Submitted:
02 March 2026
Posted:
03 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
2.1. Isolation of Murine Brain Cell Types Using MACS Technology
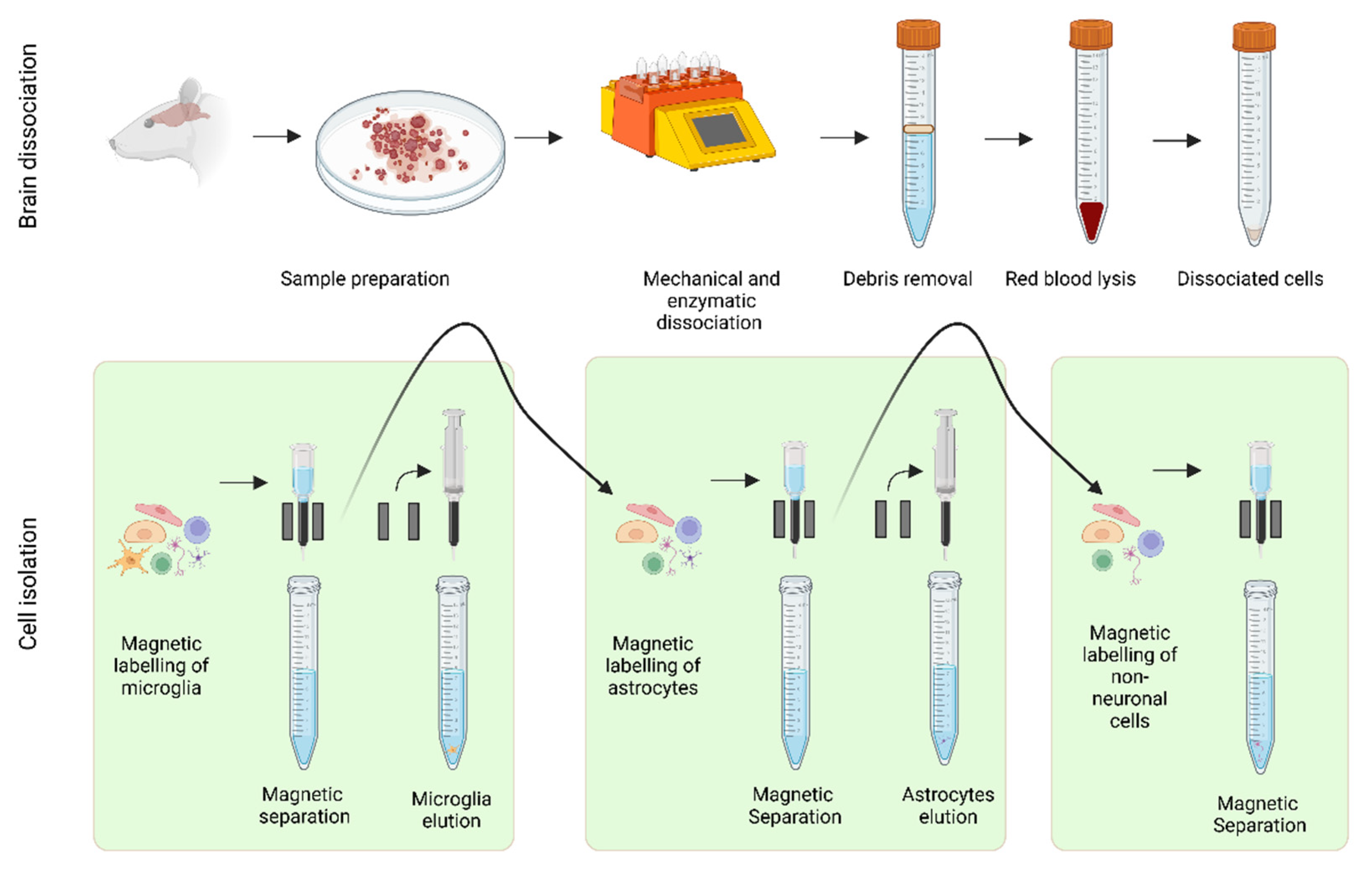
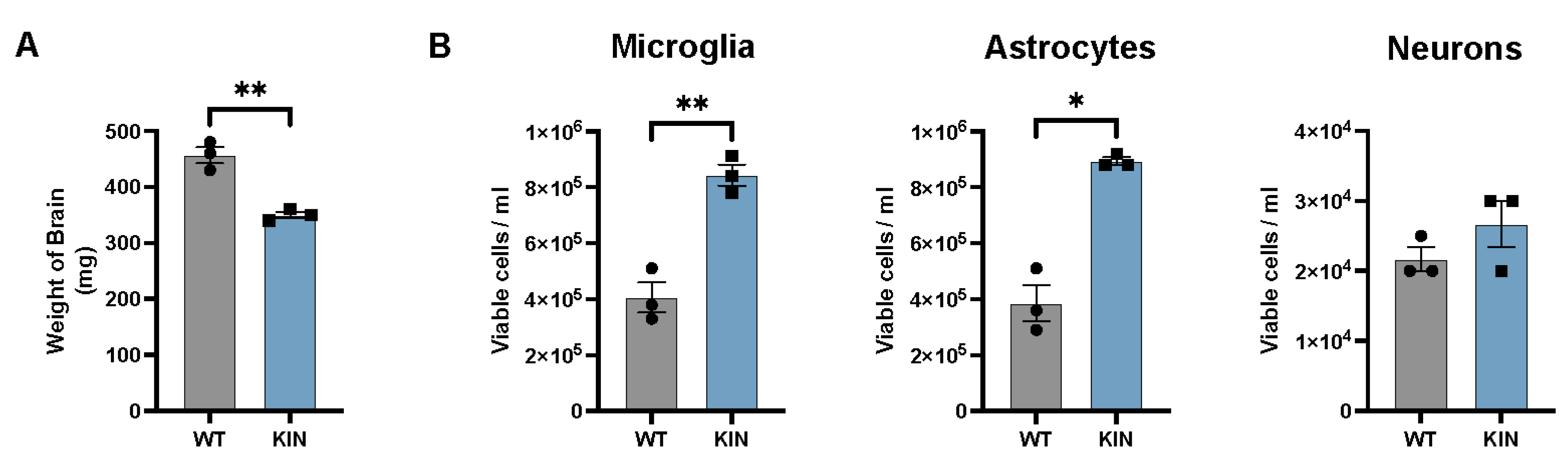
2.2. The Impact of the Atxn2-CAG100-KIN Mutation on the Brain and Its Cell Types
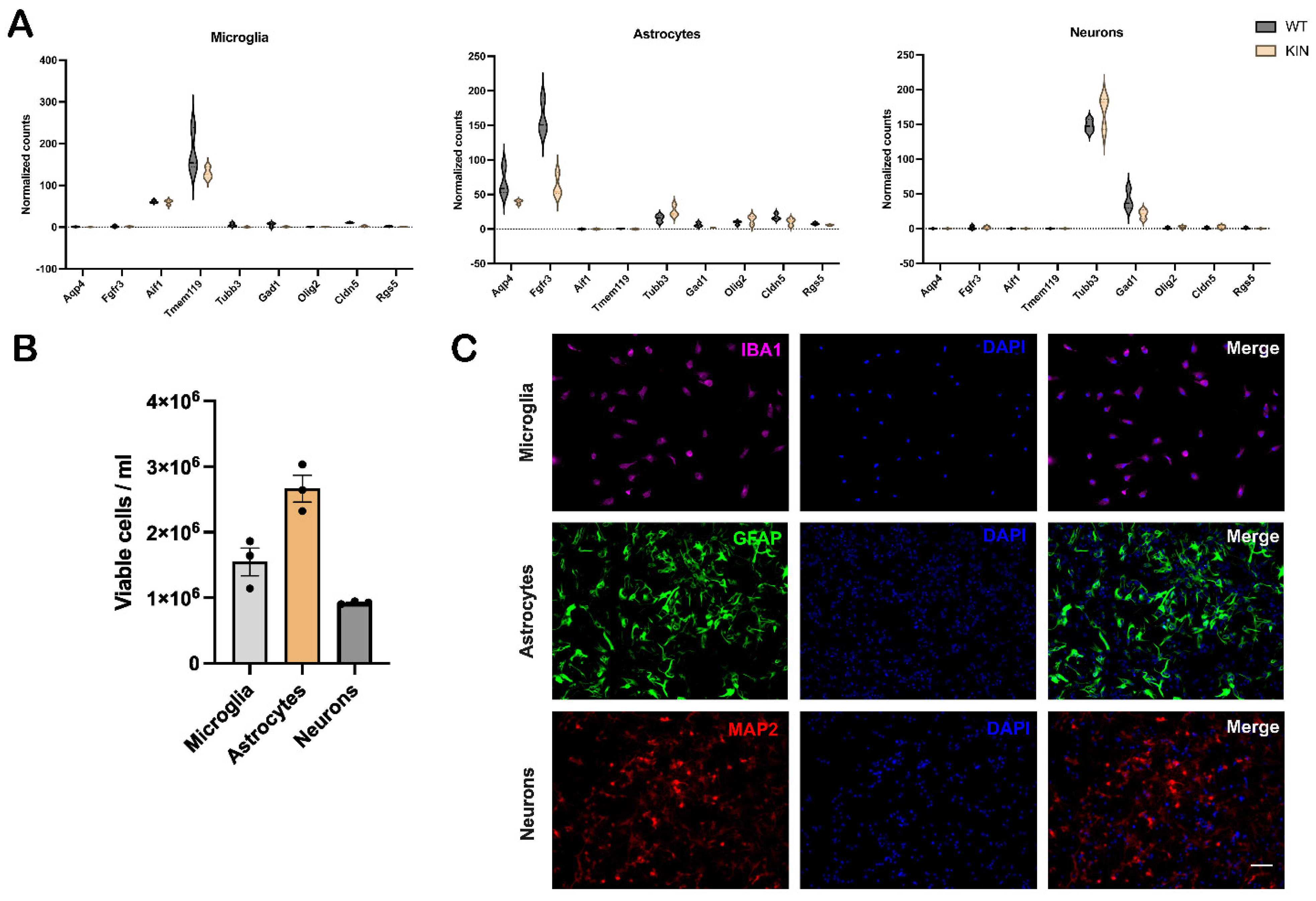
2.3. Further Purity Analyses of Isolated Individual Brain Cell Types
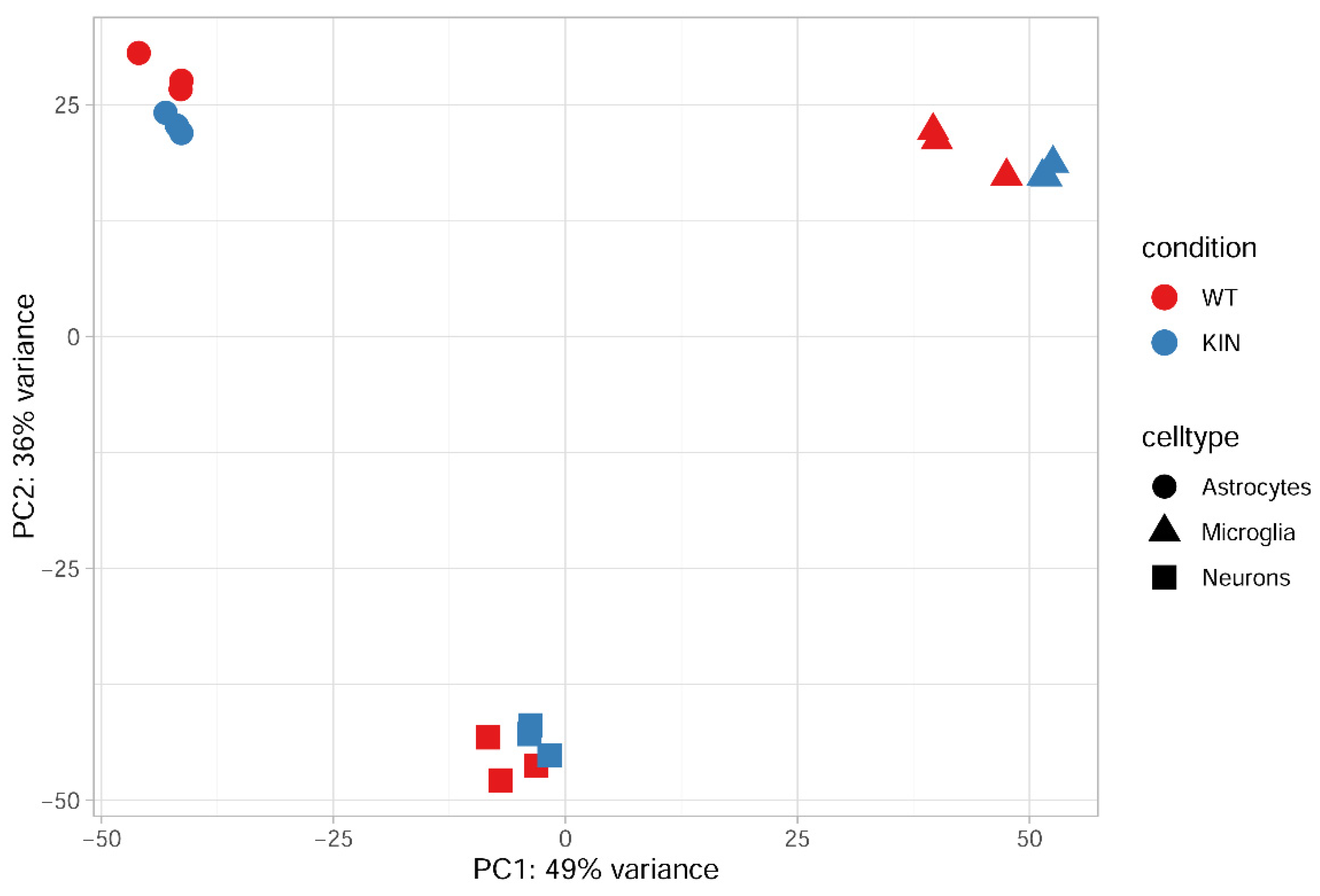
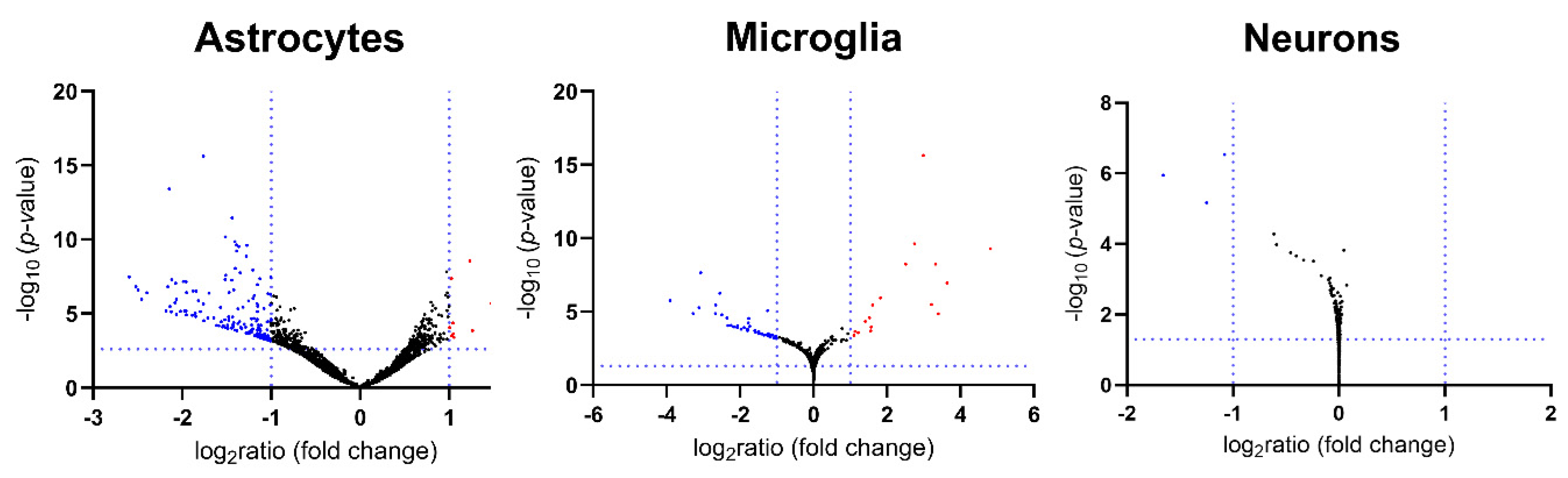
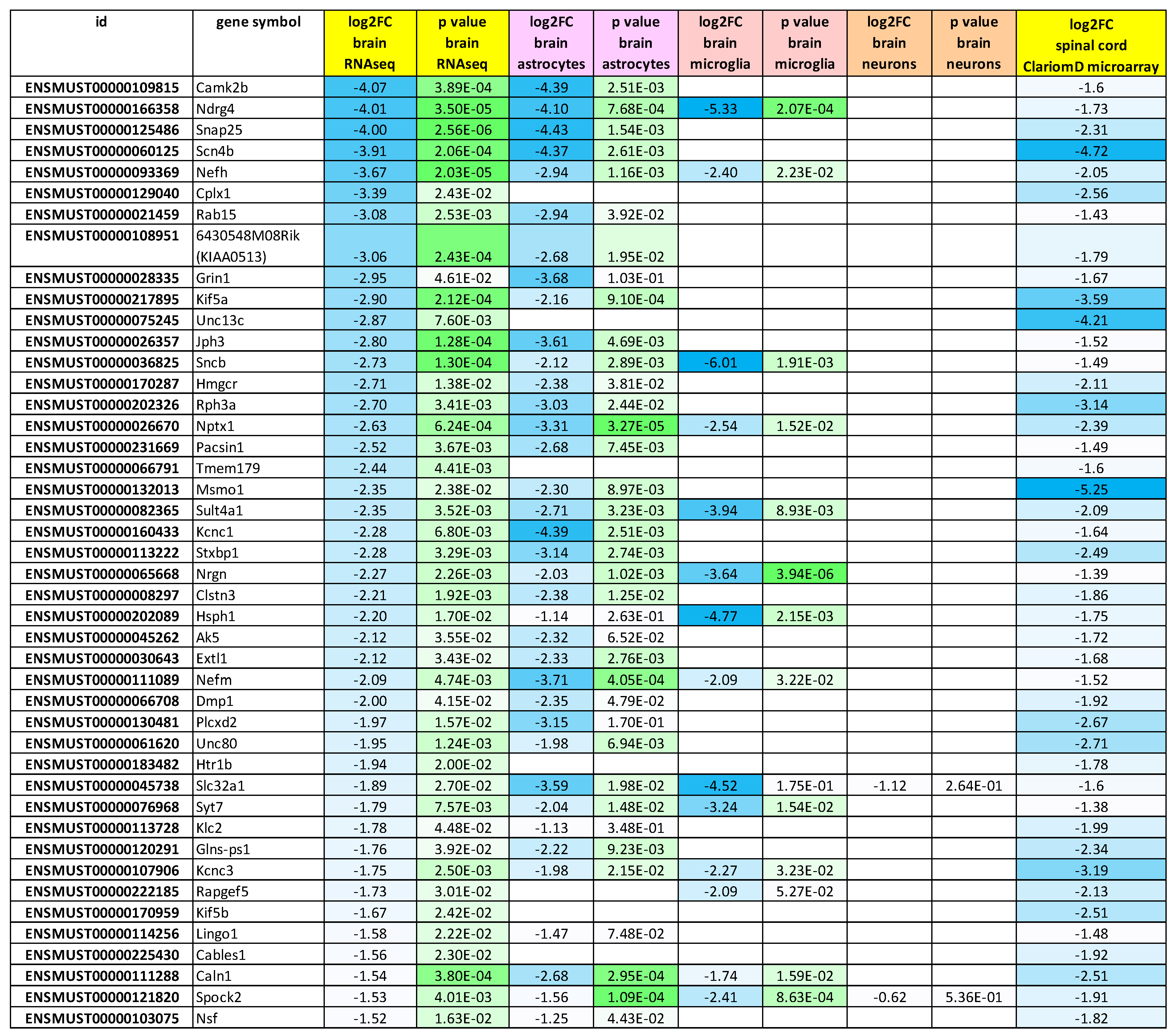
2.4. Global Volcano Plots of RNA-seq Profiles, as well as Consistency Between New RNA-seq Transcript Analysis and Previous Oligonucleotide Microarray Data Document Loss of mRNA in Synapses and Neurites from Glial Fractions, with an Elevated Neuroinflammation Signature
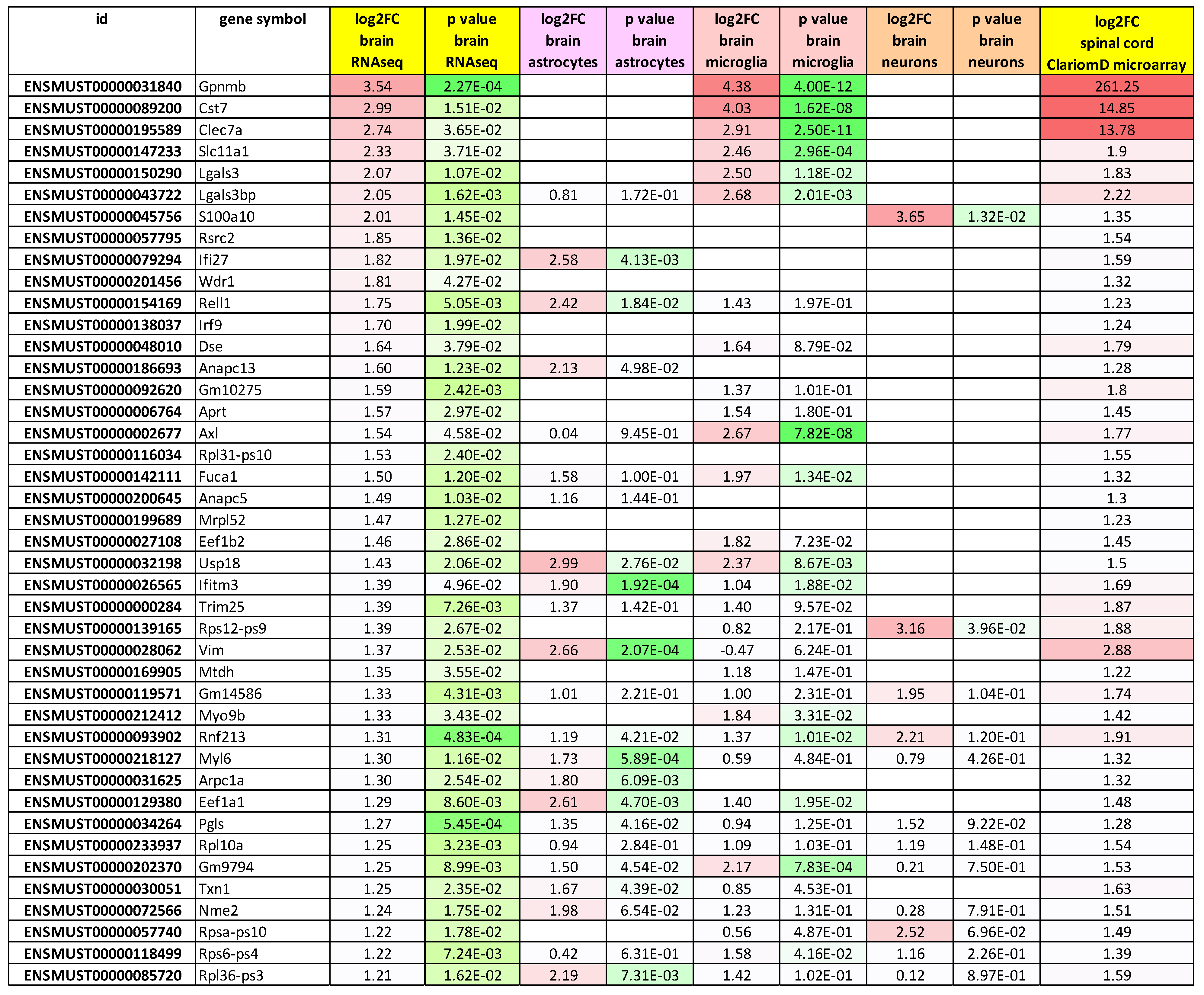 |
2.5. RNA-seq Transcript Consistency Analysis Among Glial Fractions Shows a Prominent Increase of Transcripts Encoding Cytoplasmic Ribosomal Proteins of Large and Small Subunits
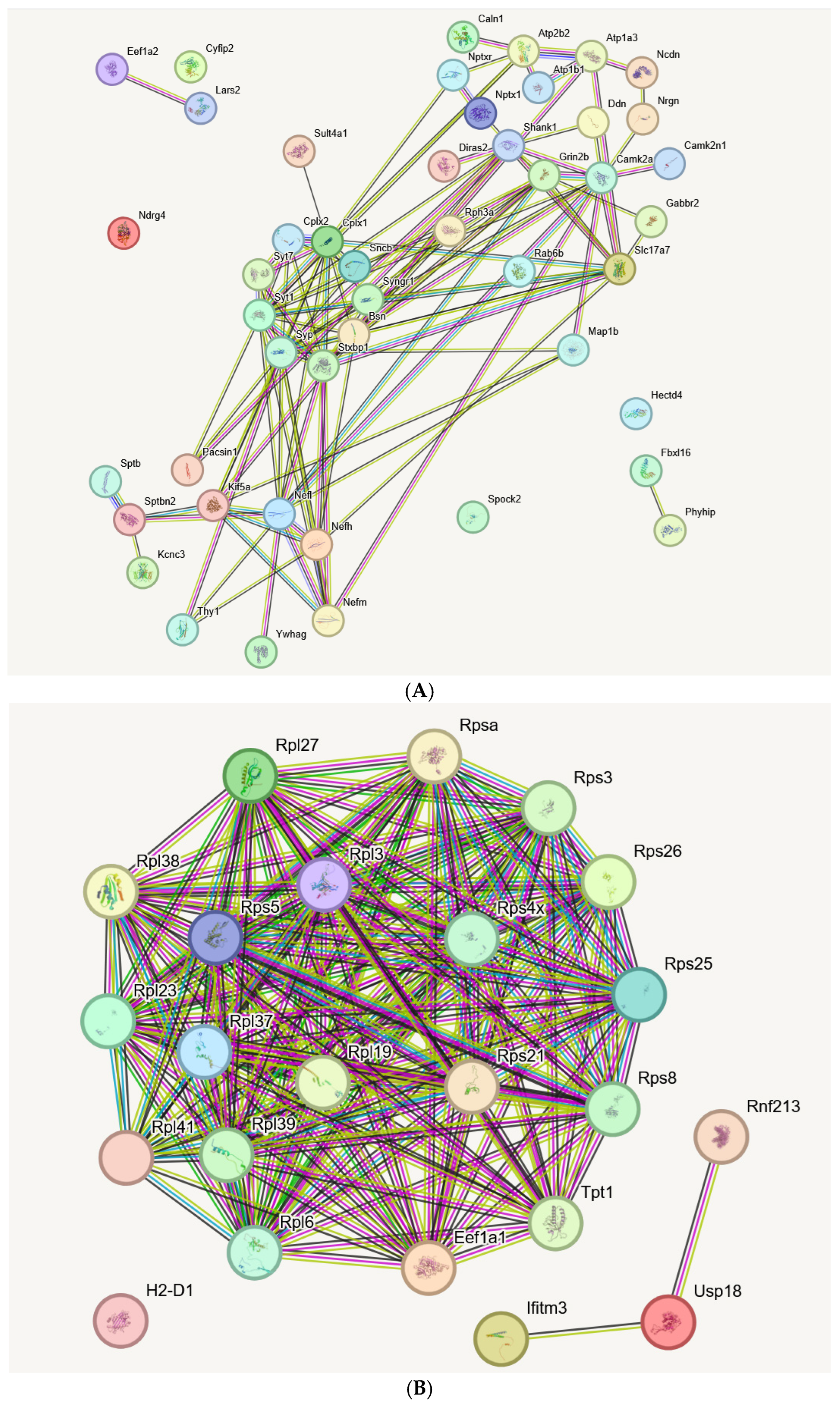
2.6. STRING Analysis of Interactions and Bioinformatics Enrichments in Glial Fractions Extends the Observations of Transcript Reduction in Synapses, Axons, and Dendrites, Versus Induction of Ribosomal and Inflammatory Factors
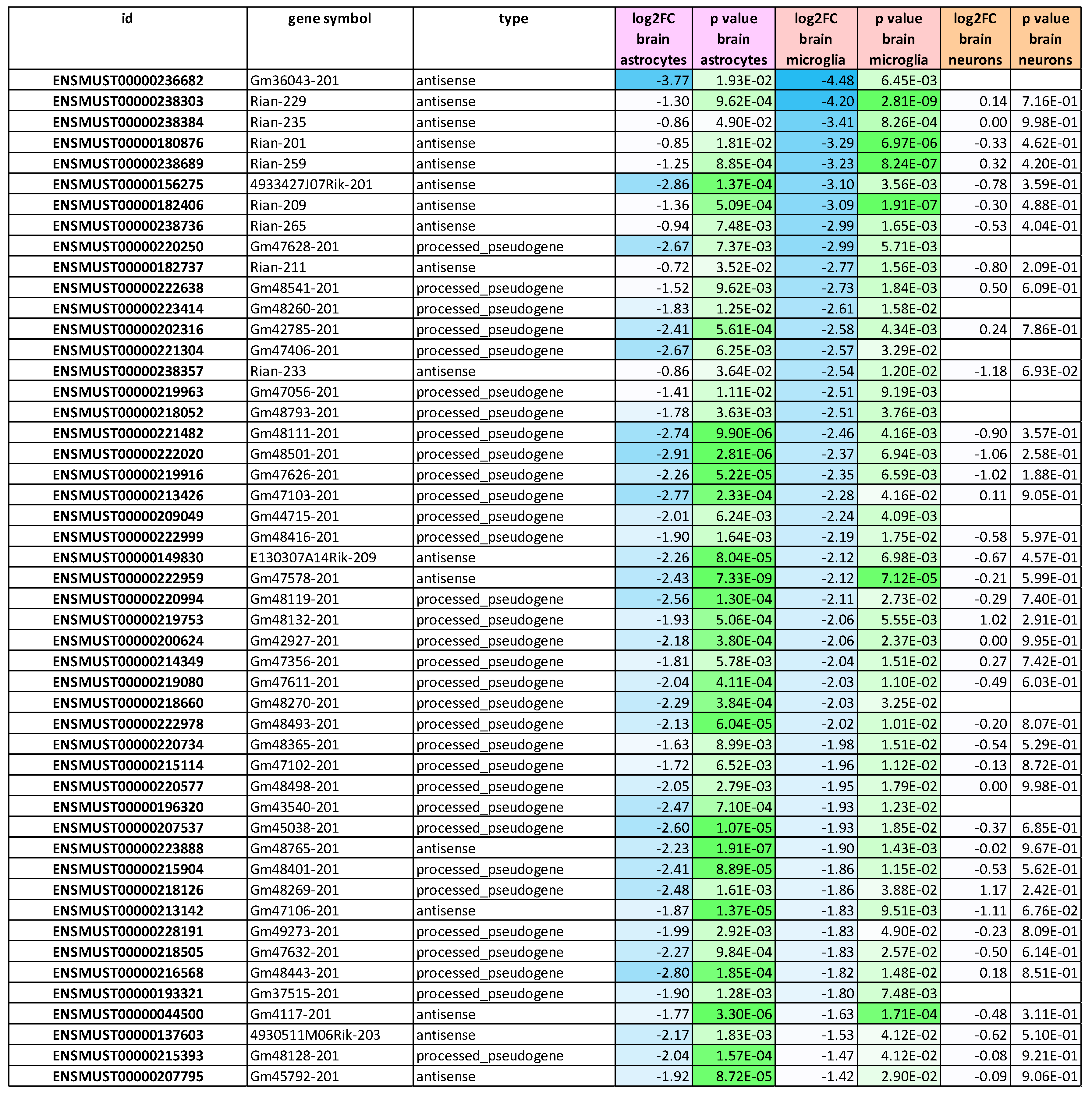
2.7. RNA-seq Consistency Analysis Among Glial Fractions for Small RNAs Finds Upregulation of All tRNAs Versus Downregulation of all Antisense RNAs and All Processed Pseudogenes
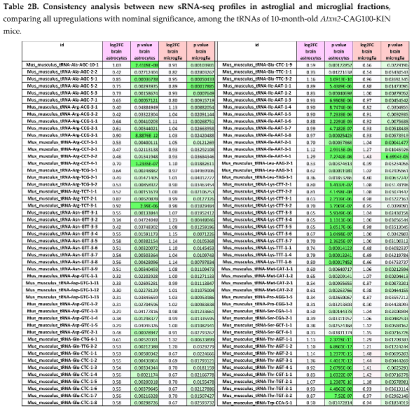
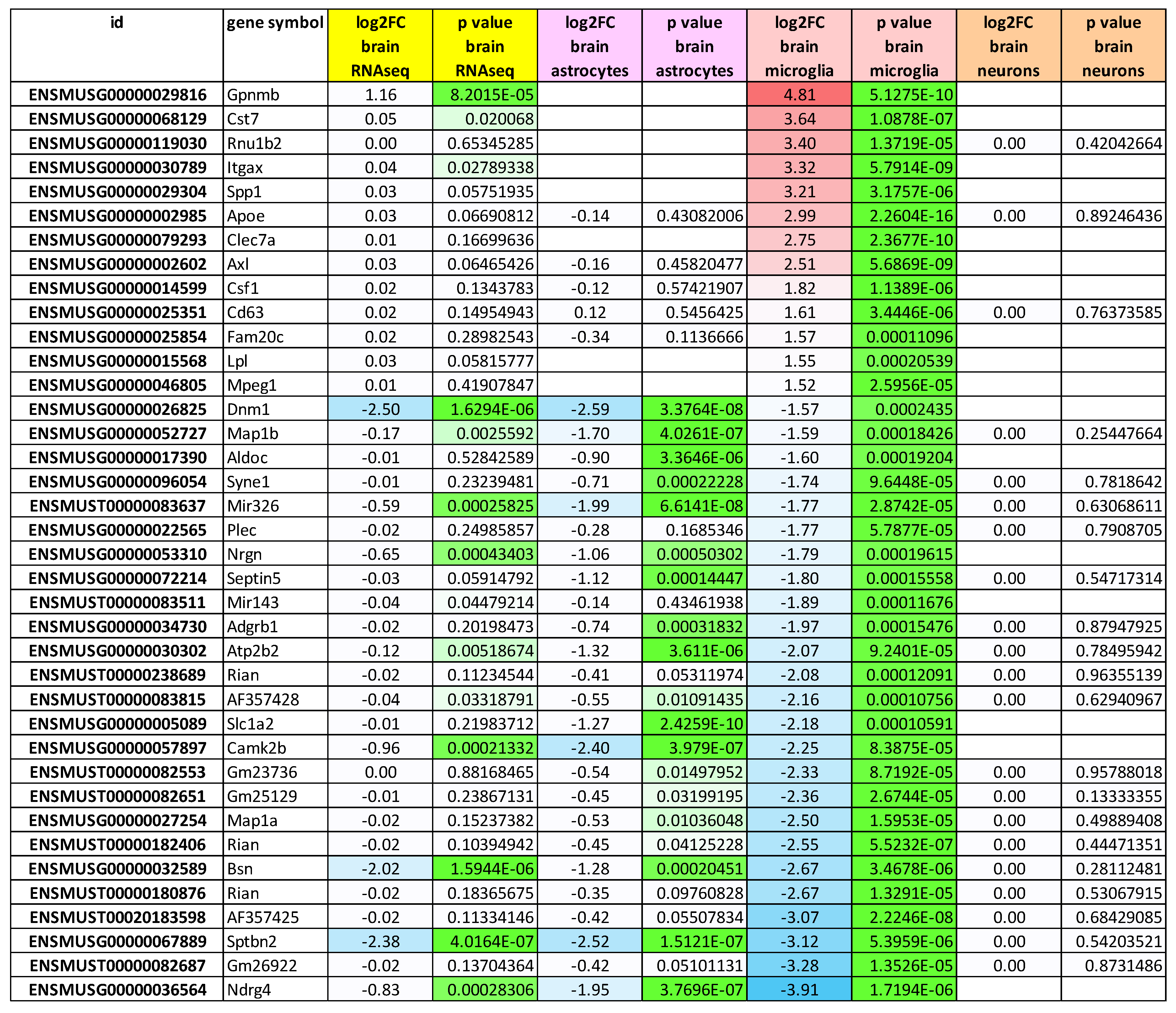
2.8. RNA-seq Analysis at the Gene Level of Microglia-Specific Effects Reveals ~8-Fold Accumulation of Uridine-Rich Small Nuclear RNA Rnu1b2
2.9. RNA-seq Analysis at the Gene Level of Astroglia-Specific Effects Documents Mainly the Neurite Retraction
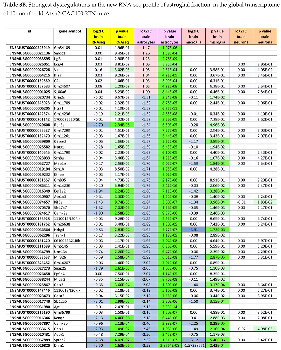
2.10. RNA-seq Analysis at the Gene Level of Neuron-Specific Effects Documents Novel DNA Effects
2.11. Systematic Interrogation of RNA-seq Findings at the Gene Level for Uridine-Rich snRNAs Shows Increases Selectively in the Astroglial Fraction that Can Be Explained by Astrogliosis with Mass Increase
3. Discussion
4. Materials and Methods
4.1. Mouse Husbandry
4.2. Genotyping
4.3. Isolation of Cell Types From The Mouse Brain
4.3.1. Tissue Dissociation
4.3.2. Debris Removal
4.3.3. Red blood Cell Removal
4.3.4. Magnetic Separation
4.3.5. Culture of Primary Cells
4.3.6. RNA Extraction, Library Preparation, RNA-seq and GSEA
4.4. Reverse Transcription-Quantitative Polymerase Chain Reaction (RT-qPCRs)
4.5. Immunohistochemistry
4.6. Quantification and Statistical Analysis
4.6.1. Statistics
4.6.2. STRING Pathway Interaction and Enrichment Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ACSA2 | Astrocyte cell surface antigen-2 |
| Adcy1 | Adenylate Cyclase 1 |
| Agt | Angiotensinogen, Serine Proteinase Inhibitor A8 |
| Aif1 | Allograft Inflammatory Factor 1 = IBA1 |
| Ald1l1 | Aldehyde Dehydrogenase 1 Family Member L1 |
| ALS13 | Amyotrophic Lateral Sclerosis type 13 |
| AMPA | Alpha-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid |
| Apoe | Apolipoprotein E |
| Atp1a3 | ATPase Na+/K+ Transporting Subunit Alpha 3 |
| Atx2 | Ataxin-2 in flies |
| Atxn2 | Ataxin-2 |
| Atxn2l | Ataxin-2-like |
| Axl | AXL Receptor Tyrosine Kinase |
| Camk2a | Calcium/Calmodulin Dependent Protein Kinase II Alpha |
| Camk2b | Calcium/Calmodulin Dependent Protein Kinase II Beat |
| Cd32 | Cluster of differentiation 32 |
| Cd63 | Cluster of differentiation 63 |
| Cd68 | Cluster of differentiation 68 |
| cDNA | Complementary DNA |
| Celsr3 | Cadherin EGF LAG Seven-Pass G-Type Receptor 3 |
| CH | Calponin-homology |
| CID3/4 | Chemical-inducible dimerization |
| Cldn5 | Claudin 5 |
| Clec7a | C-type lectin domain containing protein 7A, |
| Cplx1 | Complexin 1/2 |
| Cst7 | Cystatin-F |
| Cyfip2 | Cytoplasmic FMR1 interacting protein 2 |
| Ddx6 | DEAD-Box Helicase 6 |
| Dnm1 | Dynamin 1 |
| Eaat4 | Excitatory Amino Acid Transporter 4, Solute Carrier Family 1 Member 6 |
| EDTA | Ethylenediaminetetraacetic acid |
| Eef1a1/2 | Eukaryotic translation elongation factor 1 alpha 1/2 |
| Eftud2 | Elongation Factor Tu GTP Binding Domain Containing 2 |
| Eno2 | Enolase 2 |
| Etnppl | Ethanolamine-phosphate phospho-lyase |
| Fam20c | Family With Sequence Similarity 20, Member C |
| Fbxl16 | F-box and leucine rich repeat protein 16 |
| FC | Fold change |
| Fcrls | Fc receptor-like S |
| FDR | False Discovery Rate |
| FMRP | Fragile X Mental Retardation protein |
| Gad1/2 | Glutamate Decarboxylase 1 |
| GDP | Guanosine diphosphate |
| Gfap | Glial Fibrillary Acidic Protein |
| Gls | Glutaminase |
| GO GO-BP GO-CC GO-MF |
Gene ontology Gene ontology – biological process Gene ontology – cellular component Gene ontology – molecular function |
| Gpnmb | Glycoprotein Nmb |
| Grin1 GSEA |
Glutamate Receptor, Ionotropic, N-Methyl D-Aspartate 1 Gene Set Enrichment Analysis |
| Hexb | Hexosaminidase Subunit Beta |
| Hmgcr | 3-Hydroxy-3-Methylglutaryl-CoA Reductase |
| Hmgcs1 | 3-Hydroxy-3-Methylglutaryl-CoA Synthase 1 |
| IBA1 Ifitm3 |
Ionized calcium-binding adapter molecule 1, = AIF1 Interferon-induced transmembrane protein 3 |
| Itgax | Integrin Subunit Alpha X |
| Kcnj8 | Potassium Inwardly Rectifying Channel Subfamily J Member 8 |
| Kcnq2 | Potassium Voltage-Gated Channel Subfamily Q Member 2 |
| Kif5a | Neuronal Kinesin Heavy Chain |
| KIN | Knock-In |
| Lgals3 | Lectin, galactoside-binding, soluble factor 3 |
| Lgals3bp | Galectin-3 binding protein |
| lincRNA | Long intergenic non-coding RNA |
| Lpl | Lipoprotein Lipase |
| LSmAD MACE |
LSm-associated domain Massive analysis of cDNA ends |
| MACS | Magnetic-Activated Cell Sorting |
| Map1b | Microtubule Associated Protein 1B |
| Map2 | Microtubule Associated Protein 2 |
| mGLUR1 | Glutamate Metabotropic Receptor 1 |
| Mical2 | Microtubule Associated Monooxygenase, Calponin And LIM Domain Containing 2 |
| miRNA | MicroRNA |
| Mpeg1 | Macrophage Expressed 1 |
| Mt-tRNA | Mitochondrial transfer RNA |
| Ndrg4 | N-myc downstream-regulated gene 4 |
| Nefh | Neurofilament Heavy Chain |
| Nefl | Neurofilament Light Chain |
| Neurod2 | Neuronal Differentiation 2 |
| NGS Nptx1 |
Next generation sequencing Neuronal pentraxin 1 |
| Nufip2 | Nuclear FMR1 interacting protein 2 |
| Olfml3 | Olfactomedin Like 3 |
| Olig2 | Oligodendrocyte Transcription Factor 2 |
| P2ry12 | Purinergic Receptor P2Y12 |
| PABP | Poly(A)-binding protein |
| Pabpc1 | polyA-binding protein, cytoplasmic, type 1 |
| PAR-CLIP | Photoactivatable ribonucleoside-enhanced crosslinking and immunoprecipitation |
| PCA piRNA |
Principal component analysis Piwi-interacting RNA |
| Plca3 | = Gpnmb, Glycoprotein Nonmetastatic Melanoma Protein B |
| Ppfia4 | PPFI Scaffold Protein A4 |
| Pvalb | Parvalbumin |
| Rab15 | Ras-Related Protein Rab-15 |
| Rbfox3 | RNA Binding Fox-1 Homolog 3 |
| Rian | RNA mprinted and accumulated in nucleus |
| RNA | Ribonucleic acid |
| Rnf213 | Ring Finger Protein 213 |
| Rnu1b2 | RNA, U1 Small Nuclear 2 |
| Rph3a | Rabphilin 3A |
| rRNA | Ribonucleic acid |
| RT-qPCR | Reverse transcriptase quantitative real-time polymerase chain reaction |
| S100b | S100 Calcium Binding Protein B |
| Sall1 | Spalt Like Transcription Factor 1 |
| SCA2 | Spinocerebellar Ataxia type 2 |
| SCA5 | Spinocerebellar Ataxia Type 5 |
| SCAR14 | Spectrin Beta, Non-Erythrocytic 2 |
| scaRNA | Small Cajal body-specific RNA |
| Scn4b | Sodium voltage-gated channel beta subunit 4 |
| SEM | Standard error of the mean |
| Slc11a1 | Solute carrier family 11, member 1 |
| Slc12a5 | Solute Carrier Family 12 Member 5 |
| Slc17a7 | Vesicular glutamate transporter 1 |
| SMN1 | Survival Of Motor Neuron 1, Telomeric |
| Snap25 | Synaptosome Associated Protein 25 |
| snoRNA | Small nucleolar RNA |
| snRNA snRNP |
Small nuclear RNA Small nuclear ribonucleoprotein |
| SPARCA1 | Spectrin-associated autosomal recessive cerebellar ataxia |
| Spi1 | Spi-1 Proto-Oncogene |
| Spp1 | Secreted Phosphoprotein 1 |
| Spock2 | secreted protein acidic and rich in cysteine |
| Sptbn1 | Spectrin Beta, Non-Erythrocytic 1 |
| Sptbn2 | Spectrin Beta, Non-Erythrocytic 2 |
| sRNA | Small RNA |
| STRING | Search tool for recurring instances of neighbouring genes |
| Syne1 | Spectrin Repeat Containing Nuclear Envelope Protein 1 |
| Syt1 | Syntaxin 1 |
| Tmem119 | Transmembrane Protein 119 |
| Trank1 | Tetratricopeptide Repeat And Ankyrin Repeat Containing 1 |
| Trem2 | Triggering Receptor Expressed On Myeloid Cells 2 |
| tRNA | Transfer RNA |
| Tyrobp | TYRO Protein Tyrosine Kinase-Binding Protein |
| Unc13a | Unc-13 Homolog A |
| Usp18 | Ubiquitin Specific Peptidase 18 |
| Usp18 | Ubiquitin specific peptidase 18 |
| Vim | Vimentin |
| WT | Wildtype |
| ZFE | Central Animal Facility |
References
- Hernandez, A., et al., Genetic mapping of the spinocerebellar ataxia 2 (SCA2) locus on chromosome 12q23-q24.1. Genomics, 1995. 25(2): p. 433-5.
- Belal, S., et al., Clinical and genetic analysis of a Tunisian family with autosomal dominant cerebellar ataxia type 1 linked to the SCA2 locus. Neurology, 1994. 44(8): p. 1423-6.
- Auburger, G., et al., Autosomal dominant ataxia: genetic evidence for locus heterogeneity from a Cuban founder-effect population. Am J Hum Genet, 1990. 46(6): p. 1163-77.
- Gispert, S., et al., Chromosomal assignment of the second locus for autosomal dominant cerebellar ataxia (SCA2) to chromosome 12q23-24.1. Nat Genet, 1993. 4(3): p. 295-9.
- Wadia, N.H. and R.K. Swami, A new form of heredo-familial spinocerebellar degeneration with slow eye movements (nine families). Brain, 1971. 94(2): p. 359-74.
- Orozco Diaz, G., et al., Autosomal dominant cerebellar ataxia: clinical analysis of 263 patients from a homogeneous population in Holguin, Cuba. Neurology, 1990. 40(9): p. 1369-75.
- Estrada, R., et al., Spinocerebellar ataxia 2 (SCA2): morphometric analyses in 11 autopsies. Acta Neuropathol, 1999. 97(3): p. 306-10.
- Velazquez-Perez, L., et al., Saccade velocity is controlled by polyglutamine size in spinocerebellar ataxia 2. Ann Neurol, 2004. 56(3): p. 444-7.
- Almaguer-Mederos, L.E., et al., Estimation of the age at onset in spinocerebellar ataxia type 2 Cuban patients by survival analysis. Clin Genet, 2010. 78(2): p. 169-74.
- Almaguer-Mederos, L.E., et al., Estimation of survival in spinocerebellar ataxia type 2 Cuban patients. Clin Genet, 2013. 83(3): p. 293-4.
- Velazquez-Perez, L., et al., Corticomuscular Coherence: a Novel Tool to Assess the Pyramidal Tract Dysfunction in Spinocerebellar Ataxia Type 2. Cerebellum, 2017. 16(2): p. 602-606.
- Rodriguez-Labrada, R., et al., Sleep spindles and K-complex activities are decreased in spinocerebellar ataxia type 2: relationship to memory and motor performances. Sleep Med, 2019. 60: p. 188-196.
- Velazquez-Perez, L., et al., Early corticospinal tract damage in prodromal SCA2 revealed by EEG-EMG and EMG-EMG coherence. Clin Neurophysiol, 2017. 128(12): p. 2493-2502.
- Reetz, K., et al., Brain atrophy measures in preclinical and manifest spinocerebellar ataxia type 2. Ann Clin Transl Neurol, 2018. 5(2): p. 128-137.
- Rodriguez-Grana, T., et al., Weight loss is correlated with disease severity in Spinocerebellar ataxia type 2: a cross-sectional cohort study. Nutr Neurosci, 2022. 25(8): p. 1747-1755.
- Almaguer-Mederos, L.E., et al., Body Mass Index Is Significantly Associated With Disease Severity in Spinocerebellar Ataxia Type 2 Patients. Mov Disord, 2021. 36(6): p. 1372-1380.
- Velazquez-Perez, L., et al., Prodromal Spinocerebellar Ataxia Type 2 Subjects Have Quantifiable Gait and Postural Sway Deficits. Mov Disord, 2021. 36(2): p. 471-480.
- Velazquez-Perez, L., et al., COVID-19 Impacts the Mental Health and Speech Function in Spinocerebellar Ataxia Type 2: Evidences from a Follow-Up Study. Cerebellum, 2024. 23(3): p. 1101-1111.
- Vazquez-Mojena, Y., et al., Peripheral Inflammation Links with the Severity of Clinical Phenotype in Spinocerebellar Ataxia 2. Mov Disord, 2023. 38(5): p. 880-885.
- Rodriguez-Labrada, R., et al., Progression of Gait Ataxia in Spinocerebellar Ataxia Type 2: Clinical Utility of Digitally Measured Gait Metrics. Cerebellum, 2025. 24(6): p. 184.
- Rodriguez-Labrada, R., et al., Structural Brain Correlates of Sleep Microstructure in Spinocerebellar Ataxia Type 2 and its Role on Clinical Phenotype. Cerebellum, 2024. 23(5): p. 1839-1847.
- Schols, L., et al., Spinocerebellar ataxia type 2. Genotype and phenotype in German kindreds. Arch Neurol, 1997. 54(9): p. 1073-80.
- Lastres-Becker, I., U. Rub, and G. Auburger, Spinocerebellar ataxia 2 (SCA2). Cerebellum, 2008. 7(2): p. 115-24.
- Freund, H.J., et al., Subthalamic-thalamic DBS in a case with spinocerebellar ataxia type 2 and severe tremor-A unusual clinical benefit. Mov Disord, 2007. 22(5): p. 732-5.
- Tuin, I., et al., Stages of sleep pathology in spinocerebellar ataxia type 2 (SCA2). Neurology, 2006. 67(11): p. 1966-72.
- Auburger, G.W., Spinocerebellar ataxia type 2. Handb Clin Neurol, 2012. 103: p. 423-36.
- Hoche, F., et al., Spinocerebellar ataxia type 2 (SCA2): identification of early brain degeneration in one monozygous twin in the initial disease stage. Cerebellum, 2011. 10(2): p. 245-53.
- Rub, U., et al., Clinical features, neurogenetics and neuropathology of the polyglutamine spinocerebellar ataxias type 1, 2, 3, 6 and 7. Prog Neurobiol, 2013. 104: p. 38-66.
- Yagishita, S. and M. Inoue, Clinicopathology of spinocerebellar degeneration: its correlation to the unstable CAG repeat of the affected gene. Pathol Int, 1997. 47(1): p. 1-15.
- Mizushima, K., et al., Analysis of spinocerebellar ataxia type 2 in Gunma Prefecture in Japan: CAG trinucleotide expansion and clinical characteristics. J Neurol Sci, 1998. 156(2): p. 180-5.
- Yokota, T., et al., Electrophysiological features of central motor conduction in spinocerebellar atrophy type 1, type 2, and Machado-Joseph disease. J Neurol Neurosurg Psychiatry, 1998. 65(4): p. 530-4.
- Durr, A., et al., Autosomal dominant cerebellar ataxia type I linked to chromosome 12q (SCA2: spinocerebellar ataxia type 2). Clin Neurosci, 1995. 3(1): p. 12-6.
- Rivaud-Pechoux, S., et al., Eye movement abnormalities correlate with genotype in autosomal dominant cerebellar ataxia type I. Ann Neurol, 1998. 43(3): p. 297-302.
- Cancel, G., et al., Molecular and clinical correlations in spinocerebellar ataxia 2: a study of 32 families. Hum Mol Genet, 1997. 6(5): p. 709-15.
- Durr, A., et al., Autosomal dominant cerebellar ataxia type I in Martinique (French West Indies). Clinical and neuropathological analysis of 53 patients from three unrelated SCA2 families. Brain, 1995. 118 ( Pt 6): p. 1573-81.
- Giunti, P., et al., The role of the SCA2 trinucleotide repeat expansion in 89 autosomal dominant cerebellar ataxia families. Frequency, clinical and genetic correlates. Brain, 1998. 121 ( Pt 3): p. 459-67.
- Wadia, N., et al., A clinicogenetic analysis of six Indian spinocerebellar ataxia (SCA2) pedigrees. The significance of slow saccades in diagnosis. Brain, 1998. 121 ( Pt 12): p. 2341-55.
- Pang, J., et al., A common disease haplotype segregating in spinocerebellar ataxia 2 (SCA2) pedigrees of diverse ethnic origin. Eur J Hum Genet, 1999. 7(7): p. 841-5.
- Zhou, Y.X., et al., Spinocerebellar ataxia type 2 in China: molecular analysis and genotype-phenotype correlation in nine families. Neurology, 1998. 51(2): p. 595-8.
- Buttner, N., et al., Oculomotor phenotypes in autosomal dominant ataxias. Arch Neurol, 1998. 55(10): p. 1353-7.
- Babovic-Vuksanovic, D., et al., Spinocerebellar ataxia type 2 (SCA 2) in an infant with extreme CAG repeat expansion. Am J Med Genet, 1998. 79(5): p. 383-7.
- Koeppen, A.H., The hereditary ataxias. J Neuropathol Exp Neurol, 1998. 57(6): p. 531-43.
- Adams, C., S. Starkman, and S.M. Pulst, Clinical and molecular analysis of a pedigree of southern Italian ancestry with spinocerebellar ataxia type 2. Neurology, 1997. 49(4): p. 1163-6.
- Geschwind, D.H., et al., The prevalence and wide clinical spectrum of the spinocerebellar ataxia type 2 trinucleotide repeat in patients with autosomal dominant cerebellar ataxia. Am J Hum Genet, 1997. 60(4): p. 842-50.
- Paulson, H.L. and K.H. Fischbeck, Trinucleotide repeats in neurogenetic disorders. Annu Rev Neurosci, 1996. 19: p. 79-107.
- Subramony, S.H., Clinical aspects of hereditary ataxias. J Child Neurol, 1995. 10(5): p. 353-62.
- Sanpei, K., et al., Identification of the spinocerebellar ataxia type 2 gene using a direct identification of repeat expansion and cloning technique, DIRECT. Nat Genet, 1996. 14(3): p. 277-84.
- Imbert, G., et al., Cloning of the gene for spinocerebellar ataxia 2 reveals a locus with high sensitivity to expanded CAG/glutamine repeats. Nat Genet, 1996. 14(3): p. 285-91.
- Pulst, S.M., et al., Moderate expansion of a normally biallelic trinucleotide repeat in spinocerebellar ataxia type 2. Nat Genet, 1996. 14(3): p. 269-76.
- Elden, A.C., et al., Ataxin-2 intermediate-length polyglutamine expansions are associated with increased risk for ALS. Nature, 2010. 466(7310): p. 1069-75.
- Lee, T., et al., Ataxin-2 intermediate-length polyglutamine expansions in European ALS patients. Hum Mol Genet, 2011. 20(9): p. 1697-700.
- Gispert, S., et al., The modulation of Amyotrophic Lateral Sclerosis risk by ataxin-2 intermediate polyglutamine expansions is a specific effect. Neurobiol Dis, 2012. 45(1): p. 356-61.
- Neuenschwander, A.G., et al., Amyotrophic lateral sclerosis risk for spinocerebellar ataxia type 2 ATXN2 CAG repeat alleles: a meta-analysis. JAMA Neurol, 2014. 71(12): p. 1529-34.
- Vieira de Sa, R., et al., ATAXIN-2 intermediate-length polyglutamine expansions elicit ALS-associated metabolic and immune phenotypes. Nat Commun, 2024. 15(1): p. 7484.
- Auburger, G.W.J., et al., Bioinformatic Analyses of the Ataxin-2 Family Since Algae Emphasize Its Small Isoforms, Large Chimerisms, and the Importance of Human Exon 1B as Target of Therapies to Prevent Neurodegeneration. Int J Mol Sci, 2026. 27(3).
- Satterfield, T.F., S.M. Jackson, and L.J. Pallanck, A Drosophila homolog of the polyglutamine disease gene SCA2 is a dosage-sensitive regulator of actin filament formation. Genetics, 2002. 162(4): p. 1687-702.
- Albrecht, M., et al., Structural and functional analysis of ataxin-2 and ataxin-3. Eur J Biochem, 2004. 271(15): p. 3155-70.
- Jimenez-Lopez, D. and P. Guzman, Insights into the evolution and domain structure of Ataxin-2 proteins across eukaryotes. BMC Res Notes, 2014. 7: p. 453.
- Auburger, G., et al., Efficient Prevention of Neurodegenerative Diseases by Depletion of Starvation Response Factor Ataxin-2. Trends Neurosci, 2017. 40(8): p. 507-516.
- Boeynaems, S., et al., Poly(A)-binding protein is an ataxin-2 chaperone that regulates biomolecular condensates. Mol Cell, 2023. 83(12): p. 2020-2034 e6.
- Bakthavachalu, B., et al., RNP-Granule Assembly via Ataxin-2 Disordered Domains Is Required for Long-Term Memory and Neurodegeneration. Neuron, 2018. 98(4): p. 754-766 e4.
- Yokoshi, M., et al., Direct binding of Ataxin-2 to distinct elements in 3’ UTRs promotes mRNA stability and protein expression. Mol Cell, 2014. 55(2): p. 186-98.
- Nonhoff, U., et al., Ataxin-2 interacts with the DEAD/H-box RNA helicase DDX6 and interferes with P-bodies and stress granules. Mol Biol Cell, 2007. 18(4): p. 1385-96.
- Wang, J.Y., et al., PolyQ-expanded ataxin-2 aggregation impairs cellular processing-body homeostasis via sequestering the RNA helicase DDX6. J Biol Chem, 2024. 300(7): p. 107413.
- Damrath, E., et al., ATXN2-CAG42 sequesters PABPC1 into insolubility and induces FBXW8 in cerebellum of old ataxic knock-in mice. PLoS Genet, 2012. 8(8): p. e1002920.
- Key, J., et al., Mid-Gestation lethality of Atxn2l-Ablated Mice. Int J Mol Sci, 2020. 21(14).
- Key, J., et al., ATXN2L primarily interacts with NUFIP2, the absence of ATXN2L results in NUFIP2 depletion, and the ATXN2-polyQ expansion triggers NUFIP2 accumulation. Neurobiol Dis, 2025. 209: p. 106903.
- Key, J., et al., Conditional ATXN2L-Null in Adult Frontal Cortex CamK2a+ Neurons Does Not Cause Cell Death but Restricts Spontaneous Mobility and Affects the Alternative Splicing Pathway. Cells, 2025. 14(19).
- Lastres-Becker, I., et al., Insulin receptor and lipid metabolism pathology in ataxin-2 knock-out mice. Hum Mol Genet, 2008. 17(10): p. 1465-81.
- Lastres-Becker, I., et al., Mammalian ataxin-2 modulates translation control at the pre-initiation complex via PI3K/mTOR and is induced by starvation. Biochim Biophys Acta, 2016. 1862(9): p. 1558-69.
- Kiehl, T.R., et al., Generation and characterization of Sca2 (ataxin-2) knockout mice. Biochem Biophys Res Commun, 2006. 339(1): p. 17-24.
- Fittschen, M., et al., Genetic ablation of ataxin-2 increases several global translation factors in their transcript abundance but decreases translation rate. Neurogenetics, 2015. 16(3): p. 181-92.
- Ciosk, R., M. DePalma, and J.R. Priess, ATX-2, the C. elegans ortholog of ataxin 2, functions in translational regulation in the germline. Development, 2004. 131(19): p. 4831-41.
- Kiehl, T.R., H. Shibata, and S.M. Pulst, The ortholog of human ataxin-2 is essential for early embryonic patterning in C. elegans. J Mol Neurosci, 2000. 15(3): p. 231-41.
- Bar, D.Z., et al., Cell size and fat content of dietary-restricted Caenorhabditis elegans are regulated by ATX-2, an mTOR repressor. Proc Natl Acad Sci U S A, 2016. 113(32): p. E4620-9.
- Lim, C. and R. Allada, ATAXIN-2 activates PERIOD translation to sustain circadian rhythms in Drosophila. Science, 2013. 340(6134): p. 875-9.
- McCann, C., et al., The Ataxin-2 protein is required for microRNA function and synapse-specific long-term olfactory habituation. Proc Natl Acad Sci U S A, 2011. 108(36): p. E655-62.
- Lee, J., et al., LSM12 and ME31B/DDX6 Define Distinct Modes of Posttranscriptional Regulation by ATAXIN-2 Protein Complex in Drosophila Circadian Pacemaker Neurons. Mol Cell, 2017. 66(1): p. 129-140 e7.
- Zhang, Y., et al., A role for Drosophila ATX2 in activation of PER translation and circadian behavior. Science, 2013. 340(6134): p. 879-82.
- Kimura, Y., K. Irie, and K. Irie, Pbp1 is involved in Ccr4- and Khd1-mediated regulation of cell growth through association with ribosomal proteins Rpl12a and Rpl12b. Eukaryot Cell, 2013. 12(6): p. 864-74.
- Swisher, K.D. and R. Parker, Localization to, and effects of Pbp1, Pbp4, Lsm12, Dhh1, and Pab1 on stress granules in Saccharomyces cerevisiae. PLoS One, 2010. 5(4): p. e10006.
- Meierhofer, D., et al., Ataxin-2 (Atxn2)-Knock-Out Mice Show Branched Chain Amino Acids and Fatty Acids Pathway Alterations. Mol Cell Proteomics, 2016. 15(5): p. 1728-39.
- Seidel, G., et al., Quantitative Global Proteomics of Yeast PBP1 Deletion Mutants and Their Stress Responses Identifies Glucose Metabolism, Mitochondrial, and Stress Granule Changes. J Proteome Res, 2017. 16(2): p. 504-515.
- van de Loo, S., et al., Ataxin-2 associates with rough endoplasmic reticulum. Exp Neurol, 2009. 215(1): p. 110-8.
- Satterfield, T.F. and L.J. Pallanck, Ataxin-2 and its Drosophila homolog, ATX2, physically assemble with polyribosomes. Hum Mol Genet, 2006. 15(16): p. 2523-32.
- Dansithong, W., et al., Ataxin-2 regulates RGS8 translation in a new BAC-SCA2 transgenic mouse model. PLoS Genet, 2015. 11(4): p. e1005182.
- van de Poll, F., et al., Pbp1 associates with Puf3 and promotes translation of its target mRNAs involved in mitochondrial biogenesis. PLoS Genet, 2023. 19(5): p. e1010774.
- Lopez-Juarez, Z.M., L. Aguilar-Henonin, and P. Guzman, The ATXN2 Orthologs CID3 and CID4, Act Redundantly to In-Fluence Developmental Pathways throughout the Life Cycle of Arabidopsis thaliana. Int J Mol Sci, 2021. 22(6).
- Kaehler, C., et al., Ataxin-2-like is a regulator of stress granules and processing bodies. PLoS One, 2012. 7(11): p. e50134.
- Sen, N.E., et al., Search for SCA2 blood RNA biomarkers highlights Ataxin-2 as strong modifier of the mitochondrial factor PINK1 levels. Neurobiol Dis, 2016. 96: p. 115-126.
- Zhang, K., et al., Stress Granule Assembly Disrupts Nucleocytoplasmic Transport. Cell, 2018. 173(4): p. 958-971 e17.
- Yamagishi, R., et al., Concerted action of ataxin-2 and PABPC1-bound mRNA poly(A) tail in the formation of stress granules. Nucleic Acids Res, 2024. 52(15): p. 9193-9209.
- Koppenol, R., et al., The stress granule protein G3BP1 alleviates spinocerebellar ataxia-associated deficits. Brain, 2023. 146(6): p. 2346-2363.
- Santos, C.C., et al., Fbxo42 promotes the degradation of Ataxin-2 granules to trigger terminal Xbp1 signaling. Nat Commun, 2025. 16(1): p. 7523.
- Del Castillo, U., et al., Ataxin-2 is essential for cytoskeletal dynamics and neurodevelopment in Drosophila. iScience, 2022. 25(1): p. 103536.
- Petrauskas, A., et al., Structured and disordered regions of Ataxin-2 contribute differently to the specificity and efficiency of mRNP granule formation. PLoS Genet, 2024. 20(5): p. e1011251.
- Singh, A., et al., Antagonistic roles for Ataxin-2 structured and disordered domains in RNP condensation. Elife, 2021. 10.
- Sudhakaran, I.P., et al., FMRP and Ataxin-2 function together in long-term olfactory habituation and neuronal translational control. Proc Natl Acad Sci U S A, 2014. 111(1): p. E99-E108.
- Paul, S., et al., Cerebellar Micro-RNA Profile in a Mouse Model of Spinocerebellar Ataxia Type 2. Neurol Genet, 2024. 10(2): p. e200144.
- Sen, N.E., et al., Generation of an Atxn2-CAG100 knock-in mouse reveals N-acetylaspartate production deficit due to early Nat8l dysregulation. Neurobiol Dis, 2019. 132: p. 104559.
- Canet-Pons, J., et al., Atxn2-CAG100-KnockIn mouse spinal cord shows progressive TDP43 pathology associated with cholesterol biosynthesis suppression. Neurobiol Dis, 2021. 152: p. 105289.
- Leites, E.P. and V.A. Morais, Protocol for the isolation and culture of microglia, astrocytes, and neurons from the same mouse brain. STAR Protoc, 2024. 5(1): p. 102869.
- Flury, A., et al., Evolving insights into the identity and function of dark microglia. Trends Neurosci, 2026. 49(1): p. 20-34.
- Hume, D.A., Life without microglia. Trends Neurosci, 2025. 48(8): p. 560-569.
- Pereira-Iglesias, M., et al., Microglia as hunters or gatherers of brain synapses. Nat Neurosci, 2025. 28(1): p. 15-23.
- Arranz, A.M. and B. De Strooper, The role of astroglia in Alzheimer’s disease: pathophysiology and clinical implications. Lancet Neurol, 2019. 18(4): p. 406-414.
- Allen, N.J. and D.A. Lyons, Glia as architects of central nervous system formation and function. Science, 2018. 362(6411): p. 181-185.
- Slezak, M., F.W. Pfrieger, and Z. Soltys, Synaptic plasticity, astrocytes and morphological homeostasis. J Physiol Paris, 2006. 99(2-3): p. 84-91.
- Houmam, S., et al., Protocol to sequentially isolate mouse oligodendrocytes, microglia, endothelial cells, astrocytes, and neurons via magnetic cell sorting. STAR Protoc, 2025. 6(4): p. 104139.
- Clarke, L.E., et al., Normal aging induces A1-like astrocyte reactivity. Proc Natl Acad Sci U S A, 2018. 115(8): p. E1896-E1905.
- Bouza, A.A. and L.L. Isom, Voltage-Gated Sodium Channel beta Subunits and Their Related Diseases. Handb Exp Pharmacol, 2018. 246: p. 423-450.
- Chen, S., et al., Regulation of SPARC family proteins in disorders of the central nervous system. Brain Res Bull, 2020. 163: p. 178-189.
- Morel, M., K.N. Shah, and W. Long, The F-box protein FBXL16 up-regulates the stability of C-MYC oncoprotein by antagonizing the activity of the F-box protein FBW7. J Biol Chem, 2020. 295(23): p. 7970-7980.
- Schonkeren, S.L., et al., Nervous NDRGs: the N-myc downstream-regulated gene family in the central and peripheral nervous system. Neurogenetics, 2019. 20(4): p. 173-186.
- Lin, X., et al., LncRNA MEG8 ameliorates Parkinson’s disease neuro-inflammation through miR-485-3p/FBXO45 axis. Acta Neurol Belg, 2024. 124(2): p. 549-557.
- Genetic Modifiers of Huntington’s Disease, C., Identification of Genetic Factors that Modify Clinical Onset of Huntington’s Disease. Cell, 2015. 162(3): p. 516-26.
- Loupe, J.M., et al., Promotion of somatic CAG repeat expansion by Fan1 knock-out in Huntington’s disease knock-in mice is blocked by Mlh1 knock-out. Hum Mol Genet, 2020. 29(18): p. 3044-3053.
- McAllister, B., et al., Exome sequencing of individuals with Huntington’s disease implicates FAN1 nuclease activity in slowing CAG expansion and disease onset. Nat Neurosci, 2022. 25(4): p. 446-457.
- Kam, T.I., et al., Microglia and astrocyte dysfunction in parkinson’s disease. Neurobiol Dis, 2020. 144: p. 105028.
- Mendonca, P., E. Taka, and K.F.A. Soliman, Proteomic analysis of the effect of the polyphenol pentagalloyl glucose on proteins involved in neurodegenerative diseases in activated BV-2 microglial cells. Mol Med Rep, 2019. 20(2): p. 1736-1746.
- Capuz, A., et al., Astrocytes express aberrant immunoglobulins as putative gatekeeper of astrocytes to neuronal progenitor conversion. Cell Death Dis, 2023. 14(4): p. 237.
- Almaguer-Mederos, L.E., et al., Multiomics approach identifies SERPINB1 as candidate biomarker for spinocerebellar ataxia type 2. Sci Rep, 2025. 15(1): p. 42559.
- Scoles, D.R., et al., ALS-associated genes in SCA2 mouse spinal cord transcriptomes. Hum Mol Genet, 2020. 29(10): p. 1658-1672.
- Sen, N.E., et al., In Human and Mouse Spino-Cerebellar Tissue, Ataxin-2 Expansion Affects Ceramide-Sphingomyelin Metabolism. Int J Mol Sci, 2019. 20(23).
- Pflieger, L.T., et al., Gene co-expression network analysis for identifying modules and functionally enriched pathways in SCA2. Hum Mol Genet, 2017. 26(16): p. 3069-3080.
- Halbach, M.V., et al., Atxn2 Knockout and CAG42-Knock-in Cerebellum Shows Similarly Dysregulated Expression in Calcium Homeostasis Pathway. Cerebellum, 2017. 16(1): p. 68-81.
- Halbach, M.V., et al., Both ubiquitin ligases FBXW8 and PARK2 are sequestrated into insolubility by ATXN2 PolyQ expansions, but only FBXW8 expression is dysregulated. PLoS One, 2015. 10(3): p. e0121089.
- Almaguer-Mederos, L.E., et al., Spinal Cord Phosphoproteome of SCA2 Mouse Model Reveals Alteration of ATXN2-N-Term PRM-SH3-Actin Interactome and of Autophagy. Mol Cell Proteomics, 2025. 24(11): p. 101072.
- Diaz-Ortiz, M.E., et al., GPNMB confers risk for Parkinson’s disease through interaction with alpha-synuclein. Science, 2022. 377(6608): p. eabk0637.
- Huttenrauch, M., et al., Glycoprotein NMB: a novel Alzheimer’s disease associated marker expressed in a subset of activated microglia. Acta Neuropathol Commun, 2018. 6(1): p. 108.
- Li, X., et al., GPNMB Modulates Autophagy to Enhance Functional Recovery After Spinal Cord Injury in Rats. Cell Transplant, 2024. 33: p. 9636897241233040.
- Zhang, X., et al., Repair-associated macrophages increase after early-phase microglia attenuation to promote ischemic stroke recovery. Nat Commun, 2025. 16(1): p. 3089.
- Li, J., et al., Exploring the factors underlying remyelination arrest by studying the post-transcriptional regulatory mechanisms of cystatin F gene. J Neurochem, 2021. 157(6): p. 2070-2090.
- Rasmussen, J., et al., An early proinflammatory transcriptional response to tau pathology is age-specific and foreshadows reduced tau burden. Brain Pathol, 2022. 32(3): p. e13018.
- Zhou, H., et al., Integrating Bulk and Single-Cell Transcriptomic Data to Identify Ferroptosis-Associated Inflammatory Gene in Alzheimer’s Disease. J Inflamm Res, 2025. 18: p. 2105-2122.
- Yin, Z., et al., APOE4 impairs the microglial response in Alzheimer’s disease by inducing TGFbeta-mediated checkpoints. Nat Immunol, 2023. 24(11): p. 1839-1853.
- Arutyunov, A., et al., West Nile Virus-Induced Expression of Senescent Gene Lgals3bp Regulates Microglial Phenotype within Cerebral Cortex. Biomolecules, 2024. 14(7).
- Kohno, K., et al., A spinal microglia population involved in remitting and relapsing neuropathic pain. Science, 2022. 376(6588): p. 86-90.
- De Schepper, S., et al., Perivascular cells induce microglial phagocytic states and synaptic engulfment via SPP1 in mouse models of Alzheimer’s disease. Nat Neurosci, 2023. 26(3): p. 406-415.
- Nguyen, A.T., et al., APOE and TREM2 regulate amyloid-responsive microglia in Alzheimer’s disease. Acta Neuropathol, 2020. 140(4): p. 477-493.
- Huang, Y., et al., Microglia use TAM receptors to detect and engulf amyloid beta plaques. Nat Immunol, 2021. 22(5): p. 586-594.
- Schwabenland, M., et al., Loss of USP18 in microglia induces white matter pathology. Acta Neuropathol Commun, 2019. 7(1): p. 106.
- Wu, Z., et al., Abeta Induces Neuroinflammation and Microglial M1 Polarization via cGAS-STING-IFITM3 Signaling Pathway in BV-2 Cells. Neurochem Res, 2023. 48(9): p. 2881-2894.
- Li, W., et al., Rnf-213 Knockout Induces Pericyte Reduction and Blood-Brain Barrier Impairment in Mouse. Mol Neurobiol, 2023. 60(11): p. 6188-6200.
- Negrutskii, B.S., et al., The eEF1 family of mammalian translation elongation factors. BBA Adv, 2023. 3: p. 100067.
- Biembengut, I.V., et al., Cytoplasmic FMR1 interacting protein (CYFIP) family members and their function in neural development and disorders. Mol Biol Rep, 2021. 48(8): p. 6131-6143.
- Zhang, S., et al., The LSmAD Domain of Ataxin-2 Modulates the Structure and RNA Binding of Its Preceding LSm Domain. Cells, 2025. 14(5).
- Achsel, T., H. Stark, and R. Luhrmann, The Sm domain is an ancient RNA-binding motif with oligo(U) specificity. Proc Natl Acad Sci U S A, 2001. 98(7): p. 3685-9.
- Weichenrieder, O., RNA binding by Hfq and ring-forming (L)Sm proteins: a trade-off between optimal sequence readout and RNA backbone conformation. RNA Biol, 2014. 11(5): p. 537-49.
- Mu, L., et al., Interconversion and mechanisms between Lsm-type and Sm-type heteroheptameric rings: implications for spliceosome evolution and RNA metabolism. Nucleic Acids Res, 2025. 53(10).
- Kaida, D., et al., U1 snRNP protects pre-mRNAs from premature cleavage and polyadenylation. Nature, 2010. 468(7324): p. 664-8.
- Cohen, J.B., S.D. Broz, and A.D. Levinson, U1 small nuclear RNAs with altered specificity can be stably expressed in mammalian cells and promote permanent changes in pre-mRNA splicing. Mol Cell Biol, 1993. 13(5): p. 2666-76.
- Gadgil, A., et al., ALS-linked FUS mutants affect the localization of U7 snRNP and replication-dependent histone gene expression in human cells. Sci Rep, 2021. 11(1): p. 11868.
- Nakashima, T., et al., Diversity of U1 Small Nuclear RNAs and Diagnostic Methods for Their Mutations. Cancer Sci, 2025. 116(8): p. 2270-2280.
- Suzuki, H., et al., Recurrent noncoding U1 snRNA mutations drive cryptic splicing in SHH medulloblastoma. Nature, 2019. 574(7780): p. 707-711.
- Nojima, H. and R.D. Kornberg, Genes and pseudogenes for mouse U1 and U2 small nuclear RNAs. J Biol Chem, 1983. 258(13): p. 8151-5.
- Kato, N. and F. Harada, New U1 RNA species found in Friend SFFV (spleen focus forming virus)-transformed mouse cells. J Biol Chem, 1985. 260(12): p. 7775-82.
- Howard, E.F., et al., Functional, developmentally expressed genes for mouse U1a and U1b snRNAs contain both conserved and non-conserved transcription signals. Nucleic Acids Res, 1986. 14(24): p. 9811-25.
- Lund, E., J.E. Dahlberg, and D.J. Forbes, The two embryonic U1 small nuclear RNAs of Xenopus laevis are encoded by a major family of tandemly repeated genes. Mol Cell Biol, 1984. 4(12): p. 2580-6.
- Marzluff, W.F., et al., Isolation and characterization of two linked mouse U1b small nuclear RNA genes. Nucleic Acids Res, 1983. 11(18): p. 6255-70.
- Lund, E., B. Kahan, and J.E. Dahlberg, Differential control of U1 small nuclear RNA expression during mouse development. Science, 1985. 229(4719): p. 1271-4.
- Forbes, D.J., et al., Differential expression of multiple U1 small nuclear RNAs in oocytes and embryos of Xenopus laevis. Cell, 1984. 38(3): p. 681-9.
- Lund, E. and J.E. Dahlberg, Differential accumulation of U1 and U4 small nuclear RNAs during Xenopus development. Genes Dev, 1987. 1(1): p. 39-46.
- Liu, J.L. and J.G. Gall, U bodies are cytoplasmic structures that contain uridine-rich small nuclear ribonucleoproteins and associate with P bodies. Proc Natl Acad Sci U S A, 2007. 104(28): p. 11655-9.
- Buckingham, M. and J.L. Liu, U bodies respond to nutrient stress in Drosophila. Exp Cell Res, 2011. 317(20): p. 2835-44.
- Tsalikis, J., et al., Intracellular Bacterial Pathogens Trigger the Formation of U Small Nuclear RNA Bodies (U Bodies) through Metabolic Stress Induction. J Biol Chem, 2015. 290(34): p. 20904-20918.
- Lee, L., S.E. Davies, and J.L. Liu, The spinal muscular atrophy protein SMN affects Drosophila germline nuclear organization through the U body-P body pathway. Dev Biol, 2009. 332(1): p. 142-55.
- Patel, S.B. and M. Bellini, The assembly of a spliceosomal small nuclear ribonucleoprotein particle. Nucleic Acids Res, 2008. 36(20): p. 6482-93.
- Hamm, J., et al., The trimethylguanosine cap structure of U1 snRNA is a component of a bipartite nuclear targeting signal. Cell, 1990. 62(3): p. 569-77.
- Hyjek, M., et al., Spatial regulation of cytoplasmic snRNP assembly at the cellular level. J Exp Bot, 2015. 66(22): p. 7019-30.
- Rossoll, W., et al., Smn, the spinal muscular atrophy-determining gene product, modulates axon growth and localization of beta-actin mRNA in growth cones of motoneurons. J Cell Biol, 2003. 163(4): p. 801-12.
- Eggert, C., et al., Spinal muscular atrophy: the RNP connection. Trends Mol Med, 2006. 12(3): p. 113-21.
- Piazzon, N., et al., In vitro and in cellulo evidences for association of the survival of motor neuron complex with the fragile X mental retardation protein. J Biol Chem, 2008. 283(9): p. 5598-610.
- Baumer, D., et al., Alternative splicing events are a late feature of pathology in a mouse model of spinal muscular atrophy. PLoS Genet, 2009. 5(12): p. e1000773.
- Tapia, O., et al., Cellular bases of the RNA metabolism dysfunction in motor neurons of a murine model of spinal muscular atrophy: Role of Cajal bodies and the nucleolus. Neurobiol Dis, 2017. 108: p. 83-99.
- Beattie, C.E. and S.J. Kolb, Spinal muscular atrophy: Selective motor neuron loss and global defect in the assembly of ribonucleoproteins. Brain Res, 2018. 1693(Pt A): p. 92-97.
- Koyano, S., et al., Parallel Appearance of Polyglutamine and Transactivation-Responsive DNA-Binding Protein 43 and Their Complementary Subcellular Localization in Brains of Patients With Spinocerebellar Ataxia Type 2. J Neuropathol Exp Neurol, 2022. 81(7): p. 535-544.
- van de Warrenburg, B.P., et al., Peripheral nerve involvement in spinocerebellar ataxias. Arch Neurol, 2004. 61(2): p. 257-61.
- Abele, M., et al., Autosomal dominant cerebellar ataxia type I. Nerve conduction and evoked potential studies in families with SCA1, SCA2 and SCA3. Brain, 1997. 120 ( Pt 12): p. 2141-8.
- Shin, M.S., et al., U1-small nuclear ribonucleoprotein activates the NLRP3 inflammasome in human monocytes. J Immunol, 2012. 188(10): p. 4769-75.
- Somarelli, J.A., et al., Epitope mapping of the U1 small nuclear ribonucleoprotein particle in patients with systemic lupus erythematosus and mixed connective tissue disease. Lupus, 2011. 20(3): p. 274-89.
- Sato, T., et al., Anti-U1 RNP antibodies in cerebrospinal fluid are associated with central neuropsychiatric manifestations in systemic lupus erythematosus and mixed connective tissue disease. Arthritis Rheum, 2010. 62(12): p. 3730-40.
- Nadeu, F., et al., Disease-specific U1 spliceosomal RNA mutations in mature B-cell neoplasms. Leukemia, 2025. 39(9): p. 2076-2086.
- Cheng, Z., et al., Overexpression of U1 snRNA induces decrease of U1 spliceosome function associated with Alzheimer’s disease. J Neurogenet, 2017. 31(4): p. 337-343.
- Dong, X., et al., Small Nuclear RNAs (U1, U2, U5) in Tumor-Educated Platelets Are Downregulated and Act as Promising Biomarkers in Lung Cancer. Front Oncol, 2020. 10: p. 1627.
- Pigini, P., F.M. Giorgi, and K.B. Wee, Utargetome: A targetome prediction tool for modified U1-snRNAs to identify distal-target positions with improved selectivity. PLoS Comput Biol, 2025. 21(9): p. e1013534.
- Peretto, L., et al., Rescue of a panel of Hemophilia A-causing 5’ss splicing mutations by unique Exon-specific U1snRNA variants. Mol Med, 2025. 31(1): p. 121.
- Goncalves, M., et al., Development of Engineered-U1 snRNA Therapies: Current Status. Int J Mol Sci, 2023. 24(19).
- Romano, G., et al., Rescue of a familial dysautonomia mouse model by AAV9-Exon-specific U1 snRNA. Am J Hum Genet, 2022. 109(8): p. 1534-1548.
- Hatch, S.T., A.A. Smargon, and G.W. Yeo, Engineered U1 snRNAs to modulate alternatively spliced exons. Methods, 2022. 205: p. 140-148.
- Donega, S., et al., Rescue of common exon-skipping mutations in cystic fibrosis with modified U1 snRNAs. Hum Mutat, 2020. 41(12): p. 2143-2154.
- Breuel, S., et al., Combining Engineered U1 snRNA and Antisense Oligonucleotides to Improve the Treatment of a BBS1 Splice Site Mutation. Mol Ther Nucleic Acids, 2019. 18: p. 123-130.
- Rogalska, M.E., et al., Therapeutic activity of modified U1 core spliceosomal particles. Nat Commun, 2016. 7: p. 11168.
- Liu, S., et al., The network of protein-protein interactions within the human U4/U6.U5 tri-snRNP. RNA, 2006. 12(7): p. 1418-30.
- Zhang, X., et al., An Atomic Structure of the Human Spliceosome. Cell, 2017. 169(5): p. 918-929 e14.
- Zhang, X., et al., Structure of the human activated spliceosome in three conformational states. Cell Res, 2018. 28(3): p. 307-322.
- Yang, G., et al., Spliceosomal GTPase Eftud2 deficiency-triggered ferroptosis leads to Purkinje cell degeneration. Neuron, 2024. 112(20): p. 3452-3469 e9.
- Park, B.Y., et al., The Core Splicing Factors EFTUD2, SNRPB and TXNL4A Are Essential for Neural Crest and Craniofacial Development. J Dev Biol, 2022. 10(3).
- Boudreault, S., G. Lemay, and M. Bisaillon, U5 snRNP Core Proteins Are Key Components of the Defense Response against Viral Infection through Their Roles in Programmed Cell Death and Interferon Induction. Viruses, 2022. 14(12).
- De Arras, L., et al., Comparative genomics RNAi screen identifies Eftud2 as a novel regulator of innate immunity. Genetics, 2014. 197(2): p. 485-96.
- Lu, W., et al., Nesprin interchain associations control nuclear size. Cell Mol Life Sci, 2012. 69(20): p. 3493-509.
- Xu, S., et al., O-GlcNAcylation reduces proteome solubility and regulates the formation of biomolecular condensates in human cells. Nat Commun, 2025. 16(1): p. 4068.
- Zhang, X., et al., Syne-1 and Syne-2 play crucial roles in myonuclear anchorage and motor neuron innervation. Development, 2007. 134(5): p. 901-8.
- Yu, J., et al., KASH protein Syne-2/Nesprin-2 and SUN proteins SUN1/2 mediate nuclear migration during mammalian retinal development. Hum Mol Genet, 2011. 20(6): p. 1061-73.
- Luke, Y., et al., Nesprin-2 Giant (NUANCE) maintains nuclear envelope architecture and composition in skin. J Cell Sci, 2008. 121(11): p. 1887-98.
- Zhang, Q., et al., Nesprin-2 is a multi-isomeric protein that binds lamin and emerin at the nuclear envelope and forms a subcellular network in skeletal muscle. J Cell Sci, 2005. 118(Pt 4): p. 673-87.
- Dawe, H.R., et al., Nesprin-2 interacts with meckelin and mediates ciliogenesis via remodelling of the actin cytoskeleton. J Cell Sci, 2009. 122(Pt 15): p. 2716-26.
- Zhang, X., et al., SUN1/2 and Syne/Nesprin-1/2 complexes connect centrosome to the nucleus during neurogenesis and neuronal migration in mice. Neuron, 2009. 64(2): p. 173-87.
- Synofzik, M., et al., SYNE1 ataxia is a common recessive ataxia with major non-cerebellar features: a large multi-centre study. Brain, 2016. 139(Pt 5): p. 1378-93.
- Senghor, H.V.F., et al., SYNE1 Deficiency Manifesting Primarily With Motor Neuron Disease. Neurol Genet, 2025. 11(6): p. e200306.
- Bonnycastle, K., P.C. Kind, and M.A. Cousin, FMRP Sustains Presynaptic Function via Control of Activity-Dependent Bulk Endocytosis. J Neurosci, 2022. 42(8): p. 1618-1628.
- Ceman, S., R. Nelson, and S.T. Warren, Identification of mouse YB1/p50 as a component of the FMRP-associated mRNP particle. Biochem Biophys Res Commun, 2000. 279(3): p. 904-8.
- Malone, T.J., et al., Neuronal potassium channel activity triggers initiation of mRNA translation through binding of translation regulators. Sci Adv, 2025. 11(22): p. eadv3140.
- Zhang, Y., Y. Lee, and K. Han, Neuronal function and dysfunction of CYFIP2: from actin dynamics to early infantile epileptic encephalopathy. BMB Rep, 2019. 52(5): p. 304-311.
- Cioni, J.M., et al., Axon-Axon Interactions Regulate Topographic Optic Tract Sorting via CYFIP2-Dependent WAVE Complex Function. Neuron, 2018. 97(5): p. 1078-1093 e6.
- Santini, E., et al., Reducing eIF4E-eIF4G interactions restores the balance between protein synthesis and actin dynamics in fragile X syndrome model mice. Sci Signal, 2017. 10(504).
- Bardoni, B., A. Schenck, and J.L. Mandel, A novel RNA-binding nuclear protein that interacts with the fragile X mental retardation (FMR1) protein. Hum Mol Genet, 1999. 8(13): p. 2557-66.
- Bardoni, B., et al., NUFIP1 (nuclear FMRP interacting protein 1) is a nucleocytoplasmic shuttling protein associated with active synaptoneurosomes. Exp Cell Res, 2003. 289(1): p. 95-107.
- Taha, M.S., et al., Novel FMRP interaction networks linked to cellular stress. FEBS J, 2021. 288(3): p. 837-860.
- Jia, J., et al., Membrane Atg8ylation, stress granule formation, and MTOR regulation during lysosomal damage. Autophagy, 2023. 19(6): p. 1893-1895.
- Ramos, A., et al., The structure of the N-terminal domain of the fragile X mental retardation protein: a platform for protein-protein interaction. Structure, 2006. 14(1): p. 21-31.
- Bardoni, B., et al., 82-FIP, a novel FMRP (fragile X mental retardation protein) interacting protein, shows a cell cycle-dependent intracellular localization. Hum Mol Genet, 2003. 12(14): p. 1689-98.
- Tan, W., et al., Zfrp8 forms a complex with fragile-X mental retardation protein and regulates its localization and function. Dev Biol, 2016. 410(2): p. 202-212.
- Schenck, A., et al., A highly conserved protein family interacting with the fragile X mental retardation protein (FMRP) and displaying selective interactions with FMRP-related proteins FXR1P and FXR2P. Proc Natl Acad Sci U S A, 2001. 98(15): p. 8844-9.
- Abekhoukh, S., et al., New insights into the regulatory function of CYFIP1 in the context of WAVE- and FMRP-containing complexes. Dis Model Mech, 2017. 10(4): p. 463-474.
- Richter, J.D. and X. Zhao, The molecular biology of FMRP: new insights into fragile X syndrome. Nat Rev Neurosci, 2021. 22(4): p. 209-222.
- La Via, L., et al., Functional impact of CYFIP2 RNA editing on actin regulation, axon growth, and spinogenesis. Neurochem Int, 2025. 191: p. 106084.
- Kim, N.S., et al., CYFIP1 Dosages Exhibit Divergent Behavioral Impact via Diametric Regulation of NMDA Receptor Complex Translation in Mouse Models of Psychiatric Disorders. Biol Psychiatry, 2022. 92(10): p. 815-826.
- Majumder, P., et al., Co-regulation of mRNA translation by TDP-43 and Fragile X Syndrome protein FMRP. Acta Neuropathol, 2016. 132(5): p. 721-738.
- Di Marino, D., et al., A unique binding mode of the eukaryotic translation initiation factor 4E for guiding the design of novel peptide inhibitors. Protein Sci, 2015. 24(9): p. 1370-82.
- Genheden, M., et al., BDNF stimulation of protein synthesis in cortical neurons requires the MAP kinase-interacting kinase MNK1. J Neurosci, 2015. 35(3): p. 972-84.
- Oguro-Ando, A., et al., Increased CYFIP1 dosage alters cellular and dendritic morphology and dysregulates mTOR. Mol Psychiatry, 2015. 20(9): p. 1069-78.
- De Rubeis, S., et al., CYFIP1 coordinates mRNA translation and cytoskeleton remodeling to ensure proper dendritic spine formation. Neuron, 2013. 79(6): p. 1169-82.
- Bozdagi, O., et al., Haploinsufficiency of Cyfip1 produces fragile X-like phenotypes in mice. PLoS One, 2012. 7(8): p. e42422.
- Napoli, I., et al., The fragile X syndrome protein represses activity-dependent translation through CYFIP1, a new 4E-BP. Cell, 2008. 134(6): p. 1042-54.
- Schenck, A., et al., CYFIP/Sra-1 controls neuronal connectivity in Drosophila and links the Rac1 GTPase pathway to the fragile X protein. Neuron, 2003. 38(6): p. 887-98.
- Stubenvoll, M.D., et al., ATX-2, the C. elegans Ortholog of Human Ataxin-2, Regulates Centrosome Size and Microtubule Dynamics. PLoS Genet, 2016. 12(9): p. e1006370.
- Cha, I.J., et al., Ataxin-2 Dysregulation Triggers a Compensatory Fragile X Mental Retardation Protein Decrease in Drosophila C4da Neurons. Mol Cells, 2020. 43(10): p. 870-879.
- Ikeda, Y., et al., Spectrin mutations cause spinocerebellar ataxia type 5. Nat Genet, 2006. 38(2): p. 184-90.
- Dick, K.A., et al., Spinocerebellar ataxia type 5. Handb Clin Neurol, 2012. 103: p. 451-9.
- Yildiz Bolukbasi, E., et al., Progressive SCAR14 with unclear speech, developmental delay, tremor, and behavioral problems caused by a homozygous deletion of the SPTBN2 pleckstrin homology domain. Am J Med Genet A, 2017. 173(9): p. 2494-2499.
- Nicita, F., et al., Heterozygous missense variants of SPTBN2 are a frequent cause of congenital cerebellar ataxia. Clin Genet, 2019. 96(2): p. 169-175.
- Perkins, E.M., et al., Posterior cerebellar Purkinje cells in an SCA5/SPARCA1 mouse model are especially vulnerable to the synergistic effect of loss of beta-III spectrin and GLAST. Hum Mol Genet, 2016. 25(20): p. 4448-4461.
- Armbrust, K.R., et al., Mutant beta-III spectrin causes mGluR1alpha mislocalization and functional deficits in a mouse model of spinocerebellar ataxia type 5. J Neurosci, 2014. 34(30): p. 9891-904.
- Bar, J., et al., Periodic F-actin structures shape the neck of dendritic spines. Sci Rep, 2016. 6: p. 37136.
- Pielage, J., R.D. Fetter, and G.W. Davis, A postsynaptic spectrin scaffold defines active zone size, spacing, and efficacy at the Drosophila neuromuscular junction. J Cell Biol, 2006. 175(3): p. 491-503.
- Denha, S.A., et al., Molecular consequences of SCA5 mutations in the spectrin-repeat domains of beta-III-spectrin. J Biol Chem, 2025. 301(7): p. 110350.
- Salcedo-Sicilia, L., et al., betaIII spectrin regulates the structural integrity and the secretory protein transport of the Golgi complex. J Biol Chem, 2013. 288(4): p. 2157-66.
- Lorenzo, D.N., et al., Spectrin mutations that cause spinocerebellar ataxia type 5 impair axonal transport and induce neurodegeneration in Drosophila. J Cell Biol, 2010. 189(1): p. 143-58.
- Perkins, E., D. Suminaite, and M. Jackson, Cerebellar ataxias: beta-III spectrin’s interactions suggest common pathogenic pathways. J Physiol, 2016. 594(16): p. 4661-76.
- Gao, Y., et al., beta-III spectrin is critical for development of purkinje cell dendritic tree and spine morphogenesis. J Neurosci, 2011. 31(46): p. 16581-90.
- Machnicka, B., et al., Spectrins: a structural platform for stabilization and activation of membrane channels, receptors and transporters. Biochim Biophys Acta, 2014. 1838(2): p. 620-34.
- Hulsmeier, J., et al., Distinct functions of alpha-Spectrin and beta-Spectrin during axonal pathfinding. Development, 2007. 134(4): p. 713-22.
- Glineburg, M.R. and C. Nguyen, Diverse roles of stress-responsive RNP granules in oogenesis and infertility. Biol Reprod, 2025. 112(6): p. 1039-1053.
- van Leeuwen, W. and C. Rabouille, Cellular stress leads to the formation of membraneless stress assemblies in eukaryotic cells. Traffic, 2019. 20(9): p. 623-638.
- Ripin, N. and R. Parker, Formation, function, and pathology of RNP granules. Cell, 2023. 186(22): p. 4737-4756.
- Rub, U., et al., Anatomically based guidelines for systematic investigation of the central somatosensory system and their application to a spinocerebellar ataxia type 2 (SCA2) patient. Neuropathol Appl Neurobiol, 2003. 29(5): p. 418-33.
- Restivo, D.A., et al., Central motor conduction to lower limb after transcranial magnetic stimulation in spinocerebellar ataxia type 2 (SCA2). Clin Neurophysiol, 2000. 111(4): p. 630-5.
- Stezin, A., et al., In vivo microstructural white matter changes in early spinocerebellar ataxia 2. Acta Neurol Scand, 2021. 143(3): p. 326-332.
- Velazquez-Perez, L., et al., Abnormal corticospinal tract function and motor cortex excitability in non-ataxic SCA2 mutation carriers: A TMS study. Clin Neurophysiol, 2016. 127(8): p. 2713-2719.
- Velazquez-Perez, L., et al., Progression of corticospinal tract dysfunction in pre-ataxic spinocerebellar ataxia type 2: A two-years follow-up TMS study. Clin Neurophysiol, 2018. 129(5): p. 895-900.
- Paciorkowski, A.R., et al., Massive expansion of SCA2 with autonomic dysfunction, retinitis pigmentosa, and infantile spasms. Neurology, 2011. 77(11): p. 1055-60.
- Chen, J.L., et al., Pseudouridylation-mediated gene expression modulation. Biochem J, 2024. 481(1): p. 1-16.
- Borchardt, E.K., N.M. Martinez, and W.V. Gilbert, Regulation and Function of RNA Pseudouridylation in Human Cells. Annu Rev Genet, 2020. 54: p. 309-336.
- Luo, N., et al., Functions and therapeutic applications of pseudouridylation. Nat Rev Mol Cell Biol, 2025. 26(9): p. 691-705.
- Weissman, D. and K. Kariko, mRNA: Fulfilling the Promise of Gene Therapy. Mol Ther, 2015. 23(9): p. 1416-7.
- Weissman, D., et al., HPLC purification of in vitro transcribed long RNA. Methods Mol Biol, 2013. 969: p. 43-54.
- Becker, L.A., et al., Therapeutic reduction of ataxin-2 extends lifespan and reduces pathology in TDP-43 mice. Nature, 2017. 544(7650): p. 367-371.
- Buenaventura, R.G., et al., Sequential Isolation of Microglia and Astrocytes from Young and Aged Adult Mouse Brains for Downstream Transcriptomic Analysis. Methods Protoc, 2022. 5(5).
- van Niekerk, E.A., et al., Methods for culturing adult CNS neurons reveal a CNS conditioning effect. Cell Rep Methods, 2022. 2(7): p. 100255.
- Livak, K.J. and T.D. Schmittgen, Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods, 2001. 25(4): p. 402-8.
 |
 |
 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




