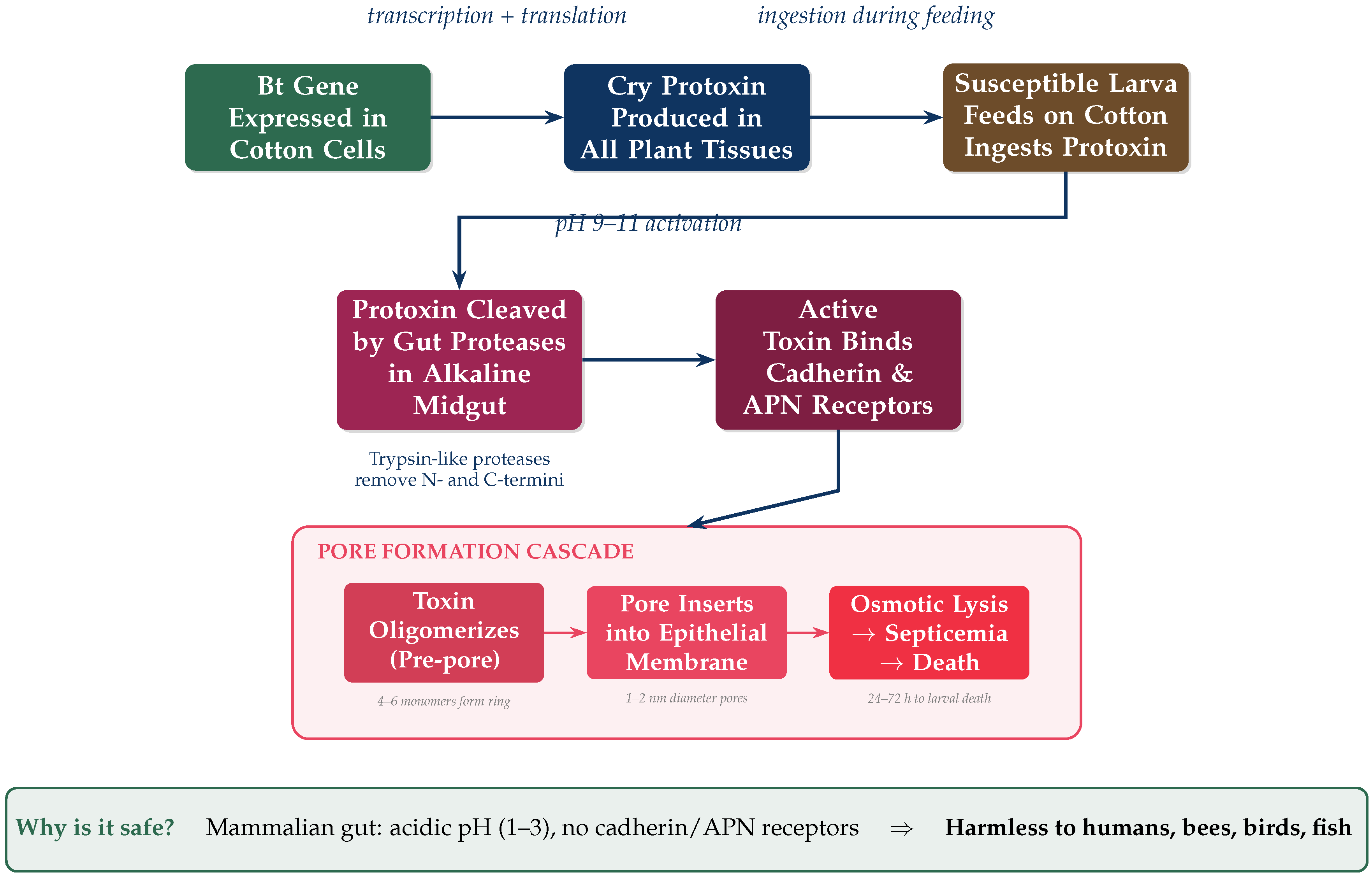
2.2. Generations of Bt Genes
| Generation | Gene(s) | Protein(s) | Target Pests | Year |
|---|---|---|---|---|
| Bt-I (Bollgard I) | cry1Ac | Cry1Ac | H. armigera, H. zea | 1996 |
| Bt-II (Bollgard II) | cry1Ac + cry2Ab | Cry1Ac + Cry2Ab | Broader Heliothinae; delays resistance | 2006 |
| Bt-III (TwinLink+) | cry1Ab + cry2Ae + vip3Aa | Cry1Ab + Cry2Ae + Vip3Aa | Very broad; multi-mode action | 2019 |
| Stacked events | cry1F + cry1Ac + HT genes | Cry1F + Cry1Ac + herbicide tol. | Bollworms + herbicide tolerance | Various |
2.3. Global Impact
3. Biochemical Basis of Cotton Fiber Pigmentation
3.1. Brown: Condensed Tannins (Proanthocyanidins)
3.2. Green: Suberin–Caffeic Acid Complexes
3.3. Pink / Red: Cyanidin-Type Anthocyanins
3.4. Blue / Purple: Delphinidin-Type Anthocyanins
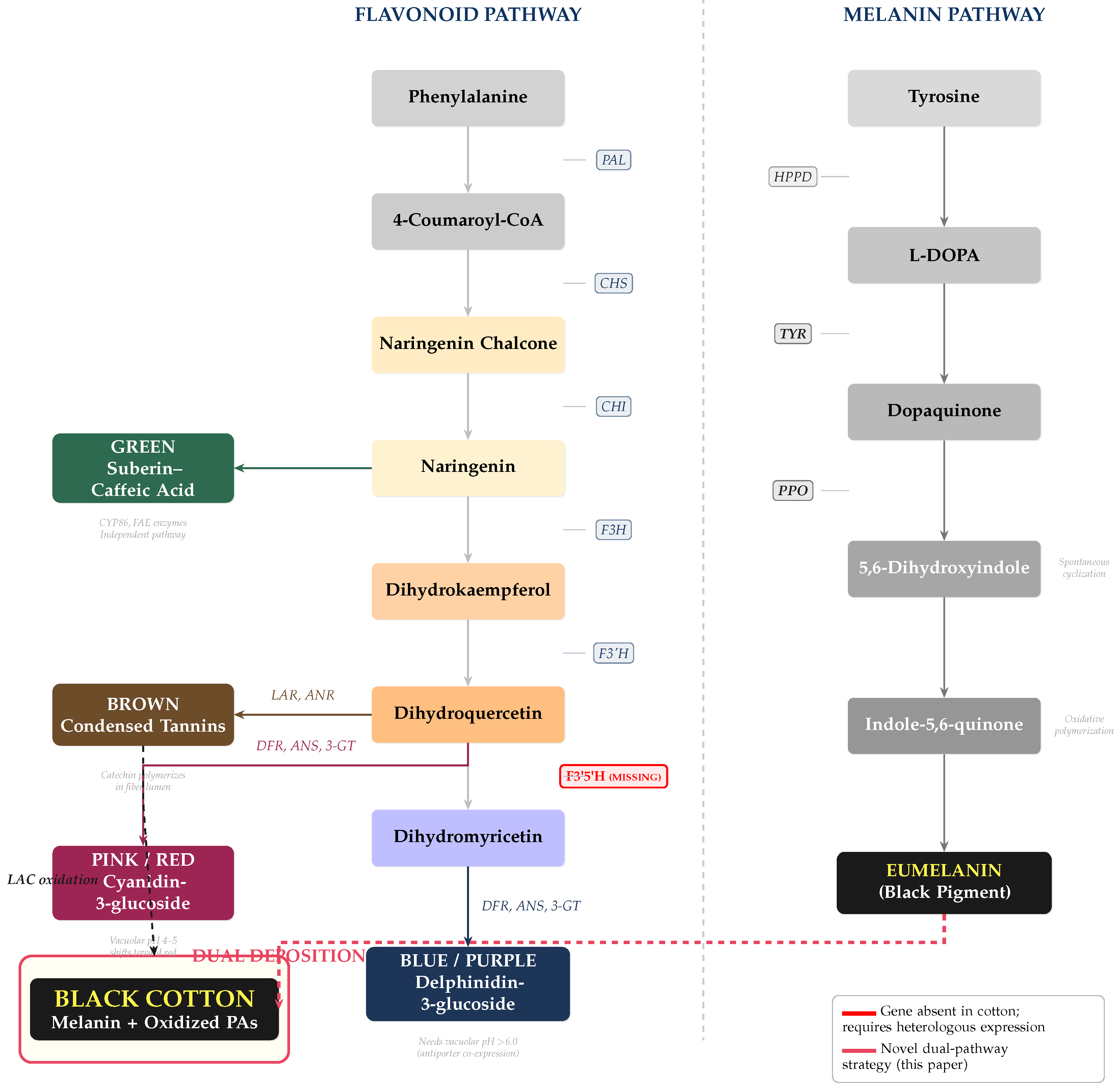
3.5. Black: Melanin + Oxidized Proanthocyanidin Hyperaccumulation

- Tyrosinase gene (TYR) from Streptomyces antibioticus (bacterial tyrosinase with high activity and thermal stability) or Agaricus bisporus (mushroom tyrosinase).
- Polyphenol oxidase (PPO) from Vitis vinifera (grape) or cotton’s own GhPPO genes, overexpressed under a fiber-specific promoter to catalyze oxidative browning of phenolic substrates toward melanin precursors.
- 4-hydroxyphenylpyruvate dioxygenase (HPPD) for enhanced tyrosine pool in fiber cells.
| Pathway | Gene | Source Organism | Function |
|---|---|---|---|
| Melanin | TYR | S. antibioticus | Tyrosinase: Tyr → DOPA → dopaquinone |
| Melanin | PPO | V. vinifera / GhPPO | Polyphenol oxidase: broad phenolic oxidation |
| Melanin | HPPD | A. thaliana | Boosts tyrosine precursor pool |
| PA accum. | TT2+TT8+TTG1 | A. thaliana / Medicago | MBW complex: activates PA biosynthesis |
| PA oxidation | LAC | T. versicolor | Laccase: oxidative tannin polymerization |
3.6. Summary: Six Target Colors
| Color | Pigment Class | Key Genes | Natural? | Status |
|---|---|---|---|---|
| White | None (bleached) | — | Yes | Global standard |
| Brown | Condensed tannins | LAR, ANR, TT2/TT8 | Yes | Commercial (niche) |
| Green | Suberin–caffeic acid | CYP86, FAE | Yes | Commercial (niche) |
| Pink | Cyanidin anthocyanins | DFR, ANS, 3-GT | Partial | Breeding / R&D |
| Blue | Delphinidin anthocyanins | F3’5’H, DFR, ANS, pH mod. | No | Transgenic R&D |
| Black | Melanin + oxidized PAs | TYR, PPO, LAC, MBW | No | Proposed (this paper) |
4. Environmental and Health Rationale
4.1. Environmental Impact of Textile Dyeing
| Parameter | White + Dyeing | Colored Cotton |
|---|---|---|
| Water use for coloring | 100–150 L/kg fabric | 0 L |
| Chemical discharge | Azo dyes, heavy metals, formaldehyde | None |
| Energy (coloring step) | High (90–100°C dye baths) | None |
| Worker health risk | Dermatitis, respiratory illness, cancer | Minimal |
| Color durability | Fades with washing | Stable or deepens |
| CO2 footprint (coloring) | 5–8 kg CO2/kg fabric | ∼0 kg |
4.2. Health and Hypoallergenic Properties
5. Genetic Design Procedures: Rigorous Laboratory Protocols
5.1. Step 1: Target Pathway Identification
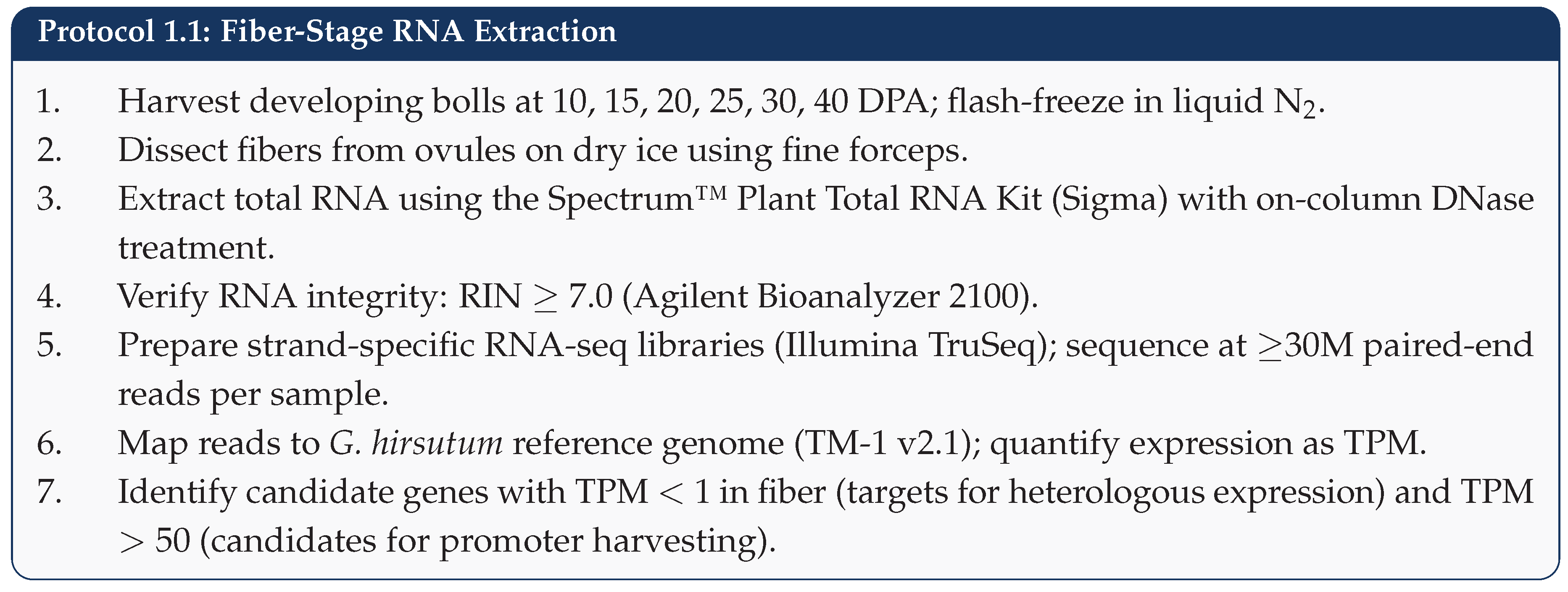

5.2. Step 2: Gene Sourcing and Synthesis


| Color | Gene(s) Needed | Donor Organism | Function |
|---|---|---|---|
| Blue | F3’5’H; pH modifier | Clitoria ternatea; Ipomoea nil | 5’-hydroxylation; vacuolar alkalinization |
| Pink | High-expression DFR, ANS | Antirrhinum majus; Gerbera hybrida | Anthocyanidin synthesis |
| Deep brown | TT2, TT8, TTG1 | Arabidopsis thaliana; Medicago truncatula | MBW transcription factor complex |
| Green | CYP86A; FAE | Cotton native genes (promoter engineering) | Suberin biosynthesis enhancement |
| Black | TYR, PPO, HPPD, MBW complex, LAC | S. antibioticus; V. vinifera; A. thaliana; T. versicolor | Melanin + hyperaccumulated oxidized PAs |
5.3. Step 3: Gene Cassette Construction


5.4. Step 4: Cotton Transformation
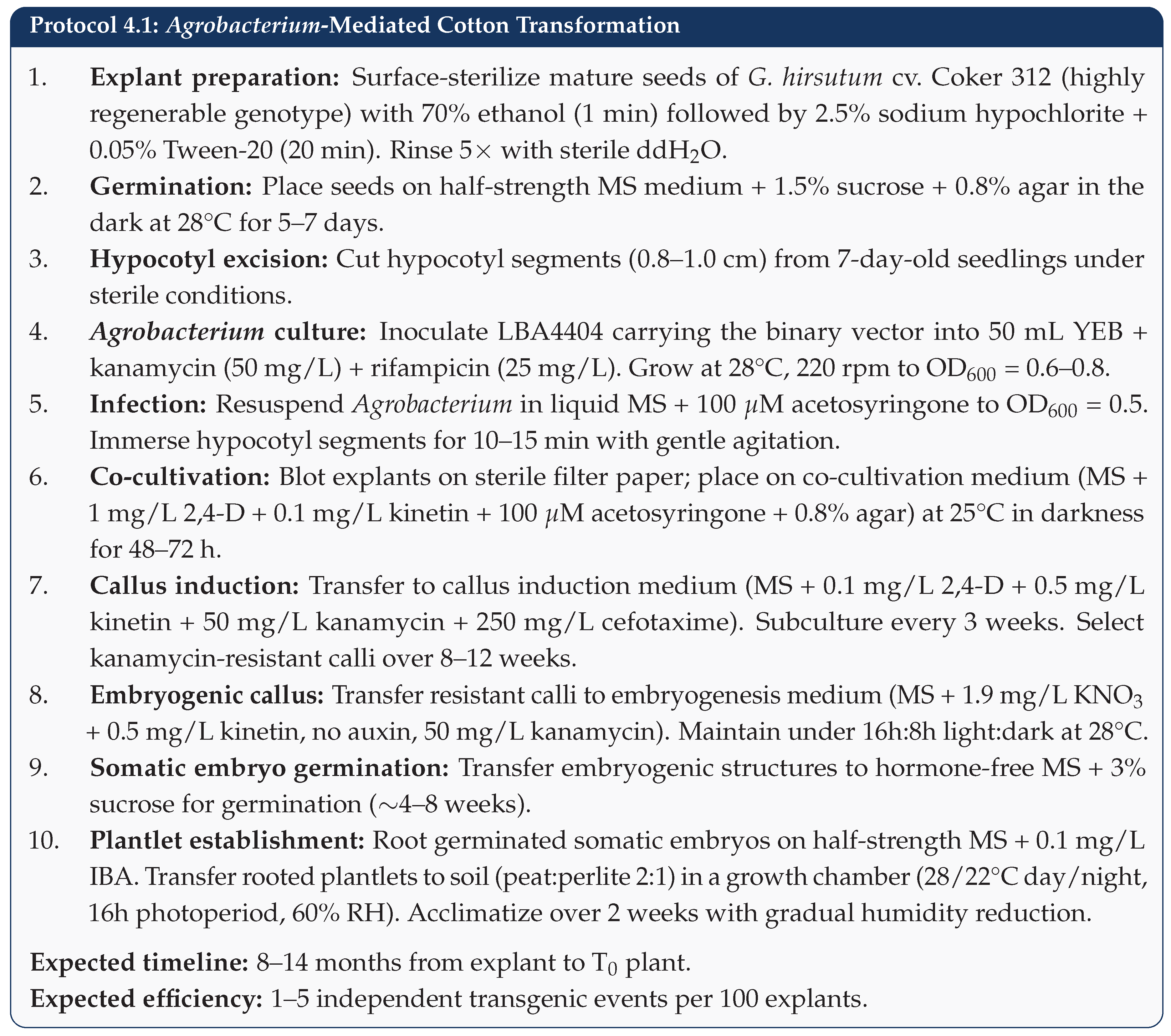

5.5. Step 5: Molecular Screening and Selection
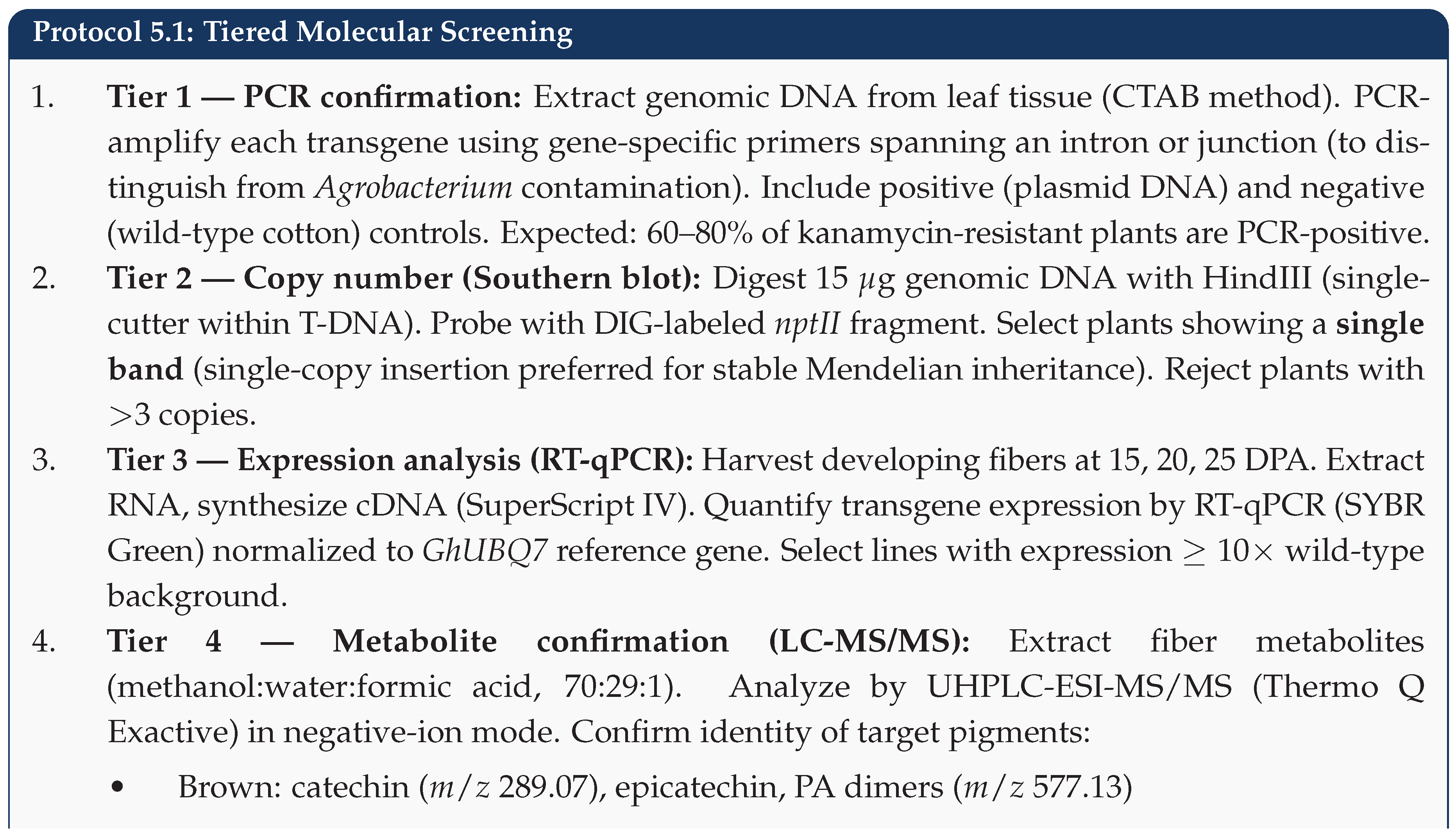
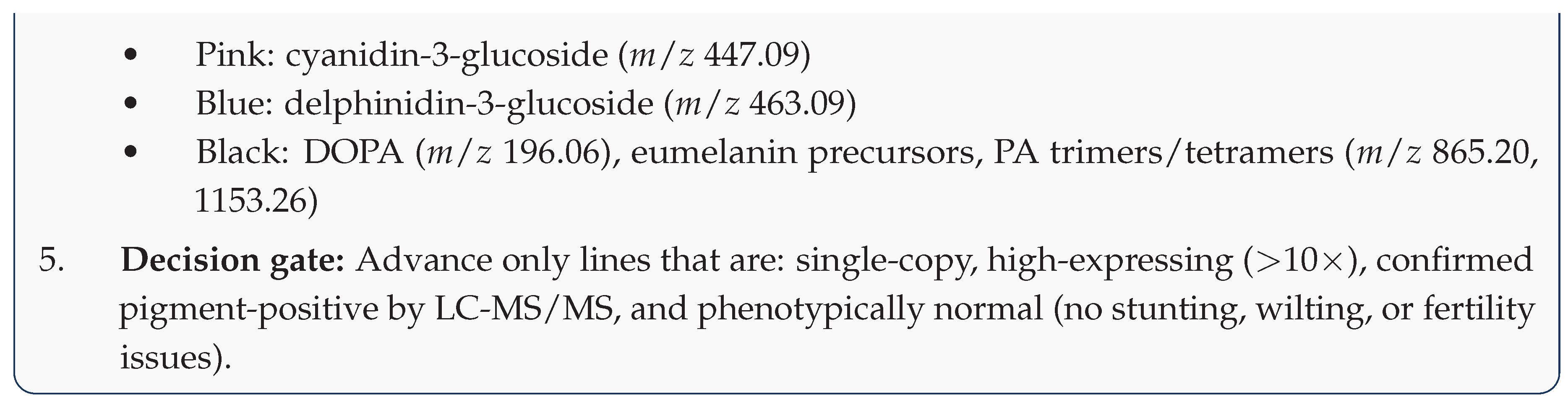


5.6. Step 6: Fiber Quality Evaluation
| Parameter | Method / Unit | Minimum | Target |
|---|---|---|---|
| Upper half mean length (UHML) | HVI / mm | 25 | >28 |
| Fiber strength (tenacity) | HVI / g/tex | 20 | >26 |
| Micronaire (fineness) | HVI / Mic units | 3.5–4.9 | 3.8–4.5 |
| Uniformity index | HVI / % | >80 | >83 |
| Color intensity | Spectrophotometer (L*a*b*) | Visually distinct | Saturated, uniform |
| Wash fastness | ISO 105-C06 (grade 1–5) | Grade 3 | Grade 4–5 |
| Light fastness | ISO 105-B02 (grade 1–8) | Grade 4 | Grade 6+ |
6. Lab to Field: The Complete Translational Pipeline
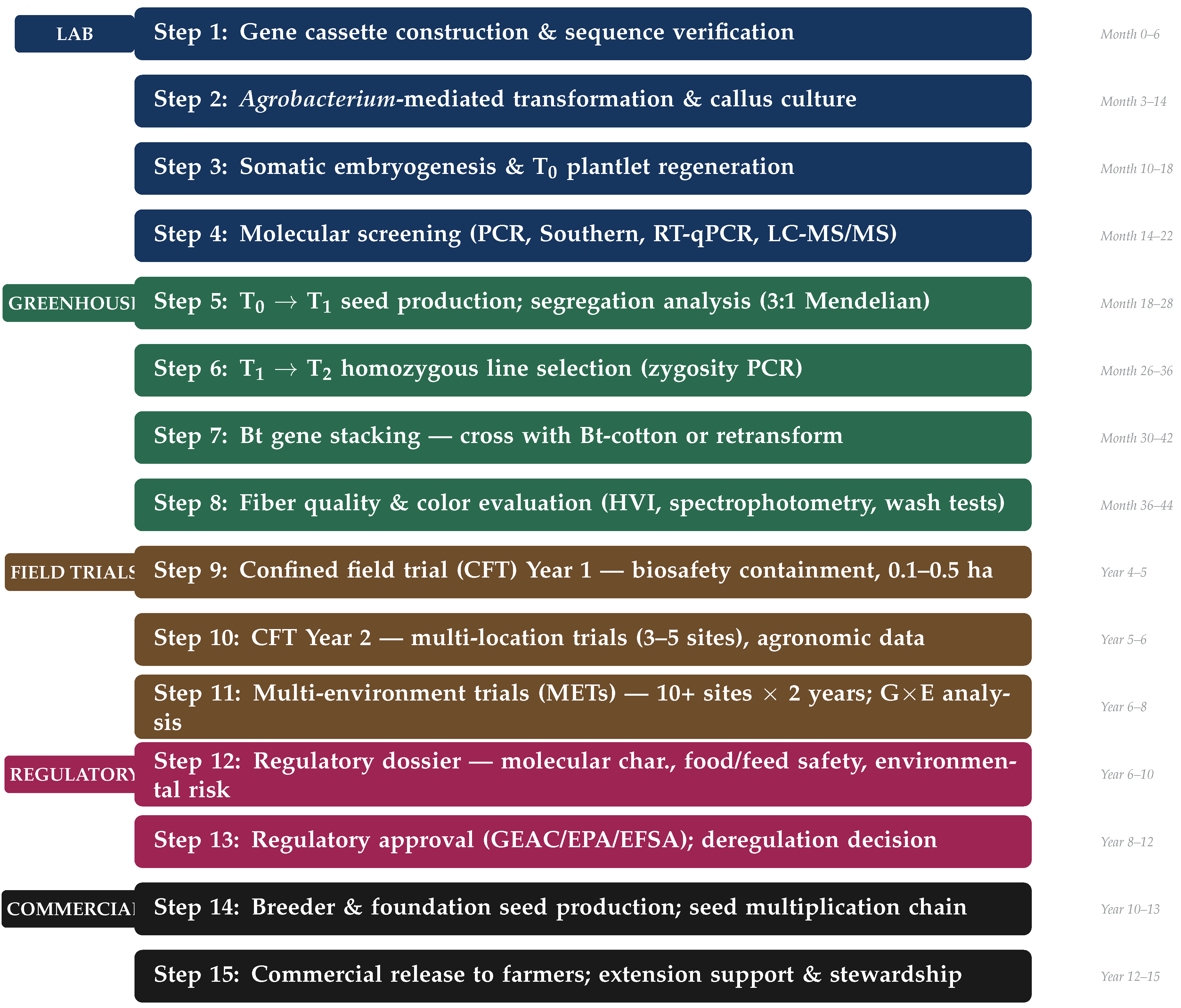


6.1. Phase I: Laboratory (Steps 1–4)
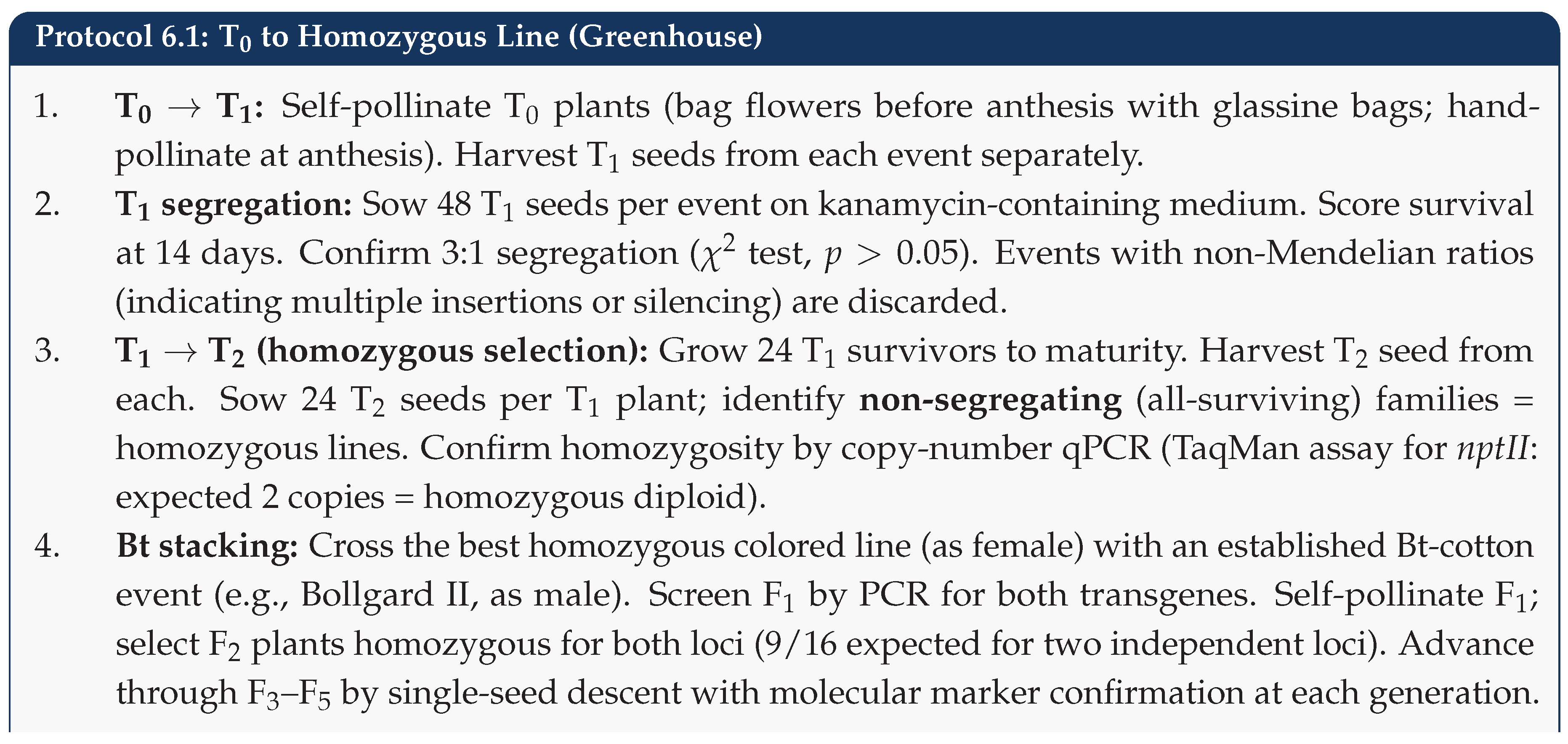
6.2. Phase II: Greenhouse Advancement (Steps 5–8)


6.3. Phase III: Contained and Multi-Environment Field Trials (Steps 9–11)

6.4. Phase IV: Regulatory Approval (Steps 12–13)
- 1.
- Molecular characterization: T-DNA insertion site (flanking sequence analysis by genome walking or WGS), copy number (Southern blot, ddPCR), absence of backbone integration, expression levels across tissues and developmental stages, transgene stability across ≥5 generations.
- 2.
- Compositional analysis: Proximate composition of seed (protein, fat, fiber, ash, moisture), fatty acid profile, amino acid profile, anti-nutritional factors (gossypol, cyclopropenoid fatty acids), compared to conventional counterpart (substantial equivalence).
- 3.
- Toxicity and allergenicity: Bioinformatic analysis (no homology to known allergens via AllergenOnline and FARRP databases), acute oral toxicity study (mouse, single dose), 90-day subchronic feeding study (rat) with transgenic cottonseed.
- 4.
- Environmental risk assessment: Gene flow analysis, non-target organism effects (beneficial insects, soil organisms), weediness potential, impact on biodiversity.
- 5.
- Fiber safety: Dermal sensitization testing (guinea pig or in-vitro), cytotoxicity of fiber extracts (ISO 10993-5).
6.5. Phase V: Seed Multiplication and Commercial Release (Steps 14–15)
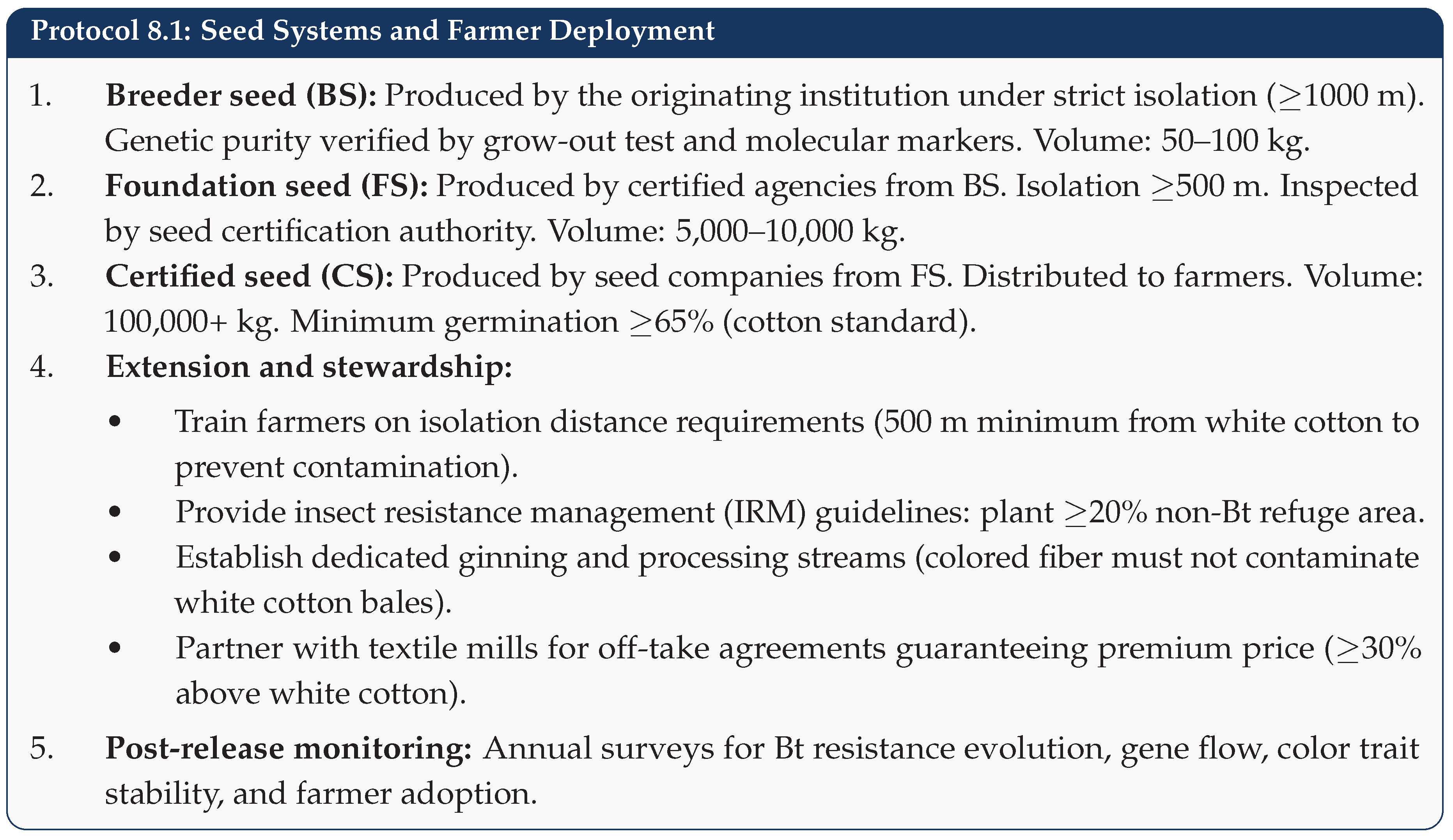

7. Breeding Techniques for Colored Cotton
7.1. Conventional Hybridization and Pedigree Selection
- 1.
- Year 1: Cross colored donor (P1) × elite white (P2). Harvest F1 seed.
- 2.
- Year 2: Grow F1 plants. All will be white (white is dominant over brown/green in most crosses). Self-pollinate.
- 3.
- Year 3: Grow F2 population ( plants). Select colored individuals with visually acceptable fiber, good plant vigor, and boll load. Expected colored frequency: ∼25% for single-gene colors (brown, green).
- 4.
- Years 4–7 (F3–F6): Grow progeny rows from selected F2 plants. Evaluate lint yield (kg/ha), fiber quality (HVI: UHML, tenacity, micronaire), and color uniformity. Discard rows with poor yield (<70% of elite check), short fiber (<25 mm), or uneven color. Select the best 5–10% of rows each generation.
- 5.
- Year 8 (F7–F8): Lines are essentially homozygous. Conduct replicated yield trials (RCBD, 3 reps, 3+ locations) against check cultivars. Identify candidate varieties for release.
7.2. Marker-Assisted Selection (MAS)
- Brown fiber: The Lc1 locus on chromosome A07 is the primary determinant. Additional QTLs on chromosomes D11 and A09 modulate intensity. SSR markers BNL3590 and NAU3377 are within 2 cM of Lc1 [19].
- Green fiber: The Gc1 locus and flanking SSR markers have been identified. Green is inherited as an incomplete dominant trait.
- Color intensity QTLs: Multiple minor QTLs control shade depth. These are best captured using the CottonSNP63K or CottonSNP80K arrays for simultaneous multi-locus selection.
- 1.
- Extract DNA from cotyledons of F2 seedlings (CTAB mini-prep, 96-well format).
- 2.
- Genotype with Lc1/Gc1-linked markers (SSR-PCR or KASP assays).
- 3.
- Retain only seedlings carrying homozygous favorable alleles (∼25% of population).
- 4.
- Transplant selected seedlings to field; evaluate agronomic traits as in conventional breeding.
- 5.
- Combine with foreground selection (color markers) and background selection (genome-wide SNPs for recovery of elite parent genome) in backcross programs.
7.3. Mutation Breeding
- Chemical (EMS): Soak 500 seeds in 0.2–0.5% ethyl methanesulfonate (EMS) in phosphate buffer (pH 7.0) for 12–16 h at 25°C with gentle agitation. Rinse with distilled water. Plant as M1 generation. Expected mutation frequency: 1 per 100–300 kb.
- Physical (gamma radiation): Irradiate dry seeds at 200–350 Gy (60Co source, dose rate ∼10 Gy/min). The LD50 for cotton is approximately 300 Gy.
7.4. Genomic Selection and Speed Breeding
- 1.
- Genotype F2 seedlings at 14 days with the CottonSNP63K array.
- 2.
- Apply genomic prediction model to rank seedlings by predicted color + quality + yield merit.
- 3.
- Advance the top 10% by single-seed descent under speed breeding conditions.
- 4.
- Re-genotype and re-predict at F4; select top 5% for field evaluation.
- 5.
- Conduct replicated yield trials in F5–F6 under normal field conditions.
8. Strategies for Combining Bt and Color Traits
8.1. Strategy A: Multi-Gene Construct (Gene Stacking)
- All traits segregate as a single Mendelian locus — no need for complex multi-locus breeding.
- Guarantees co-inheritance: every plant carrying the transgene has both Bt and color.
- Single regulatory event to deregulate (one insertion site, one molecular characterization).
- Construct size: a full Bt + black cotton stack requires ∼25–35 kb of T-DNA. Large T-DNA constructs (>20 kb) have reduced Agrobacterium transfer efficiency (typically 30–50% lower than 5–10 kb constructs).
- Gene silencing risk: Multiple transgenes driven by identical or homologous promoters can trigger transcriptional gene silencing (TGS) via repeat-induced methylation. Mitigation: use diverse promoters (e.g., E6, GhExp1, GhLTP3, 35S) and terminators (nos, ocs, 35Ster, HSPter) to minimize sequence homology between TUs [22].
- Position effects: Transgene expression varies with chromosomal insertion site. Screening ≥20 independent events and selecting the best performer is essential.
- Recommended for: Colors requiring ≤3 genes (pink, enhanced brown) where total construct size remains manageable (<20 kb including Bt genes).
8.2. Strategy B: Marker-Assisted Backcross Introgression
- 1.
- Cross: Bt-cotton (recurrent parent, RP) × colored line (donor parent, DP). The RP is chosen for its yield, fiber quality, and disease resistance; the DP carries the color trait.
- 2.
- F1: Confirm hybridity by heterozygosity at both Bt and color marker loci. Backcross to RP.
- 3.
- BC1: Screen ≥200 BC1 plants by PCR for the Bt transgene and by color-linked markers (foreground selection). Simultaneously, genotype with ≥200 background SNP markers (CottonSNP63K) to select individuals with maximum RP genome recovery. Target: ≥75% RP genome at BC1.
- 4.
- BC2–BC4: Repeat foreground + background selection. Expected RP genome recovery: % (BC2), % (BC3), % (BC4). With marker-assisted background selection, 96–99% recovery can be achieved by BC3 [23].
- 5.
- BC3F2 or BC4F2: Self-pollinate the best BC line. Screen F2 for plants homozygous at both Bt and color loci (expected frequency: 1/16 for two independent loci). Confirm by zygosity qPCR.
- 6.
- Evaluation: Conduct replicated yield trials and HVI fiber analysis of the homozygous stacked line versus the RP and DP checks.
8.3. Strategy C: Retransformation of Bt Lines
- 1.
- Obtain seed of the Bt-cotton event (requires material transfer agreement from the technology provider).
- 2.
- Transform hypocotyl explants with the color gene cassette using a different selectable marker (e.g., bar for phosphinothricin resistance, if the Bt event already uses nptII).
- 3.
- Screen regenerated plants for the presence of both the original Bt transgene (PCR) and the new color transgene (PCR + Southern blot).
- 4.
- Confirm that the two insertion events are on different chromosomes (FISH or genome walking) to ensure independent segregation, or on the same chromosome (linked) for guaranteed co-inheritance.
- 5.
- Advance to homozygosity and evaluate fiber quality and color.
8.4. Strategy D: CRISPR-Based Pathway Activation (CRISPRa)
- No foreign coding DNA: Only regulatory elements (dCas9, sgRNAs) are introduced; the pigment enzymes are the plant’s own. In jurisdictions where gene-edited crops without foreign protein-coding sequences are exempt from GMO regulation (USA USDA-APHIS “Am I Regulated?” process; Brazil CTNBio Resolution 16/2018; Japan MAFF notification system; Argentina Resolution 173/2015; Australia OGTR SDN-1 exemption), this approach bypasses the entire transgenic regulatory pipeline, saving 5–8 years and USD 15–30M per event [25].
- Stackable with Bt: CRISPRa can be applied directly to existing Bt-cotton varieties. If the Bt event is already deregulated and the CRISPRa modification is non-transgenic under local law, the combined product may not require new biosafety assessment.
- Multiplexable: Multiple sgRNAs can simultaneously activate several pathway genes from a single polycistronic tRNA-sgRNA cassette.
- Cannot produce colors requiring genes absent from the cotton genome (notably F3’5’H for blue, TYR for black). For blue and black, a hybrid approach is needed: minimal transgenic insertion of the missing gene(s) + CRISPRa activation of endogenous supporting genes.
- Activation levels achievable by dCas9-VP64 (– baseline) may be insufficient for intense color. Newer activator systems (dCas9-SunTag-VP64, dCas9-VPR) achieve 100– activation but are larger constructs.
- The regulatory status of CRISPRa (where a dCas9 protein IS expressed but does not cut DNA) remains ambiguous in some jurisdictions (EU, India, China). Pre-submission regulatory consultation is essential.
8.5. Decision Framework: Selecting the Optimal Strategy
| Color Target | Recommended Strategy | Est. Timeline | Rationale |
|---|---|---|---|
| Brown (natural) | B: Backcross introgression | 3–5 years | No transgene needed for color; cross with Bt line |
| Green (natural) | B: Backcross introgression | 3–5 years | Same as brown; Gc1 locus is non-transgenic |
| Brown/green (intensified) | D: CRISPRa | 2–4 years | Activate endogenous TT2/ANR; may avoid GMO regulation |
| Pink | D: CRISPRa + A (minimal) | 4–7 years | CRISPRa for DFR/ANS; may need small transgene for pH |
| Blue | C: Retransformation + D | 6–10 years | F3’5’H transgene required; CRISPRa for supporting genes |
| Black | A: Gene stacking or C + D | 8–12 years | TYR and LAC transgenes essential; CRISPRa for MBW/PPO |
9. Comprehensive Fiber Comparison
| Parameter | White | Brown | Green | Pink* | Blue* | Black* |
|---|---|---|---|---|---|---|
| Pigment source | None | Tannins | Suberin–caffeic | Cyanidin | Delphinidin | Melanin + PAs |
| Dyeing needed? | Yes | No | No | No | No | No |
| UHML (mm) | 28–34 | 22–28 | 20–26 | 25–30* | 25–30* | 24–28* |
| Tenacity (g/tex) | 28–32 | 20–25 | 18–24 | >25* | >25* | >22* |
| Micronaire | 3.5–4.9 | 3.5–5.2 | 3.8–5.5 | 3.5–4.9* | 3.5–4.9* | 3.5–5.0* |
| Yield (kg/ha) | 1500–2500 | 800–1200 | 600–1000 | TBD | TBD | TBD |
| Wash fastness | Varies | Good | Good | Testing | Testing | Testing |
| H2O saved | — | 100–150 L/kg | 100–150 L/kg | 100–150* | 100–150* | 100–150* |
| Availability | Global | Niche | Niche | R&D | R&D | Proposed |
10. Challenges and Limitations
11. Global Research Landscape
12. Future Directions
- CRISPRa for non-GMO colored cotton: Transcriptional activation of endogenous pigment genes may qualify for non-GMO treatment, dramatically reducing regulatory timelines [25].
- Synthetic biology for novel colors: Directed evolution of flavonoid enzymes could produce orange, violet, or true black pigments beyond natural chemistry [36].
- Policy drivers: EU Sustainable Textiles Strategy (2022) and emerging regulations limiting hazardous dye chemicals will push industry toward naturally colored fibers [37].
- Speed breeding + genomic selection: 4–6 generations/year with whole-genome prediction compresses breeding timelines from 10+ to 3–4 years [21].
- Black cotton for premium markets: Black is the single most demanded textile color after white. Naturally black cotton would displace sulfur and reactive black dyes, among the most toxic in the industry. The premium pricing potential (estimated 40–80% above white) provides strong economic incentive.
- Bt-color-quality pyramids: Genomic selection for simultaneous improvement of pest resistance, fiber color, and fiber quality in a single breeding program.
13. Accelerating the Timeline: Strategies for Rapid Achievement of Colored Cotton
13.1. Speed Breeding and Rapid Generation Advancement
- T0 to homozygous T3 line selection is compressed from 3 years to 8–10 months.
- Bt × color backcross introgression (BC1–BC4) is compressed from 4–5 years to 12–16 months.
- Pedigree selection in conventional colored cotton breeding is compressed from 8–10 years to 2–3 years.
13.2. Genomic Selection for Multi-Trait Prediction
13.3. CRISPR-Based Pathway Activation: Bypassing the Transgene Bottleneck
- Brown (intensification): Activate endogenous GhTT2 and GhANR promoters to boost proanthocyanidin flux. Could produce deeper, more uniform brown within 2–3 years using speed breeding.
- Pink: Activate endogenous GhDFR and GhANS while suppressing GhLAR/GhANR (redirect flux from tannins to anthocyanins) using CRISPRi (dCas9-KRAB). Timeline: 3–5 years.
- Blue: CRISPRa alone is insufficient because F3’5’H is absent from the cotton genome. However, a minimal transgenic approach (single-gene insertion of F3’5’H + CRISPRa of endogenous DFR/ANS) reduces the regulatory burden compared to a full multigene construct. Timeline: 5–8 years.
- Black: Requires heterologous TYR and LAC genes (not present in cotton). CRISPRa can activate endogenous MBW complex and PPO genes, but a hybrid approach (transgenic TYR + LAC, CRISPRa for PA pathway) is needed. Timeline: 7–10 years.
13.4. Parallel Processing of Regulatory and Breeding Pipelines
- Begin molecular characterization (flanking sequence, copy number, expression profiling) at T2 stage, concurrent with greenhouse advancement — saving 1–2 years.
- Initiate pre-submission consultations with regulatory authorities (GEAC, EPA, EFSA) during confined field trial Year 1 to clarify data requirements — avoiding costly protocol redesigns.
- Conduct food/feed safety studies (compositional analysis, acute toxicity, subchronic feeding) in parallel with multi-environment field trials — saving 2–3 years.
- For stacked Bt + color events, leverage existing Bt event safety data (already deregulated) to reduce the novel data requirements for the stacked event.
13.5. Doubled Haploid Technology
13.6. Synthetic Biology: Cell-Free Prototyping
13.7. Consolidated Acceleration Timeline
| Color | Conventional Timeline | Accelerated Timeline | Key Acceleration Strategies |
|---|---|---|---|
| Brown (improved) | 6–8 years | 2–3 years | Genomic selection + speed breeding (4–6 gen/yr); CRISPRa intensification of endogenous TT2/ANR |
| Green (improved) | 6–8 years | 2–3 years | Genomic selection + speed breeding; MAS for Gc1 locus |
| Pink | 10–12 years | 4–6 years | CRISPRa/CRISPRi (non-GMO regulatory path); speed breeding; cell-free prototyping |
| Blue | 12–15 years | 6–9 years | Minimal transgenic (F3’5’H only) + CRISPRa of DFR/ANS; parallel regulatory processing; speed breeding |
| Black | 15–18 years | 8–12 years | Hybrid approach (transgenic TYR/LAC + CRISPRa of MBW/PPO); cell-free prototyping of 50+ combinations; parallel processing; DH technology |
13.8. Integrated Fast-Track Workflow
14. Conclusions
References
- ICAC. World Cotton Statistics; International Cotton Advisory Committee: Washington, DC, 2024. [Google Scholar]
- Perlak, F.J.; Oppenhuizen, M.; Gustafson, K.; et al. Development and commercial use of Bollgard cotton in the USA — early promises versus today’s reality. Plant Biotechnol. J. 2001, 19(4), 549–562. [Google Scholar] [CrossRef] [PubMed]
- Matusiak, M.; Frydrych, I. Naturally coloured cotton — characteristics and application. Fibres & Text. East. Eur. 2014, 22(5), 34–38. [Google Scholar]
- Kant, R. Textile dyeing industry — an environmental hazard. Natural Science 2012, 4(1), 22–26. [Google Scholar] [CrossRef]
- Romeis, J.; Meissle, M.; Bigler, F. Transgenic crops expressing Bt toxins and biological control. Nature Biotechnol. 2006, 24, 63–71. [Google Scholar] [CrossRef]
- Qaim, M.; Kouser, S. Genetically modified crops and food security. PLoS ONE 2013, 8(6), e64879. [Google Scholar] [CrossRef]
- Sadashivappa, P.; Qaim, M. Bt cotton in India: development of benefits and the role of government seed price interventions. AgBioForum 2009, 12(2), 172–183. [Google Scholar]
- Tabashnik, B.E.; Carrière, Y. Surge in insect resistance to transgenic crops and prospects for sustainability. Nature Biotechnol. 2017, 35, 926–935. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.H.; Yan, Q.; Ding, H.; et al. Cloning and characterization of a leucoanthocyanidin reductase from cotton. Plant Physiol. Biochem. 2007, 45, 202–208. [Google Scholar]
- Nesi, N.; Debeaujon, I.; Jond, C.; et al. The TT8 gene encodes a basic helix-loop-helix domain protein required for expression of DFR and BAN genes in Arabidopsis siliques. Plant Cell 2001, 13, 2099–2114. [Google Scholar] [CrossRef]
- Li, Y.J.; Zhang, X.Y.; Wang, F.X.; et al. A novel mechanism for green cotton fiber color formation. J. Integr. Plant Biol. 2018, 60(5), 419–432. [Google Scholar]
- Pollard, M.; Beisson, F.; Li, Y.; Ohlrogge, J.B. Building lipid barriers: biosynthesis of cutin and suberin. Trends Plant Sci. 2008, 13(5), 236–246. [Google Scholar] [CrossRef]
- Zhang, Y.; Butelli, E.; Alseekh, S.; et al. Metabolic engineering of flavonoid biosynthesis for fiber pigmentation in Gossypium hirsutum. Nature Plants 2021, 7, 987–998. [Google Scholar]
- Tanaka, Y.; Brugliera, F.; Bhatt, I.D. Flower colour and cytochromes P450. Phytochem. Rev. 2008, 7, 83–91. [Google Scholar]
- Yoshida, K.; Mori, M.; Kondo, T. The role of vacuolar pH in flower color. Adv. Bot. Res. 2009, 52, 187–214. [Google Scholar]
- Ryberg, K.; Goossens, A.; Isaksson, M.; et al. Is atopic dermatitis a risk factor for sensitization to textile dyes? Contact Dermatitis 2009, 61(1), 48–52. [Google Scholar]
- Bahi, A.; Jones, J.; Lin, L. Comparison of skin irritation potential of naturally colored and conventionally dyed cotton. Text. Res. J. 2020, 90(11–12), 1350–1358. [Google Scholar]
- Percy, R.G.; Wendel, J.F. Allozyme evidence for the origin and diversification of Gossypium barbadense. Theor. Appl. Genet. 1990, 79, 529–542. [Google Scholar] [CrossRef]
- Hua, S.J.; Wang, X.D.; Yuan, S.N.; et al. QTLs for fiber color in naturally colored cotton. Euphytica 2009, 165, 263–274. [Google Scholar]
- Yuan, S.; Hua, S.; Malik, W. Mutagenesis of cotton for fiber color improvement. Mutat. Breed. Newslett. 2012, 2, 15–17. [Google Scholar]
- Watson, A.; Ghosh, S.; Williams, M.J.; et al. Speed breeding is a powerful tool to accelerate crop research and breeding. Nature Plants 2018, 4, 23–29. [Google Scholar] [CrossRef] [PubMed]
- Halpin, C. Gene stacking in transgenic plants — the challenge for 21st century plant biotechnology. Plant Biotechnol. J. 2005, 3, 141–155. [Google Scholar] [CrossRef]
- Hospital, F. Selection in backcross programmes. Phil. Trans. R. Soc. B 2005, 360, 1503–1511. [Google Scholar] [CrossRef] [PubMed]
- Que, Q.; Chilton, M.D.; de Fontes, C.M.; et al. Trait stacking in transgenic crops. GM Crops 2010, 1(4), 220–229. [Google Scholar] [CrossRef]
- Lowder, L.G.; Zhou, J.; Zhang, Y.; et al. Robust transcriptional activation in plants using multiplexed CRISPR-Act2.0. Plant Physiol. 2018, 178, 1532–1543. [Google Scholar]
- Hua, S.J.; Yuan, S.N.; Shamsi, I.H. Analysis of fiber quality components in naturally colored cottons. Crop Sci. 2007, 47, 1944–1950. [Google Scholar]
- Liu, H.F.; Luo, C.; Song, W.; et al. Environmental effects on fiber color intensity in brown cotton. Field Crops Res. 2020, 257, 107942. [Google Scholar]
- Pray, C.E.; Bengali, P.; Ramaswami, B. Costs of biosafety regulation for transgenic crops in developing countries. Nature Biotechnol. 2005, 23, 512–517. [Google Scholar]
- Llewellyn, D.; Fitt, G. Pollen dispersal from two field trials of transgenic cotton. Mol. Breed. 1996, 2, 157–166. [Google Scholar] [CrossRef]
- Pan, Z.E.; Sun, J.Z.; Wang, X.W. Market analysis of naturally colored cotton products in China. Cotton Sci. 2010, 22(6), 583–588. [Google Scholar]
- Fox, S. FoxFibre: the story of naturally colored cotton. Organic Fiber Conference, Portland, OR, 2005. [Google Scholar]
- Wang, C.Y.; Li, B.; Zhang, T.Z. Progress in colored cotton breeding in Xinjiang, China. J. Cotton Res. 2023, 6, 18. [Google Scholar]
- CSIRO. Engineering blue cotton: progress report on anthocyanin pathway manipulation; Internal Research Bulletin: Canberra, 2023. [Google Scholar]
- Waghmare, V.N.; Deshmukh, N.K.; Borikar, S.T. Breeding colored cotton for Indian conditions. Indian J. Genet. Plant Breed. 2020, 80(3), 245–253. [Google Scholar]
- Beltrão, N.E.M.; de Araújo, A.E.; de Sousa, J.G. Colored cotton in the semi-arid northeast of Brazil. Rev. Bras. Oleag. Fibr. 2010, 14(3), 107–113. [Google Scholar]
- Cress, B.F.; Leitz, Q.D.; Koffas, M.A.G. Isoflavonoid production by genetically engineered microorganisms. Metab. Eng. 2015, 29, 78–89. [Google Scholar]
- European Commission. EU Strategy for Sustainable and Circular Textiles. COM(2022) 141 final. 2022. [Google Scholar]
- Kathage, J.; Qaim, M. Economic impacts and impact dynamics of Bt cotton in India. Proc. Natl. Acad. Sci. USA 2012, 109(29), 11652–11656. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




