1. Introduction
Carpal tunnel syndrome (CTS) is the most prevalent entrapment neuropathy and results from compression of the median nerve as it traverses the carpal tunnel [
1]. Clinically, it manifests with pain, numbness, and tingling in the hand, frequently impairing fine motor function and daily activities [
2]. Beyond diagnosis, accurate assessment of CTS severity is clinically relevant, as it directly influences treatment decisions along the diagnostic–therapeutic pathway, particularly when weighing conservative management against surgical decompression. In current clinical practice, the diagnosis and grading of CTS severity primarily rely on nerve conduction studies (NCS), which remain the reference standard for quantifying neuropathic involvement [
3]. Nevertheless, evidence comparing surgical and non-surgical treatments has shown variable results, and it remains unclear which patients derive the greatest benefit from operative intervention [
4]. This variability suggests that stratification strategies based predominantly on clinical presentation and electrodiagnostic severity may be insufficient to fully capture the complexity of CTS. In particular, potentially relevant structural and morphological information, not reflected by NCS alone, may contribute to improved patient selection and more individualized treatment decision-making.
Ultrasound (US) has emerged as a valuable complementary imaging modality for CTS because it enables real-time visualization of the median nerve and surrounding structures, is widely accessible, and is cost-effective compared with electrodiagnostic studies [
5]. Median nerve cross-sectional area (CSA) is the most widely validated sonographic parameter for the diagnosis of CTS [
6]. However, although CSA provides a useful morphological indicator, it does not always capture the full spectrum of structural alterations relevant to CTS. Subtle changes in internal fascicular pattern, echotexture, or epineural definition may occur without a measurable increase in CSA, whereas variations in nerve size can also arise from non-pathological factors such as patient anthropometry or anatomical diversity. Consequently, reliance on gross morphological measurements alone may overlook clinically meaningful tissue changes and contributes to variability between examiners, complicating the establishment of robust and reproducible diagnostic pathways [
7].
To address the limitations of qualitative interpretation, quantitative ultrasound (QUS) approaches have been proposed to characterize internal nerve architecture, echotexture, and border definition [
8]. These techniques aim to reduce subjectivity and capture subtle structural alterations associated with oedema, fibrosis, or axonal degeneration. Yet, most available QUS methods either rely on manual segmentation or analyze only broad regional image statistics, without explicitly modelling the individual diagnostic features that clinicians routinely assess, such as internal tissue quality or clarity of the epineural border [
9]. Consequently, despite their potential, existing QUS approaches have not fully resolved the challenge of objective, reproducible CTS evaluation.
These shortcomings have driven increasing interest in computational methods capable of learning diagnostic patterns directly from ultrasound images, without relying on manual measurements or predefined feature sets. Artificial intelligence, particularly deep learning (DL) using convolutional neural networks (CNNs), has rapidly advanced the field of musculoskeletal ultrasound image analysis [
10]. Early work in this area relied on multi-stage, feature-engineered systems for nerve identification [
11]. The shift toward modern DL architectures, inspired by hierarchical feature learning in biological systems, has enabled automatic learning of complex sonographic patterns without handcrafted feature engineering [
12]. Among these architectures, CNNs have demonstrated exceptional performance in visual recognition tasks and have been increasingly applied to peripheral nerve imaging [
13].
Nevertheless, while DL offers a powerful framework to overcome the subjectivity and limited reproducibility of conventional and quantitative ultrasound analysis, its application to CTS remains incomplete. Most existing models focus narrowly on nerve localization or segmentation, or on binary diagnostic classification, and therefore do not address the clinically essential task of assigning severity grades [
14]. This limitation is particularly relevant from an orthopedic and surgical perspective, as CTS severity plays a central role in guiding treatment strategies, including the choice between conservative management and surgical decompression. Therefore, there remains a clear need for fully automated, operator-independent systems capable not only of identifying and segmenting the median nerve, but also of classifying CTS severity using clinically interpretable categories that may better support treatment stratification. Accordingly, the present study aimed to develop and validate convolutional neural network models capable of automatically classifying CTS severity from standard B-mode ultrasound images. Using electrodiagnostic testing as the reference standard and incorporating both transverse and longitudinal imaging planes, the proposed framework seeks to provide a reproducible, scalable, and clinically interpretable tool for automated CTS severity assessment with potential relevance for clinical decision-making.
2. Materials and Methods
2.1. Study Design
A cross-sectional analytical study was conducted to develop and validate a machine-learning–based approach for the automated diagnostic evaluation of the median nerve in the context of CTS using B-mode ultrasound imaging. The investigation focused on predicting neurophysiological diagnostic labels from ultrasound examinations obtained at the level of the carpal tunnel. Ultrasound acquisition and neurophysiological testing were performed during the same clinical visit, with nerve conduction studies conducted immediately after completion of the ultrasound examination, thereby minimizing potential temporal variability in median nerve status. All study procedures adhered to the principles of the Declaration of Helsinki and received approval from the local ethics committee (Comité de Ética de la Investigación Clínica de Aragón, PI23-437).
2.2. Participants
Patients with a clinical suspicion of CTS were consecutively referred to the Neurophysiology Service at Hospital Clínico Universitario Lozano Blesa (Zaragoza, Spain) for specialized evaluation. These individuals, previously assessed in primary care or specialty consultations, were considered for inclusion based on their referral for suspected CTS. Eligible participants were adults aged 18 years or older undergoing NCS for suspected CTS. Exclusion criteria included the inability to cooperate with ultrasound or neurophysiological testing, as well as comorbidities that could interfere with accurate diagnostic assessment.
A total of 50 participants were included, contributing 94 wrists to the study dataset. The sample comprised 13 men (25 wrists) and 37 women (69 wrists), with a mean age of 55.9±13.8 years (range: 20–90). All participants were right-hand dominant. Based on combined clinical and neurophysiological criteria [
15], 30 wrists (31.9%) were classified as normal, 34 (36.2%) as mild CTS, 13 (13.8%) as moderate CTS, and 17 (18.1%) as severe CTS. Demographic and clinical characteristics are shown in
Table 1.
2.3. Ultrasound Acquisition
All ultrasound examinations were performed by an examiner with more than fifteen years of experience in musculoskeletal imaging, who was blinded to the clinical and neurophysiological findings. Scans were acquired using a VScan Air ultrasound system (General Electric, Chicago, IL, USA) with standardized software settings and fixed parameters for depth, gain, and focal zones to ensure consistency across patients. Ultrasound acquisition and electrodiagnostic testing were conducted during the same clinical visit, with nerve conduction studies performed immediately after completion of the ultrasound examination, minimizing potential temporal variability in median nerve status.
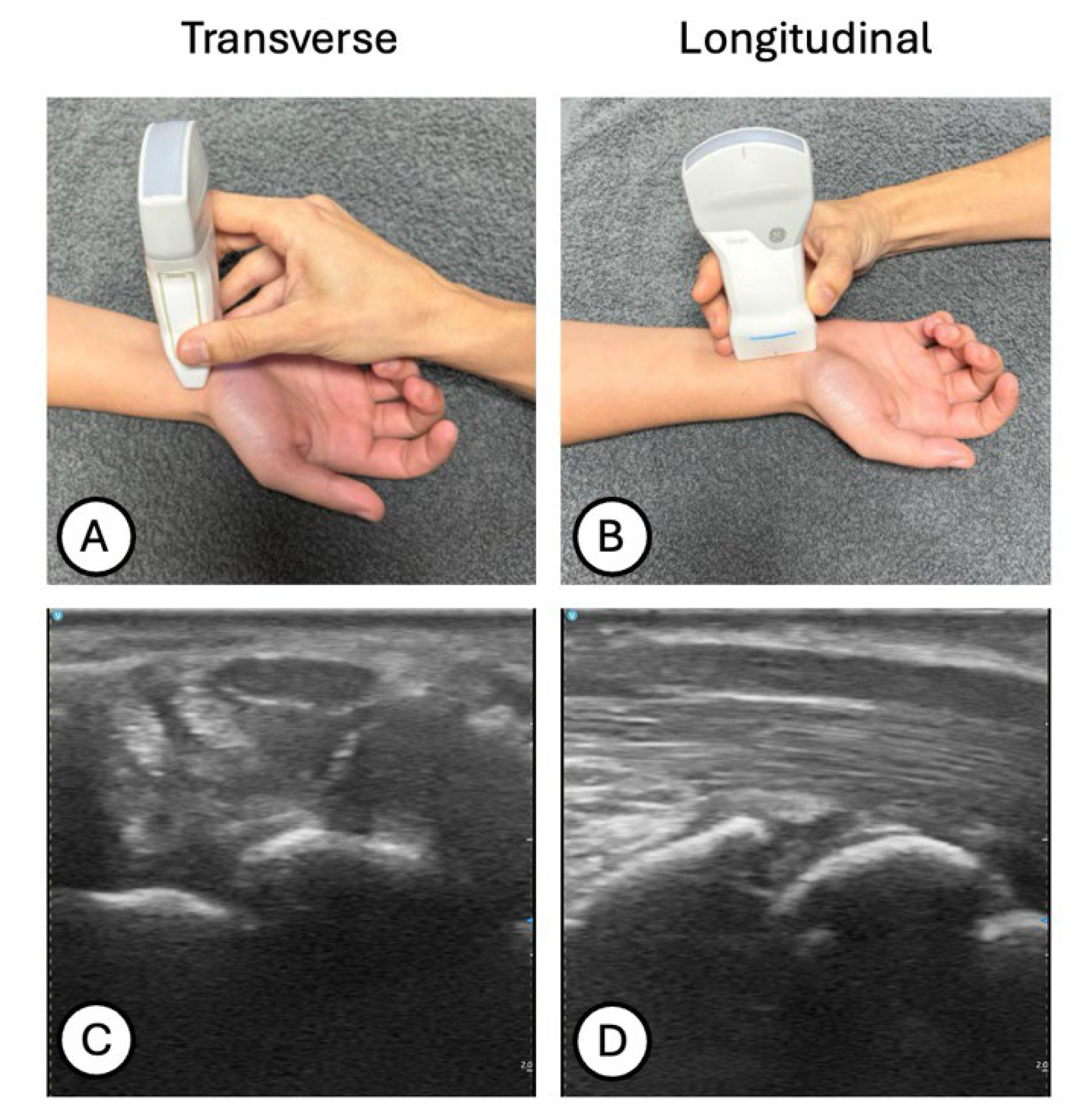
Participants were seated with the forearm relaxed in supination and the wrist maintained in a neutral position to avoid artificial deformation of the median nerve. The transducer was placed perpendicular to the skin surface to minimize anisotropy, applying only its own weight without additional compression. For each wrist, transverse scans were obtained at the proximal carpal tunnel, specifically at the level where the lunate bone appeared most superficial (
Figure 1A,C). Subsequently, longitudinal scans were acquired along the course of the median nerve at the proximal carpal tunnel, with the transducer aligned parallel to the nerve fibers to visualize its fibrillar architecture (
Figure 1B,D).
2.4. Ground Truth Definition
All diagnostic labels used for training and validating the machine-learning model were derived exclusively from standardized electrodiagnostic testing. A blinded neurophysiologist performed NCS on all participants and classified each wrist into four CTS severity categories according to the internal diagnostic criteria of the Neurophysiology Service at Hospital Clínico Universitario Lozano Blesa, which are based on the principles of the modified Canterbury scale [
15]. This scale integrates distal motor latency, sensory nerve conduction velocity, and sensory amplitude to establish the degree of median nerve impairment. Wrists were classified as normal when all electrodiagnostic parameters fell within normal limits; as mild CTS when distal motor latency was below 4 ms and sensory conduction velocity was reduced while sensory amplitude remained preserved; as moderate CTS when distal motor latency ranged between 4 and 6.5 ms with reduced sensory conduction velocity and normal sensory amplitude; and as severe CTS when distal motor latency exceeded 6.5 ms and both sensory conduction velocity and sensory amplitude were reduced. These electrodiagnostic categories constituted the ground truth for the supervised learning framework. Each wrist was assigned a single diagnostic label based on the NCS results, and these labels were directly paired with the corresponding ultrasound images. No subjective grading of ultrasound features was incorporated into the reference standard, ensuring full independence between the predictor (ultrasound images) and the outcome variable (neurophysiological diagnosis).
2.5. Data Preprocessing
2.5.1. Frame Extraction
Ultrasound videos of the wrist were exported in anonymized high-resolution video format and processed using custom Python scripts based on the OpenCV library. Individual frames were extracted from each B-mode video at a sampling rate of five frames per second to maximize data diversity while maintaining computational efficiency. All frames were saved as high-quality .png images, preserving spatial resolution and gray-scale information. This procedure yielded a total of 11,895 ultrasound images across transverse and longitudinal views of the median nerve.
2.5.2. Annotation Workflow
Following frame extraction, all ultrasound images were anonymized and reviewed by an experienced musculoskeletal sonographer to ensure that each frame provided a clear and clinically interpretable depiction of the median nerve at the level of the carpal tunnel. Frames with motion artefacts, suboptimal probe alignment, insufficient visualization of the nerve, or structures outside the diagnostic region were excluded to maintain dataset consistency.
For every retained image, the corresponding electrodiagnostic classification (normal, mild, moderate, or severe CTS) was assigned as the reference label. These diagnostic labels, derived from nerve conduction studies, were linked to each image through structured metadata files, enabling a precise pairing between ultrasound frames and their ground truth categories. This curated annotation workflow ensured that the supervised learning model was trained exclusively on valid images accompanied by reliable electrodiagnostic outcomes.
2.5.3. Dataset Composition
After quality filtering and expert verification, the final diagnostic dataset comprised 2,518 ultrasound images: 1,360 transverse and 1,158 longitudinal views of the median nerve. All images were converted to grayscale, resized to 256 × 256 pixels, expanded to a single-channel tensor format, and normalized to the [0, 1] intensity range. Each image was then paired with the corresponding electrodiagnostic label (normal, mild, moderate, or severe CTS) derived from nerve conduction studies, yielding a curated database of ultrasound–neurophysiology pairs. This dataset was randomly divided into training (75%) and validation (25%) subsets, maintaining the proportional representation of CTS severity grades and image orientations in both splits.
2.6. Machine Learning Framework
A deep-learning classification framework was developed to automatically determine CTS severity from B-mode ultrasound images. The objective of the system was to approximate the ground-truth electrodiagnostic diagnosis (normal, mild, moderate, or severe CTS) by learning discriminative patterns in the morphology and echotexture of the median nerve. To maximize diagnostic specificity, the model was trained using cropped images centred on the median nerve, obtained through a segmentation-assisted preprocessing pipeline. No manual annotations or handcrafted measurements were required, enabling a fully automated and operator-independent diagnostic workflow.
2.7. Model Architecture
Two parallel convolutional neural networks were developed, one for transverse (transverse model) and one for longitudinal (longitudinal model) ultrasound images. Both networks shared the same general design principles (compact architectures, limited depth, and high dropout regularization) reflecting the reduced spatial complexity and homogeneous appearance of the cropped nerve-focused images.
2.7.1. Transverse Model Architecture
The transverse model began with a sequence of convolutional layers (16–64 filters, 3×3 kernel, ReLU activation, he_normal initialization), each followed by batch normalization and max-pooling, progressively encoding spatial relationships within the nerve cross-section. The feature extractor was followed by a dense classification block comprising layers of 256, 72, and 16 neurons, each regularized with dropout (0.6) to control overfitting. The output layer consisted of four neurons with softmax activation, corresponding to the four CTS diagnostic categories.
2.7.2. Longitudinal Model Architecture
The longitudinal model employed a lighter feature extractor due to the lower structural complexity of sagittal nerve representations. The network consisted of convolutional blocks with 16 and 32 filters (3×3 kernel, ReLU activation) followed by max-pooling. The dense block included a 256-neuron layer with dropout (0.6), directly connected to a four-neuron softmax output layer. This streamlined architecture reduced the number of trainable parameters and improved generalization on the longitudinal dataset.
2.8. Training Configuration
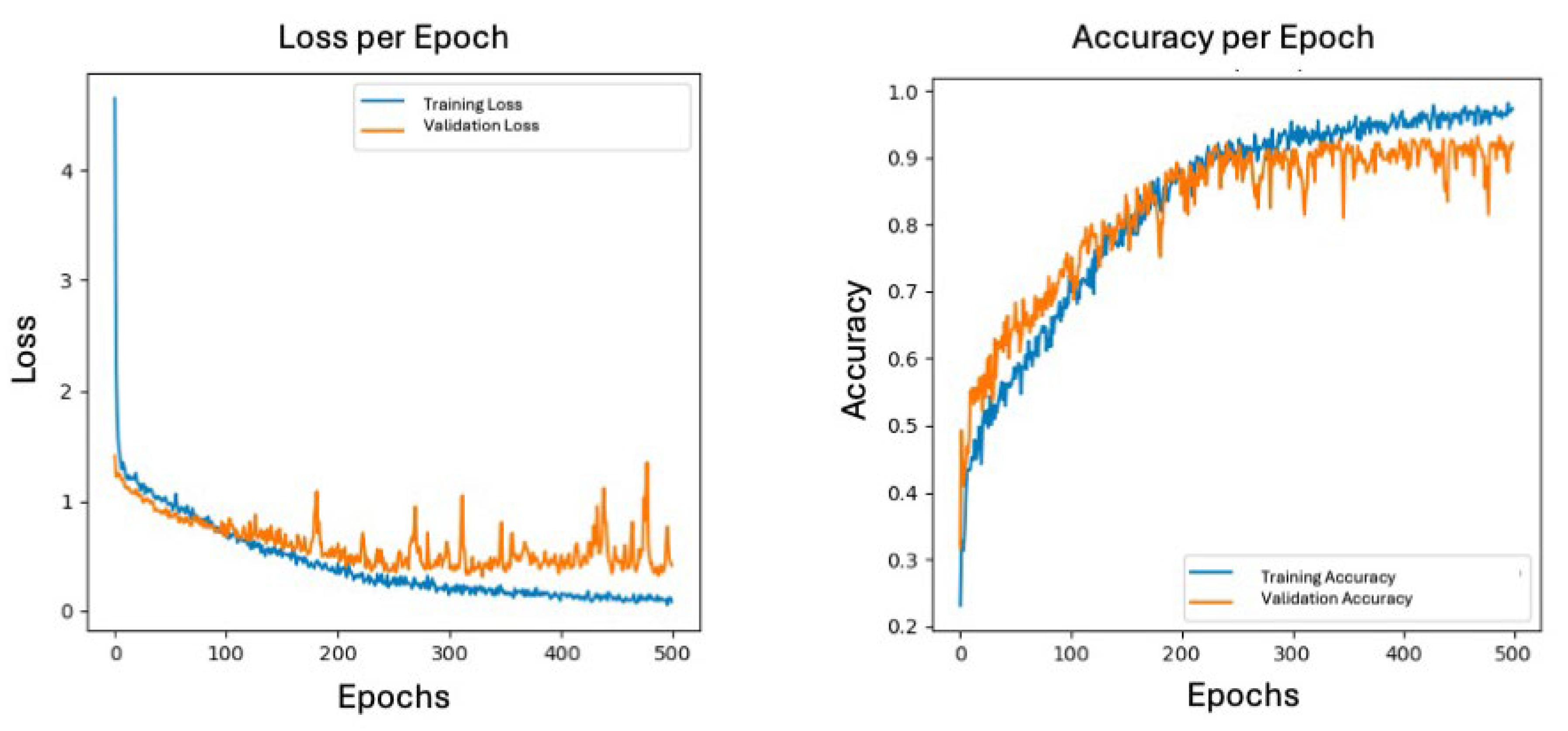
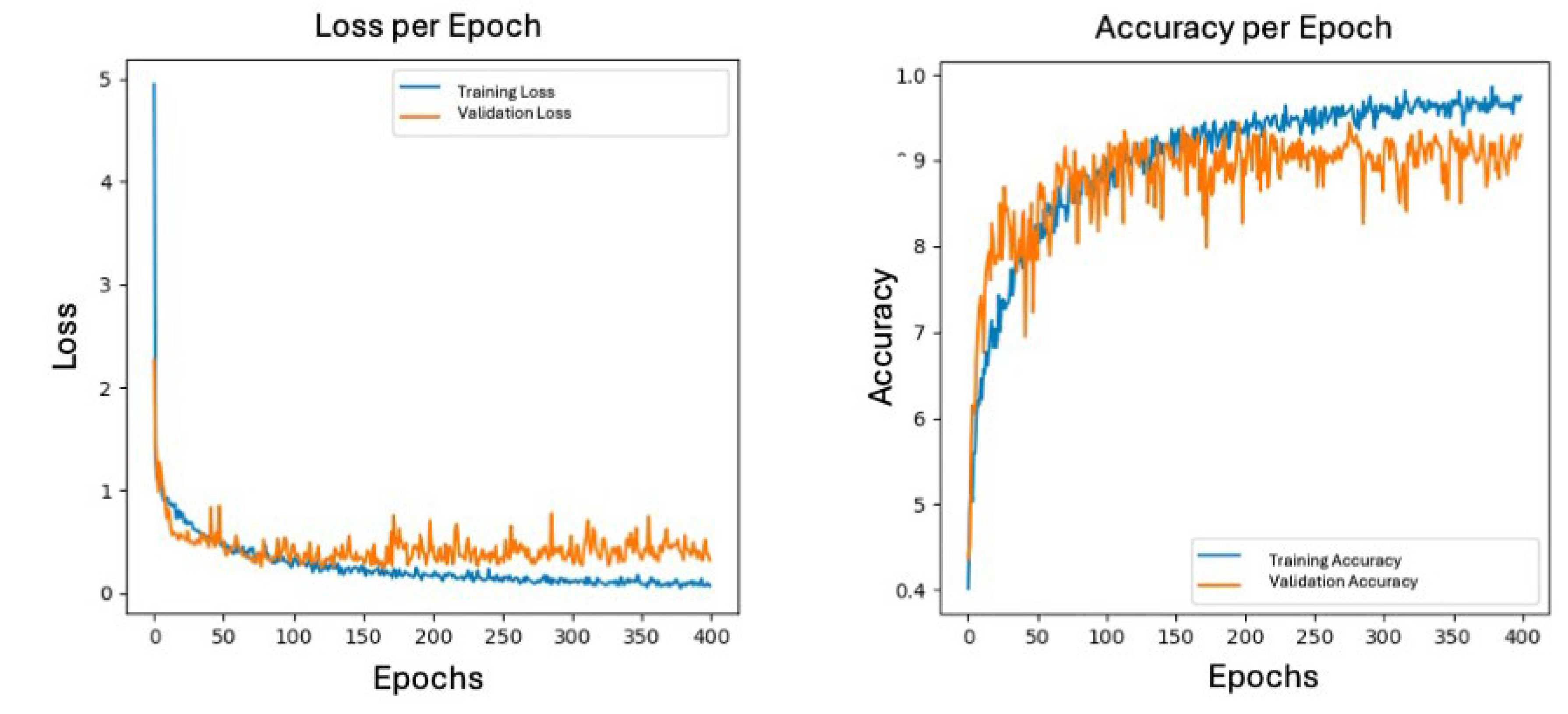
All cropped nerve images were normalized to the [0–1] intensity range and used as input for both diagnostic networks. The dataset was split into 75% for training and 25% for validation, ensuring balanced representation of all four CTS severity categories across both orientations. To improve generalization while preserving anatomical fidelity, a conservative augmentation protocol was applied, consisting of small rotations and horizontal flips only. Model training was performed using the Adam optimizer and categorical cross-entropy loss. Batch sizes were adjusted to the characteristics of each dataset (30 for the transverse model and 20 for the longitudinal model). The networks were trained for 500 epochs (transverse) and 400 epochs (longitudinal), with dropout serving as the primary regularization mechanism to mitigate overfitting. All experiments were implemented in TensorFlow/Keras and executed on GPU-accelerated environments. This training configuration yielded efficient and discriminative classifiers capable of capturing subtle morphological and textural changes associated with progressive CTS severity.
2.9. Evaluation Metrics
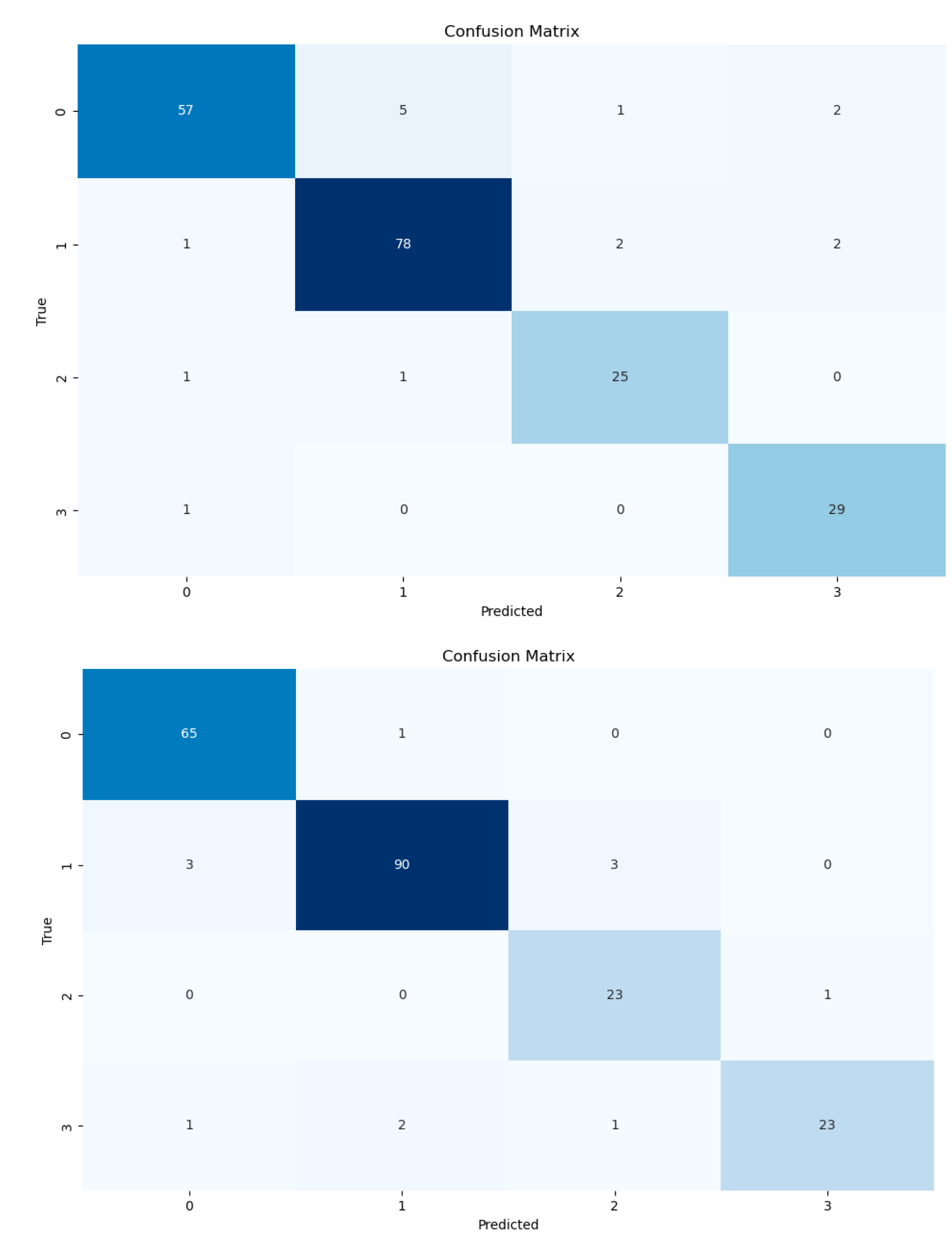
Model performance was assessed using standard metrics for multiclass classification, including overall accuracy, categorical cross-entropy loss, and class-wise precision. Accuracy quantified the proportion of correctly classified images across the four diagnostic categories (normal, mild, moderate, and severe CTS), while the loss function reflected the discrepancy between predicted class probabilities and the ground-truth electrodiagnostic labels. Confusion matrices were generated for each network using the validation dataset to characterize class-specific behavior by detailing true positives, false positives, and false negatives. This analysis enabled identification of classes prone to misclassification, particularly among adjacent severity levels. Softmax outputs were treated as probability distributions across the four classes, with the highest-probability label assigned as the model prediction. Accuracy and loss curves across epochs were visually inspected to evaluate convergence stability and detect early signs of overfitting.
2.10. Statistical Analysis
All statistical procedures were performed in Python (version 3.9) using TensorFlow/Keras and standard scientific computing libraries. Model performance was summarized descriptively using accuracy, categorical cross-entropy loss, and class-wise precision values obtained from the validation dataset. Confusion matrices were computed to quantify class-specific prediction behavior and to evaluate the distribution of errors across CTS severity categories. Training and validation curves were visually reviewed to confirm stable learning dynamics, appropriate convergence, and absence of harmful overfitting. Given the methodological nature of the study, focused on model development rather than group comparison, no inferential statistical testing was conducted. Results were interpreted in terms of classification consistency, error patterns, and the ability of each model to discriminate among clinically relevant severity levels of CTS.
4. Discussion
This study provides new evidence supporting the application of deep learning to the automated severity assessment of carpal tunnel syndrome using B-mode ultrasound imaging. The proposed multiclass convolutional neural networks, developed separately for transverse and longitudinal views, demonstrated strong classification performance, achieving overall accuracies of 0.92 and 0.94, respectively. These results highlight the capacity of modern neural architectures to detect and interpret subtle variations in median nerve morphology, echotexture, and cross-sectional configuration, hallmarks of CTS severity, within the inherent variability of musculoskeletal ultrasound.
Artificial intelligence is increasingly used across orthopedics, neurology, and musculoskeletal medicine, especially for diagnostic tasks involving medical imaging [
16]. Prior literature has largely focused on applications such as fracture detection, implant identification, and anatomical segmentation, where machine learning approaches have shown considerable promise in improving diagnostic accuracy, efficiency, and workflow reproducibility across a wide range of medical imaging modalities, including Magnetic Resonance Imaging Computed Tomography, radiographs, and ultrasound [
17,
18]. However, within the context of peripheral nerve disorders, artificial intelligence models remain scarce, particularly those applied to ultrasound imaging, where operator dependency and subtle morphological variability pose additional challenges. Early computer-aided approaches for nerve identification relied on multi-stage pipelines that required extensive preprocessing, handcrafted feature extraction, and manual feature selection, limiting their practicality as end-to-end diagnostic systems [
19]. In contrast, modern deep learning techniques, particularly CNNs, which automatically learn hierarchical image features, have transformed biomedical image analysis and become foundational tools for radiological interpretation [
20,
21]. CNN-based models have already demonstrated strong performance in identifying neural and vascular structures across several anatomical regions [
22,
23].
Within carpal tunnel syndrome specifically, a recent systematic review and meta-analysis summarized the performance of deep learning algorithms for automatic localization and segmentation of the median nerve at the carpal tunnel level, reporting acceptable precision, recall, and Dice coefficients for nerve identification on ultrasound [
14]. However, the scope of these studies has largely been restricted to nerve-centric tasks, such as localization, segmentation, or the extraction of geometric descriptors, and has not addressed the clinically critical challenge of CTS severity grading aligned with electrodiagnostic standards. Previous deep learning approaches have primarily focused on delineating the nerve and quantifying features such as cross-sectional area, circularity, or motion-related displacement, without directly translating these representations into clinically interpretable severity categories [
24,
25].
In this context, the present work expands the existing literature by moving beyond localization and segmentation to evaluate whether convolutional neural networks can assign CTS severity grades (normal, mild, moderate, severe) directly from raw B-mode ultrasound images, without manual feature engineering. Across the included studies in prior literature, sample sizes ranged from 6 to 103 participants and image acquisition relied predominantly on high-end ultrasound systems (e.g., ACUSON S2000, MyLab Class C, Philips iE33, Aplio 500), introducing substantial technical heterogeneity and limiting insight into real-world applicability. By contrast, the present study contributes a cohort of 50 participants and 94 wrists, positioning it among the larger datasets used to date for deep-learning research in CTS ultrasound. Although the distribution of electrodiagnostic categories was not fully balanced, the dataset captures the full clinical spectrum of CTS severity, which is essential for developing and evaluating a multiclass classification framework. Notably, the overall accuracies achieved by our transverse and longitudinal models (0.92 and 0.94, respectively) are comparable in magnitude to the pooled accuracy reported for localization and segmentation tasks in previous work [
14], despite the greater complexity inherent to multiclass severity grading relative to purely geometric or binary endpoints. Furthermore, all ultrasound examinations were acquired using a handheld, wireless ultrasound device, rather than laboratory-grade systems. This design choice enhances the translational relevance of the proposed approach, supporting the feasibility of automated CTS severity assessment in portable, scalable, and resource-constrained clinical settings such as primary care, occupational health, and outpatient orthopaedic practice. To our knowledge, this is the first study to validate deep-learning models for CTS severity assessment using images exclusively obtained from a handheld ultrasound system.
From a technical perspective, prior deep-learning research in CTS ultrasound has focused almost entirely on nerve-centric tasks rather than diagnostic classification. The reviewed models encompassed U-Net, DeepNerve, Mask R-CNN, DeepSL, ResNet-based architectures, and feature pyramid networks, among others, and were primarily designed for tasks of localization, segmentation, or tracking of the median nerve during motion [
24,
26,
27,
28,
29]. Pooled estimates from this meta-analysis showed high performance for nerve-centric tasks, with precision and recall around 0.92–0.94, accuracy near 0.92, and a summarized Dice coefficient of approximately 0.90 [
14]. These results underscore the technical maturity of segmentation models for identifying the nerve on ultrasound images. Our study aligns with this line of work in that it also relies on nerve-centered inputs (obtained through a segmentation-assisted preprocessing pipeline) yet it diverges from the prevailing literature in the specific task being addressed. Whereas most published models focus on structural delineation or tracking, the present work evaluates whether CNNs can leverage those nerve-focused representations to perform multiclass diagnostic classification aligned with electrodiagnostic severity grades.
From a clinical perspective, the automated assessment of CTS severity presented in this study may offer relevant advantages in real-world decision-making. In daily orthopedic and hand surgery practice, severity grading is frequently used to support treatment selection and to contextualize clinical findings, particularly when determining the appropriateness and timing of surgical decompression. Surveys among hand surgeons have demonstrated substantial variability in both the use of non-surgical treatments and the reliance on diagnostic testing, with severity grading commonly cited as a key justification for ordering electrodiagnostic studies prior to or during surgical decision-making [
30]. In this context, an automated, operator-independent portable ultrasound- based severity classifier may help reduce inter-observer variability, support standardized patient stratification, and provide rapid, non-invasive morphological assessment aligned with electrodiagnostic severity grades, thereby complementing clinical evaluation and electrophysiology, facilitating baseline documentation and longitudinal monitoring, and helping optimize treatment selection while avoiding unnecessary delays or prolonged ineffective conservative care.
Despite these encouraging findings, several limitations should be acknowledged. The dataset was derived from a single center, using a single handheld ultrasound system and a single highly experienced examiner, which limits the ability to assess model robustness under multi-device or multi-operator conditions. Similar constraints have been noted in previous deep learning studies on median nerve ultrasound, where most models were trained and tested on images acquired from a single machine [
14]. In addition, many prior frameworks relied on manually defined regions of interest and were not fully end-to-end, increasing dependence on expert labelling and potentially limiting scalability [
29,
31]. Although the present approach employs a segmentation-assisted preprocessing pipeline to focus on the nerve, external validation on independent cohorts (including multi-center, multi-device, and multi-operator datasets) will be essential to confirm generalizability. Furthermore, a notable finding of this study was the differential performance across severity categories: while normal, mild, and moderate cases were classified with high precision, performance for severe CTS was lower in the longitudinal model. This behavior is consistent with the relative scarcity of high-severity cases, which constrained the model’s exposure to extreme pathology and limited its ability to fully capture the most pronounced sonographic abnormalities. Similar data-imbalance issues have been widely reported in deep learning applications to nerve ultrasound [
14] and should be interpreted as a dataset-related limitation rather than a model-specific shortcoming. Increasing the representation of clinically confirmed severe CTS would likely improve calibration across the full pathological spectrum. Finally, this study focused exclusively on B-mode imaging; future work integrating complementary sonographic parameters, such as cross-sectional area measurements, elastography, or radiomic descriptors, may further enhance diagnostic depth and strengthen the clinical relevance of automated severity assessment frameworks.
Author Contributions
For research articles with several authors, a short paragraph specifying their individual contributions must be provided. The following statements should be used “Conceptualization, M.M.H., M.M.U. and E.B.G.; methodology, M.M.U., A.A.C., E.G.G. and M.M.U.; software, A.A.C., M.G.L. and M.G.D.; validation, A.A.C., M.G.L. and M.G.D.; formal analysis, A.A.C., M.G.D. and M.M.H.; investigation, I.R.A., G.C.; data curation, I.R.A, G.C. and E.B.G.; writing—original draft preparation, M.M.U. and M.M.H.; writing—review and editing, E.B.G.; supervision, M.M.U. and M.M.H; project administration, M.M.U. All authors have read and agreed to the published version of the manuscript.