Submitted:
06 March 2026
Posted:
06 March 2026
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
2. Materials and Methods
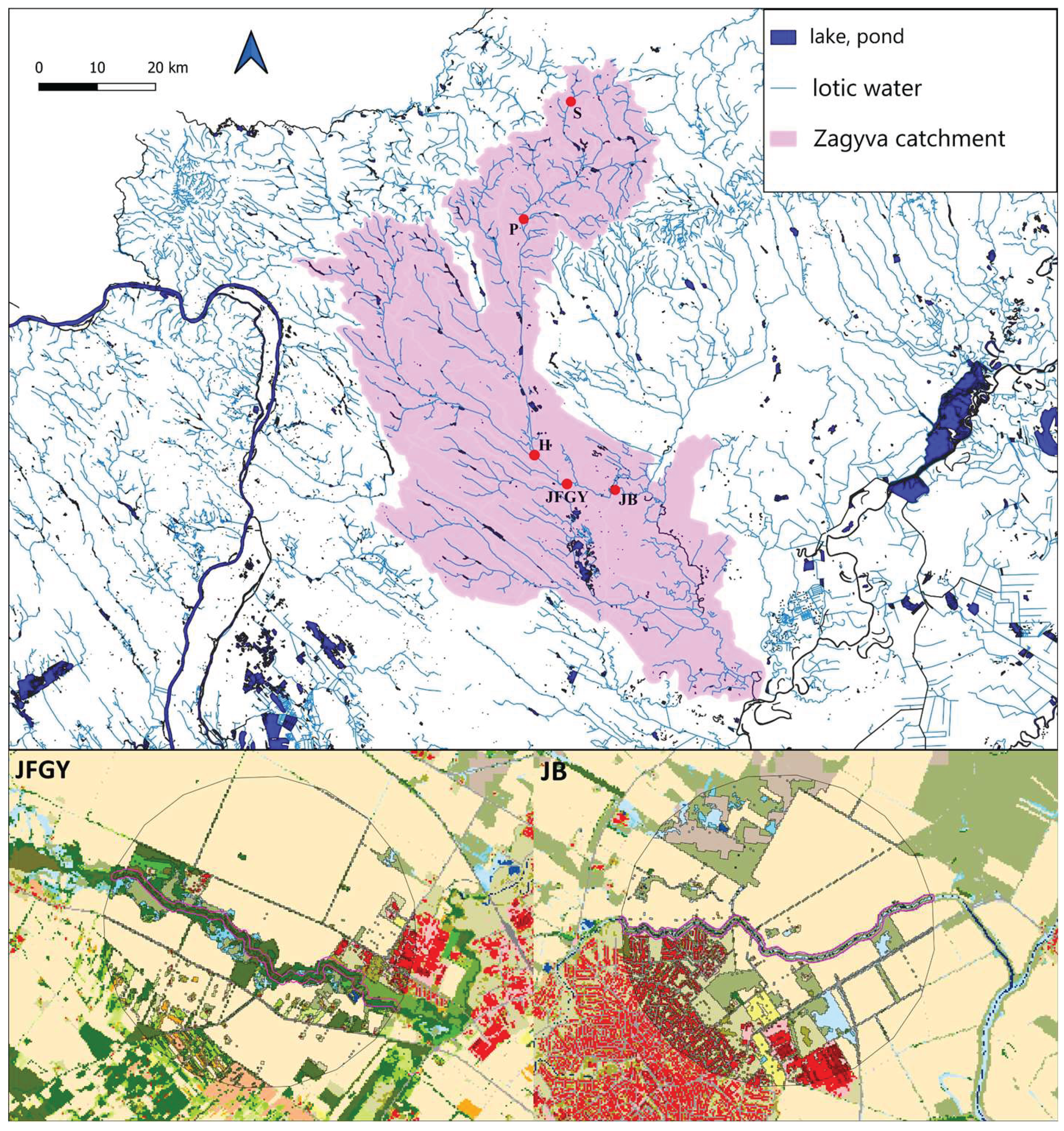
2.1. Study area
2.2. Sample collection
2.3. Bacteriological investigation
2.4. Data analysis
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AMI | amikacin |
| AMR | Antimicrobial Resistance |
| AMP/SAM | ampicillin/sulbactam |
| ARG | Antimicrobial Resistance Gene |
| AST | Antimicrobial Susceptibility Testing |
| AZT | aztreonam |
| B-PEN | benzylpenicillin |
| CFP | cefepime |
| CIP | ciprofloxacin |
| CLIN | clindamycin |
| COL | colistin |
| CZD | ceftazidime |
| ERY | erythromycin |
| FOM | fosfomycin |
| FOX | cefoxitin |
| FUS | fusidic acid |
| GEN | gentamicin |
| GN | Gram-negative |
| GP | Gram-positive |
| HPC | Heterotrophic Plate Count |
| IMI | imipenem |
| JB | Jászberény (study site) |
| JFGY | Jászfelsőszentgyörgy (study site) |
| LEV | levofloxacin |
| LIN | linezolid |
| MAR | Multiple Antibiotic Resistance |
| MDR | Multidrug Resistance |
| MER | meropenem |
| MOX | moxifloxacin |
| OXA | oxacillin |
| PCA | Plate Count Agar |
| PIP | piperacillin |
| RIF | rifampicin |
| SD | Standard Deviation |
| SXT | trimethoprim/sulfamethoxazole |
| TEI | teicoplanin |
| TET | tetracycline |
| TIC/CLA | ticarcillin/clavulanic acid |
| TIGE | tigecycline |
| TOM | tobramycin |
| TRIM | trimethoprim |
| TZP | piperacillin/tazobactam |
| VAN | vancomycin |
References
- Huang, J.; Zhou, N.; Cheng, Z.; Chen, Y.; Li, M.; Lv, C.; Guo, X.; Li, Q.; Yao, Y.; Dong, K.; Zhu, Y. Chromosomally located blaCMH in Enterobacter cloacae complex across human-bird-environment interfaces: A one-health perspective. Sci. Total. Environ. 2024, 954, 176486. [Google Scholar] [CrossRef]
- Doyle, C.; Wall, K.; Fanning, S.; McMahon, B. J. Making sense of sentinels: wildlife as the One Health bridge for environmental antimicrobial resistance surveillance. J. Appl. Microbiol. 2025, 136, lxaf017. [Google Scholar] [CrossRef] [PubMed]
- Kim, D. W.; Cha, C. J. Antibiotic resistome from the One-Health perspective: understanding and controlling antimicrobial resistance transmission. Exp. Mol. Med. 2021, 53, 301–309. [Google Scholar] [CrossRef] [PubMed]
- Zanardi, G.; Iemmi, T.; Spadini, C.; Taddei, S.; Cavirani, S.; Cabassi, C. S. Wild micromammals as bioindicators of antibiotic resistance in ecopathology in Northern Italy. Animals 2020, 10, 1184. [Google Scholar] [CrossRef]
- Piotrowska, M.; Popowska, M. The prevalence of antibiotic resistance genes among Aeromonas species in aquatic environments. Ann. Microbiol. 2014, 64, 921–934. [Google Scholar] [CrossRef]
- Grilli, G.; Rapi, M. C.; Musa, L.; Di Giacinto, G.; Passamonti, F.; Raimondi, S.; Cianca, O.; Franciosini, M. P. Looking for ESKAPE bacteria: Occurrence and phenotypic antimicrobial resistance profiles in wild birds from Northern and Central Italy sites. Antibiotics 2025, 14, 1025. [Google Scholar] [CrossRef]
- Cai, F.; Calisi, R. M. Seasons and neighborhoods of high lead toxicity in New York City: the feral pigeon as a bioindicator. Chemosphere 2016, 161, 274–279. [Google Scholar] [CrossRef]
- Haynes, P. J.; Inglis, I. R.; Isaacson, T. J.; Fryday, S. L. Woodpigeon Columba palumbus movements in eastern England. Bird Study 2023, 50, 33–38. [Google Scholar] [CrossRef]
- Schumm, Y.R.; Masello, J.F.; Cohou, V.; Mourguiart, P.; Metzger, B.; Rösner, S.; Quillfedt, P. Should I stay or should I fly? Migration phenology, individual-based migration decision and seasonal changes in foraging behaviour of Common Woodpigeons. Sci. Nat. 2022, 109, 44. [Google Scholar] [CrossRef]
- Rapi, MC; Filipe, J; Filippone Pavesi, L; Raimondi, S; Addis, MF; Franciosini, MP; Grilli, G. Resisting the final line: Phenotypic detection of resistance to last-resort antimicrobials in Gram-negative bacteria isolated from wild birds in Northern Italy. Animals 2025, 15, 2289. [Google Scholar] [CrossRef]
- Pollack, L.; Ondrasek, N. R.; Calisi, R. Urban health and ecology: the promise of an avian biomonitoring tool. Curr. Zool. 2017, 63, 205–212. [Google Scholar] [CrossRef] [PubMed]
- Martín-Vélez, V.; Sánchez, M.I.; Lovas-Kiss, Á.; Hortas, F.; Green, A. J. Dispersal of aquatic invertebrates by lesser black-backed gulls and white storks within and between inland habitats. Aquat. Sci. 2022, 84, 10. [Google Scholar] [CrossRef]
- Luna, Á.; Pomeda-Gutiérrez, F.; Galán Díaz, J. Feeding ecology of the common wood pigeon (Columba palumbus) in a major European city. R. Soc. Open Sci. 2024, 11, 1231721. [Google Scholar] [CrossRef] [PubMed]
- BirdLife International. Species factsheet: Common Woodpigeon Columba palumbus. Available online: https://datazone.birdlife.org/species/factsheet/common-woodpigeon-columba-palumbus (accessed on 28 December 2025).
- Negrier, C.; Fantinati, M.; Jouglar, J. Y.; Lyazrhi, F.; Cohou, V.; Priymenko, N. Dietary regimen of the woodpigeon (Columba palumbus). J. Anim. Physiol. Anim. Nutr. 2021, 105, 376–384. [Google Scholar] [CrossRef]
- Haraszthy, L. Common Wood Pigeon Columba palumbus Linnaeus, 1758. in: Breeding biology of Hungarian nesting birds (in Jungarian). Haraszthy, L., Ed.; Pro Vértes Nonprofit Zrt.: Csákvár, Hungary, 2019; Volume 1, pp. 201–206. [Google Scholar]
- Schuster, S. Verhaltensänderungen bei Ringeltauben Columba palumbus im Voralpenraum. Ornithol. Jh. Baden-Württemb. 2017, 31, 100–110. [Google Scholar]
- Bozó, L.; Csörgő, T. Changes in spring arrival dates of Central European bird species over the past 100 years. Acta Zool. Acad. Sci. Hung. 2020, 66, 283–298. [Google Scholar] [CrossRef]
- Juhász, L. Synecological analysis of the ornithofauna of the city of Debrecen (in Hungarian). Puszta 1985, 3, 37–52. [Google Scholar]
- Lykov, E. L. Urban population of the Common Wood Pigeon in Krasnodar. Ornis Hung. 2024, 32, 38–153. [Google Scholar] [CrossRef]
- Illés, A.; Nagy, E.; Nagy, R. R.; Tari, T. The environmental impact of the feral pigeon (Columba livia f. domestica) in the historic city centre of Sopron. Analecta Tech. Szeged. 2023, 17, 69–76. [Google Scholar] [CrossRef]
- Radimersky, T.; Frolkova, P.; Janoszowska, D.; Dolejska, M.; Svec, P.; Roubalova, E.; Cikova, P.; Cizek, A.; Literak, I. Antibiotic resistance in faecal bacteria (Escherichia coli, Enterococcus spp.) in feral pigeons. J. Appl. Microbiol. 2010, 109, 1687–1695. [Google Scholar] [CrossRef]
- Sausse, C.; Lévy, M. Bird damage to sunflower: International situation and prospects. OCL, 2021, 28, 34. [Google Scholar] [CrossRef]
- European Council. Directive 2009/147/EC of the European Parliament and of the Council of 30 November 2009 on the conservation of wild birds. Available online: https://eur-lex.europa.eu/eli/dir/2009/147/oj/eng (accessed on 04 January 2026).
- Butkauskas, D.; Švažas, S.; Bea, A.; Prakas, P.; Olano, I.; Grishanov, G.; Mischenko, A.; Kozulin, A.; Stanevičius, V.; Báldi, A.; et al. Designation of flyways and genetic structure of Woodpigeon Columba palumbus in Europe and Morocco. Eur. J. Wildl. Res. 2019, 65, 91. [Google Scholar] [CrossRef]
- Ahmad, S.; Lohiya, S.; Taksande, A.; Meshram, R. J.; Varma, A.; Vagha, K. A comprehensive review of innovative paradigms in microbial detection and antimicrobial resistance: Beyond traditional cultural methods. Cureus 2024, 16, e61476. [Google Scholar] [CrossRef]
- Klümper, U.; Gionchetta, G.; Catão, E.; Bellanger, X.; Dielacher, I.; Elena, A. X.; Fang, P.; Galazka, S.; Goryluk-Salmonowicz, A.; Kneis, D.; et al. Environmental microbiome diversity and stability is a barrier to antimicrobial resistance gene accumulation. Commun. Biol. 2024, 7, 706. [Google Scholar] [CrossRef]
- Gensberger, E. T.; Gössl, E. M.; Antonielli, L.; Sessitsch, A.; Kostić, T. Effect of different heterotrophic plate count methods on the estimation of the composition of the culturable microbial community. PeerJ. 2015, 3, e862. [Google Scholar] [CrossRef]
- Allen, M. J.; Edberg, S. C.; Reasoner, D. J. Heterotrophic plate count bacteria—what is their significance in drinking water? Int. J. Food Microbiol. 2004, 92, 265–274. [Google Scholar] [CrossRef] [PubMed]
- Falcone-Dias, M.F.; Farache Filho, A. Quantitative variations in heterotrophic plate count and in the presence of indicator microorganisms in bottled mineral water. Food Control 2013, 31, 90–96. [Google Scholar] [CrossRef]
- Bitar, M. R.; Pinto, V. D.; Moreira, L. M.; Ribeiro, S. P. Gram-negative bacteria associated with a dominant arboreal ant species outcompete phyllosphere-associated bacteria species in a tropical canopy. Oecologia 2021, 195, 959–970. [Google Scholar] [CrossRef]
- Biswas, R.; Halder, U.; Kabiraj, A.; Mondal, A.; Bandopadhyay, R. Overview on the role of heavy metals tolerance on developing antibiotic resistance in both Gram-negative and Gram-positive bacteria. Arch. Microbiol. 2021, 203, 2761–2770. [Google Scholar] [CrossRef]
- Tóthmérész, B. DivOrd 1.50: a program for diversity ordering. Tiscia 1993, 27, 33–44. [Google Scholar]
- Chao, A.; Ricotta, C. Quantifying evenness and linking it to diversity, beta diversity, and similarity. Ecology 2019, 100, e02852. [Google Scholar] [CrossRef] [PubMed]
- Tóthmérész, B. Comparison of different methods for diversity ordering. J. Veg. Sci. 1995, 6, 283–290. [Google Scholar] [CrossRef]
- Sebők, R.; Prikler, B.; Dobolyi, C.; Szoboszlay, S.; Kriszt, B.; Szabó, I. Effects of treated wastewater discharge into a stream on microscopic fungal communities of microplastic surfaces. In 19th International Congress of the Hungarian Society for Microbiology, Márialigeti, K.; Dobay, O. (Eds.); Acta Microbiol. Immunol. Hung. 2023, 70 (Suppl 1), 53–88. [CrossRef]
- Sebők, B. The social changes of Szolnok County in the period of socialist industrialization. Paideia 2024, 14, 147–157. [Google Scholar] [CrossRef]
- Balla, A.; Moshen, A.; Kiss, T. Microplastic clouds in rivers: spatiotemporal dynamics of microplastic pollution in a fluvial system. Environ. Sci. Eur. 2024, 36, 143. [Google Scholar] [CrossRef]
- Huallacháin, D.Ó.; Dunne, J. Analysis of biometric data to determinethe sex of woodpigeons Columba palumbus. Ringing Migr. 2010, 25, 29–32. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and environment for statistical computing, R Foundation for Statistical Computing, Vienna, Austria, 2024; (https://www.R-project.org/).
- Oksanen, J.; Blanchet, F. G.; Kindt, R.; Legendre, P.; Minchin, P. R.; O'Hara, R. B.; Simpson, G. L.; Sólymos, P.; Stevens, M. H. H.; Wagner, H. vegan: Community Ecology Package Package v2.7-2, [R package]. 2025. (https://CRAN.R-project.org/package=vegan).
- Sievert, C. Interactive web-based data visualization with R, plotly, and shiny; Chapman and Hall/CRC: Florida, USA, 2020. [Google Scholar]
- Wickham, H. ggplot2: Elegant graphics for data analysis; Springer-Verlag: New York, USA, 2016. [Google Scholar]
- Carranza, M.L.; Acosta, A.; Ricotta, C. Analyzing landscape diversity in time: The use of Renyi's generalized entropy function. Ecol. Indic. 2007, 7, 05–510. [Google Scholar] [CrossRef]
- Hammer, Ø.; Harper, D.A.T.; Ryan, P.D. PAST: Paleontological Statistics Software Package for education and data analysis. Palaeontol. Electron. 2001, 4, 9. [Google Scholar]
- Conzett, P.; Dijkstra Haugstvedt, N. DataverseNO README File Template - General (v2.4). Zenodo. 2024. [Google Scholar] [CrossRef]
- Kristich, C.J.; Rice, L.B.; Arias, C.A. Enterococcal infection—treatment and antibiotic resistance. In Enterococci: From commensals to leading causes of drug resistant infection; Gilmore, M.S., Ed.; Massachusetts Eye and Ear Infirmary: Boston, USA, 2014. [Google Scholar]
- Radisic, V.; Grevskott, D. H.; Junghardt, N.; Øvreås, L.; Marathe, N. P. Multidrug-resistant Enterococcus faecium strains enter the Norwegian marine environment through treated sewage. Microbiology Open 2024, 13, e1397. [Google Scholar] [CrossRef]
- Oladipo, A. O.; Oladipo, O. G.; Bezuidenhout, C. C. Bezuidenhout, Multi-drug resistance traits of methicillin-resistant Staphylococcus aureus and other Staphylococcal species from clinical and environmental sources. J. Water Health 2019, 17, 930–943. [Google Scholar] [CrossRef] [PubMed]
- Abdullahi, I. N.; Lozano, C.; Höfle, Ú.; Cardona-Cabrera, T.; Zarazaga, M.; Torres, C. Antimicrobial resistome of coagulase-negative staphylococci from nasotracheal cavities of nestlings of Ciconia ciconia in Southern Spain: detection of mecC-SCCmec type-XI-carrying S. lentus. Comp. Immunol. Microbiol. Infect. Dis. 2023, 99, 102012. [Google Scholar] [CrossRef] [PubMed]
- Lewandowska, W.; Mahillon, J.; Drewnowska, J. M.; Swiecicka, I. Insight into the phylogeny and antibiotic resistance of Pseudomonas spp. originating from soil of the Białowieża National Park in Northeastern Poland. Front. Microbiol. 2025, 16, 1454510. [Google Scholar] [CrossRef]
- Santoro, D. O.; Romao, C. M.; Clementino, M. M. Decreased aztreonam susceptibility among Pseudomonas aeruginosa isolates from hospital effluent treatment system and clinical samples. Int. J. Environ. Health Res. 2012, 22, 560–570. [Google Scholar] [CrossRef]
- Flores Ribeiro, A.; Bodilis, J.; Alonso, L.; Buquet, S.; Feuilloley, M.; Dupont, J. P.; Pawlak, B. Occurrence of multi-antibiotic resistant Pseudomonas spp. in drinking water produced from karstic hydrosystems. Sci. Total Environ. 2014, 490, 370–378. [Google Scholar] [CrossRef]
- Cooper, G. L.; Louie, A.; Baltch, A. L.; Chu, R. C.; Smith, R. P.; Ritz, W. J.; Michelsen, P. Influence of zinc on Pseudomonas aeruginosa susceptibilities to imipenem. J. Clin. Microbiol. 1993, 31, 2366–2370. [Google Scholar] [CrossRef] [PubMed]
- Poirel, L.; Palmieri, M.; Brilhante, M.; Masseron, A.; Perreten, V.; Nordmann, P. PFM-like enzymes are a novel family of subclass B2 metallo-β-lactamases from Pseudomonas synxantha belonging to the Pseudomonas fluorescens complex. Antimicrob. Agents Chemother. 2020, 64, e01700-19. [Google Scholar] [CrossRef]
- Dehbanipour, R.; Maleki, V. T. Z.; Ghalavand, Z. Colistin heteroresistance, mechanisms, diagnostic methods, and therapeutic options: A review. Germs 2015, 15, 166. [Google Scholar] [CrossRef]
- Doijad, S. P.; Gisch, N.; Frantz, R.; Kumbhar, B. V.; Falgenhauer, J.; Imirzalioglu, C.; Falgenhauer, L.; Mischnik, A.; Rupp, J.; Behnke, M.; et al. Resolving colistin resistance and heteroresistance in Enterobacter species. Nat. Commun. 2023, 14, 140. [Google Scholar] [CrossRef]
- Ozer, G.; Ergene, A.; Icgen, B. Biochemical and molecular characterization of strontium-resistant environmental isolates of Pseudomonas fluorescens and Sphingomonas paucimobilis. Geomicrobiol. J. 2013, 30, 381–390. [Google Scholar] [CrossRef]
- Ryan, M. P.; Adley, C. C. Sphingomonas paucimobilis: a persistent Gram-negative nosocomial infectious organism. J. Hosp. Infect. 2010, 75, 153–157. [Google Scholar] [CrossRef]
- Kremer, P.; Hamstead, Z.; Haase, D.; McPhearson, T.; Frantzeskaki, N.; Andersson, E.; Kabisch, N.; Larondelle, N.; Rall, E. L.; Voigt, A.; et al. Key insights for the future of urban ecosystem services research. Ecol. Soc. 2016, 21, 29. [Google Scholar] [CrossRef]
- Alberti, M. The effects of urban patterns on ecosystem function. Int. Reg. Sci. Rev. 2005, 28, 168–192. [Google Scholar] [CrossRef]
- Clark, J. S.; Dietze, M.; Chakraborty, S.; Agarwal, P. K.; Ibanez, I.; LaDeau, S.; Wolosin, M. Resolving the biodiversity paradox. Ecol. Lett. 2007, 10, 647–659. [Google Scholar] [CrossRef] [PubMed]
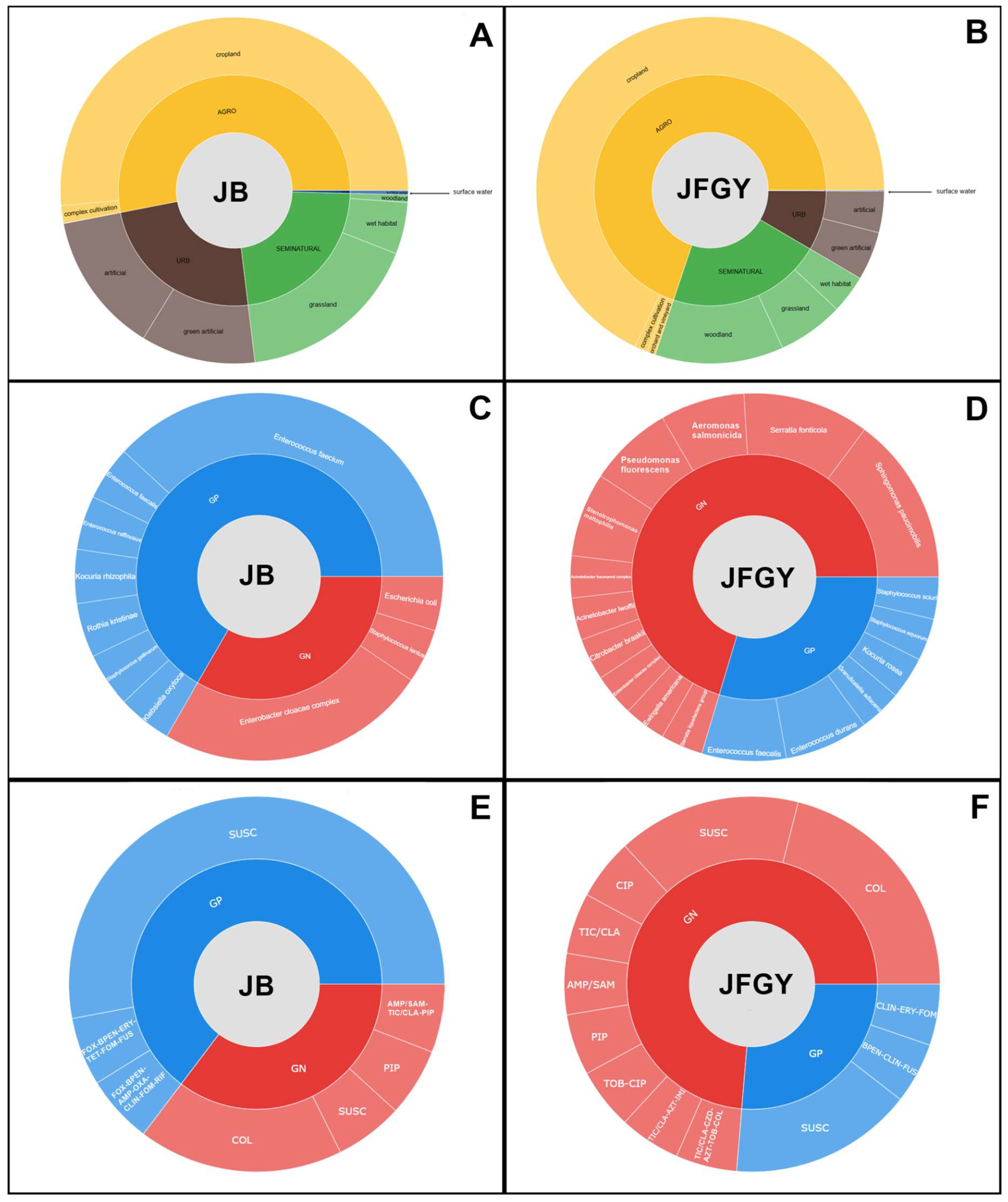
| Study site | Bacterial species | AMR profile |
| JB | Staphylococcus gallinarum (GP)1 | FOX2, B-PEN, ERY, TET3, FOM, FUS, RIF (I)4 |
| Mammaliicoccus lentus (GP) | FOX, B-PEN, OXA, CLIN, FOM, RIF | |
| Klebsiella oxytoca (GN) | AMP, TIC/CLA, PIP | |
| JFGY | Staphylococcus equorum (GP) | ERY, CLIN, FOM |
| Mammaliicoccus sciuri (GP) | B-PEN, CLIN, FUS | |
| Acinetobacter baumannii Complex (GN) | PIP | |
| Acinetobacter lwoffii (GN) | AMP (I) | |
| Aeromonas hydrophila (GN) | CIP | |
| Citrobacter braakii (GN) | AMP/SAM | |
| Pseudomonas fluorescens (GN) | TIC/CLA, FOX, AZT, IMI, AMI, TOB, COL, MER (I), GEN (I) | |
| Serratia fonticola (GN) | COL | |
| Serratia liquefaciens Group (GN) | COL | |
| Sphingomonas paucimobilis (GN) | TOB, CIP, AMI (I), GEN (I) | |
| Both5 | Enterobacter cloacae Complex (GN) | COL |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




