1. Introduction
Staphylococcus aureus, including both methicillin-sensitive (MSSA) and methicillin-resistant (MRSA), remains a significant pathogen in European healthcare settings [
1]. These bacteria can cause a range of infections, from minor skin and soft tissue infections to severe diseases like bacteremia, endocarditis, and osteomyelitis. The transmission of MRSA within healthcare facilities, especially in intensive care units, remains a persistent challenge [
2,
3]. While the incidence of hospital-acquired MRSA infections has witnessed a decline in certain European regions, thanks to effective infection control measures, community-acquired MRSA continues to pose a substantial public health threat [
4,
5,
6].
The occurrence of hospital-acquired
S. aureus infection adversely affects patient prognosis [
7,
8]. The outcomes are even worse if the pathogen is methicillin-resistant [
9]. The nosocomial MRSA infections pose also significant economical burned [
10,
11]. Multi-resistance, where bacteria develop resistance to multiple antibiotic classes, has been surging. The overuse of antibiotics, both in medical practice and agriculture, are cardinal drivers of this phenomenon. The European Centre for Disease Prevention and Control (ECDC) has been monitoring antimicrobial resistance trends and consistently warning about the consequences of antibiotic overuse [
12]. European initiatives against the threat of antibiotic resistance have not been confined to surveillance. Research and development efforts are ongoing to discover novel antibiotics and alternative therapeutic strategies [
13,
14].
The resistance patterns vary not only among and within different bacterial species but also across geographical regions. Consequently, robust surveillance of various bacterial pathogens is essential for developing effective strategies and interventions to combat the emergence and spread of antibiotic resistance [
15]. Slovakia (Central Europe) faces significant challenges related to antimicrobial resistance. Among 204 countries, Slovakia ranks 56th in age-standardized mortality rate per 100,000 population associated with AMR [
16]. Slovakia has consistently exceeded the EU average. At the country level, eight countries in EU had increases in hospital antibiotic consumption between 2019 and 2020, and 11 had increases between 2020 and 2021. Five countries (Bulgaria, Croatia, Greece, Portugal, and Slovakia) had higher hospital consumption in 2021 than in 2019 [
12].
The COVID-19 pandemic substantially influenced the epidemiology of
Staphylococcus aureus, including both MRSA and MSSA, although reported effects have been heterogeneous across settings. While some regions observed a temporary decline in community-associated
S. aureus transmission following the implementation of non-pharmaceutical interventions (e.g., enhanced hygiene and reduced social contact) [
17], multiple surveillance studies reported increased rates of hospital-onset MRSA bacteremia during the pandemic, particularly among patients with COVID-19 and in intensive care settings [
18,
19]. In parallel, increased empirical use of broad-spectrum antibiotics and disruptions in antimicrobial stewardship programs may have contributed to selective pressure favoring antimicrobial resistance [
20,
21]. Overall, the pandemic appears to have differentially affected MRSA/MSSA prevalence and multidrug resistance patterns in community versus hospital environments, with long-term consequences still requiring further evaluation.
An increasing number of diverse data sources are now being integrated for epidemiological analysis, allowing researchers to combine evidence from multiple origins [
22]. Incorporating real-time information and big data analytics into public health systems strengthens preventive strategies and enables more effective outbreak control measures. By leveraging these tools, public health organizations can improve forecasting and management of such events, thereby mitigating their impact on human lives and the economy [
23].
Systematically collecting, analyzing data and regular data monitoring allows for the identification of emerging resistance patterns, high-risk areas, specific multidrug resistant (MDR) strains and the effect of epidemiological approach. However, the efforts to combat AMR are challenged by the poor availability of reliable data, issues of integration, privacy, scalability, and data quality [
24,
25], particularly from low and middle-income countries.
2. Results
This section presents the results of the longitudinal analysis of S. aureus derived from routine laboratory diagnostics and hospital activity data. First, the analytical framework developed for standardized data processing and surveillance is described to provide context for the subsequent findings. This is followed by an overview of the study dataset and temporal changes in hospital activity. Finally, incidence trends of S. aureus are reported after normalization to hospitalizations, stratified by resistance phenotype, specimen classification, and department category.
2.1. KNIME-Based Workflow for Longitudinal Surveillance
To enable standardized and reproducible analysis of
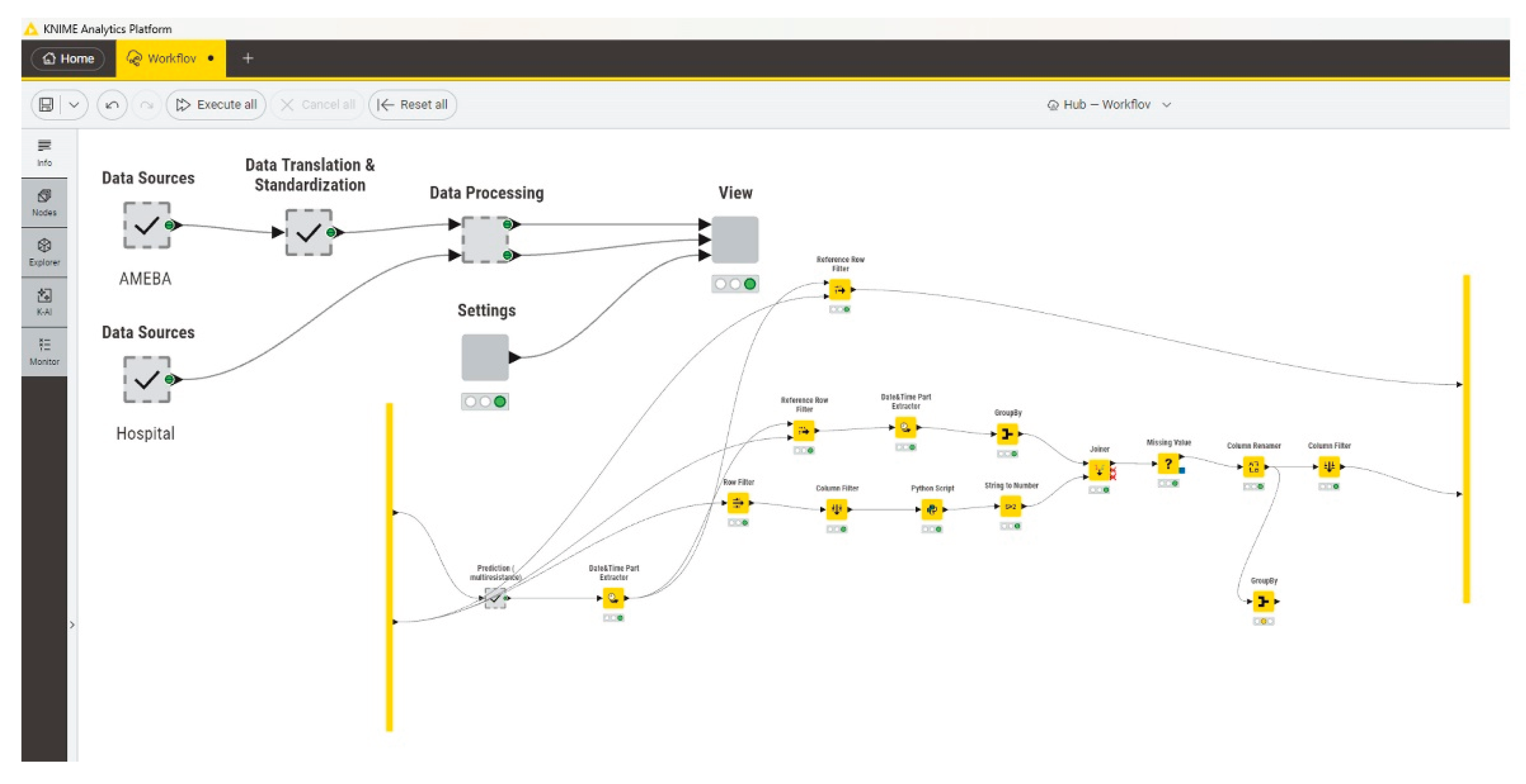
S. aureus burden from routine laboratory diagnostics, a comprehensive analytical workflow was developed within the KNIME Analytics Platform (
Figure 1). The workflow integrates microbiological data with hospital activity data and applies a unified preprocessing strategy across the entire study period.
The analytical pipeline performs automated data cleaning, deduplication, and harmonization of antimicrobial susceptibility results in accordance with EUCAST criteria. Isolates were subsequently classified by resistance phenotype, with particular focus on multidrug-resistant (MDR) S. aureus and methicillin-resistant S. aureus (MRSA). The assignment of isolates to colonization or infection categories was performed automatically within the KNIME workflow based on predefined classification rules. These rules were developed and validated by infectious disease specialists and hospital epidemiologists, who defined the criteria for assigning individual specimen types and clinical contexts to colonization or infection. Once implemented, the workflow applied these expert-derived rules consistently across the entire dataset, ensuring reproducible and unbiased classification.
Similarly, clinical departments were clustered into surgical and non-surgical categories using expert-defined institutional criteria. These categorization rules, reflecting differences in patient populations, care pathways, and exposure to invasive procedures, were encoded directly into the KNIME workflow. This approach enabled automated, standardized department-level stratification while preserving clinical relevance.
A central feature of the workflow is automated temporal aggregation and normalization of resistance incidence metrics to completed hospitalizations, ensuring robust comparability across calendar years and department types despite substantial changes in hospital activity and capacity over time. All steps of data translation, standardization, filtering, and aggregation were implemented using KNIME Base nodes complemented by selected extensions, including KNIME AI Extension, KNIME Excel Support, KNIME Expressions, KNIME Javasnippet, KNIME JSON-Processing, KNIME Math Expression (JEP), KNIME Python Integration, KNIME Quick Forms, KNIME Views, and Vernalis KNIME Nodes. Together, these components form a fully reproducible analytical pipeline that underpins all downstream results presented in this study (
Figure 1).
2.2. Characteristics of the Study Dataset and Hospital Activity
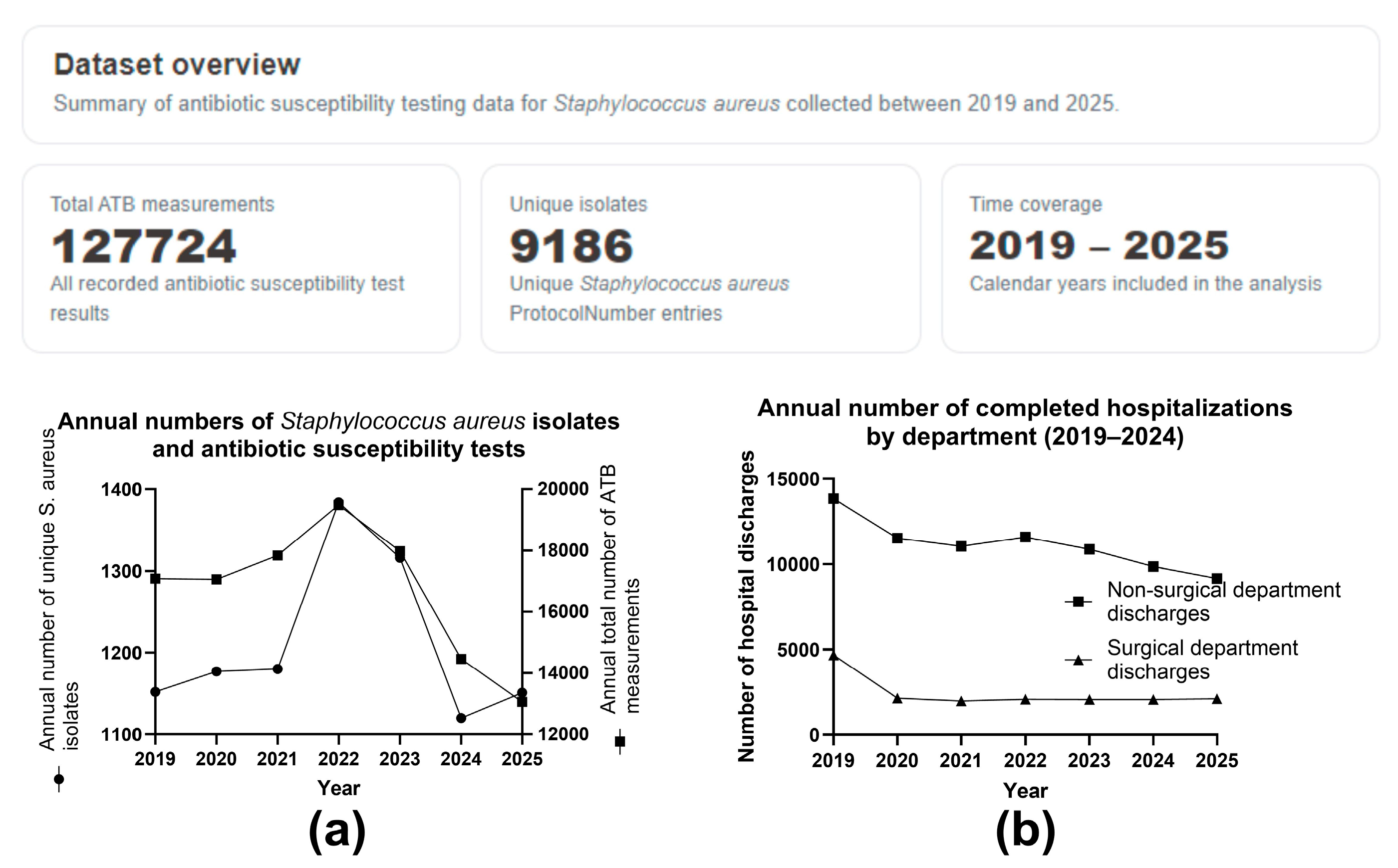
After data cleaning, expert-rule–based classification, and deduplication implemented within the KNIME analytical workflow, the final dataset comprised 127,724 antibiotic susceptibility testing (ATB) measurements corresponding to 9,186 unique
Staphylococcus aureus isolates collected between 2019 and 2025 (
Figure 2). Because multiple antibiotics were routinely tested per isolate, several ATB measurements were available for each strain.
As shown in
Figure 2A, the annual number of unique
S. aureus isolates remained relatively stable between 2019 and 2021, followed by a pronounced increase in 2022, when both the number of isolates and the total number of ATB measurements reached their maximum. In subsequent years, a marked decline was observed, with the lowest values recorded in 2024, followed by a partial rebound in 2025. A closely corresponding temporal pattern was observed for the total number of ATB measurements, indicating that changes in laboratory workload largely mirrored fluctuations in isolate numbers rather than alterations in diagnostic strategy.
Despite these variations in absolute counts, the overall structure of antimicrobial susceptibility testing remained comparable across calendar years, reflecting stable laboratory protocols and consistent diagnostic practices. Observed year-to-year differences were therefore primarily driven by changes in sample throughput and clinical activity rather than by methodological shifts.
Over the same period, hospital activity—expressed as the annual number of completed hospitalizations—showed a pronounced decline after 2019 and remained consistently lower throughout 2020–2024 across both non-surgical and surgical departments (
Figure 2B). The most substantial decrease occurred between 2019 and 2020, followed by a prolonged period of reduced but relatively stable hospitalization numbers. Non-surgical departments consistently accounted for the majority of hospital discharges, while surgical departments exhibited a parallel trend at substantially lower absolute volumes.
Given these substantial fluctuations in hospital activity over time, all resistance incidence metrics presented in subsequent analyses were normalized to 1,000 completed hospitalizations, enabling robust temporal comparisons and meaningful evaluation of differences between surgical and non-surgical departments independent of underlying changes in patient throughput.
2.3. Incidence of Staphylococcus aureus Stratified by Resistance Phenotype, Specimen Type, and Department
Figure 3 and
Figure 4 summarize the temporal trends in the incidence
of Staphylococcus aureus normalized to 1,000 completed hospitalizations, stratified by resistance phenotype (MRSA vs. MSSA), multidrug resistance status, specimen classification (colonization vs. infection), and department category (surgical vs. non-surgical). Across all panels, incidence rates differed substantially between resistance phenotypes and clinical settings, with consistently higher values observed in surgical departments compared with non-surgical departments and marked differences between colonization and infection dynamics.
2.3.1. MRSA Incidence Trends
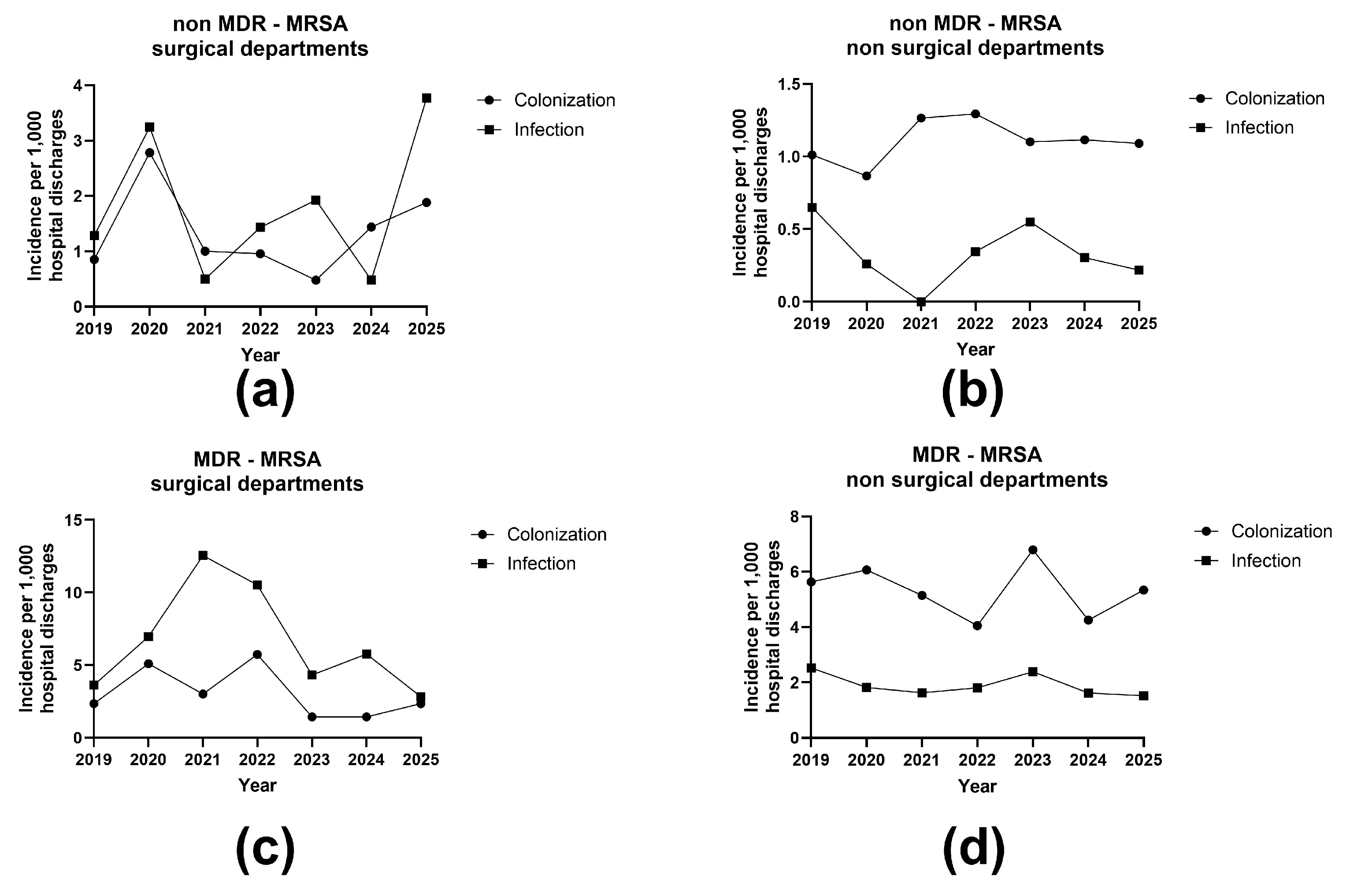
As shown in
Figure 3, MRSA incidence exhibited pronounced temporal variability and clear differences between surgical and non-surgical departments. In surgical departments, MDR MRSA incidence was consistently higher than non-MDR MRSA, particularly for infections, with peak infection incidence observed between 2020 and 2022. Following this period, MDR MRSA infection incidence declined markedly in 2023 and 2024, with a slight increase again in 2025. Colonization incidence in surgical departments followed a broadly similar pattern but remained consistently lower than infection incidence.
In non-surgical departments, overall MRSA incidence was substantially lower than in surgical settings. MDR MRSA colonization predominated over infection throughout the study period, with relatively stable colonization rates and lower, fluctuating infection incidence. Non-MDR MRSA in non-surgical departments showed low and relatively stable incidence for both colonization and infection, with only minor year-to-year variation.
Across both department categories, MRSA infection incidence demonstrated greater temporal fluctuation than colonization, while surgical departments consistently exhibited higher incidence rates than non-surgical departments for both MDR and non-MDR MRSA.
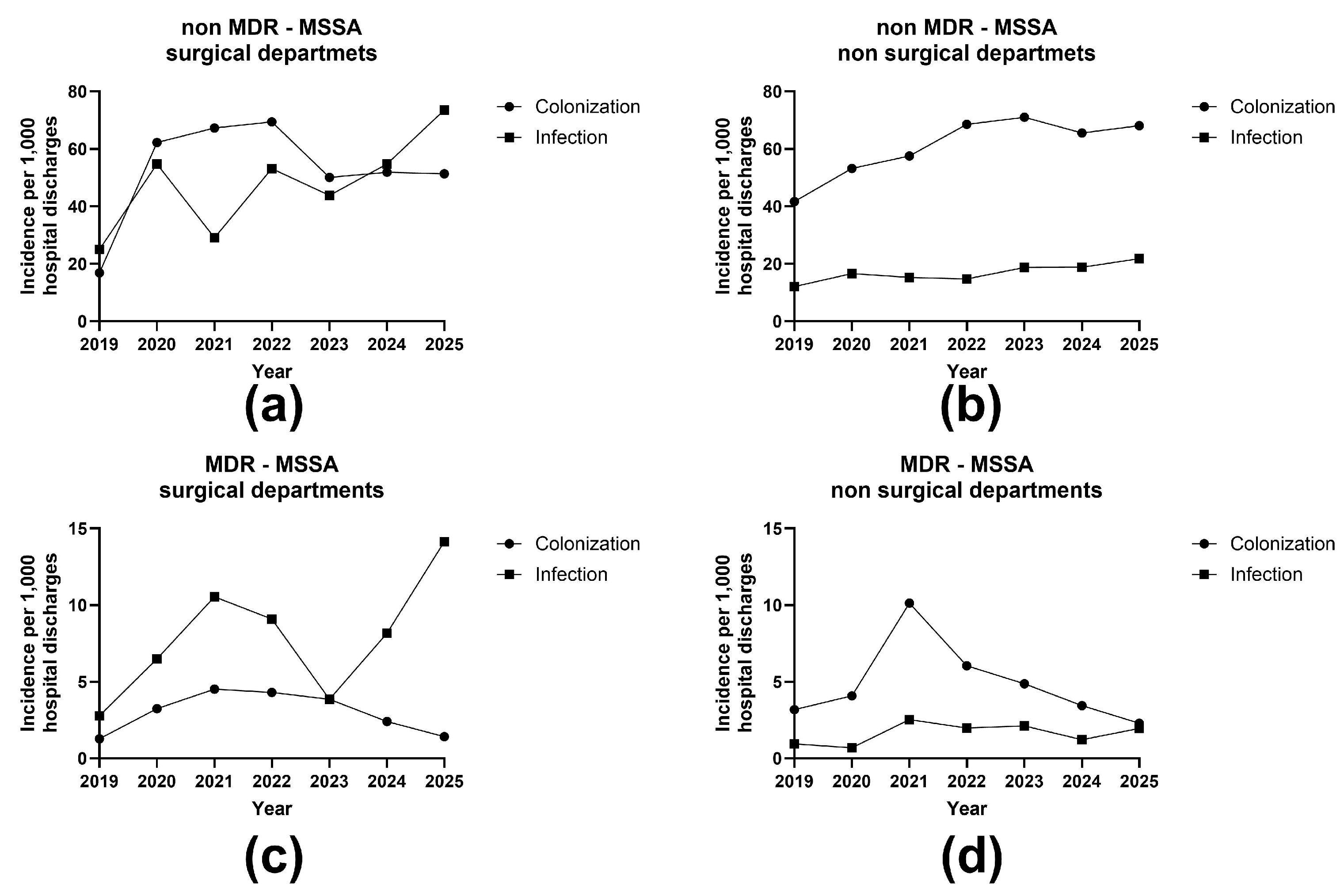
2.3.2. MSSA Incidence Trends
Figure 4 illustrates markedly higher overall incidence rates for MSSA compared with MRSA across all strata. In surgical departments, non-MDR MSSA colonization showed high and relatively stable incidence throughout the study period, while infection incidence displayed greater variability, with a notable increase in 2025. MDR MSSA incidence in surgical departments was substantially lower than non-MDR MSSA but demonstrated pronounced fluctuations over time, particularly for infections, which increased again toward the end of the study period.
In non-surgical departments, non-MDR MSSA colonization incidence increased steadily from 2019 through 2023 and remained elevated thereafter, whereas infection incidence remained consistently lower but showed a gradual upward trend over time. MDR MSSA incidence in non-surgical departments was low for both colonization and infection, with a transient peak in colonization around 2021 followed by a gradual decline in subsequent years.
Overall, MSSA incidence was dominated by colonization rather than infection, particularly in non-surgical departments, and by non-MDR strains across both department categories. Surgical departments consistently exhibited higher MSSA incidence than non-surgical departments, with greater temporal variability observed for infection compared with colonization.
3. Discussion
This study reveals a sustained increase in the incidence of multidrug-resistant
Staphylococcus aureus (MDR
S. aureus), including methicillin-resistant
S. aureus (MRSA), that began during the COVID-19 pandemic and continued into the post-pandemic period. Notably, this increase persisted beyond the acute phases of the pandemic, suggesting that the effects of COVID-19 on hospital epidemiology and antimicrobial resistance (AMR) dynamics were not short-lived. While much of the previous research has concentrated on the early stages of the pandemic, our findings indicate that the pandemic’s impact on MDR
S. aureus and MRSA may endure for years, underscoring their long-term clinical and organizational relevance [
26,
27,
28].
The data highlights a significant and prolonged rise in MDR
S. aureus incidence, especially MRSA, which began during the COVID-19 pandemic and has persisted post-pandemic. MRSA bacteremia and other healthcare-associated infections (HAIs) notably increased in 2020 compared to pre-pandemic levels, with a marked surge in cases across U.S. hospitals during the pandemic’s peak. This rise was strongly associated with the overwhelming burden of COVID-19 hospitalizations. Similarly, other research has reported a substantial link between the COVID-19 surge and increased MRSA and other multidrug-resistant pathogens, such as vancomycin-resistant enterococci (VRE). While earlier studies predominantly focused on the acute pandemic period, our analysis suggests that the effects of COVID-19 on antimicrobial resistance have extended far beyond that phase. In particular, MDR
S. aureus infections are expected to remain a significant concern for healthcare systems. These findings stress the necessity for continued vigilance and adaptation of infection control measures to combat the ongoing challenge of AMR [
26,
28].
Our results show that the increase in MDR
S. aureus (including MRSA) did not revert to pre-pandemic baseline levels immediately following the resolution of major COVID-19 waves. These extended temporal trends support the hypothesis that disruptions to infection control practices and hospital ecosystems during the pandemic could have long-lasting effects on pathogen dynamics, even after broad infection control measures have been relaxed or adjusted. Notably, retrospective studies show significant shifts in hospital-acquired infection patterns and MDRO rates during the COVID-19 period compared to previous years, including a reduction in MRSA rates during the pandemic compared with pre-pandemic levels [
29]. Furthermore, the infection prevention measures introduced during the pandemic had varying effects on HAIs across different settings and over time, emphasizing that the broader epidemiological impact of COVID-19-related measures extends beyond short-term fluctuations and can influence HAI ecology in complex ways [
30].
The observed increase in MDR
S. aureus, particularly MRSA, aligns with the strain placed on healthcare systems during the pandemic, which likely contributed to higher infection rates. With hospitals overwhelmed by surges of critically ill patients requiring extended hospitalization and intensive care, the risk of healthcare-associated infections (HAIs)—especially those caused by contact-transmitted pathogens like MRSA—increased significantly. The incidence of MRSA surged during the pandemic, particularly in intensive care settings. This intense healthcare burden likely led to compromised infection prevention and control (IPC) measures, further contributing to transmission, especially in environments with high patient density and intensive care needs [
26,
27].
In contrast to studies that primarily focus on the early pandemic period, our findings demonstrate that the elevated incidence of MDR
S. aureus and MRSA continued even after the acute COVID-19 waves subsided. This reinforces the idea that the pandemic caused long-lasting disruptions in hospital microbial ecology. Despite efforts to return to routine IPC practices, factors such as workforce shortages and delayed recovery in antimicrobial stewardship programs have likely played a role in the sustained higher incidence of MRSA. Our data suggest that the effects of the pandemic on MRSA will continue into the post-pandemic period, highlighting the need for ongoing vigilance in infection control and antimicrobial resistance management [
31].
The reorganization of healthcare during the COVID-19 pandemic further exacerbated existing vulnerabilities in infection prevention and control (IPC). The rapid establishment of COVID-19 units, extensive patient cohorting, and the redeployment of healthcare workers across departments disrupted established workflows and reduced local expertise in IPC. Although enhanced use of personal protective equipment (PPE) was effective in controlling SARS-CoV-2 transmission, these measures were primarily designed to prevent respiratory spread and did not necessarily improve control over contact-transmitted pathogens like MRSA. In many healthcare settings, resources for IPC and isolation capacity were diverted to COVID-19 patients, potentially diminishing the effectiveness of MRSA-focused prevention strategies [
32]. This trend was evident in our study, which observed a significant increase in MDR
S. aureus incidence in the aftermath of the pandemic’s acute phases.
Antibiotic prescribing practices during the pandemic also played a crucial role in shaping the dynamics of antimicrobial resistance (AMR). National and international guidelines consistently recommended that antibiotics should be reserved for patients with clear evidence of bacterial co-infection, with the aim of avoiding prophylactic use in COVID-19 cases [
33]. Despite these recommendations, substantial overuse of antibiotics was documented, particularly in the early stages of the pandemic. This was likely due to diagnostic uncertainty, the severity of disease, and the overwhelming clinical pressure faced by healthcare teams, all of which led to an increase in unnecessary antibiotic prescriptions. Our findings align with this, showing that such antibiotic overuse likely exerted selective pressure on
S. aureus populations, promoting the persistence of multidrug-resistant strains. The widespread use of antibiotics during this period may have contributed to the sustained elevated incidence of MRSA, which is expected to remain a significant issue for healthcare settings long after the resolution of the pandemic [
34,
35].
The widespread antibiotic exposure during the COVID-19 pandemic likely exerted substantial selective pressure, favoring the persistence of multidrug-resistant
S. aureus (MDR
S. aureus) and methicillin-resistant
S. aureus (MRSA). This antibiotic pressure may have facilitated prolonged colonization by resistant strains, increasing the risk of progression from colonization to infection, especially in patients with extended hospital stays, invasive devices, and severe illness. These effects are unlikely to resolve quickly, even after antibiotic prescribing practices return to pre-pandemic norms, providing a plausible explanation for the continued high incidence of MDR
S. aureus and MRSA observed into the post-pandemic period [
36].
While many healthcare systems have attempted to reinstate routine infection prevention and control (IPC) and antimicrobial stewardship activities following the acute phases of the pandemic, the recovery has been uneven and delayed. Workforce shortages, staff fatigue, and backlogs of deferred care have persisted, contributing to the ongoing challenges in infection control. In line with these observations, large surveillance studies have shown that hospital-onset MRSA bloodstream infections remained elevated beyond the first year of the pandemic [
36,
37]. This supports the idea that the disruptions caused by the pandemic may have long-lasting consequences for the epidemiology of antimicrobial resistance, including sustained MRSA burden.
Hospital-acquired MRSA infections have significant clinical and organizational consequences. As with previous studies, our analysis shows that MRSA infections are associated with increased morbidity and mortality, extended hospital stays, greater utilization of last-line antimicrobial agents, and elevated healthcare costs compared to infections caused by methicillin-susceptible
S. aureus (MSSA) [
38,
39]. Additionally, at the institutional level, sustained increases in MRSA burden can strain patient flow, necessitate enhanced isolation measures, and consume limited hospital resources. These issues are especially impactful during periods of healthcare system recovery, such as post-crisis or pandemic situations [
39].
Our findings confirm that hospital-specific surveillance is essential to detecting and addressing localized increases in MRSA incidence that may not be reflected in broader population-level surveillance. Despite the general decline in MRSA bloodstream infections reported across Europe in recent years, as highlighted in European surveillance data [
40], our study observed more significant fluctuations in MRSA trends at the hospital level. This underscores the importance of interpreting hospital-level data with a tailored focus on healthcare activity, as broader population metrics may fail to capture critical localized patterns. By standardizing surveillance to hospital activity, our analysis reveals emerging vulnerabilities in care delivery that aggregate population-level data might obscure, further emphasizing the necessity of targeted institutional surveillance to guide effective infection control strategies.
4. Materials and Methods
Data Source
The primary data source was comprised records from the laboratory information system (LIS). These unstructured diagnostic data included microorganism identification by MALDI TOF-MS method and antibiotic susceptibility testing. All results were interpreted in accordance with the latest EUCAST guidelines. The unstructured records were transformed into structured data using the Ameba software.
Database
The data used in this study originated from the expert medical information system AMEBA, which has been operational since 2018 and is actively used by major university hospitals in Bratislava and Martin, Slovakia. AMEBA is an advanced software platform designed for real-time integration, interpretation, and visualization of microbiological laboratory results. It enables interdisciplinary collaboration among hospital epidemiologists, clinical pharmacologists, and treating physicians by offering consolidated overviews, resistance reports, and automated alerts on relevant findings.
The system collects unstructured diagnostic data from microbiological laboratories (including MALDI-TOF MS identification and antibiotic susceptibility tests), which are then processed using machine learning algorithms to extract relevant information. The structured data is stored securely in a cloud-based Microsoft Azure environment, enabling high availability and scalability for multi-center epidemiological analyses.
Interpretation of laboratory results is aligned with current EUCAST criteria and further enhanced by referencing national eHealth codelists (číselníky národného systému eZdravie). Data processing pipelines include filtering department identifiers (P codes), transformation into structured form, and deduplication routines. All data were collected and maintained in Slovak and English to enable consistency in international reporting and future interoperability.
Data Cleaning & Preprocessing
Microbiological datasets were processed using the Konstanz Information Miner (KNIME) Analytics Platform (v5.4.3). Data integration pipelines were developed to harmonize sample metadata, antimicrobial susceptibility profiles, and hospital department identifiers across a five-year retrospective window. Preprocessing included dataset merging, standardization of susceptibility interpretations according to EUCAST criteria, classification of ward types via P codes, and consistency checks for duplicate or incomplete records. All KNIME workflows used in this phase are provided as Supplementary Material (File S1) to ensure full reproducibility.
Classification of isolates as originating from surgical versus non-surgical wards, as well as the interpretation of results in the context of infection versus colonization, were based on definitions and operational criteria established by a multidisciplinary team of infectious disease specialists and hospital epidemiologists. These definitions reflect routine clinical and epidemiological practice within the institution. Given the retrospective design, the study deliberately leveraged available real-world data to evaluate their suitability and validity for longitudinal surveillance and epidemiological analyses.
Data Analysis and Visualization
The processed dataset was analyzed using KNIME through a series of visual and statistical workflow components. The core analyses focused on comparing counts of multiresistant versus susceptible Staphylococcus isolates over time, stratified by predefined time periods). Aggregation and grouping operations were applied to compute frequency distributions, followed by visualization using categorical bar plots to illustrate temporal trends.
Visual outputs were generated directly within KNIME using bar chart nodes configured with “TimePeriod” as the category axis and multiple frequency columns (e.g., “Multiresistant+Count(Pathogen)”, “Susceptible+Count(Pathogen)”). These plots supported rapid interpretation of resistance evolution across clinical departments. All workflow logic, including node configurations and output mappings, is available as Supplementary Material (File S1) to ensure full reproducibility and transparency.