1. Introduction
Human papillomavirus (HPV) is a primary etiological agent for cervical cancer and a significant contributor to several head and neck cancers [
1]. The oncogenic potential of HPV is largely driven by its early proteins, E6 and E7, which aberrantly interact with host cellular machinery. Specifically, E6 targets and degrades the tumor suppressor protein p53, while E7 inactivates the retinoblastoma protein (Rb), collectively leading to cell cycle dysregulation, evasion of apoptosis, and genomic instability, thereby promoting oncogenesis [
2]. A comprehensive understanding of how E6/E7 precisely reshapes the host cell’s gene regulatory networks is paramount for elucidating HPV’s carcinogenic mechanisms and developing effective therapeutic strategies [
3]. Studies involving expression profiling of mRNA have been instrumental in dissecting the complex functional networks of genes regulated by HPV E6 and E7 proteins in host cells, providing crucial insights into their oncogenic roles [
4].
Despite significant advancements in high-throughput sequencing technologies, which can comprehensively capture E6/E7-induced transcriptional perturbations, existing computational methods still face considerable challenges in deciphering complex gene-gene regulatory relationships and inferring deep biological mechanisms. Traditional approaches, such as differential expression analysis [
5], gene enrichment analysis [
6], or even graph neural network-based regulatory network predictions [
7], often provide lists of statistically significant genes or pathways. However, they typically fall short in offering a chain-of-thought, interpretable biological reasoning that explains why certain genes or pathways are affected and their specific contribution to cancer progression. This lack of explicit, mechanistic interpretability hinders a holistic understanding of the intricate biological processes at play.
Figure 1.
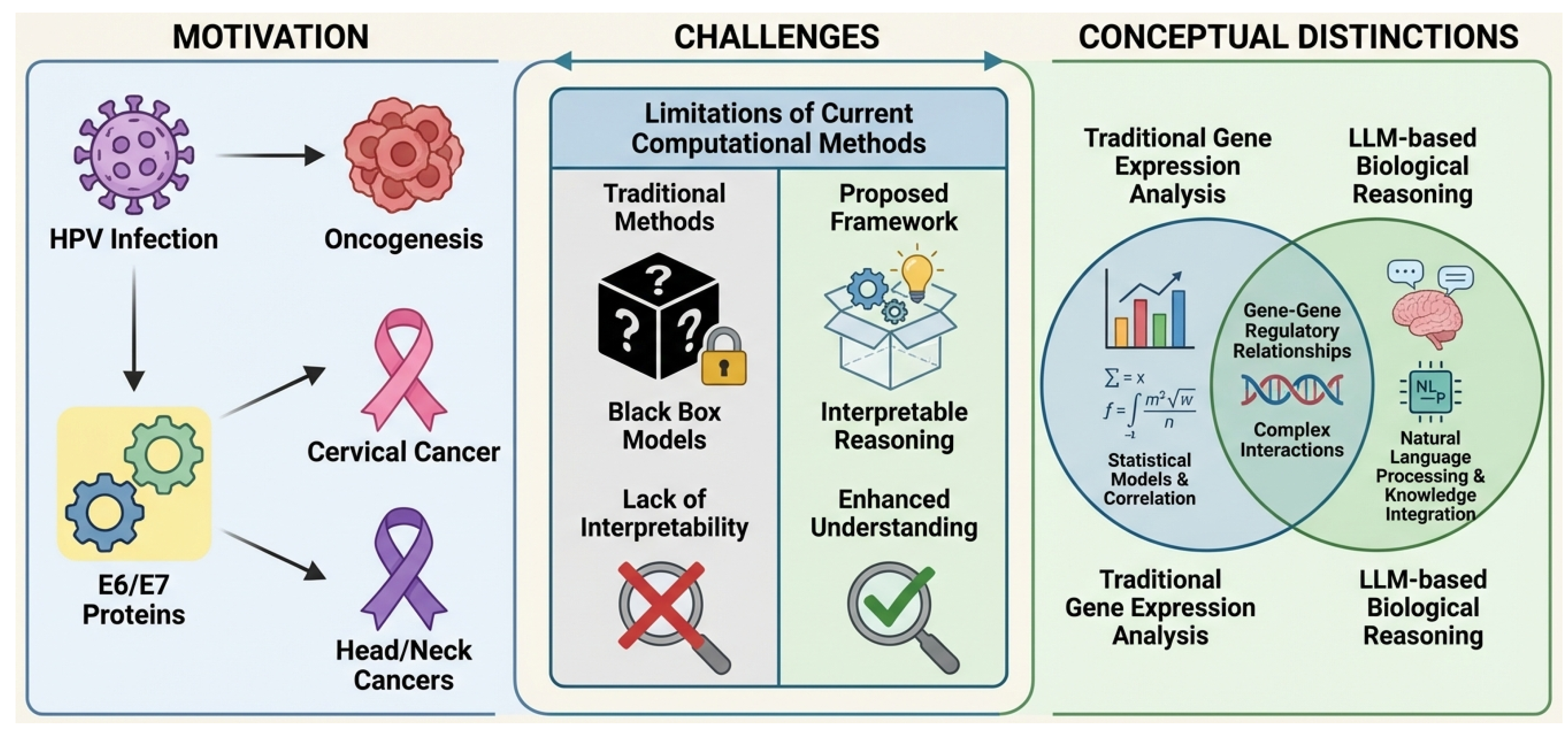
Illustrative Framework: Motivation, Challenges, and Conceptual Distinctions of OncoReasoner. The figure highlights the motivation stemming from HPV E6/E7 oncogenesis leading to cervical and head/neck cancers. It then details the challenges of current computational methods, which suffer from a lack of interpretability, contrasting them with the interpretable reasoning offered by our proposed framework. Finally, it illustrates the conceptual distinction of OncoReasoner by integrating traditional gene expression analysis with LLM-based biological reasoning to comprehensively understand gene-gene regulatory relationships and complex interactions.
Figure 1.
Illustrative Framework: Motivation, Challenges, and Conceptual Distinctions of OncoReasoner. The figure highlights the motivation stemming from HPV E6/E7 oncogenesis leading to cervical and head/neck cancers. It then details the challenges of current computational methods, which suffer from a lack of interpretability, contrasting them with the interpretable reasoning offered by our proposed framework. Finally, it illustrates the conceptual distinction of OncoReasoner by integrating traditional gene expression analysis with LLM-based biological reasoning to comprehensively understand gene-gene regulatory relationships and complex interactions.
To bridge this critical gap, we propose a novel framework, OncoReasoner, designed for interpretable regulatory network inference driven by viral oncogenic proteins. For the first time, OncoReasoner integrates advanced biological expression analysis with the powerful reasoning capabilities of large language models (LLMs) to construct HPV E6/E7-driven regulatory networks and provide highly interpretable biological explanations of underlying oncogenic mechanisms. Our research extends beyond merely predicting regulatory relationships; it leverages the semantic understanding and knowledge integration capabilities of LLMs to deeply interpret the biological significance behind E6/E7-induced transcriptomic perturbations.
The proposed OncoReasoner framework comprises three core modules: an Expression Encoder, a Bio-LLM Reasoning Module, and a Graph Refinement Module. The Expression Encoder transforms raw gene expression profiles into high-dimensional semantic embeddings, providing a quantitative biological signal. The Bio-LLM Reasoning Module, fine-tuned on biomedical data, then takes these perturbation signals and, using its vast internal knowledge base, generates chain-of-thought biological explanations of likely oncogenic mechanisms. Finally, the Graph Refinement Module, utilizing graph neural networks and biological prior knowledge, refines and ensures the biological consistency and accuracy of the LLM-inferred regulatory relationships.
For experimental validation, we utilize gene expression data predominantly from the Gene Expression Omnibus (GEO) database, including specific datasets focusing on HPV E6/E7 functional studies (e.g., GSE6791), and RNA-seq data from cervical cancer patients within The Cancer Genome Atlas (TCGA) [
5]. Ground truth regulatory networks are compiled from established biological databases such as KEGG HPV carcinogenesis pathways [
6], Reactome [
8], and STRING [
5]. Furthermore, a rich corpus of biomedical literature from PubMed and protein interaction evidence from BioGRID [
6] is used to instruct-tune the LLM component.
Our evaluation encompasses three distinct tasks: differential gene expression classification, regulatory network edge prediction, and crucially, functional pathway reasoning and mechanistic interpretability. Through comprehensive comparisons with various baseline methods, including traditional statistical approaches (e.g., DESeq2 + KEGG), gene set enrichment analysis (GSEA), standalone graph neural networks, and biomedical BERT classifiers, OncoReasoner consistently demonstrates superior performance. Notably, in the core task of functional pathway reasoning, OncoReasoner achieves significantly higher accuracy in identifying correct pathways and provides qualitatively superior, expert-rated mechanistic explanations (e.g., an average explanation score of 4.5 out of 5), validating its unique advantage in combining biological data with LLM-driven mechanistic interpretation.
Our main contributions are summarized as follows:
We introduce OncoReasoner, a novel and comprehensive framework that uniquely integrates biological expression analysis, large language models, and graph neural networks to infer interpretable gene regulatory networks driven by viral oncogenic proteins.
We develop a sophisticated three-stage training strategy designed to maximize the synergistic potential of each module, ensuring effective gene expression encoding, precise LLM instruction tuning for biological reasoning, and robust graph consistency training.
We significantly advance the state-of-the-art in mechanistic interpretability for cancer biology, providing chain-of-thought explanations for oncogenic perturbations that surpass existing methods in accuracy and biological depth.
2. Related Work
2.1. Large Language Models in Biomedical Research and Explainable AI
Large Language Models (LLMs) show transformative potential in biomedical research, with surveys highlighting their broad applicability and future directions [
9]. While LLMs promise to accelerate knowledge extraction, scientific discovery, and clinical decision support, their deployment in critical environments demands reliability, safety, and explainability. This subsection reviews LLM applications in biomedical research and the evolving Explainable AI (XAI) landscape. Foundation Models impact information access in scientific and biomedical applications; [
10] highlights generative AI’s role in information synthesis while emphasizing concerns like hallucination for reliable biomedical outputs.
To integrate domain-specific knowledge, research improves LLM handling of structured data. For Biomedical Knowledge Extraction, [
11] introduces SKILL, infusing structured knowledge from KGs into LLMs via training on factual triples, improving question-answering. [
12] further supports structured knowledge construction with a Knowledge Graph Completion method using a linear temporal regularizer and multi-vector embeddings for temporal KGs. Despite their promise, LLM application in healthcare requires significant safety and trustworthiness. [
13] evaluates ChatGPT toxicity, showing persona assignments amplify harmful outputs and biases, underscoring the need for robust AI safety guardrails, especially for patient interaction.
Robust Explainable AI (XAI) methods are indispensable for responsibly integrating LLMs into biomedical research and clinical practice, providing transparency and understanding of model decisions. For XAI in biomedical applications, precise representations are crucial. [
14] proposes SapBERT, a self-alignment pretraining scheme leveraging biomedical ontologies like UMLS to generate fine-grained entity representations, enhancing medical entity linking and improving AI reliability and interpretability. Understanding LLM internal reasoning is vital for explainability. [
15] investigates Chain-of-Thought Reasoning, finding that for effective CoT, the
relevance and
correct ordering of reasoning steps are more critical than factual validity, offering insights into LLM thought processes. In summary, LLM integration into biomedical research balances advanced capabilities with critical needs for safety, trustworthiness, and explainability. Current efforts focus on enhancing LLMs with structured knowledge, robust representations, and investigating reasoning mechanisms for transparent systems. Our work aims to contribute to this evolving landscape by...
2.2. Computational Approaches for Gene Regulatory Network Inference and Pathway Analysis
Computational approaches are indispensable for unraveling gene regulatory networks (GRNs) and their functional implications via pathway analysis, providing critical tools for interpreting high-throughput biological data and generating hypotheses. A foundational aspect involves reconstructing regulatory relationships, with advances in computational methods for
Gene Regulatory Network (GRN) inference being crucial for understanding cellular processes, aiming for scalable, accurate reconstruction from high-dimensional data [
16]. Inference relies on robust data processing, making effective
Transcriptomic Data Analysis fundamental for revealing gene expression patterns. Methods focus on extracting insights from RNA-seq data to identify differentially expressed genes or co-expression modules [
17].
These computational approaches are widely applied across biomedical domains, from analyzing gene regulation in comorbid pain disorders [
18] to bioinformatics analysis of temporomandibular joint stem cell responses [
19]. Identifying significant changes in gene activity,
Differential Gene Expression analysis is a cornerstone of pathway analysis, highlighting genes with altered activity across conditions. Computational tools are essential for robust detection and biological interpretation [
20]. Beyond individual gene analysis, interpreting gene expression changes in a broader biological context is paramount. Comprehensive
Biological Pathway Analysis integrates computational approaches to understand functional implications of molecular changes, interpreting omics data within known pathways to identify perturbed processes [
21]. Complementing this,
Gene Set Enrichment Analysis (GSEA) identifies pathways or functions collectively enriched in differentially expressed genes, interpreting the systemic biological impact of expression changes by considering gene groups [
22].
Modern computational biology increasingly leverages machine learning. GRN’s complex topological structure lends itself to graph representation, making
Graph Neural Networks (GNNs) powerful for analyzing their properties and dynamics; GNNs model intricate gene relationships and infer interactions by learning from network structures [
23]. Specifically,
Graph Attention Networks (GATs), a specialized GNN form, are useful for GRN inference and analysis, allowing differential weighting of neighboring nodes to prioritize influential regulatory interactions based on learned importance [
24]. Finally, incorporating
Prior Biological Knowledge is critical for improving GRN inference and pathway analysis accuracy and interpretability. This involves leveraging databases of known interactions, annotations, or curated pathways to guide/validate predictions, grounding data-driven insights in biological context [
25].
3. Method
We propose OncoReasoner, a novel and comprehensive framework designed for interpretable regulatory network inference driven by viral oncogenic proteins. At its core, OncoReasoner integrates advanced biological expression analysis with the robust reasoning capabilities of large language models (LLMs) and graph neural networks (GNNs) to dissect Human Papillomavirus (HPV) E6/E7-induced transcriptomic perturbations. The framework is structured into three main modules: an Expression Encoder, a Bio-LLM Reasoning Module, and a Graph Refinement Module, which are synergistically trained through a multi-stage process to ensure both predictive accuracy and biological interpretability.
3.1. Model Architecture
The OncoReasoner framework operates as a multi-modal deep learning system, meticulously designed to achieve viral oncogenic protein-driven regulatory network inference and functional pathway prediction. Each module contributes a distinct analytical capability, working in concert to process diverse biological data types and generate mechanistic insights.
Figure 2.
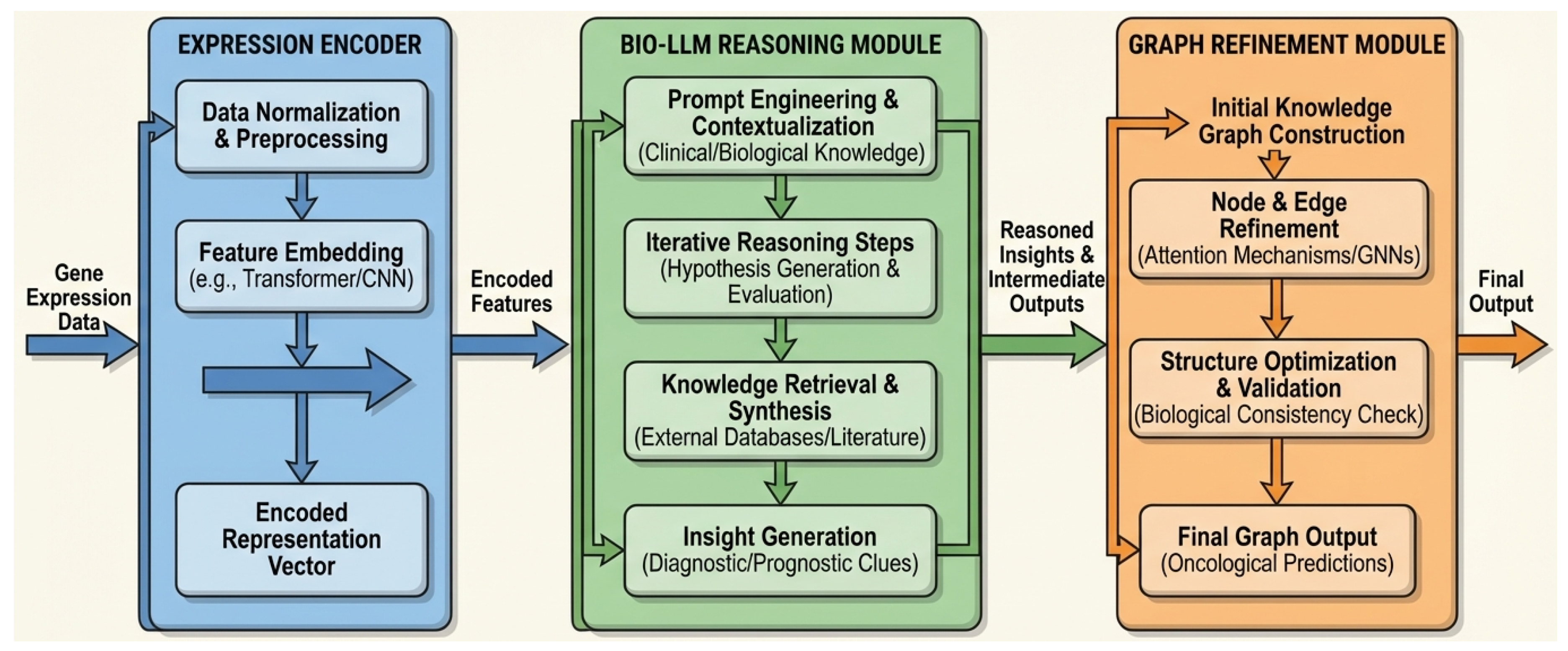
Overall architecture of the OncoReasoner framework. Gene expression data is initially processed by the Expression Encoder to generate embedded representations. These encoded features are then fed into the Bio-LLM Reasoning Module, which performs context-aware biological interpretation and generates reasoned insights. Finally, the Graph Refinement Module leverages these insights and graph neural networks (GNNs) to construct and refine a biologically consistent regulatory network, yielding oncological predictions.
Figure 2.
Overall architecture of the OncoReasoner framework. Gene expression data is initially processed by the Expression Encoder to generate embedded representations. These encoded features are then fed into the Bio-LLM Reasoning Module, which performs context-aware biological interpretation and generates reasoned insights. Finally, the Graph Refinement Module leverages these insights and graph neural networks (GNNs) to construct and refine a biologically consistent regulatory network, yielding oncological predictions.
3.1.1. Expression Encoder
The Expression Encoder module is responsible for transforming raw gene expression profiles, such as those derived from RNA-seq or microarray data, into high-dimensional semantic embeddings. This conversion is critical for capturing the intrinsic features of gene sequences and their quantitative expression patterns, providing a rich biological signal for subsequent mechanistic reasoning.
For a given gene , we consider its sequence information (e.g., promoter region, coding sequence) and its measured expression level . We leverage powerful pre-trained biological sequence models, such as Nucleotide Transformer v2 (NT-v2) or DNABERT-2, which are adept at encoding genomic and transcriptomic information into a continuous vector space. The encoder E processes the input gene sequence and expression data to generate a compact, meaningful representation. Each gene’s expression state and sequence context are thus represented as an embedding vector, , where D is the embedding dimension.
The encoding function for a gene
can be formally expressed as:
where
denotes the sequence information of gene
, and
represents its scalar expression value. The output embeddings
capture complex relationships and latent biological properties that are crucial for downstream analysis by the LLM.
3.1.2. Bio-LLM Reasoning Module
This module constitutes the core innovation of OncoReasoner, providing a powerful engine for context-aware biological interpretation. We employ large language models that have been specifically fine-tuned on extensive biomedical domain data, such as LLaMA3-8B or BioMedLM.
The Bio-LLM Reasoning Module receives processed gene expression perturbation information from the Expression Encoder. This typically includes lists of significantly up-regulated or down-regulated genes under the influence of viral oncogenes like E6/E7, often presented alongside their embedding vectors and associated metadata.
Given a structured query comprising a list of differentially expressed genes (DEGs) and a specific biological prompt (e.g., "Explain the likely oncogenic mechanisms induced by HPV E6/E7 affecting these genes"), the LLM accesses its vast internal biomedical knowledge base. It then performs sophisticated reasoning to generate detailed, chain-of-thought biological explanations. This process involves:
- 1.
Gene-Pathway Association: Identifying known pathways associated with the input DEGs.
- 2.
Interaction Inference: Deducing potential molecular interactions, including protein-protein interactions, gene regulations, and their consequences.
- 3.
Oncogenic Mechanism Elucidation: Explaining how these interactions contribute to specific cancer hallmarks (e.g., p53 pathway disruption, cell cycle deregulation, apoptosis evasion).
This capability allows
OncoReasoner to provide highly interpretable and contextually rich insights into complex biological phenomena, generating textual outputs
from input DEGs
and prompt
P:
The outputs
typically include identified pathways, key molecular players, and their specific contributions to cancer progression, presented as natural language explanations.
3.1.3. Graph Refinement Module
To ensure the biological consistency and accuracy of the regulatory relationships implicitly or explicitly inferred by the Bio-LLM Reasoning Module, we integrate a Graph Refinement Module. This module employs graph neural networks (GNNs), specifically a Graph Attention Network (GAT), to process and refine a preliminary regulatory network.
The preliminary network, potentially derived from parsing the LLM’s textual output into a graph structure, or directly predicted by an LLM-guided process, provides an initial topology. The GAT then learns to represent the network structure and node features, leveraging existing biological prior knowledge networks such as STRING and KEGG. The input to the GAT consists of the gene embeddings as node features and an initial adjacency matrix representing inferred or known relationships.
For each node
i, the GAT computes new node features
by aggregating information from its neighbors
(where
is the set of neighbors of node
i) through an attention mechanism:
where
are the input features (e.g.,
),
is a learnable weight matrix,
is an activation function, and
are attention coefficients computed as:
Here,
is a learnable attention vector, and ∥ denotes concatenation. By analyzing the topological structure, learned gene embeddings, and integrating prior knowledge, the GAT identifies and corrects inconsistent or weakly supported regulatory edges. This process refines the initial network, thereby enhancing the reliability and biological plausibility of the final inferred network. The output of this module is a refined adjacency matrix
representing the inferred regulatory network, where
if a refined regulatory relationship from gene
i to gene
j exists, and 0 otherwise.
3.2. Three-Stage Training Strategy
To effectively harness the capabilities of each module and ensure their synergistic operation, OncoReasoner employs a meticulously designed three-stage training strategy. This progressive approach allows for specialized pre-training and fine-tuning, leading to a robust and interpretable framework.
3.2.1. Stage 1: Expression Pretraining
The primary objective of this stage is to train the Expression Encoder to effectively capture and represent gene expression patterns. This is achieved through a self-supervised task analogous to masked language modeling (MLM) for sequences. During training, for a given gene expression profile (where N is the total number of genes), a random portion of the gene expression values, specifically those in a masked set , are masked. The model is then tasked with predicting these masked values based on the unmasked genes where .
The encoder
E learns to generate contextualized embeddings that allow for accurate reconstruction of masked information. The training objective is to minimize the Mean Squared Error (MSE) between the predicted and ground truth expression values for the masked genes:
where
represents the expression profile with masked values,
f is a prediction head (e.g., a linear layer) operating on the encoder’s output for masked genes,
denotes the predicted expression value for the masked gene
i, and
represents its true expression value. This stage ensures that the embeddings
are rich in information regarding gene function and regulatory context.
3.2.2. Stage 2: LLM Instruction Tuning
This stage focuses on fine-tuning the Bio-LLM Reasoning Module to enable it to provide precise, accurate, and highly interpretable reasoning in response to biological queries. We construct a specialized instruction dataset comprising triplets of input gene lists, specific biological questions (e.g., related to oncogenic pathways, mechanism of action for viral proteins), and corresponding chain-of-thought explanations derived from extensive biomedical literature and expert knowledge.
For instance, an input triplet might be:
Input Gene List: {CDKN1A ↑, BAX ↓, TP53 ↓}
Question: "What oncogenic pathway is likely affected by E6/E7 and how?"
Output Explanation (Ground Truth): "The observed up-regulation of CDKN1A, coupled with down-regulation of BAX and TP53, strongly indicates disruption of the p53 tumor suppressor pathway. HPV E6 protein is known to target p53 for proteasomal degradation, leading to reduced expression and impaired transcriptional activity of p53 target genes like CDKN1A (p21) and BAX. This results in evasion of apoptosis and uncontrolled cell proliferation, characteristic of oncogenesis."
Efficient fine-tuning techniques, such as
LoRA (Low-Rank Adaptation), are employed to adapt the large language model to this biomedical reasoning task without full retraining. The objective is to minimize the negative log-likelihood of the ground truth output tokens given the input prompt and gene list:
where
T is the length of the ground truth explanation sequence,
is the
t-th token in the explanation,
are preceding tokens, and
represents the LLM’s parameters. This optimizes its ability to generate high-quality, mechanistic explanations that align with expert biological understanding.
3.2.3. Stage 3: Graph Consistency Training
The final stage aims to integrate the LLM’s reasoning outputs with the graph structure information, thereby optimizing both the accuracy of regulatory network prediction and its biological consistency. This involves training the entire framework to predict the existence and direction of regulatory relationships between gene pairs. This task can be framed as a binary classification problem for each potential edge in the network.
An additional consistency loss term is incorporated to penalize discrepancies or inconsistencies within the inferred network structure, drawing upon established biological prior knowledge networks . This loss term can encourage adherence to known network motifs, ensure smoothness of gene embeddings on the graph, or enforce agreement with high-confidence interactions from prior databases. For example, might quantify the divergence between the predicted adjacency matrix and a prior knowledge graph, or penalize violations of expected network properties.
The total loss function for this stage is a weighted sum of the Binary Cross Entropy (BCE) loss for edge prediction and a consistency loss:
The BCE loss is calculated for all possible edges
:
Here,
represents the predicted probability of a regulatory edge existing between gene
i and gene
j, and
is the ground truth label (0 or 1).
ensures that the inferred network adheres to biological constraints and topological properties, while
is a hyperparameter balancing the importance of edge prediction accuracy and network consistency. This multi-objective optimization refines the network, ensuring that the LLM’s mechanistic interpretations are grounded in a coherent and biologically plausible regulatory graph.
4. Experiments
In this section, we detail the experimental setup, including the datasets used, the evaluation tasks designed to assess our framework, the baseline models for comparison, and the metrics employed. We then present a comprehensive performance comparison of OncoReasoner against these baselines, followed by an ablation study to validate the contributions of each proposed module. Finally, we include results from a human evaluation of the interpretability of our framework’s mechanistic explanations.
4.1. Experimental Setup
4.1.1. Data Collection and Preprocessing
Our experimental validation leverages a diverse set of biological data, meticulously curated to support the multi-modal nature of OncoReasoner.
Expression Data We primarily sourced gene expression profiles from the Gene Expression Omnibus (GEO) database, including specific datasets critical for studying HPV E6/E7 function, such as GSE6791. Additionally, we utilized RNA-seq data from cervical cancer patients available through The Cancer Genome Atlas (TCGA) [
5]. These data were processed to construct comparative datasets, including HPV-positive versus HPV-negative samples, and samples with E6 or E7 overexpression compared to their respective controls, enabling us to thoroughly capture E6/E7-induced transcriptomic perturbations.
Regulatory Network Ground Truth To establish ground truth for regulatory relationships, we integrated information from several authoritative biological databases. This included known gene-gene interactions from the KEGG HPV carcinogenesis pathway [
6], the Reactome database [
8], and the STRING database [
5]. This information was compiled into binary labels indicating the presence or absence of a regulatory edge between gene pairs, serving as crucial ground truth for training and evaluating Task 2.
Textual Data For the instruction tuning of the Bio-LLM Reasoning Module, we compiled a rich corpus of biomedical text. This involved filtering abstracts from the PubMed literature database using keywords related to HPV, p53, E6/E7, and other relevant biological concepts. Furthermore, evidence text from the BioGRID protein interaction database [
6] was incorporated to provide rich biological knowledge and diverse reasoning examples for the LLM.
4.1.2. Evaluation Tasks
We designed three distinct evaluation tasks to comprehensively assess the capabilities of OncoReasoner:
Task 1: Differential Gene Expression Classification This task evaluates the model’s ability to accurately classify whether a given gene expression profile represents an HPV-induced transcriptional perturbation. The input consists of a gene expression vector, along with information indicating the presence or absence of E6/E7 activity. The output is a binary classification: whether the perturbation is HPV-induced.
Task 2: Regulatory Network Edge Prediction The goal of this task is to assess the model’s proficiency in predicting the existence and direction (activation or inhibition) of regulatory relationships between gene pairs. The inputs include a pair of genes (Gene A and Gene B), their respective expression change patterns, and relevant textual information (e.g., literature abstracts). The model’s output is a classification indicating whether a regulatory relationship exists and, if so, its direction.
Task 3: Functional Pathway Reasoning (Core Innovation) This is the central task, designed to evaluate OncoReasoner’s capacity for providing interpretable biological mechanism reasoning for E6/E7-induced transcriptomic perturbations. The input to the model is a list of genes that are significantly up-regulated or down-regulated under the influence of E6/E7. The expected output is a chain-of-thought explanation generated by the Bio-LLM Reasoning Module, detailing the involved biological pathways (e.g., p53 pathway, cell cycle regulation, apoptosis evasion) and their specific mechanistic contributions.
4.1.3. Baseline Models
To provide a rigorous comparison, we evaluated OncoReasoner against several representative baseline methods:
Traditional Statistical Methods We used DESeq2 [
5] for differential expression analysis, followed by gene set enrichment using KEGG pathways, as a benchmark for pathway identification.
Gene Set Enrichment Analysis (GSEA) This method [
6] was used to assess the enrichment of predefined gene sets (pathways) within the ranked list of differentially expressed genes.
Graph Neural Network (GNN) Only A standalone Graph Attention Network (GAT) was implemented for direct regulatory network prediction, using gene expression features and prior network information, but without the LLM reasoning component.
Biomedical BERT Classifier We employed a BioBERT-based classifier, fine-tuned on biomedical literature, to directly predict regulatory relationships or functional pathways from textual descriptions and gene lists.
Expression-based Machine Learning Classifier A Random Forest classifier was trained on gene expression profiles to perform classification tasks such as Task 1.
GPT-3.5 (LLM Only) For Task 3, we also included a general-purpose Large Language Model, GPT-3.5, as a baseline to evaluate its ability to perform biological reasoning without specific domain fine-tuning or integration with expression data encoding.
4.1.4. Evaluation Metrics
The performance of all models was quantified using standard metrics tailored to each task:
Task 1 Metrics For differential gene expression classification, we reported Accuracy, F1-score, and Area Under the Receiver Operating Characteristic curve (AUROC).
Task 2 Metrics For regulatory network edge prediction, Precision, Recall, and F1-score were used to evaluate the accuracy of predicting edges.
Task 3 Metrics For the core functional pathway reasoning task, we evaluated two key aspects: the percentage of Correct Pathway Identification (Correct Pathway %) and a Mechanism Explanation Score (ranging from 0 to 5), which was determined by expert human evaluators assessing the depth, accuracy, and coherence of the LLM-generated mechanistic explanations.
4.2. Performance Comparison
Table 1 summarizes the performance of
OncoReasoner and all baseline methods across the three evaluation tasks. The results demonstrate the superior capabilities of our proposed framework.
As shown in
Table 1,
OncoReasoner consistently outperforms all baselines across all applicable tasks. For Task 1, differential gene expression classification,
OncoReasoner achieved an AUROC of 0.95, indicating a strong ability to identify HPV-induced transcriptional perturbations. In Task 2, regulatory network edge prediction, our framework yielded an F1-score of 0.80, surpassing the standalone GNN. This highlights the benefit of integrating LLM-derived insights into the network refinement process. The most significant advancements are observed in Task 3, our core innovation of functional pathway reasoning.
OncoReasoner achieved a remarkable 91% correct pathway identification rate, outperforming both specialized (BioBERT, DESeq2+KEGG, GSEA) and general-purpose (GPT-3.5) LLM baselines. Furthermore, the expert-rated Mechanism Explanation Score of 4.5 out of 5 for
OncoReasoner signifies a substantial improvement in the quality and interpretability of biological mechanistic explanations, validating its unique advantage in combining advanced biological expression analysis with LLM-driven reasoning.
4.3. Ablation Study
To understand the individual contributions of each core module within
OncoReasoner, we conducted an ablation study. We evaluated simplified versions of our framework by systematically removing or replacing one or more key components. The results, presented in
Table 2, underscore the importance of each module to the overall performance and interpretability.
Removing the Expression Encoder (e.g., using raw expression values or simpler embeddings) led to a noticeable drop in performance across all tasks. The AUROC for Task 1 decreased to 0.89, and the F1-score for Task 2 dropped to 0.72. More importantly, the Correct Pathway % for Task 3 fell to 85% and the Mechanism Explanation Score to 3.8. This highlights the critical role of the sophisticated expression embeddings generated by models like NT-v2 or DNABERT-2 in providing rich, context-aware biological signals for downstream reasoning.
Disabling the Bio-LLM Reasoning Module significantly impaired the framework’s core interpretability features. While Task 1 and Task 2 showed a moderate decrease (AUROC of 0.92, F1 of 0.76), Task 3’s Correct Pathway % dropped to 78%, and no mechanism explanation score could be provided, underscoring that the LLM is indispensable for generating chain-of-thought biological reasoning. This confirms that the LLM is not merely an auxiliary component but central to the framework’s innovative interpretive capabilities.
The absence of the Graph Refinement Module also resulted in a performance decrement, with the F1-score for Task 2 dropping to 0.77 and Task 3’s Correct Pathway % to 89%. This indicates that the GAT, by integrating prior knowledge and refining LLM-inferred relationships, plays a crucial role in ensuring the biological consistency and accuracy of the predicted regulatory networks and subsequently improving the overall reasoning quality. The minor drop in Mechanism Explanation Score (4.3 vs. 4.5) suggests that a more accurate underlying graph facilitates clearer and more coherent explanations.
Finally, using only the Expression Encoder without subsequent reasoning or graph modules naturally limited its application, primarily performing simple classification tasks, affirming that the full integration of modules is necessary for comprehensive analysis. In summary, the ablation study unequivocally demonstrates that all three modules of OncoReasoner contribute synergistically, with the Bio-LLM Reasoning Module being paramount for interpretability, and the Expression Encoder and Graph Refinement Module providing essential biological grounding and structural coherence.
4.4. Human Evaluation of Interpretability
To thoroughly evaluate the qualitative aspects of OncoReasoner’s core contribution – interpretable biological mechanism reasoning – we conducted a human evaluation for Task 3 outputs. A panel of five expert biologists and oncologists independently reviewed a randomized set of mechanistic explanations generated by OncoReasoner, BioBERT, and GPT-3.5. Each expert scored the explanations on a scale of 0 to 5, based on criteria such as biological accuracy, depth of reasoning, coherence, and clarity of the chain-of-thought. The scores were then averaged to produce the final Mechanism Explanation Score.
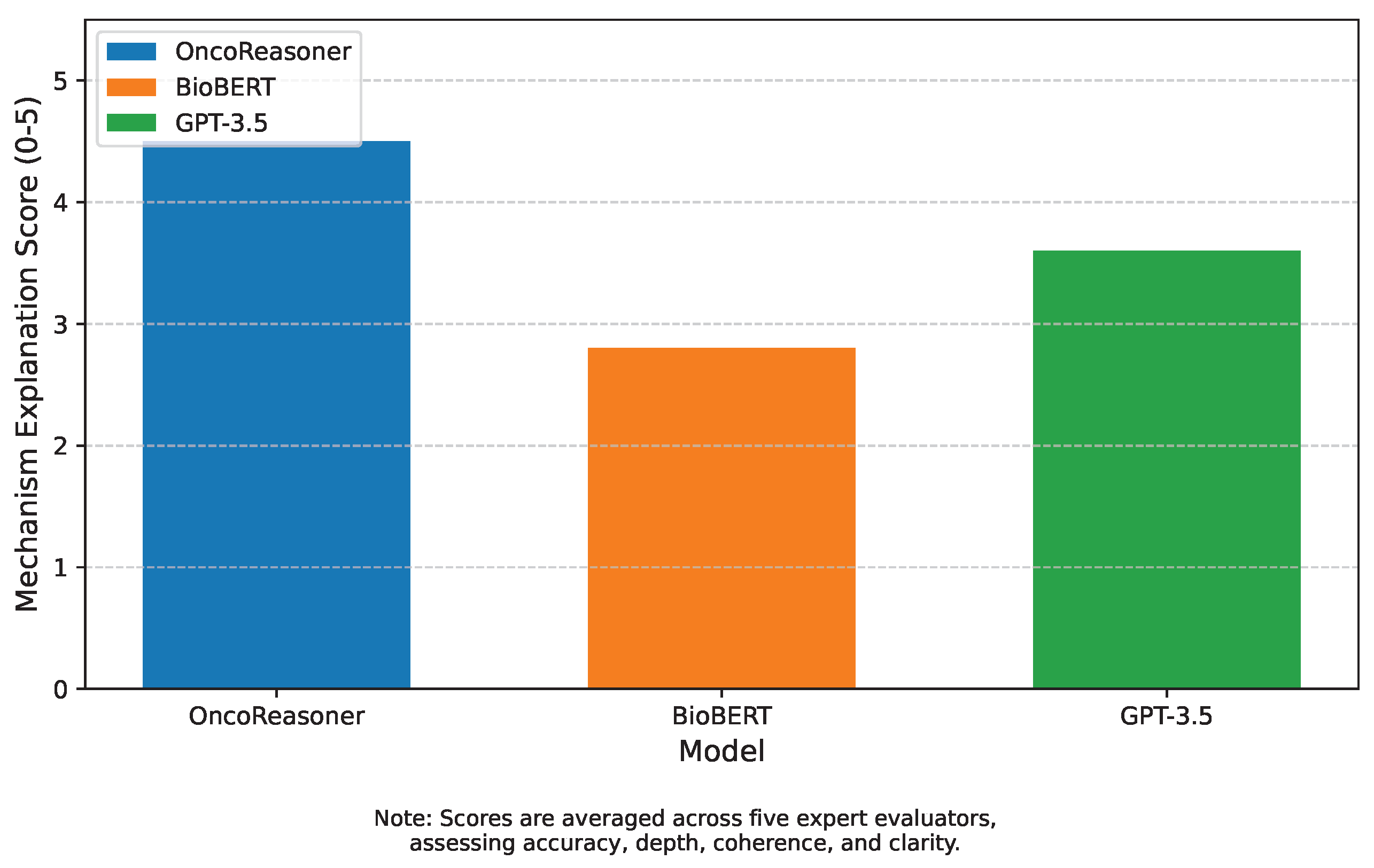
Figure 3 presents the results of this human evaluation.
OncoReasoner achieved an impressive average score of 4.5, significantly higher than both BioBERT (2.8) and GPT-3.5 (3.6). Experts noted that
OncoReasoner’s explanations were consistently more accurate, provided deeper insights into molecular mechanisms, and articulated a clearer chain-of-thought from observed transcriptomic perturbations to specific oncogenic processes. For instance, explanations from
OncoReasoner often precisely linked gene up/down-regulation to the direct targets of E6/E7 (e.g., p53 degradation, Rb inactivation), and subsequently elucidated the downstream consequences on cell cycle progression, apoptosis evasion, and genomic instability. In contrast, BioBERT explanations were often superficial or fragmented, while GPT-3.5, though generally coherent, sometimes lacked the precise biological nuance or made less direct connections specific to HPV E6/E7 mechanisms due to its general-purpose training. These human evaluation results strongly corroborate our quantitative findings, affirming
OncoReasoner’s ability to not only predict but also intelligently interpret complex biological phenomena in a highly comprehensible and biologically sound manner, marking a significant step forward in explainable AI for cancer research.
5. Conclusion
The intricate mechanisms of HPV E6/E7 oncogenesis necessitate interpretable computational tools to decipher complex gene regulatory networks. We introduced OncoReasoner, a novel framework designed to bridge this interpretability gap by integrating an Expression Encoder, a Bio-LLM Reasoning Module, and a Graph Refinement Module. This multi-modal architecture, coupled with a three-stage training strategy, enables OncoReasoner to not only predict but also profoundly interpret the transcriptomic consequences of viral activity. Our extensive validation demonstrated superior performance across differential gene expression and regulatory network prediction tasks. Crucially, in functional pathway reasoning, OncoReasoner achieved 91% accuracy in pathway identification and a remarkable expert-rated Mechanism Explanation Score of 4.5 out of 5, providing unprecedented deep, accurate, and highly interpretable biological explanations. This represents a significant advancement in explainable AI for biomedical research, offering powerful insights into oncogenic mechanisms, identifying potential therapeutic targets, and holding broad applicability beyond HPV studies. Future work will explore multimodal data integration and clinical validation.
References
- Cheng, L.; Li, X.; Bing, L. Is GPT-4 a Good Data Analyst? In Proceedings of the Findings of the Association for Computational Linguistics: EMNLP 2023. Association for Computational Linguistics, 2023, pp. 9496–9514. [CrossRef]
- Iida, H.; Thai, D.; Manjunatha, V.; Iyyer, M. TABBIE: Pretrained Representations of Tabular Data. In Proceedings of the Proceedings of the 2021 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies. Association for Computational Linguistics, 2021, pp. 3446–3456. [CrossRef]
- Hedderich, M.A.; Lange, L.; Adel, H.; Strötgen, J.; Klakow, D. A Survey on Recent Approaches for Natural Language Processing in Low-Resource Scenarios. In Proceedings of the Proceedings of the 2021 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies. Association for Computational Linguistics, 2021, pp. 2545–2568. [CrossRef]
- Dai, R.; Tao, R.; Li, X.; Shang, T.; Zhao, S.; Ren, Q. Expression profiling of mRNA and functional network analyses of genes regulated by human papilloma virus E6 and E7 proteins in HaCaT cells. Frontiers in Microbiology 2022, 13, 979087.
- Li, R.; Chen, H.; Feng, F.; Ma, Z.; Wang, X.; Hovy, E. Dual Graph Convolutional Networks for Aspect-based Sentiment Analysis. In Proceedings of the Proceedings of the 59th Annual Meeting of the Association for Computational Linguistics and the 11th International Joint Conference on Natural Language Processing (Volume 1: Long Papers). Association for Computational Linguistics, 2021, pp. 6319–6329. [CrossRef]
- Lyu, H.; Jiang, S.; Zeng, H.; Xia, Y.; Wang, Q.; Zhang, S.; Chen, R.; Leung, C.; Tang, J.; Luo, J. LLM-Rec: Personalized Recommendation via Prompting Large Language Models. In Proceedings of the Findings of the Association for Computational Linguistics: NAACL 2024. Association for Computational Linguistics, 2024, pp. 583–612. [CrossRef]
- Yang, X.; Feng, S.; Zhang, Y.; Wang, D. Multimodal Sentiment Detection Based on Multi-channel Graph Neural Networks. In Proceedings of the Proceedings of the 59th Annual Meeting of the Association for Computational Linguistics and the 11th International Joint Conference on Natural Language Processing (Volume 1: Long Papers). Association for Computational Linguistics, 2021, pp. 328–339. [CrossRef]
- Li, J.; Xu, K.; Li, F.; Fei, H.; Ren, Y.; Ji, D. MRN: A Locally and Globally Mention-Based Reasoning Network for Document-Level Relation Extraction. In Proceedings of the Findings of the Association for Computational Linguistics: ACL-IJCNLP 2021. Association for Computational Linguistics, 2021, pp. 1359–1370. [CrossRef]
- Zhou, Y.; Zheng, H.; Chen, D.; Yang, H.; Han, W.; Shen, J. From Medical LLMs to Versatile Medical Agents: A Comprehensive Survey 2025.
- Ahuja, K.; Diddee, H.; Hada, R.; Ochieng, M.; Ramesh, K.; Jain, P.; Nambi, A.; Ganu, T.; Segal, S.; Ahmed, M.; et al. MEGA: Multilingual Evaluation of Generative AI. In Proceedings of the Proceedings of the 2023 Conference on Empirical Methods in Natural Language Processing. Association for Computational Linguistics, 2023, pp. 4232–4267. [CrossRef]
- Moiseev, F.; Dong, Z.; Alfonseca, E.; Jaggi, M. SKILL: Structured Knowledge Infusion for Large Language Models. In Proceedings of the Proceedings of the 2022 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies. Association for Computational Linguistics, 2022, pp. 1581–1588. [CrossRef]
- Xu, C.; Chen, Y.Y.; Nayyeri, M.; Lehmann, J. Temporal Knowledge Graph Completion using a Linear Temporal Regularizer and Multivector Embeddings. In Proceedings of the Proceedings of the 2021 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies. Association for Computational Linguistics, 2021, pp. 2569–2578. [CrossRef]
- Deshpande, A.; Murahari, V.; Rajpurohit, T.; Kalyan, A.; Narasimhan, K. Toxicity in chatgpt: Analyzing persona-assigned language models. In Proceedings of the Findings of the Association for Computational Linguistics: EMNLP 2023. Association for Computational Linguistics, 2023, pp. 1236–1270. [CrossRef]
- Liu, F.; Shareghi, E.; Meng, Z.; Basaldella, M.; Collier, N. Self-Alignment Pretraining for Biomedical Entity Representations. In Proceedings of the Proceedings of the 2021 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies. Association for Computational Linguistics, 2021, pp. 4228–4238. [CrossRef]
- Wang, B.; Min, S.; Deng, X.; Shen, J.; Wu, Y.; Zettlemoyer, L.; Sun, H. Towards Understanding Chain-of-Thought Prompting: An Empirical Study of What Matters. In Proceedings of the Proceedings of the 61st Annual Meeting of the Association for Computational Linguistics (Volume 1: Long Papers). Association for Computational Linguistics, 2023, pp. 2717–2739. [CrossRef]
- Kurtic, E.; Campos, D.; Nguyen, T.; Frantar, E.; Kurtz, M.; Fineran, B.; Goin, M.; Alistarh, D. The Optimal BERT Surgeon: Scalable and Accurate Second-Order Pruning for Large Language Models. In Proceedings of the Proceedings of the 2022 Conference on Empirical Methods in Natural Language Processing. Association for Computational Linguistics, 2022, pp. 4163–4181. [CrossRef]
- Piper, A.; So, R.J.; Bamman, D. Narrative Theory for Computational Narrative Understanding. In Proceedings of the Proceedings of the 2021 Conference on Empirical Methods in Natural Language Processing. Association for Computational Linguistics, 2021, pp. 298–311. [CrossRef]
- Tao, R.; Liu, S.; Maltezos, H.; Tao, F. Gene Regulation in Comorbid Migraine and Myogenic Temporomandibular Disorder Pain. Genes 2025, 16, 1435.
- Lou, Y.; Tao, R.; Weng, X.; Sun, S.; Yang, Y.; Ying, B. Bioinformatics analysis of synovial fluid-derived mesenchymal stem cells in the temporomandibular joint stimulated with IL-1β. Cytotechnology 2023, 75, 325–334.
- Feng, S.Y.; Gangal, V.; Wei, J.; Chandar, S.; Vosoughi, S.; Mitamura, T.; Hovy, E. A Survey of Data Augmentation Approaches for NLP. In Proceedings of the Findings of the Association for Computational Linguistics: ACL-IJCNLP 2021. Association for Computational Linguistics, 2021, pp. 968–988. [CrossRef]
- Han, C.; Fan, Z.; Zhang, D.; Qiu, M.; Gao, M.; Zhou, A. Meta-Learning Adversarial Domain Adaptation Network for Few-Shot Text Classification. In Proceedings of the Findings of the Association for Computational Linguistics: ACL-IJCNLP 2021. Association for Computational Linguistics, 2021, pp. 1664–1673. [CrossRef]
- Zhang, W.; Deng, Y.; Liu, B.; Pan, S.; Bing, L. Sentiment Analysis in the Era of Large Language Models: A Reality Check. In Proceedings of the Findings of the Association for Computational Linguistics: NAACL 2024. Association for Computational Linguistics, 2024, pp. 3881–3906. [CrossRef]
- Ribeiro, L.F.R.; Zhang, Y.; Gurevych, I. Structural Adapters in Pretrained Language Models for AMR-to-Text Generation. In Proceedings of the Proceedings of the 2021 Conference on Empirical Methods in Natural Language Processing. Association for Computational Linguistics, 2021, pp. 4269–4282. [CrossRef]
- Shi, W.; Li, F.; Li, J.; Fei, H.; Ji, D. Effective Token Graph Modeling using a Novel Labeling Strategy for Structured Sentiment Analysis. In Proceedings of the Proceedings of the 60th Annual Meeting of the Association for Computational Linguistics (Volume 1: Long Papers). Association for Computational Linguistics, 2022, pp. 4232–4241. [CrossRef]
- Gui, L.; Wang, B.; Huang, Q.; Hauptmann, A.; Bisk, Y.; Gao, J. KAT: A Knowledge Augmented Transformer for Vision-and-Language. In Proceedings of the Proceedings of the 2022 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies. Association for Computational Linguistics, 2022, pp. 956–968. [CrossRef]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).