Submitted:
01 March 2026
Posted:
03 March 2026
Read the latest preprint version here
Abstract
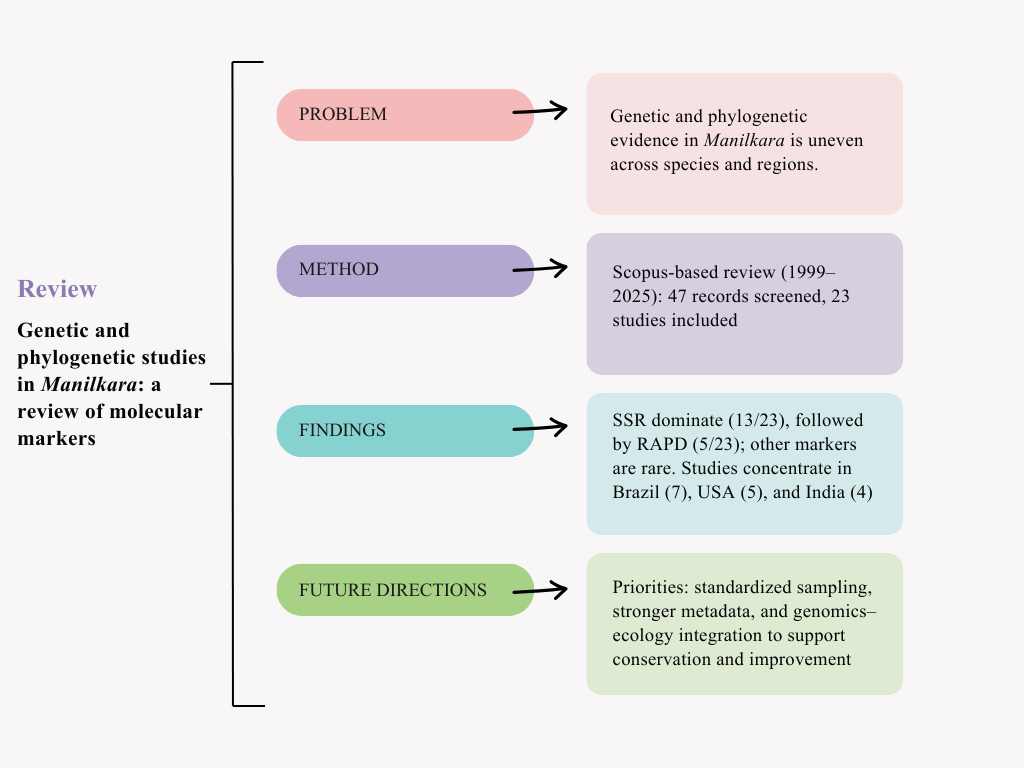
Keywords:
1. Introduction
2. Materials and Methods
2.1. Study Design
2.2. Bibliographic Search and Selection of Articles
2.3. Eligibility Criteria
2.3.1. Inclusion Criteria
2.3.2. Exclusion Criteria
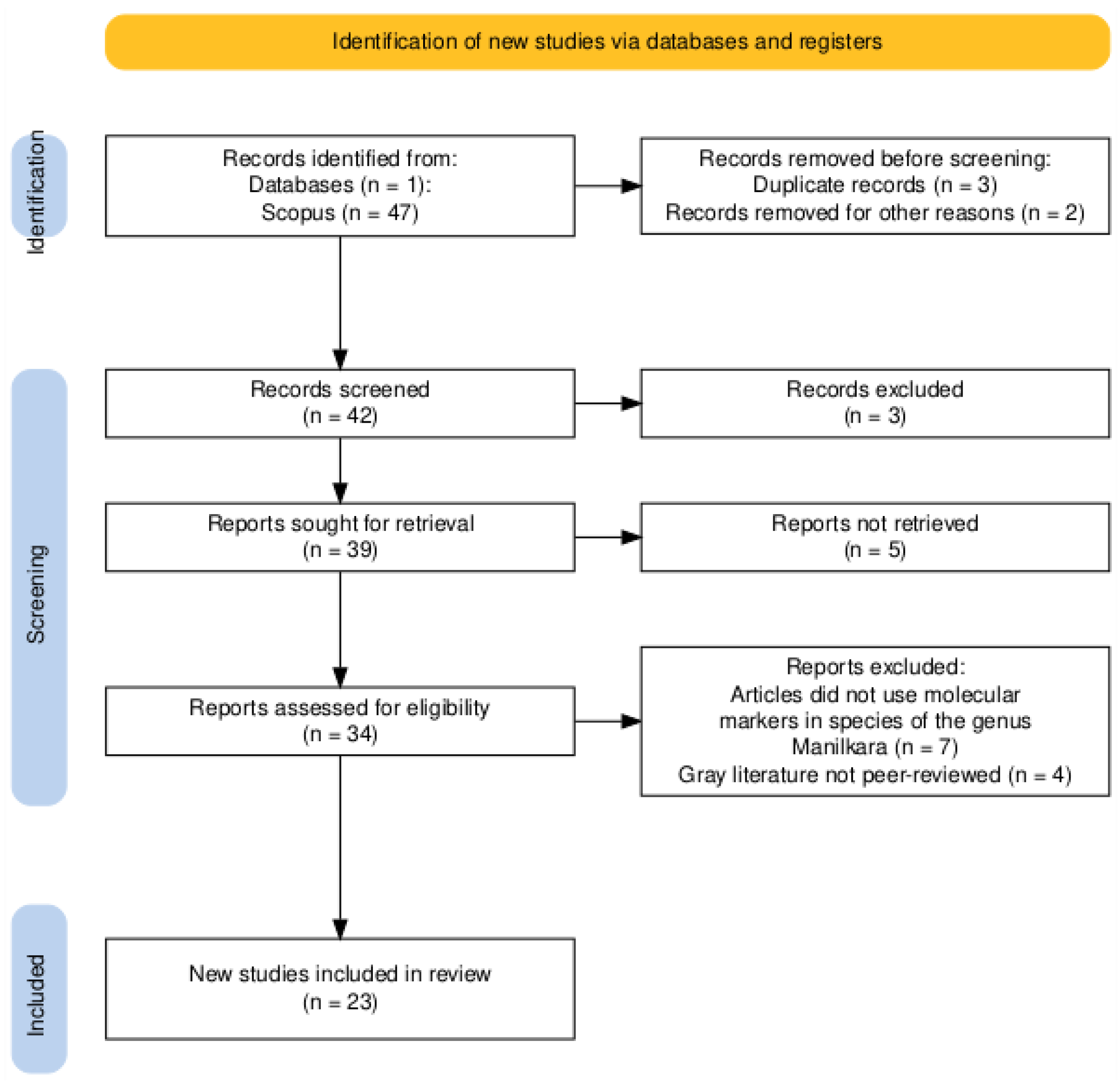
2.4. Study Selection
2.5. Data Extraction
3. Results
3.1. Study Characteristics
3.2. Synthesis of the Results
3.3. Utilization of Molecular Marker Systems
3.3.1. Random Amplification of Polymorphic DNA (RAPD)
3.3.2. Start Codon-Targeted Polymorphism (SCoT)
3.3.3. Simple Sequence Repeats (SSRs)
3.3.4. Chloroplast Simple Sequence Repeat (cpSSRs) Microsatellites
3.3.5. Internal Transcribed Spacer (ITS)
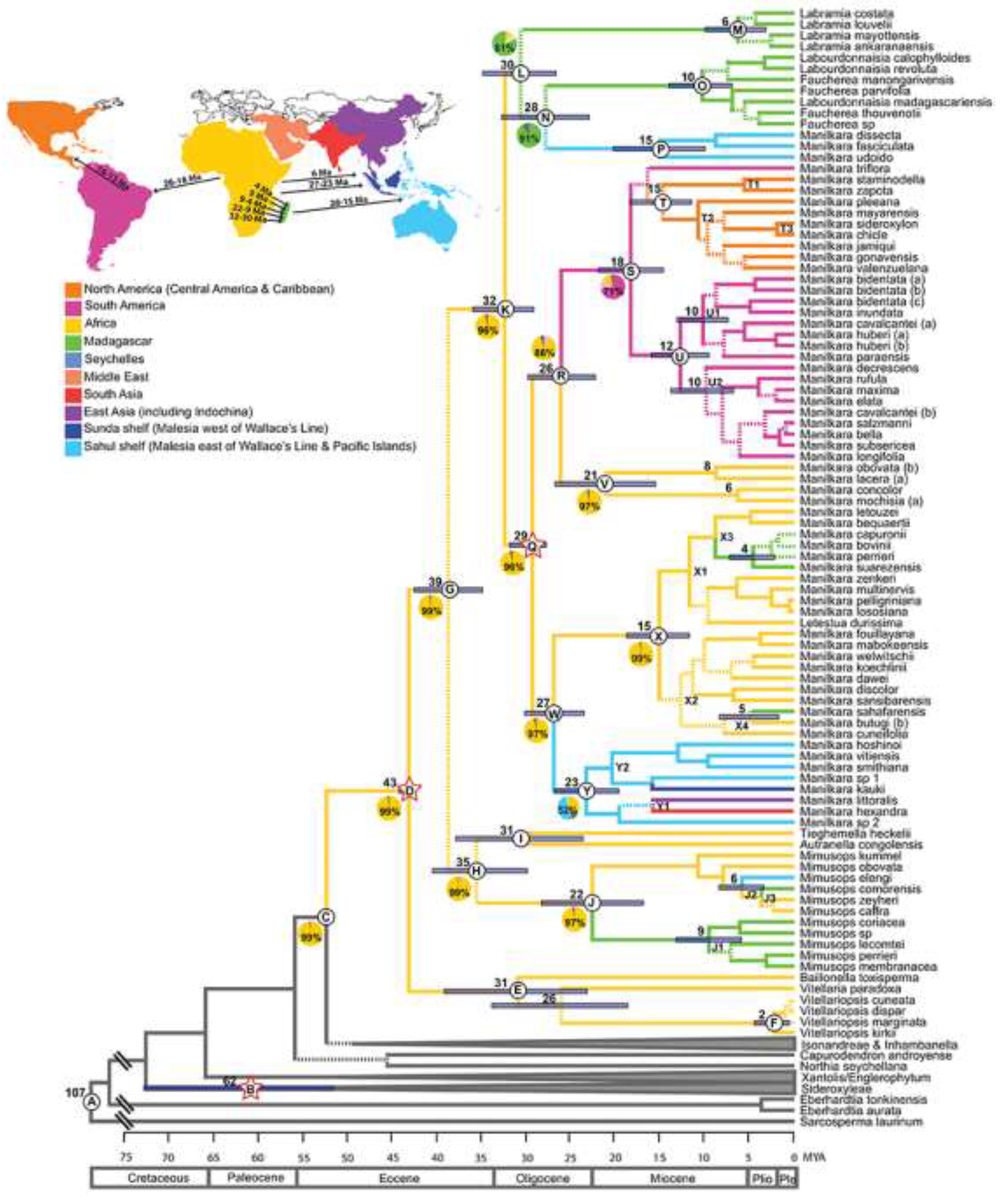
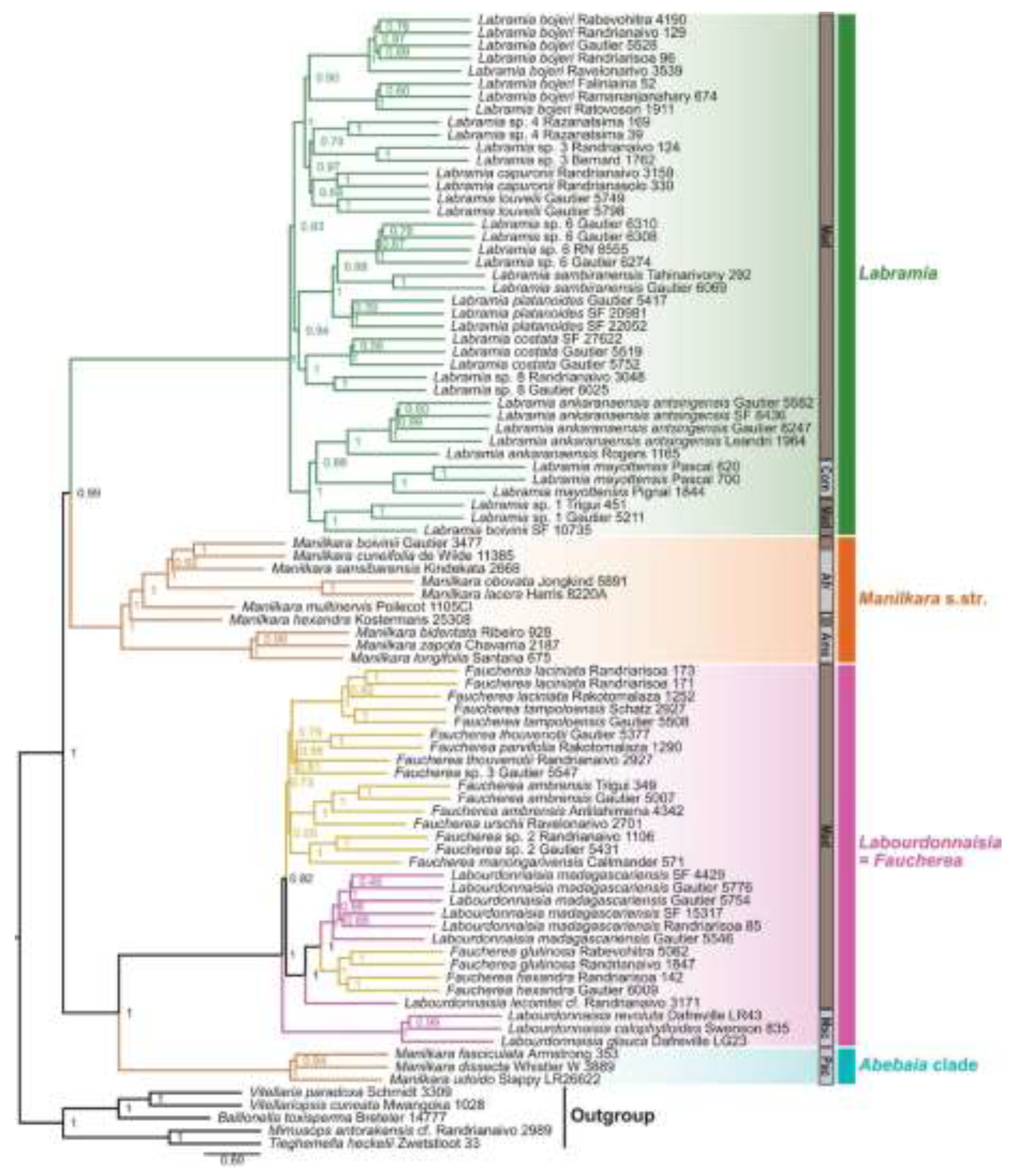
4. Discussion
5. Future Directions
6. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Aguilar, R.; Quesada, M.; Ashworth, L.; Herrerias-Diego, Y.; Lobo, J. Genetic consequences of habitat fragmentation in plant populations: susceptible signals in plant traits and methodological approaches. Molecular Ecology 2008, 17, 5177–5188. [Google Scholar] [CrossRef] [PubMed]
- Aitken, S.N.; Jordan, R.; Tumas, H.R. Conserving Evolutionary Potential: Combining Landscape Genomics with Established Methods to Inform Plant Conservation. Annual Review of Plant Biology 2024, 75, 707–736. [Google Scholar] [CrossRef] [PubMed]
- Alseekh, S.; Klemmer, A.; Yan, J.; Guo, T.; Fernie, A.R. Embracing plant plasticity or robustness as a means of ensuring food security. Nat Commun 2025, 16, 461. [Google Scholar] [CrossRef] [PubMed]
- Alves, S.I.A.; Dantas, C.W.D.; Macedo, D.B.; Ramos, R.T.J. What are microsatellites and how to choose the best tool: a user-friendly review of SSR and 74 SSR mining tools. Front. Genet. 2024, 15. [Google Scholar] [CrossRef]
- Amiteye, S. Basic concepts and methodologies of DNA marker systems in plant molecular breeding. Heliyon 2021, 7. [Google Scholar] [CrossRef]
- Amom, T.; Nongdam, P. The use of molecular marker methods in plants: A review. Int. J. Curr. Res. Rev. 2017, 9. [Google Scholar] [CrossRef]
- Amom, T.; Tikendra, L.; Apana, N.; Goutam, M.; Sonia, P.; Koijam, A.S.; Potshangbam, A.M.; Rahaman, H.; Nongdam, P. Efficiency of RAPD, ISSR, iPBS, SCoT and phytochemical markers in the genetic relationship study of five native and economical important bamboos of North-East India. Phytochemistry 2020, 174, 112330. [Google Scholar] [CrossRef]
- Amom, T.; Tikendra, L.; Rahaman, H.; Potshangbam, A.; Nongdam, P. Evaluation of genetic relationship between 15 bamboo species of North-East India based on ISSR marker analysis. Mol Biol Res Commun 2018, 7, 7–15. [Google Scholar] [CrossRef]
- Arias, R.S.; Ballard, L.L.; Duke, M.V.; Simpson, S.A.; Liu, X.F.; Orner, V.A.; Sobolev, V.S.; Scheffler, B.E.; Martinez-Castillo, J. Development of nuclear microsatellite markers to facilitate germplasm conservation and population genetics studies of five groups of tropical perennial plants with edible fruits and shoots: rambutan (Nephelium lappaceum L.), sapodilla (Manilkara zapota (L.) P. Royen), lychee (Litchi chinensis Sonn.), mangosteen (Garcinia mangostana Linn. and Garcinia cochinchinensis (Lour.) Choisy) and bamboo (Bambusa vulgaris Schrad. ex J.C. Wendl and Guadua angustifolia Kunth). Genet Resour Crop Evol 2020, 67, 1715–1731. [Google Scholar] [CrossRef]
- Armstrong, K.E.; Stone, G.N.; Nicholls, J.A.; Valderrama, E.; Anderberg, A.A.; Smedmark, J.; Gautier, L.; Naciri, Y.; Milne, R.; Richardson, J.E. Patterns of diversification amongst tropical regions compared: a case study in Sapotaceae. Front. Genet. 2014, 5. [Google Scholar] [CrossRef]
- Avise, J.C. Molecular Markers, Natural History and Evolution; Springer Science & Business Media, 2012. [Google Scholar]
- Azevedo, V.C.R.; Kanashiro, M.; Ciampi, A.Y.; Grattapaglia, D. Genetic structure and mating system of Manilkara huberi (Ducke) A. Chev., a heavily logged Amazonian timber species. Journal of Heredity 2007, 98, 646–654. [Google Scholar] [CrossRef]
- Azevedo, V.C.R.; Kanashiro, M.; Grattapaglia, D.; Ciampi, A.Y. CpDNA variability in Manilkara huberi, a species under sustainable management in the Brazilian Amazon. Pesquisa Agropecuaria Brasileira 2008, 43, 859–867. [Google Scholar] [CrossRef]
- Azevedo, V.C.R.; Vinson, C.C.; Ciampi, A.Y. Twelve microsatellite loci in Manilkara huberi (Ducke) Standl (Sapotaceae), an Amazonian timber species. Molecular Ecology Notes 2005, 5, 13–15. [Google Scholar] [CrossRef]
- Beckers, V.; Jepma, P.J.; Kokshoorn, S.; Wilkie, P.; Hendriks, K.; Smets, E.; Lens, F. Wood anatomy helps solve challenging taxonomic relationships in the tropical Sapotaceae. 2025. [Google Scholar] [CrossRef]
- Betts, M.G.; Wolf, C.; Pfeifer, M.; Banks-Leite, C.; Arroyo-Rodríguez, V.; Ribeiro, D.B.; Barlow, J.; Eigenbrod, F.; Faria, D.; Fletcher, R.J.; Hadley, A.S.; Hawes, J.E.; Holt, R.D.; Klingbeil, B.; Kormann, U.; Lens, L.; Levi, T.; Medina-Rangel, G.F.; Melles, S.L.; Mezger, D.; Morante-Filho, J.C.; Orme, C.D.L.; Peres, C.A.; Phalan, B.T.; Pidgeon, A.; Possingham, H.; Ripple, W.J.; Slade, E.M.; Somarriba, E.; Tobias, J.A.; Tylianakis, J.M.; Urbina-Cardona, J.N.; Valente, J.J.; Watling, J.I.; Wells, K.; Wearn, O.R.; Wood, E.; Young, R.; Ewers, R.M. Extinction filters mediate the global effects of habitat fragmentation on animals. Science 2019, 366, 1236–1239. [Google Scholar] [CrossRef] [PubMed]
- Bidyananda, N.; Jamir, I.; Nowakowska, K.; Varte, V.; Vendrame, W.A.; Devi, R.S.; Nongdam, P. Plant Genetic Diversity Studies: Insights from DNA Marker Analyses. International Journal of Plant Biology 2024, 15, 607–640. [Google Scholar] [CrossRef]
- Bjornson, S.; Verbruggen, H.; Upham, N.S.; Steenwyk, J.L. Reticulate evolution: Detection and utility in the phylogenomics era. Mol Phylogenet Evol 2024, 201, 108197. [Google Scholar] [CrossRef]
- Bock, D.G.; Cai, Z.; Elphinstone, C.; González-Segovia, E.; Hirabayashi, K.; Huang, K.; Keais, G.L.; Kim, A.; Owens, G.L.; Rieseberg, L.H. Genomics of plant speciation. Plant Comm 2023, 4. [Google Scholar] [CrossRef]
- Burnham, J.F. Scopus database: a review. Biomedical Digital Libraries 2006, 3, 1. [Google Scholar] [CrossRef]
- Cao, H.X.; Vu, G.T.H.; Gailing, O. From Genome Sequencing to CRISPR-Based Genome Editing for Climate-Resilient Forest Trees. International Journal of Molecular Sciences 2022, 23, 966. [Google Scholar] [CrossRef]
- Caradus, J.R. Is co-existence and/or containment of genetically modified plants possible, and is it important? New Zealand Journal of Agricultural Research 2025, 68, 1329–1380. [Google Scholar] [CrossRef]
- Cardi, T.; Murovec, J.; Bakhsh, A.; Boniecka, J.; Bruegmann, T.; Bull, S.E.; Eeckhaut, T.; Fladung, M.; Galovic, V.; Linkiewicz, A.; Lukan, T.; Mafra, I.; Michalski, K.; Kavas, M.; Nicolia, A.; Nowakowska, J.; Sági, L.; Sarmiento, C.; Yıldırım, K.; Zlatković, M.; Hensel, G.; Van Laere, K. CRISPR/Cas-mediated plant genome editing: outstanding challenges a decade after implementation. Trends in Plant Science 2023, 28, 1144–1165. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, C. da S.; Ballesteros-Mejia, L.; Ribeiro, M.C.; Côrtes, M.C.; Santos, A.S.; Collevatti, R.G. Climatic stability and contemporary human impacts affect the genetic diversity and conservation status of a tropical palm in the Atlantic Forest of Brazil. Conserv Genet 2017, 18, 467–478. [Google Scholar] [CrossRef]
- Chadegani, A.A.; Salehi, H.; Yunus, M.M.; Farhadi, H.; Fooladi, M.; Farhadi, M.; Ebrahim, N.A. A Comparison between Two Main Academic Literature Collections: Web of Science and Scopus Databases. Asian Social Science 2013, 9, p18. [Google Scholar] [CrossRef]
- Chambers, E.A.; Bishop, A.P.; Wang, I.J. Individual-based landscape genomics for conservation: An analysis pipeline. Mol Ecol Resour 2025, 25, e13884. [Google Scholar] [CrossRef]
- Chen, S.; Yao, H.; Han, J.; Liu, C.; Song, J.; Shi, L.; Zhu, Y.; Ma, X.; Gao, T.; Pang, X.; Luo, K.; Li, Y.; Li, X.; Jia, X.; Lin, Y.; Leon, C. Validation of the ITS2 Region as a Novel DNA Barcode for Identifying Medicinal Plant Species. PLOS ONE 2010, 5, e8613. [Google Scholar] [CrossRef]
- Cheng, T.; Xu, C.; Lei, L.; Li, C.; Zhang, Y.; Zhou, S. Barcoding the kingdom Plantae: new PCR primers for ITS regions of plants with improved universality and specificity. Molecular Ecology Resources 2016, 16, 138–149. [Google Scholar] [CrossRef]
- Choi, Y.; Ha, Y.-H.; Choi, J. Genetic diversity assessment of a plant for forest restoration on the Korean Peninsula: A case study of Lespedeza cuneata G. Don (Fabaceae). Journal of Asia-Pacific Biodiversity 2024, 17, 35–42. [Google Scholar] [CrossRef]
- Christe, C.; Boluda, C.G.; Koubínová, D.; Gautier, L.; Naciri, Y. New genetic markers for Sapotaceae phylogenomics: More than 600 nuclear genes applicable from family to population levels. Molecular Phylogenetics and Evolution 2021, 160, 107123. [Google Scholar] [CrossRef]
- Chung, M.Y.; Merilä, J.; Li, J.; Mao, K.; López-Pujol, J.; Tsumura, Y.; Chung, M.G. Neutral and adaptive genetic diversity in plants: An overview. Front. Ecol. Evol. 2023, 11. [Google Scholar] [CrossRef]
- Collard, B.C.Y.; Mackill, D.J. Start Codon Targeted (SCoT) Polymorphism: A Simple, Novel DNA Marker Technique for Generating Gene-Targeted Markers in Plants. Plant Mol Biol Rep 2009, 27, 86–93. [Google Scholar] [CrossRef]
- Coutinho, J.P.; Carvalho, A.; Martín, A.; Lima-Brito, J. Molecular characterization of Fagaceae species using inter-primer binding site (iPBS) markers. Mol Biol Rep 2018, 45, 133–142. [Google Scholar] [CrossRef] [PubMed]
- Dauphin, B.; Rellstab, C.; Wüest, R.O.; Karger, D.N.; Holderegger, R.; Gugerli, F.; Manel, S. Re-thinking the environment in landscape genomics. Trends in Ecology & Evolution 2023, 38, 261–274. [Google Scholar] [CrossRef]
- De Faria, A.D.; Pirani, J.R.; Ribeiro, J.E.L.D.S.; Nylinder, S.; Terra-Araujo, M.H.; Vieira, P.P.; Swenson, U. Towards a natural classification of Sapotaceae subfamily Chrysophylloideae in the Neotropics. Botanical Journal of the Linnean Society 2017, 185, 27–55. [Google Scholar] [CrossRef]
- Dirzo, R.; Raven, P.H. Global State of Biodiversity and Loss. Annual Review of Environment and Resources 2003, 28, 137–167. [Google Scholar] [CrossRef]
- Dong, Y.; Yao, H.; Lin, Z.; Zhang, L.; Bao, Z. Characterization of 62 polymorphic EST-SSR markers in the blood clam (Tegillarca granosa) and their cross-amplification in Scapharca subcrenata. Conservation Genet Resour 2012, 4, 991–997. [Google Scholar] [CrossRef]
- Doyle, J.J.; Doyle, J.L. Isolation of plant DNA from fresh tissue; Focus., 1990. [Google Scholar]
- Drew, B.T.; Ruhfel, B.R.; Smith, S.A.; Moore, M.J.; Briggs, B.G.; Gitzendanner, M.A.; Soltis, P.S.; Soltis, D.E. Another Look at the Root of the Angiosperms Reveals a Familiar Tale. Syst Biol 2014, 63, 368–382. [Google Scholar] [CrossRef]
- Duarte, D.; Jurcic, E.J.; Dutour, J.; Villalba, P.V.; Centurión, C.; Grattapaglia, D.; Cappa, E.P. Genomic selection in forest trees comes to life: unraveling its potential in an advanced four-generation Eucalyptus grandis population. Front. Plant Sci. 2024, 15. [Google Scholar] [CrossRef]
- El-Kassaby, Y.A.; Cappa, E.P.; Chen, C.; Ratcliffe, B.; Porth, I.M. Efficient genomics-based ‘end-to-end’ selective tree breeding framework. Heredity 2024, 132, 98–105. [Google Scholar] [CrossRef]
- Feng, J.; Dan, X.; Cui, Y.; Gong, Y.; Peng, M.; Sang, Y.; Ingvarsson, P.K.; Wang, J. Integrating evolutionary genomics of forest trees to inform future tree breeding amid rapid climate change. Plant Communications 2024, 5, 101044. [Google Scholar] [CrossRef]
- Ganzhorn, S.M.; Thomas, W.W.; Gaiotto, F.A.; Lewis, J.D. Spatial genetic structure of Manilkara maxima (Sapotaceae), a tree species from the Brazilian Atlantic forest. Journal of Tropical Ecology 2015, 31, 437–447. [Google Scholar] [CrossRef]
- Gautier, L. Sapotáceas. In La historia natural de Madagascar; Goodman, JP, Bensted, SM, Eds.; The University of Chicago Press: Chicago., 2003. [Google Scholar]
- Gautier, L.; Lachenaud, O.; van der Burgt, X.; Kenfack, D. Five new species of Englerophytum K. Krause (Sapotaceae) from central Africa; Candollea, 2016. [Google Scholar]
- GBIF.org. accessed. Global Biodiversity Information Facility [WWW Document], 2026; (accessed on 2.24.26). [CrossRef]
- Gentry, A.H. A field guide to the families and genera of woody plants of northeast South America; Conservation International.: Washington, D.C, 1993. [Google Scholar]
- González-Hernández, D.; García-Pérez, E.; Guntin-Marey, P. Genetic characterization of Manilkara zapota from veracruz, Mexico, with SSR markers. Agrociencia 2012, 46, 663–675. [Google Scholar]
- Goraniya, S.; Tusamda, N.; Shirolkar, A.R.; Rao, G.; Murthy, S.N.; Pawar, S.D. Molecular analysis of Manilkara hexandra Roxb. and Averrhoa carambola using RAPD markers helps to understand genetic variations. International Journal of Pharmacy and Pharmaceutical Sciences 2013, 5, 626–628. [Google Scholar]
- Govaerts, R.; Frodin, D.G.; Pennington, T.D. World Checklist and Bibliography of Sapotaceae; Royal Botanic Gardens, 2001. [Google Scholar]
- Grattapaglia, D. Twelve Years into Genomic Selection in Forest Trees: Climbing the Slope of Enlightenment of Marker Assisted Tree Breeding. Forests 2022, 13, 1554. [Google Scholar] [CrossRef]
- Haasl, R.J.; Payseur, B.A. Multi-locus inference of population structure: a comparison between single nucleotide polymorphisms and microsatellites. Heredity 2011, 106, 158–171. [Google Scholar] [CrossRef] [PubMed]
- Haddaway, N.R.; Page, M.J.; Pritchard, C.C.; McGuinness, L.A. PRISMA2020: An R package and Shiny app for producing PRISMA 2020-compliant flow diagrams, with interactivity for optimised digital transparency and Open Synthesis. Campbell Systematic Reviews 2022, 18, e1230. [Google Scholar] [CrossRef]
- Hasan, N.; Choudhary, S.; Naaz, N.; Sharma, N.; Laskar, R.A. Recent advancements in molecular marker-assisted selection and applications in plant breeding programmes. Journal of Genetic Engineering and Biotechnology 2021, 19, 128. [Google Scholar] [CrossRef]
- Heaton, H.J.; Whitkus, R.; Gómez-Pompa, A. Extreme ecological and phenotypic differences in the tropical tree chicozapote (Manilkara zapota (L.) P. Royen) are not matched by genetic divergence: a random amplified polymorphic DNA (RAPD) analysis. Molecular Ecology 1999, 8, 627–632. [Google Scholar] [CrossRef]
- Heuertz, M.; Carvalho, S.B.; Galindo, J.; Rinkevich, B.; Robakowski, P.; Aavik, T.; Altinok, I.; Barth, J.M.I.; Cotrim, H.; Goessen, R.; González-Martínez, S.C.; Grebenc, T.; Hoban, S.; Kopatz, A.; McMahon, B.J.; Porth, I.; Raeymaekers, J.A.M.; Träger, S.; Valdecantos, A.; Vella, A.; Vernesi, C.; Garnier-Géré, P. The application gap: Genomics for biodiversity and ecosystem service management. Biological Conservation 2023, 278, 109883. [Google Scholar] [CrossRef]
- Hibbins, M.S.; Hahn, M.W. Phylogenomic approaches to detecting and characterizing introgression. Genetics 2022, 220, iyab173. [Google Scholar] [CrossRef]
- Hoelzel, A.R. Where to now with the evolutionarily significant unit? Trends in Ecology & Evolution 2023, 38, 1134–1142. [Google Scholar] [CrossRef]
- Hollingsworth, P.M.; Graham, S.W.; Little, D.P. Choosing and Using a Plant DNA Barcode. PLOS ONE 2011, 6, e19254. [Google Scholar] [CrossRef] [PubMed]
- Ijaz, S.; Iqbal, J.; Abbasi, B.A.; Yaseen, T.; Rehman, S.; Kazi, M.; Ansari, L.; Mahmood, T. Role of OMICS-Based Technologies in Plant Sciences. In OMICs-Based Techniques for Global Food Security; 2024; pp. 45–66. [Google Scholar] [CrossRef]
- Islam, K.K.; Saifullah, M.; Mahboob, M.G.; Jewel, K.N.-E.-A.; Ashraf, S.M.K.; Hyakumura, K. Restoring Soil Fertility, Productivity and Biodiversity through Participatory Agroforestry: Evidence from Madhupur Sal Forest, Bangladesh. Land 2024, 13, 326. [Google Scholar] [CrossRef]
- Kassaye, M.; Emiru, E.; Derebe, Y.; Tsega, A. Carbon stock dynamics in Ethiopian forests; a systematic review for sustainable forest management towards climate change mitigation. Trees, Forests and People 2025, 20, 100841. [Google Scholar] [CrossRef]
- Koch, M.; DeMond, J.; Pence, M.G.; Schaefer, E.A.; Rudgers, G. One risk assessment for genetically modified plants. Front. Bioeng. Biotechnol. 2025, 13. [Google Scholar] [CrossRef]
- Kremer, A.; Ronce, O.; Robledo-Arnuncio, J.J.; Guillaume, F.; Bohrer, G.; Nathan, R.; Bridle, J.R.; Gomulkiewicz, R.; Klein, E.K.; Ritland, K.; Kuparinen, A.; Gerber, S.; Schueler, S. Long-distance gene flow and adaptation of forest trees to rapid climate change. Ecology Letters 2012, 15, 378–392. [Google Scholar] [CrossRef]
- Kümpers, B.M.C.; Richardson, J.E.; Anderberg, A.A.; Wilkie, P.; Ronse De Craene, L.P. The significance of meristic changes in the flowers of Sapotaceae. Botanical Journal of the Linnean Society 2016, 180, 161–192. [Google Scholar] [CrossRef]
- Laurance, W.F.; Sayer, J.; Cassman, K.G. Agricultural expansion and its impacts on tropical nature. Trends in Ecology & Evolution 2014, 29, 107–116. [Google Scholar] [CrossRef]
- Lebedev, V.G.; Lebedeva, T.N.; Chernodubov, A.I.; Shestibratov, K.A. Genomic Selection for Forest Tree Improvement: Methods, Achievements and Perspectives. Forests 2020, 11, 1190. [Google Scholar] [CrossRef]
- Li, C.; Li, B.; Zhao, W.; Jiang, J.; Tang, J. Forest tree breeding under the global environmental change: Challenges and opportunities. Trees, Forests and People 2025, 20, 100867. [Google Scholar] [CrossRef]
- Liu, J.; Ren, S.-N.; Li, K.-X.; Xu, T.; Niu, Y.-F.; Shi, C. The complete chloroplast genome sequence of Manilkara zapota (Linn.) van Royen. Mitochondrial DNA Part B 2019, 4, 2127–2128. [Google Scholar] [CrossRef] [PubMed]
- López-Alvarado, J.; Sáez, L.; Filigheddu, R.; Garcia-Jacas, N.; Susanna, A. The limitations of molecular markers in phylogenetic reconstruction: The case of Centaurea sect. Phrygia (Compositae). TAXON 2014, 63, 1079–1091. [Google Scholar] [CrossRef]
- Lynch, M.; Milligan, B.G. Analysis of population genetic structure with RAPD markers. Molecular Ecology 1994, 3, 91–99. [Google Scholar] [CrossRef] [PubMed]
- Malik, S.K.; Kumar, S.; Choudhary, R.; Kole, P.R.; Chaudhury, R.; Bhat, K.V. Assessment of genetic diversity in khirni [Manilkara hexandra (Roxb.) Dubard]: An important underutilized fruit species of India using random amplified polymorphic DNA markers. Indian Journal of Horticulture 2013, 70, 18–25. [Google Scholar]
- Martínez-Natarén, D.A.; Parra-Tabla, V.; Munguía-Rosas, M.A. Genetic diversity and structure of the tree Manilkara zapota in a naturally fragmented tropical forest. Journal of Tropical Ecology 2017, 33, 285–294. [Google Scholar] [CrossRef]
- McCauley, D.E.; Sundby, A.K.; Bailey, M.F.; Welch, M.E. Inheritance of chloroplast DNA is not strictly maternal in Silene vulgaris (Caryophyllaceae): evidence from experimental crosses and natural populations. American Journal of Botany 2007, 94, 1333–1337. [Google Scholar] [CrossRef]
- Mendonça, E.G.; de Souza, A.M.; Vieira, F. de A.; Estopa, R.A.; Reis, C.A.F.; de Carvalho, D. Using Random Amplified Polymorphic DNA to Assess Genetic Diversity and Structure of Natural Calophyllum brasiliense (Clusiaceae) Populations in Riparian Forests. International Journal of Forestry Research 2014, 2014, 305286. [Google Scholar] [CrossRef]
- Moore, M.J.; Soltis, P.S.; Bell, C.D.; Burleigh, J.G.; Soltis, D.E. Phylogenetic analysis of 83 plastid genes further resolves the early diversification of eudicots. Proceedings of the National Academy of Sciences 2010, 107, 4623–4628. [Google Scholar] [CrossRef]
- Moraes, R.C.S.; Vivas, C.V.; Oliveira, F.A.; Menezes, I.P.P.; van den Berg, C.; Gaiotto, F.A. Microsatellite markers for an endemic Atlantic Forest tree, Manilkara multifida (Sapotaceae). AoB PLANTS 2013, 5, plt006. [Google Scholar] [CrossRef]
- Mora-Poblete, F.; Mieres-Castro, D.; do Amaral Júnior, A.T.; Balach, M.; Maldonado, C. Integrating deep learning for phenomic and genomic predictive modeling of Eucalyptus trees. Industrial Crops and Products 2024, 220, 119151. [Google Scholar] [CrossRef]
- Munzinger, J.; Swenson, U. Three new species of Planchonella Pierre (Sapotaceae) with a dichotomous and online key to the genus in New Caledonia. In Adansonia, séries, 3. ed.; 2009. [Google Scholar]
- Nadeem, M.A.; Nawaz, M.A.; Shahid, M.Q.; Doğan, Y.; Comertpay, G.; Yıldız, M.; Hatipoğlu, R.; Ahmad, F.; Alsaleh, A.; Labhane, N.; Özkan, H.; Chung, G.; Baloch, F.S. DNA molecular markers in plant breeding: current status and recent advancements in genomic selection and genome editing. Biotechnology & Biotechnological Equipment 2018, 32, 261–285. [Google Scholar] [CrossRef]
- Naidoo, S.; Slippers, B.; Plett, J.M.; Coles, D.; Oates, C.N. The Road to Resistance in Forest Trees. Front. Plant Sci. 2019, 10. [Google Scholar] [CrossRef] [PubMed]
- Nilkanta, H.; Amom, T.; Tikendra, L.; Rahaman, H.; Nongdam, P. ISSR Marker Based Population Genetic Study of Melocanna baccifera (Roxb.) Kurz: A Commercially Important Bamboo of Manipur, North-East India. Scientifica 2017, 3757238. [Google Scholar] [CrossRef]
- Olivares, I.; Tusso, S.; José Sanín, M.; de La Harpe, M.; Loiseau, O.; Rolland, J.; Salamin, N.; Kessler, M.; Shimizu, K.K.; Paris, M. Hyper-Cryptic radiation of a tropical montane plant lineage. Molecular Phylogenetics and Evolution 2024, 190, 107954. [Google Scholar] [CrossRef]
- Panchariya, D.C.; Dutta, P.; Ananya; Mishra, A.; Chawade, A.; Nayee, N.; Azam, S.; Gandham, R.K.; Majumdar, S.; Kushwaha, S.K. Genetic marker: a genome mapping tool to decode genetic diversity of livestock animals. Front Genet 2024, 15, 1463474. [Google Scholar] [CrossRef]
- Pennington, T. Flora of Ecuador 80. 152. Sapotaceae; Department of Plant and Environmental Sciences, Göteborg University: Gothenburg, 2007. [Google Scholar]
- Pennington, T. Flora da Reserva Ducke, Amazonas, Brasil: Sapotaceae, Rodriguésia. ed.; 2006. [Google Scholar]
- Pennington, T.D. Sapotaceae. In Nuevo catálogo de la flora vascular de venezuela; Hokche, 0., Berry, P., Huber, O., Lasser, Dr. Tobías, Eds.; Fundación Instituto Botánico de Venezuela: Caracas, 2008. [Google Scholar]
- Pennington, T.D. The genera of Sapotaceae; New York Botanical Garden e Kew., United Kingdom, 1991. [Google Scholar]
- Pennington, T.D. Flora Neotropica Monograph 52 Sapotaceae; New York Botanical Garden for the Organization for Flora Neotropica: New York, USA, 1990. [Google Scholar]
- Pezzini, F.F.; Ferrari, G.; Forrest, L.L.; Hart, M.L.; Nishii, K.; Kidner, C.A. Target capture and genome skimming for plant diversity studies. Appl Plant Sci 2023, 11, e11537. [Google Scholar] [CrossRef]
- Phang, A.; Niissalo, M.A.; Herrera-Alsina, L.; Pezzini, F.F.; Hollands, R.; Burslem, D.F.R.P.; Ruhsam, M.; Middleton, D.J.; Wilkie, P.; Khew, G.S. Novel phylogenomic and biogeographic insights into Palaquium and tribe Isonandreae (Sapotaceae). Molecular Phylogenetics and Evolution 2025, 212, 108403. [Google Scholar] [CrossRef]
- Phillips, H.R.P.; Newbold, T.; Purvis, A. Land-use effects on local biodiversity in tropical forests vary between continents. Biodivers Conserv 2017, 26, 2251–2270. [Google Scholar] [CrossRef]
- Pither, R.; Shore, J.S.; Kellman, M. Genetic diversity of the tropical tree Terminalia amazonia (Combretaceae) in naturally fragmented populations. Heredity 2003, 91, 307–313. [Google Scholar] [CrossRef]
- POWO. Plants of the World Online [WWW Document]. Plants of the World Online. 2025. Available online: https://powo.science.kew.org/ (accessed on 7.9.25).
- Randriarisoa, A.; Naciri, Y.; Armstrong, K.; Boluda, C.G.; Dafreville, S.; Pouchon, C.; Gautier, L. One in, one out: Generic circumscription within subtribe Manilkarinae (Sapotaceae). TAXON 2023, 72, 98–125. [Google Scholar] [CrossRef]
- Rathva, H.; Pandey, A.K.; Suthar, K.; Suthar, H.; Chakote, A.; Singh, Diwakar; Ahlawat, T.; Parmar, V.; Dhiman, V.K.; Pandey, H.; Singh, Devendra. Genetic relatedness analysis in sapota using SSR markers. Ecological Genetics and Genomics 2024, 31, 100234. [Google Scholar] [CrossRef]
- Reyes-Valdés, M.H.; Santacruz-Varela, A.; Martínez, O.; Simpson, J.; Hayano-Kanashiro, C.; Cortés-Romero, C. Analysis and Optimization of Bulk DNA Sampling with Binary Scoring for Germplasm Characterization. PLOS ONE 2013, 8, e79936. [Google Scholar] [CrossRef] [PubMed]
- Rosenberger, K.; Schumacher, E.; Brown, A.; Hoban, S. Proportional sampling strategy often captures more genetic diversity when population sizes vary. Biological Conservation 2021, 261, 109261. [Google Scholar] [CrossRef]
- Sari, V.K.; Murti, R.H. AN EFFECTIVE METHOD FOR DNA EXTRACTION OF MATURE LEAF OF SAPODILLA (Manilkara zapota (L.) van Royen). AGRIVITA Journal of Agricultural Science 2015, 37, 18–23. [Google Scholar] [CrossRef]
- Sari, V.K.; Wulandari, R.A.; Murti, R.H. Study on Diversity of Sapodilla (Manilkara zapota) by Molecular Marker in the Special Region of Yogyakarta. AGRIVITA Journal of Agricultural Science 2018, 40, 295–303. [Google Scholar] [CrossRef]
- Sathanandam, P.N.; Patil, P.; Rekha, A.; Muthuvel, I.; Patel, A.R.; Boggala, R.B.; Shirol, A.M.; Ravishankar, K.V. Development and characterization of microsatellite markers, genetic diversity and population structure analysis in Sapota (Manilkara zapota (L.) P. Royen). Genet Resour Crop Evol 2022, 69, 2787–2801. [Google Scholar] [CrossRef]
- Savolainen, O.; Lascoux, M.; Merilä, J. Ecological genomics of local adaptation. Nat Rev Genet 2013, 14, 807–820. [Google Scholar] [CrossRef]
- Scaketti, M.; Sujii, P.S.; Alves-Pereira, A.; Schwarcz, K.D.; Francisconi, A.F.; Moro, M.S.; Martins, K.K.M.; de Jesus, T.A.; de Souza, G.B.F.; Zucchi, M.I. Sample Size Impact (SaSii): An R script for estimating optimal sample sizes in population genetics and population genomics studies. PLOS ONE 2025, 20, e0316634. [Google Scholar] [CrossRef]
- Scott, D.J.; Broyd, T.; Ma, L. Exploratory literature review of blockchain in the construction industry. Automation in Construction 2021, 132, 103914. [Google Scholar] [CrossRef]
- Sebbenn, A.M.; Degen, B.; Azevedo, V.C.R.; Silva, M.B.; de Lacerda, A.E.B.; Ciampi, A.Y.; Kanashiro, M.; Carneiro, F. da S.; Thompson, I.; Loveless, M.D. Modelling the long-term impacts of selective logging on genetic diversity and demographic structure of four tropical tree species in the Amazon forest. Forest Ecology and Management 2008, 254, 335–349. [Google Scholar] [CrossRef]
- Sembada, A.A.; Faizal, A.; Sulistyawati, E. Photosynthesis efficiency as key factor in decision-making for forest design and redesign: A systematic literature review. Ecological Frontiers 2024, 44, 1128–1139. [Google Scholar] [CrossRef]
- Shaw, J.; Lickey, E.B.; Schilling, E.E.; Small, R.L. Comparison of whole chloroplast genome sequences to choose noncoding regions for phylogenetic studies in angiosperms: the tortoise and the hare III. American Journal of Botany 2007, 94, 275–288. [Google Scholar] [CrossRef] [PubMed]
- Silva-Junior, J.A.; de Souza França, D.; Moraes, R.C.S.; Gaiotto, F.A. Development of microsatellite markers for Manilkara maxima T.D. Penn. (Sapotaceae) and their use in conservation genetics. Mol Biol Rep 2016, 43, 451–455. [Google Scholar] [CrossRef] [PubMed]
- Sosa, C.C.; Arenas, C.; García-Merchán, V.H. Human Population Density Influences Genetic Diversity of Two Rattus Species Worldwide: A Macrogenetic Approach. Genes 2023, 14, 1442. [Google Scholar] [CrossRef]
- Spök, A.; Sprink, T.; Allan, A.C.; Yamaguchi, T.; Dayé, C. Towards social acceptability of genome-edited plants in industrialised countries? Emerging evidence from Europe, United States, Canada, Australia, New Zealand, and Japan. Front Genome Ed 2022, 4, 899331. [Google Scholar] [CrossRef]
- Sthapit, S.R.; Crain, J.; Larson, S.; Anderson, J.A.; Bajgain, P.; DeHaan, L.R.; Poland, J. A low-coverage skim-sequencing and imputation pipeline for genomic selection. Plant Genome 2025, 18, e70139. [Google Scholar] [CrossRef]
- Swenson, U.; Anderberg, A.A. Phylogeny, character evolution, and classification of Sapotaceae (Ericales). Cladistics 2005, 21, 101–130. [Google Scholar] [CrossRef]
- Swenson, U.; Lepschi, B.; Lowry, P.P., II; Terra-Araujo, M.H.; Santos, K.; Nylinder, S.; Alves-Araújo, A. Reassessment of generic boundaries in Neotropical Chrysophylloideae (Sapotaceae): Eleven reinstated genera and narrowed circumscriptions of Chrysophyllum and Pouteria. TAXON 2023, 72, 307–359. [Google Scholar] [CrossRef]
- Swenson, U.; Lowry, P.P., II; Cronholm, B.; Nylinder, S. Resolving the relationships of the enigmatic Sapotaceae genera Beauvisagea and Boerlagella, and the position of Planchonella suboppositifolia. TAXON 2020, 69, 998–1015. [Google Scholar] [CrossRef]
- Swenson, U.; Munzinger, J. Five new species and a systematic synopsis of Pycnandra (Sapotaceae), the largest endemic genus in New Caledonia. In Australian Systematic Botany; 2016. [Google Scholar]
- Swenson, U.; Munzinger, J. Revision of Pichonia (Sapotaceae) in New Caledonia. Australian Systematic Botany 2012. [Google Scholar] [CrossRef]
- Swenson, U.; Munzinger, J.; Bartish, I. Molecular phylogeny of Planchonella (Sapotaceae) and eight new species from New Caledonia; Taxon, 2007. [Google Scholar]
- Tautz, D. Hypervariability of simple sequences as a general source for polymorphic DNA markers. Nucleic Acids Research 1989, 17, 6463–6471. [Google Scholar] [CrossRef]
- Taylor, P.D.; Fahrig, L.; Henein, K.; Merriam, G. Connectivity Is a Vital Element of Landscape Structure. Oikos 1993, 68, 571–573. [Google Scholar] [CrossRef]
- Thompson, K.M.; Culley, T.M.; Zumberger, A.M.; Lentz, D.L. Genetic variation and structure in the neotropical tree, Manilkara zapota (L) P. Royen (Sapotaceae) used by the ancient Maya. Tree Genetics & Genomes 2015, 11, 40. [Google Scholar] [CrossRef]
- Vanijajiva, O. Start codon targeted (SCoT) polymorphism reveals genetic diversity of Manilkara in Thailand. Biodiversitas Journal of Biological Diversity 2020, 21. [Google Scholar] [CrossRef]
- Wang, H.; Bernardo, A.; St. Amand, P.; Bai, G.; Bowden, R.L.; Guttieri, M.J.; Jordan, K.W. Skim exome capture genotyping in wheat. The Plant Genome 2023, 16, e20381. [Google Scholar] [CrossRef]
- Wang, Huaying; Yang, B.; Wang, Huan; Xiao, H. Impact of different numbers of microsatellite markers on population genetic results using SLAF-seq data for Rhododendron species. Sci Rep 2021, 11, 8597. [Google Scholar] [CrossRef]
- Wang, Y.; Ding, C.; Hermoso, V.; Ji, S.; Du, T.; Chen, J.; Ding, L.; Tao, J. Using Marxan to optimize the geographical and environmental representativeness of biodiversity sampling sites. Biological Conservation 2024, 294, 110591. [Google Scholar] [CrossRef]
- Wang, B. 王博; Jia, P. 贾鹏; Gao, S. 高胜寒; Zhao, H. 赵焕焕; Zheng, G. 郑高洋; Xu, L. 许林峰; Ye, K. 叶凯. Long and Accurate: How HiFi Sequencing is Transforming Genomics. genom. proteom. bioinform. 2025, 23, qzaf003. [Google Scholar] [CrossRef]
- Waqar, Z.; Moraes, R.C.S.; Benchimol, M.; Morante-Filho, J.C.; Mariano-Neto, E.; Gaiotto, F.A. Gene Flow and Genetic Structure Reveal Reduced Diversity between Generations of a Tropical Tree, Manilkara multifida Penn., in Atlantic Forest Fragments. Genes 2021a, 12, 2025. [Google Scholar] [CrossRef]
- Waqar, Z.; Moraes, R.C.S.; Benchimol, M.; Morante-Filho, J.C.; Mariano-Neto, E.; Gaiotto, F.A. Gene Flow and Genetic Structure Reveal Reduced Diversity between Generations of a Tropical Tree, Manilkara multifida Penn., in Atlantic Forest Fragments. Genes 2021b, 12, 2025. [Google Scholar] [CrossRef]
- Wei, X.; Jiang, M. Meta-analysis of genetic representativeness of plant populations under ex situ conservation in contrast to wild source populations. Conservation Biology 2021, 35, 12–23. [Google Scholar] [CrossRef]
- WFO World Flora Online [WWW Document]. 2025. Available online: https://www.worldfloraonline.org/ (accessed on 8.12.25).
- White, G.M.; Boshier, D.H.; Powell, W. Increased pollen flow counteracts fragmentation in a tropical dry forest: An example from Swietenia humilis Zuccarini. Proceedings of the National Academy of Sciences 2002, 99, 2038–2042. [Google Scholar] [CrossRef]
- Williams, J.G.K.; Kubelik, A.R.; Livak, K.J.; Rafalski, J.A.; Tingey, S.V. DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Research 1990, 18, 6531–6535. [Google Scholar] [CrossRef]
- Xu, R.; Li, C. A Review of High-Throughput Field Phenotyping Systems: Focusing on Ground Robots. Plant Phenomics 2022. [Google Scholar] [CrossRef]
- Xu, Y.; Zhang, X.; Li, H.; Zheng, H.; Zhang, J.; Olsen, M.S.; Varshney, R.K.; Prasanna, B.M.; Qian, Q. Smart breeding driven by big data, artificial intelligence, and integrated genomic-enviromic prediction. Mol. Plant 2022, 15, 1664–1695. [Google Scholar] [CrossRef]
- Young, A.; Boyle, T.; Brown, T. The population genetic consequences of habitat fragmentation for plants. Trends in Ecology & Evolution 1996, 11, 413–418. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



