Submitted:
03 March 2026
Posted:
04 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. DNA Extraction
2.3. Primer Design and Synthesis
2.4. Primer Screening
- Preliminary screening: Primers were initially tested via 2% agarose gel electrophoresis using genomic DNA from 8 randomly selected mandarin accessions. Each PCR reaction was performed in duplicate to verify reproducibility. Primers that yielded clear, non-smearing, and reproducible bands were selected for secondary screening.
- Secondary screening: Candidate primers from the preliminary step were further evaluated using 6% denaturing polyacrylamide gel electrophoresis (PAGE) to assess polymorphism. Each reaction was repeated twice, and primers exhibiting distinct polymorphic bands among the test accessions were retained.
- Final validation: Primers with satisfactory performance in secondary screening were subjected to final validation using SSR fluorescent capillary electrophoresis (FCE). This step confirmed their polymorphism, amplicon stability, and suitability for large-scale fingerprinting analysis of the 69 mandarin accessions.
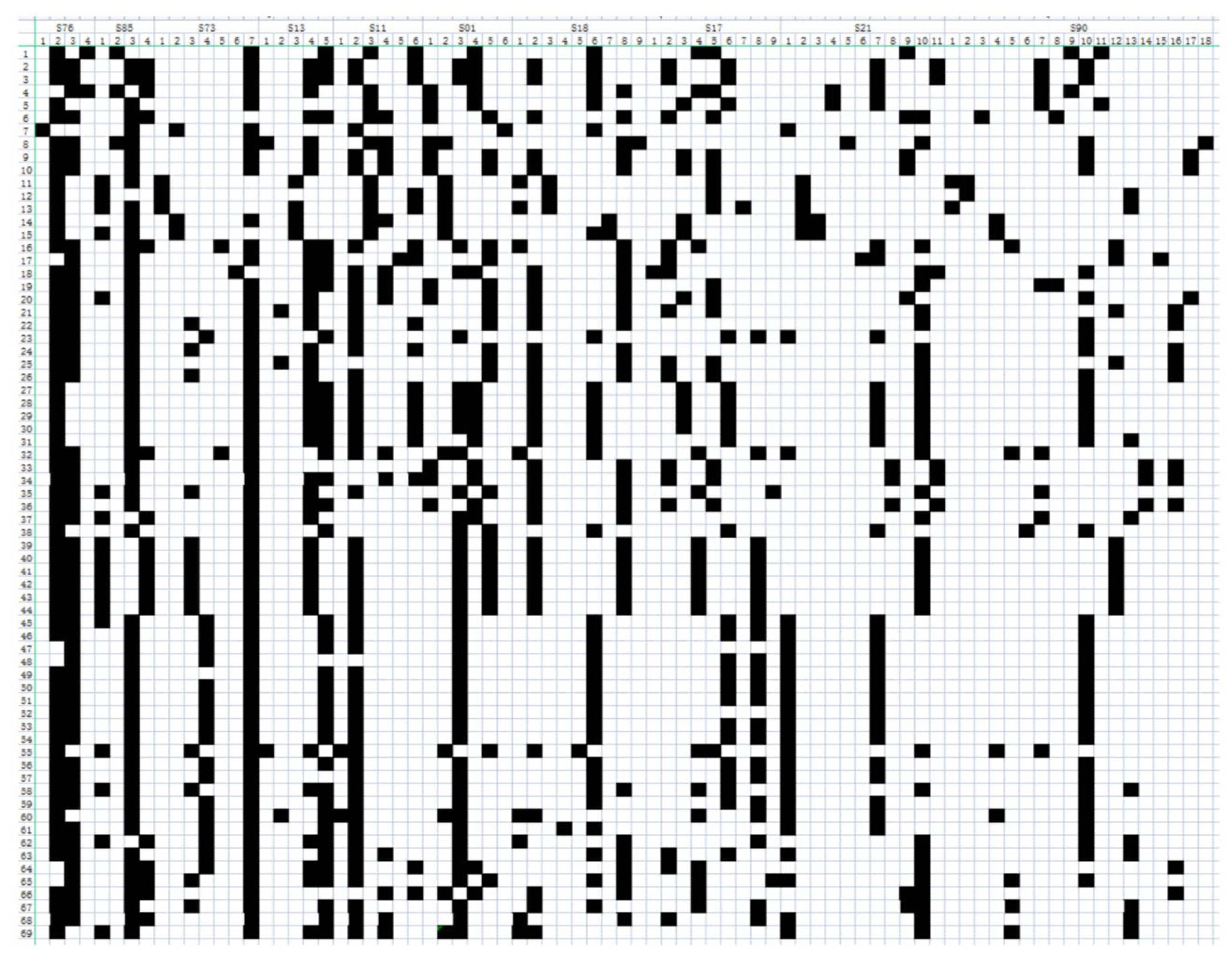
2.5. Fingerprint Construction
2.6. Molecular Identity Card Construction
- For each primer pair, the fragment sizes of amplified alleles were first sorted in ascending order;
- Unique band patterns (genotypes) among the 69 accessions were encoded sequentially using Arabic numerals 1–9;
- When the number of unique band patterns exceeded 9, uppercase English letters (A, B, C,…) were used to represent the 10th, 11th, 12th, and subsequent patterns;
- Null alleles (no amplification products) were denoted as "0".
3. Results and Analysis
3.1. Primer Design and Screening
3.2. Genetic Diversity Analysis of 69 Mandarin Accessions
3.2.1. Polymorphism Evaluation of SSR Primers
3.2.2. SSR Characteristic Fingerprint Information of 69 Mandarin Varieties
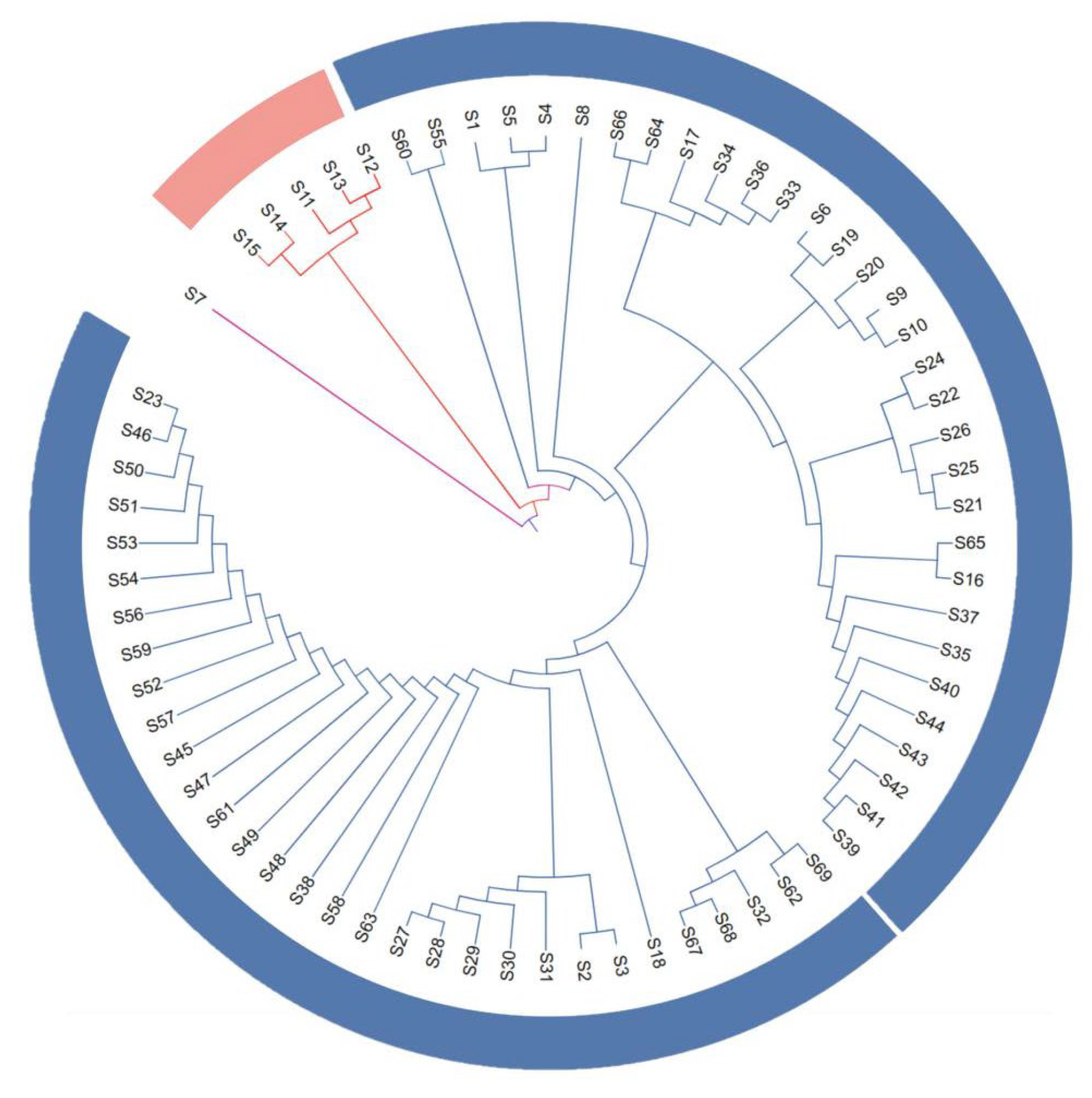
3.2.3. Cluster Analysis
3.3. Construction of DNA Fingerprint and Molecular Identity Card
3.3.1. DNA Fingerprint Construction
3.3.2. Varietal Discrimination Using Core Primer Combinations
3.3.3. Molecular Identity Card Construction
4. Discussion
4.1. Genetic Diversity of Mandarin Germplasms
4.2. Insights from Cluster Analysis
- ‘Yinduyeju’ formed a distinct single-accession group. Previous studies have debated its taxonomic status: Swingle [29] proposed it is a hybrid, while Yang et al. [30] clustered it with true citrons based on cpInDel markers, and Li et al. [31] suggested it is a primitive mandarin species. In contrast, ‘Mangshanyeju’ and ‘Mangshan wild mandarin’ did not cluster together, indicating a distant genetic relationship—consistent with the view that ‘Mangshan wild mandarin’ is more primitive than ‘Mangshanyeju’. Additionally, Tachibana orange (native to Taiwan, China, and Japan) did not cluster with ‘Yinduyeju’ and was separated from other cultivated and wild accessions, which contradicts the findings of Xie et al. [32]. This discrepancy may be attributed to material errors during multi-location transfers, requiring further verification with authenticated germplasms.
- ‘Mangshanyeju’ clustered with ‘Nieduyeju’ but separated from ‘Guposhanyeju’, indicating a closer genetic relationship between the former two. Liu et al. [33] identified ‘Mangshanyeju’ as the most ancient type among five wild mandarin types distributed in the Lingnan Mountains using SSR markers, and Zeng et al. [34] also considered it more primitive than Tachibana orange and ‘Daoxianyeju’. Shi [1989] noted high similarity between ‘Guposhanyeju’ and ‘Daoxianyeju’, the latter of which is regarded as a progenitor of mandarins due to its distant genetic relationship with most cultivated accessions [34].
- ‘Guposhanchougan’ clustered with ‘Mangshanyegan’ (both pointed-leaf and round-leaf types), consistent with the pollen morphology-based clustering results of Wu et al. [3]. ‘Cenxisuanju’, ‘Guangxihongpisuanju’, and ‘Hongpisuanju’ formed a distinct subgroup, separate from ‘Hezhouyeju’ and ‘Huangpisuanju’—indicating genetic relatedness among the former three, which aligns with molecular marker-based clustering by Liu et al. [35]. ‘Hezhouyeju’ and ‘Huangpisuanju’ co-clustered, suggesting a potential genetic relationship that requires further validation via genomic analyses. In contrast, ‘Shagan’ and ‘Biangan’ did not cluster together, which contradicts Wu et al.’s [3] pollen morphology results, necessitating additional research to resolve this inconsistency.
- Accessions with highly similar genetic backgrounds (e.g., bud sports) clustered closely, which is consistent with previous studies: ‘Shatangju’, early-ripening ‘Shatangju’, ‘Bayueju’, ‘Denglongju’, and ‘Jinkui tangerine’ co-grouped, matching Yan et al.’s [36] SRAP marker clustering; ‘Chuntian tangerine’, ‘Mingliutianju’, and ‘Yingxinju’ clustered with ‘Biangan’, while ‘Shagan’ and ‘Gonggan’ formed a separate subgroup; ‘Guangxiju’ clustered with Ponkan, and ‘Wogan’ grouped with ‘W. Murcott’—all suggesting potential kinship.
- Wild-cultivated germplasm relationships were also revealed: Group I contained Indian wild mandarin as a distinct lineage; Group II included wild wrinkled-skin mandarin and ‘Mangshan wild mandarin’, consistent with Wu et al. [3]; Group III comprised ‘Guposhan wild Yuanju’, ‘Mangshan wild mandarin’, ‘Niedu wild mandarin’, ‘Shengshan wild mandarin’, and the hybrid ‘Tsunoka’—the latter two may carry wild genetic components. ‘Hezhou wild mandarin’ and ‘Biangan’ co-clustered with ‘Shatangju’ and Ponkan, demonstrating kinship. However, definitive cultivated-wild relationships require advanced genomic sequencing.
4.3. Value of DNA Fingerprints and Molecular Identity Cards
4.4. Limitations and Future Perspectives
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Shen, Y.G.; Zhu, F.N.; Lu, J.Q.; Chen, J.Y.; Zhou, M.; Wang, Q.; Yang, H.Y. Advances in research on quality characters and processing eligibility of Citrus reticulata. Acta Agriculturae Universitatis Jiangxiensis 2017, 39(04), 669–677. [Google Scholar]
- Wu, X.X.; Chen, C.W.; Liu, P.; Tang, Y.; Deng, C.L. Genetic Evolution and Taxonomic Status Analysis of Wild Citrus Resources Based on Resequencing in Guposhan Mountain in Guangxi Province. Acta Horticulturae Sinica 2022, 49(02), 407–415. [Google Scholar]
- Wu, X.X. Analysis of genetic Diversity of Native Citrus Germplasm Resources in Guangxi Province by Palynology and SNP; Guangxi normal university: Guangxi, 2018. [Google Scholar]
- Green, R.M.; Vardi, A.; Galun, E. The plastome of citrus physical map, variation among Citrus cultivars and species and comparison with related genera. Theoretical and Applied Genetics 1986, 72(2), 170–177. [Google Scholar] [CrossRef]
- Leng, X.P.; Li, H.R.; Zhong, G.Y. Employment of a new strategy for identification of looseskin mandarin (Citrus reticulata Blanco) cultivars using RAPD markers. Romanian Biotechnological Letters 2012, 17(2), 7073–7083. [Google Scholar]
- Yu, Q.M.; Li, G.G.; Xu, R.W.; Peng, Z.X.; Yuan, Z.Y.; Li, W.Y.; Tian, J.; Zeng, J.W.; Peng, S.A.; Xu, J. Physiological Mechanisms for the Phenomenon of“Blue albedo”Fruits of Citrus reticulata in Guangxi. Acta Horticulturae Sinica 2020, 47(06), 1172–1182. [Google Scholar]
- Zhang, S.l.; Li, Y.; Li, Y.; Zhang, Y.Q.; Hao, Y.B.; Hou, Z.X.; Qi, J.X. Development of SSR Markers for and Fingerprinting of Walnut Genetic Resources. Forests 2024, Vol.15(3), 405. [Google Scholar] [CrossRef]
- Snježana, B.; Aleš, V.; Kristina, B.L.; Đani, B. Genotyping of Croatian Olive Germplasm with Consensus SSR Markers. Horticulturae 2024, Vol.10(4), 417. [Google Scholar]
- Cao, Z.; Song, C.R.; Chen, D.; Chen, X.F. Analysis of Genetic Diversity of Cherry Germplasm Resources in Shandong Yantai Based on SSR Technology. Guangdong Agricultural Sciences 2025, 52(1), 24–36. [Google Scholar]
- Liu, X.F.; Yu, B.; Ren, G.P.; Yu, C.R.; Liu, C.; Sun, Y.B.; Zhong, R.H.; Feng, E.Y. Construction of Fingerprint Map of 12 Varieties Based on SSR Markers and Identification of Hybrid F1 Progenies in Gerbera hybrida. Guangdong Agricultural Sciences 2023, 50(9), 16–24. [Google Scholar]
- Nie, X.H.; Wang, Z.H.; Liu, N.W.; Song, L.; Yan, B.Q.; Xing, Y.; Zhang, Q.; Fang, K.F.; Zhao, Y.L.; Chen, X.; et al. Fingerprinting 146 Chinese chestnut (Castanea mollissima Blume) accessions and selecting a core collection using SSR markers. Journal of Integrative Agriculture 2021, 20(5), 1277–1286. [Google Scholar] [CrossRef]
- Hao, L.; Zhai, Y.G.; Zhang, G.S.; Lu, D.Y.; Huang, H.G. Efficient Fingerprinting of the Tetraploid Salix psammophila Using SSR Markers. Forests 2020, 11(2), 176. [Google Scholar] [CrossRef]
- Li, X.; Zheng, B.; Xu, W.T.; Ma, X.W.; Wang, S.B.; Qian, M.J.; Wu, H.X. Identification of F1 Hybrid Progenies in Mango Based on Fluorescent SSR Markers. Horticulturae 2022, 8(1122), 1122. [Google Scholar] [CrossRef]
- Wu, X.X.; Wu, S.M.; Chen, C.W.; Lou, B.H.; Tang, Y.; Feng, J.; Fang, H.M.; Deng, C.L. Research progress on fingerprinting of citrus. China Fruits 2024, 07, 16–24. [Google Scholar]
- Wu, S.M.; Lou, B.H.; Chen, C.W.; Tang, Y.; Deng, C.L.; Wu, X.X. Establishment of molecular identity of 22 pomelo varieties using fluorescent labeled SSR markers. Journal of Fruit Science 2023, 40(04), 605–614. [Google Scholar]
- Rohini, M.R.; Sankaran, M.; Rajkumar, S.; Prakash, K.; Gaikwad, A.; Chaudhury, R.; Malik, S.K. Morphological characterization and analysis of genetic diversity and population structure in citrus × jambhiri Lush using SSR markers. Genetic Resources and Crop Evolution 2020, 67(1), 1259–1275. [Google Scholar] [CrossRef]
- Yan, W.; Youheng, Q.; Wen, H. Genetic diversity of pummelo (Citrus grandis osbeck) germplasms in sichuan Basin inferred from SSR markers. AIP Conference Proceedings 2019, 2079(1), 02004-1-02004-6. [Google Scholar]
- Lidija, B.; Slavojka, M.; Natasa, S.; Teija, T.R.; Branka, J. Identification of citruses from montenegrobased on microsatellite clustering analyses. Erwerbs Obstbau 2020, 62(3), 1–8. [Google Scholar]
- Fang, D. DNA fingerprinting inspection technique for citrus nursery tree purity and Genuineness; Southwest University: Chongqing, 2011. [Google Scholar]
- Zeng, T. Construction of Fingerprinting and Analysis of Genetic Diversity with SSR Markers for Nanfeng Tangerine; Jiangxi agricultural university, 2012. [Google Scholar]
- Lei, T.G.; He, Y.R.; Wu, X.; Yao, L.X.; Peng, A.H.; Xu, L.Z.; Liu, X.F.; Chen, C.S. Construction of DNA Fingerprinting Database of Citrus Cultivars (Lines). Scientia Agricultura Sinica 2009, 42(08), 2852–2861. [Google Scholar]
- Li, Y.; Ma, X.F.; Tang, H.; Li, N.; Jiang, D.; Long, G.Y.; Li, D.Z.; Niu, Y.; Han, R.X.; Deng, Z.N. SSR Markers Screening for Identification of Citrus Cultivar and Construction of DNA Fingerprinting Library. Scientia Agricultura Sinica 2018, 51(15), 149–159. [Google Scholar]
- Biswas, M.K.; Xu, Q.; Mayer, C.; Deng, X.X. Genome wide characterization of short tandem repeat markers in sweet orange (Citrus sinensis). PloS One 2014, 9(8), 1–12. [Google Scholar] [CrossRef]
- Chen, Y.; Yu, X.; Cao, L.; LeiT, G.; Zhou, X.; Zhang, X.Y.; Peng, L.Z.; Lu, Z.M. Identification of CRIC 5 by using SSR Molecular markers. South China Fruits 2018, 47(03), 1–4. [Google Scholar]
- Hong, L.; Andrew, W.M. Extracting DNA from cambium tissue for analysis of grape rootstocks. HortScience 1997, 32(7), 1264–1266. [Google Scholar]
- Jason, D.Z.; April, N.; Sara, M.; Joseph, P.; David, N.; Nahla, B. A new SSR fingerprinting set and its comparison to existing SSR and SNP based genotyping platforms to manage Pyrus germplasm resources. Tree Genetics and Genomes 2020, 16(5), 1–10. [Google Scholar]
- Raja, W.H.; Yousuf, N.; Qureshi, I. Morpho-molecular characterization and genetic diversity analysis across wild apple (Malus baccata) accessions using simple sequence repeat markers. South African Journal of Botany 2022, 145, 378–385. [Google Scholar] [CrossRef]
- Li, Y.K.; Zhang, P.X.; Chachar, S.; Xu, J.C.; Y, Y.; Guang, C.F. A comprehensive evaluationof genetic diversity in persimmon (Diospyros kaki Thunb.) germplasms based on largescale morphological traits and SSR markers. Scientia Horticulturae 2023, 313, 111866. [Google Scholar] [CrossRef]
- Yan, J.W.; Wu, X.X.; Tang, Y.; Chen, C.W.; Deng, C.L. Analysis of Genetic Diversity of 11 Citrus Germplasm Resources by SRAP Molecular Markers. Molecular Plant Breeding 2021, 19(02), 664–671. [Google Scholar]
- Swingle, W.T.; Reece, P.C. The botany of citrus and its wild relatives; University of California, 1967. [Google Scholar]
- Wang, J.; Gong, G.Z.; Peng, Z.C.; Li, Y.B.; Wang, Y.B.; Hong, Q.B. Genetic and Phylogenetic Relationships Among Citrus and Its Close and Distant Relatives Based on COS Marker. Scientia Agricultura Sinica 2017, 50(02), 320–331. [Google Scholar]
- Li, Y.Z. Research on the genetic diversity and phylogenetic relationship of loose skin mandarins(Citrus reticulate Blanco); Huazhong Agricultural University, 2006. [Google Scholar]
- Xie, R.J. Taxonomic and phylogenetic relationships among the genera of the true citrus fruit trees group(Aurantioideae, Rutaceae),based on AFLP markers; Soutewest University, 2008. [Google Scholar]
- Liu, Y.; Wu, B.; Liu, D.C.; Sun, Z.H. On genetic diversity of Jiangxi native citrus and its wild varieties based on SSR markers. Acta Agriculturae Universitatis Jiangxiensis 2005, 04, 486–490. [Google Scholar]
- Liu, T.; Deng, C.L.; Cheng, Y.F.; Liu, Q.J.; Chen, C.W.; Liu, B.H.; Yi, H.L. Analyzing genetic diversity of citrus germplasm in Guangxi Province with SSR and SRAP markers. 2016; 35, 02. [Google Scholar]
- Zeng, B.Q. Genetic diversity of mandarin investigated by molecular markers; Hunan Agricultural University, 2009. [Google Scholar]
- Le, T.N.; Miyazaki, Y.; Takuno, S. Epigenetic regulation of intragenic transposable elements impacts gene transcription in Arabidopsis thaliana. Nucleic Acids Research 2015, 43(8), 3911–3921. [Google Scholar] [CrossRef]
- Ke, L.J.; Yu, H.W.; Xu, H.D.; Xie, Z.Z.; Deng, X.X.; Xu, Q. Identification the Citrus Bud Mutants by Transposon Display Technology. Acta Horticulturae Sinica 2017, 44(06), 1207–1216. [Google Scholar]
- Zhu, Y.S.; Zhang, Y.F.; Cheng, L.; Yang, S.N.; Zhao, W.T.; Jiang, D. Identification of 60 Citrus Accessions Using Target SSR-seq Technology. Scientia Agricultura Sinica 2022, 55(22), 4458–4472. [Google Scholar]
- Gao, Y.; Wang, K.; Wang, D.J.; Gong, X.; Liu, L.J.; Liu, F.Z. Molecular ID Establishment of Apple Cultivars by TP-M13-SSR. Acta Horticulturae Sinica 2016, 43(01), 25. [Google Scholar]
- Tang, Y.J.; Luo, S.X.; Huang, G.D.; Song, E.L.; Li, R.W.; Zhao, Y.; Zhang, Y.; Mo, Y.L.; Tang, Y.Y. Genetic Diversity Analysis and Molecular ID Construction of Mango Germplasm Based on SSR Fluorescence Markers. Chinese Journal of Tropical Crops 2023, 44(11), 2292–2304. [Google Scholar]

| Code | Accession name | Scientific Name | Code | Accession name | Scientific Name |
|---|---|---|---|---|---|
| 1 | Guposhanyeju | Sinocitrus chuana | 36 | Yingxinju | C. reticulata |
| 2 | Xinganyeju | C. reticulata | 37 | Pixeju | C. reticulata |
| 3 | Daoxianyeju | C. daoxianensis | 38 | Bendizao | C. reticulata |
| 4 | Mangshanyeju | C. reticulata | 39 | Penggan No79-2 | C. reticulata |
| 5 | Niuduyeju | C. reticulata | 40 | Dong No13penggan | C. reticulata |
| 6 | Hezhouyeju | C. reticulata | 41 | Shi18penggan | C. reticulata |
| 7 | Yinduyeju | C. indica | 42 | Taitianpenggan | C. reticulata |
| 8 | Lihuaju | C. tachibana | 43 | Xinshengxipenggan No3 | C. reticulata |
| 9 | Cengxisuanju | C. reticulata | 44 | Wuhepenggan | C. reticulata |
| 10 | Guangxihongpisuanju | C. reticulata | 45 | Dafen No4 | C. reticulata |
| 11 | Guposhanchougan No2 | C. reticulata | 46 | Rinan No1 | C. reticulata |
| 12 | Guposhanchougan No5 | C. reticulata | 47 | Dapu No5 | C. reticulata |
| 13 | Guposhanchougan No6 | C. reticulata | 48 | Miyamoto | C. reticulata |
| 14 | Yuanyemangshanyegan | C. mangshanesis | 49 | Miyagawa | C. reticulata |
| 15 | Jianyemangshanyegan | C. mangshanesis | 50 | Hashikawa | C. reticulata |
| 16 | Shagan | C. nobilis Lour | 51 | Xingjin | C. reticulata |
| 17 | Biangan | C. reticulata | 52 | Yoshida | C. reticulata |
| 18 | Banyeshenggan | C. reticulata | 53 | Ichibun | C. reticulata |
| 19 | Huangpisuanju | C. reticulata | 54 | Yamasitabeni | C. reticulata |
| 20 | Hongpisuanju | C. reticulata | 55 | Katsuyamano | C. reticulata |
| 21 | Shatangju | C. reticulata | 56 | Ueno | C. reticulata |
| 22 | Zaoshushatangju | C. reticulata | 57 | Dajin No4 | C. reticulata |
| 23 | Yamada | C. reticulata | 58 | Zaoxiang | C. reticulata |
| 24 | Bayueju | C. reticulata | 59 | Sakikubo | C. reticulata |
| 25 | Denglongju | C. reticulata | 60 | Jinzhixiang | C. reticulata |
| 26 | Jinkuimiju | C. reticulata | 61 | Youliang | C. reticulata |
| 27 | Nanfengmiju1 | C. reticulata | 62 | Chunjian | C. reticulata |
| 28 | Nanfengmiju2 | C. reticulata | 63 | Nanxiang | C. reticulata |
| 29 | Tezaoshumiju | C. reticulata | 64 | Murcott | C. reticulata |
| 30 | Liuchengmiju | C. reticulata | 65 | Gonggan | C. reticulata |
| 31 | Guijuyihao | C. reticulata | 66 | Wogan | C. reticulata |
| 32 | Clementine | C. reticulata | 67 | Huangmeiren | C. reticulata |
| 33 | Mingliutianju | C. reticulata | 68 | Aiyuan No38 | C. reticulata |
| 34 | Chuntianju | C. reticulata | 69 | Mingrijian | C. reticulata |
| 35 | Guangxiju | C. reticulata |
| Primer | Amplified Bands | Na | Ne | I | Ho | He | PIC | Size Range(bp) |
|---|---|---|---|---|---|---|---|---|
| S17 | 19 | 9 | 6.089 | 1.884 | 0.833 | 0.836 | 0.814 | 166~200 |
| S81 | 15 | 10 | 5.678 | 1.916 | 0.507 | 0.824 | 0.802 | 167~192 |
| S90 | 30 | 18 | 4.784 | 2.109 | 0.529 | 0.791 | 0.776 | 148~202 |
| S21 | 19 | 11 | 4.535 | 1.780 | 0.638 | 0.78 | 0.751 | 244~269 |
| S70 | 18 | 14 | 4.411 | 1.854 | 0.783 | 0.773 | 0.747 | 218~248 |
| S01 | 13 | 6 | 3.492 | 1.462 | 0.493 | 0.714 | 0.677 | 282~307 |
| S11 | 10 | 6 | 2.346 | 1.16 | 0.492 | 0.574 | 0.539 | 172~191 |
| S13 | 8 | 6 | 2.537 | 1.111 | 0.381 | 0.606 | 0.527 | 230~251 |
| S28 | 9 | 6 | 2.836 | 1.239 | 0.319 | 0.647 | 0.582 | 308~322 |
| S18 | 15 | 9 | 3.749 | 1.517 | 0.681 | 0.733 | 0.689 | 230~249 |
| S23 | 12 | 8 | 2.260 | 1.248 | 0.435 | 0.558 | 0.535 | 139~189 |
| S71 | 13 | 6 | 3.614 | 1.438 | 0.681 | 0.723 | 0.681 | 211~237 |
| S73 | 9 | 7 | 2.130 | 1.141 | 0.529 | 0.53 | 0.503 | 344~369 |
| S76 | 6 | 4 | 2.073 | 0.81 | 0.696 | 0.518 | 0.411 | 168~186 |
| S74 | 8 | 6 | 2.383 | 1.151 | 0.338 | 0.58 | 0.538 | 182~203 |
| S82 | 17 | 9 | 3.084 | 1.437 | 0.368 | 0.676 | 0.622 | 122~158 |
| S84 | 10 | 8 | 2.557 | 1.225 | 0.471 | 0.609 | 0.561 | 265~292 |
| S85 | 8 | 4 | 1.883 | 0.889 | 0.406 | 0.469 | 0.432 | 125~137 |
| Primer | Number | Accession Name | Idiotype | Primer | Number | Accession Name | Idiotype | Primer | Number | Accession Name | Idiotype |
|---|---|---|---|---|---|---|---|---|---|---|---|
| S01 | 2 | Yinduyeju | 307/307 | S70 | 7 | Guposhanyeju | 230/232 | S84 | 3 | Guposhanchougan No5 | 170/170 |
| Lihuaju | 282/283 | Hezhouyeju | 226/240 | Yuanyemangshanyegan | 279/279 | ||||||
| S11 | 2 | Yinduyeju | 179/179 | Yinduyeju | 228/228 | Jianyemangshanyegan | 277/285 | ||||
| Biangan | 188/191 | Lihuaju | 224/224 | S90 | 19 | Guposhanyeju | 165/171 | ||||
| S13 | 2 | Lihuaju | 230/245 | Huangpisuanju | 234/248 | Mangshanyeju | 161/165 | ||||
| Yinduyeju | 251/251 | Guijuyihao | 230/244 | Nieduyeju | 161/171 | ||||||
| S17 | 5 | Biangan | 170/170 | Aiyuan No38 | 220/226 | Hezhouyeju | 152/164 | ||||
| Banyeshenggan | 166/170 | S71 | 4 | Mangshanyeju | 219/234 | Lihuaju | 166/202 | ||||
| Zaoxiang | 180/184 | Yinduyeju | 211/237 | Guposhanchougan No2 | 148/150 | ||||||
| Wogan | 180/180 | Clementine | 237/237 | Guposhanchougan No5 | 150/173 | ||||||
| Aiyuan No38 | 170/198 | Gonggan | 219/219 | Guposhanchougan No6 | 148/173 | ||||||
| S18 | 6 | Lihuaju | 247/249 | S74 | 2 | Yinduyeju | 191/191 | Shagan | 158/172 | ||
| Guposhanchougan No5 | 233/233 | Huangpisuanju | 182/203 | Biangan | 172/177 | ||||||
| Yuanyemangshanyegan | 246/246 | S81 | 5 | Guposhanyeju | 169/174 | Huangpisuanju | 161/164 | ||||
| Jianyemangshanyegan | 243/246 | Mangshanyeju | 173/176 | Kelimandingju | 158/161 | ||||||
| Katsuyamano | 231/239 | Hezhouyeju | 169/178 | Guangxiju | 161/161 | ||||||
| Youliang | 237/243 | Yinduyeju | 170/170 | Pixeju | 161/173 | ||||||
| S21 | 5 | Yinduyeju | 244/244 | Bendizao | 178/187 | Bendizao | 159/166 | ||||
| Lihuaju | 251/267 | S82 | 13 | Nieduyeju | 140/147 | Katsuyamano | 157/161 | ||||
| Biangan | 255/259 | Yinduyeju | 147/147 | Jinzhixiang | 157/166 | ||||||
| Banyeshenggan | 267/269 | Lihuaju | 122/150 | Gonggan | 158/166 | ||||||
| Bendizao | 259/259 | Yuanyemangshanyegan | 140/153 | Aiyuan No38 | 173/173 | ||||||
| S23 | 6 | Guposhanyeju | 163/163 | Jianyemangshanyegan | 153/153 | S76 | 2 | Yinduyeju | 168/168 | ||
| Nieduyeju | 163/169 | Huangpisuanju | 137/137 | Mangshanyeju | 180/186 | ||||||
| Hezhouyeju | 169/189 | Kelimandingju | 134/150 | S85 | 4 | Guposhanyeju | 129/129 | ||||
| Yinduyeju | 139/139 | Pixeju | 134/134 | Mangshanyeju | 129/137 | ||||||
| Lihuaju | 163/177 | Katsuyamano | 122/140 | Lihuaju | 129/133 | ||||||
| Murcott | 169/169 | Gonggan | 134/140 | Guposhanchougan No5 | 125/125 | ||||||
| S28 | 1 | Yinduyeju | 312/132 | Wogan | 134/158 | ||||||
| S73 | 2 | Jianyemangshanyegan | 345/345 | Huangmeiren | 128/158 | ||||||
| Banyeshenggan | 366/366 | Aiyuan No38 | 128/150 |
| Primer Combination | Number of Varieties Identified | Differentiation Rate (%) |
|---|---|---|
| S90 | 20 | 28.99 |
| S90+S18 | 23 | 33.33 |
| S90+S11 | 23 | 33.33 |
| S90+S17 | 24 | 34.78 |
| S90+S01 | 24 | 34.78 |
| S90+S73 | 21 | 30.43 |
| S90+S21 | 21 | 30.43 |
| S90+S85 | 22 | 31.88 |
| S90+S11+S13 | 23 | 33.33 |
| S90+S76+S11+S17+S13 | 24 | 34.78 |
| Total | 45 | 65.22 |
| Code | S76 | S85 | S73 | S13 | S11 | S01 | S18 | S17 | S21 | S90 |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 168/168 | 125/125 | 344/344 | 230/230 | 172/179 | 282/283 | 230/231 | 166/170 | 224/259 | 148/150 |
| 2 | 174/174 | 125/133 | 345/345 | 230/245 | 178/185 | 282/296 | 230/233 | 170/170 | 224/267 | 148/173 |
| 3 | 174/180 | 125/137 | 345/369 | 239/245 | 179/179 | 282/299 | 230/243 | 170/180 | 244/244 | 150/173 |
| 4 | 174/186 | 129/129 | 348/348 | 239/248 | 179/185 | 283/283 | 230/247 | 170/182 | 244/259 | 152/164 |
| 5 | 180/180 | 129/133 | 348/369 | 242/242 | 179/191 | 283/290 | 231/239 | 170/184 | 244/267 | 157/157 |
| 6 | 180/186 | 129/137 | 356/369 | 242/248 | 182/182 | 283/296 | 231/243 | 170/198 | 246/246 | 157/161 |
| 7 | 133/133 | 359/369 | 245/245 | 182/185 | 283/299 | 231/247 | 172/172 | 246/248 | 157/166 | |
| 8 | 133/137 | 366/366 | 245/248 | 182/191 | 290/290 | 233/233 | 172/182 | 249/259 | 158/161 | |
| 9 | 369/369 | 248/248 | 185/191 | 290/296 | 237/243 | 172/184 | 251/267 | 158/166 | ||
| A | 251/251 | 188/191 | 290/299 | 243/243 | 180/180 | 255/259 | 158/172 | |||
| B | 296/296 | 243/246 | 180/182 | 259/259 | 158/173 | |||||
| C | 299/299 | 243/247 | 180/184 | 259/267 | 159/166 | |||||
| D | 307/307 | 246/246 | 180/198 | 259/269 | 161/161 | |||||
| E | 247/247 | 180/200 | 261/269 | 161/164 | ||||||
| F | 247/249 | 182/182 | 263/263 | 161/165 | ||||||
| G | 182/190 | 263/267 | 161/166 | |||||||
| H | 184/184 | 267/267 | 161/171 | |||||||
| I | 184/198 | 267/269 | 161/173 | |||||||
| J | 198/198 | 269/269 | 165/171 | |||||||
| K | 166/166 | |||||||||
| M | 166/173 | |||||||||
| N | 166/179 | |||||||||
| P | 166/182 | |||||||||
| Q | 166/202 | |||||||||
| R | 172/172 | |||||||||
| S | 172/177 | |||||||||
| T | 172/179 | |||||||||
| U | 173/173 | |||||||||
| V | 176/179 | |||||||||
| W | 179/179 |
| Germless Name | Molecular ID | Germless Name | Molecular ID |
|---|---|---|---|
| Guposhanyeju | 44998BABFJ | Yingxinju | 37980274EV |
| Xinganyeju | 38985965DG | Pixeju | 33970970HI |
| Daoxianyeju | 38985965DG | Bendizao | 279808AHBC |
| Mangshanyeju | 669762CB8F | Penggan No79-2 | 33573A7DHR |
| Niuduyeju | 279062A98H | Dong No13penggan | 33573A7DHR |
| Hezhouyeju | 38087374G4 | Shi No18penggan | 33573A7DHR |
| Yinduyeju | 173A3DA030 | Taitianpenggan | 33573A7DHR |
| Lihuaju | 359271F09Q | Xinshengxi No3penggan | 33573A7DHR |
| Cengxisuanju | 37974378FP | Wuhepenggan | 33573A7DHR |
| Guangxihongpisuanju | 37974378FP | Dafen No4 | 326938AI4K |
| Guposhanchougan No2 | 2215642F61 | Rinan No1 | 376938AI4K |
| Guposhanchougan No5 | 2110848F63 | Dapu No5 | 576938A04K |
| Guposhanchougan No6 | 2215842G62 | Miyamoto | 576008AI4K |
| Yuanyemangshanyegan | 273574D775 | Miyagawa | 379938AI4K |
| Jianyemangshanyegan | 222564B775 | Hashikawa | 376938AI4K |
| Shagan | 38785A43CA | Xingjin | 376938AI4K |
| Biangan | 5798ACE2AS | Yoshida | 376938A04K |
| Banyeshenggan | 37884971IK | Ichibun | 376938AI4K |
| Huangpisuanju | 3798437FHE | Yamasitabeni | 376938AI4K |
| Hongpisuanju | 32974378FP | Katsuyamano | 2253175B56 |
| Shatangju | 37933C74HT | Shangye | 376938AI4K |
| Zaoshushatangju | 37575C70HN | Dajin No4 | 376038AI4K |
| Yamada | 376938AI4K | Zaoxiang | 325838CC5M |
| Bayueju | 37575C70HN | Sakikubo | 376938AI4K |
| Denglongju | 37930C74HT | Jinzhixiang | 3764151D47 |
| Jinkuimiju | 37573C74HN | Youliang | 376938904K |
| Nanfengmiju1 | 27985969CK | Chunjian | 3368384JHM |
| Nanfengmiju2 | 27985969CK | Nanxiang | 376948C55M |
| Tezaoshumiju | 27985969CK | Murcott | 586859E3HW |
| Liuchengmiju | 27985969CK | Gonggan | 58583ACE59 |
| Guijuyihao | 27985B6HCM | Wogan | 3809967AGW |
| Clementine | 3879453D58 | Huangmeiren | 3759386DGB |
| Mingliutianju | 37900274EV | Aiyuan No38 | 389948465U |
| Chuntianju | 37989274EV | Mingrijian | 229845105B |
| Guangxiju | 32573A7EHD |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.





