Submitted:
25 February 2026
Posted:
27 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
- Accuracy and Trustworthiness: Medical LLMs rely solely on static training data, which may become outdated as medical knowledge evolves [14], increasing the risk of incorrect or misleading outputs [15]. By retrieving content from validated clinical guidelines, databases, and scientific literature, RAG systems ground generated responses in up-to-date and trustworthy evidence, significantly reducing hallucinations [16,17]. This capability is particularly important for aligning recommendations with national and regional clinical guidelines, thereby improving compliance and reducing legal risk.
- Evidence-Based Responses: Medical LLMs typically generate recommendations without explicit source attribution, limiting transparency and clinician trust [18]. RAG systems combine generative AI with retrieved evidence, enabling responses that are explicitly grounded in and traceable to clinical sources, a critical requirement for clinical validity and acceptance in healthcare settings [19].
- Scalability and Flexibility: Training medical LLMs for each domain is resource-intensive and inefficient for cross-domain use. RAG systems instead retrieve relevant information from specialized knowledge bases, allowing efficient support across multiple medical domains and guideline sets without retraining.
- Reduced Computational Costs: Medical LLMs require heavy computational resources to generate responses, particularly as model sizes increase, leading to slower response times and higher infrastructure costs [20]. RAG systems improve efficiency by retrieving only relevant information prior to generation, reducing inference time and infrastructure requirements.
2. Related Work
2. Materials and Methods
2.1. Case Study Selection
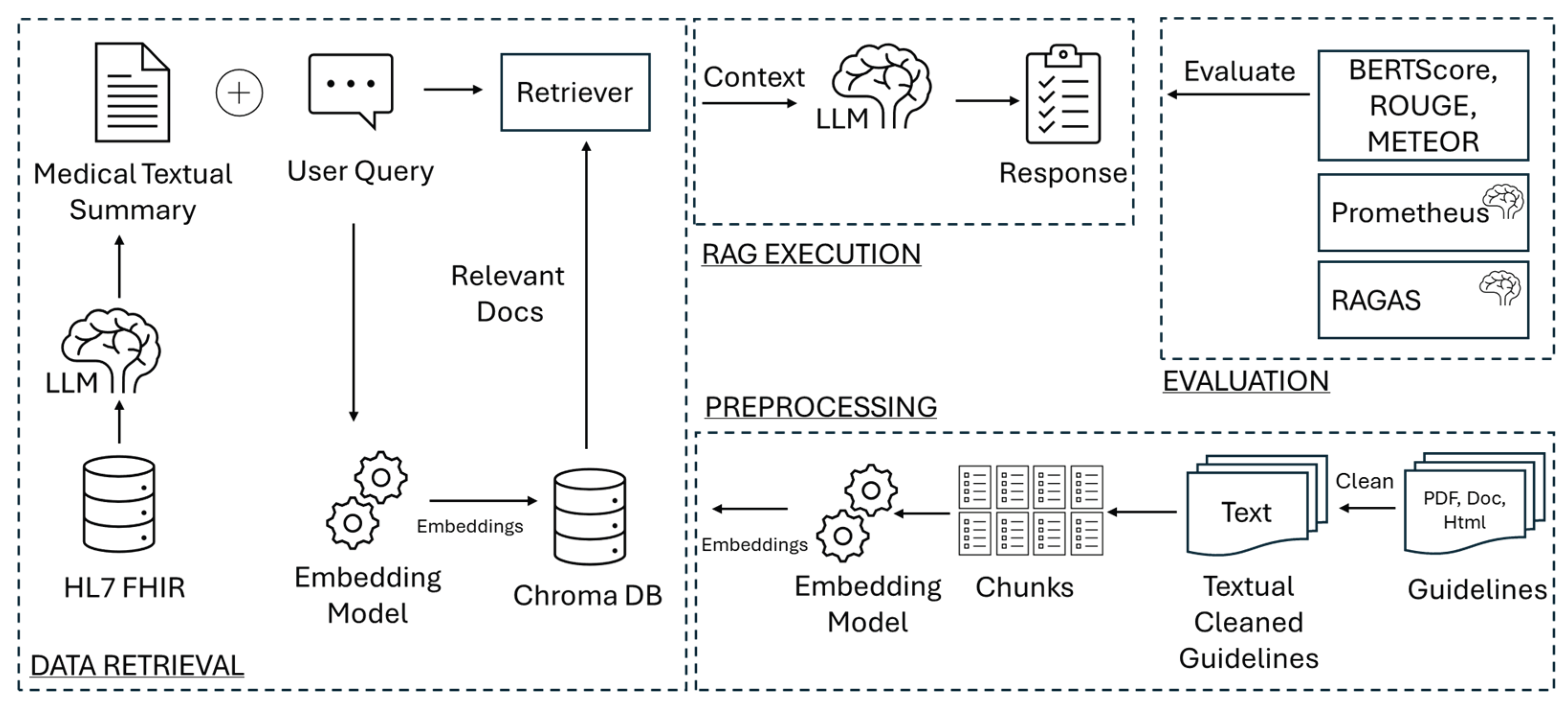
2.2. System Architecture
- Preprocessing: This step is required to prepare the guideline text to be processed by the RAG framework. It includes segmenting the text into smaller chunks and generating embeddings from these chunks to populate the vector database, which serves as an input to the RAG system.
- Data Retrieval and Query Processing: In this phase, patient context data is retrieved from the FHIR server and pre-processed to create a medical summary, which is integrated into the RAG system as input. SMART on FHIR (Substitutable Medical Applications and Reusable Technologies on Fast Healthcare Interoperability Resources) specifications are employed at this stage. SMART on FHIR is a widely adopted framework that enhances interoperability in healthcare by providing standardized, secure, and scalable solutions for accessing and exchanging clinical data [26]. Unlike proprietary or isolated solutions, SMART on FHIR fosters a robust ecosystem where applications can interact with any FHIR-compliant server, ensuring consistent access to patient data across platforms. This framework is particularly valuable for clinical decision support systems like FHIR-RAG-MEDS, where real-time access to patient-specific data is crucial for generating accurate, evidence-based recommendations. Next, the user’s plain text query (e.g., ‘What should be the approach to pharmacological treatment for Patient X?’) is merged with the generated medical summary text (e.g., ‘Patient X is an 85-year-old female patient with cognitive impairment and a history of injurious falls. She is diagnosed with hypertension’) and processed to extract embeddings, which are stored in the vector database.
- RAG Execution: This step integrates the generative capabilities of LLMs with the retrieval of specific, up-to-date information from the vector database containing pre-processed guidelines. It matches the embeddings extracted from the patient’s medical summary and the user query with the guideline embeddings in the vector storage to identify the closest matching vectors. These matching vectors, along with the medical summary, are then used by the LLM to generate a response to the clinician’s query.
2.2.1. Preprocessing
2.2.2. Data Retrieval and Integration with Existing EHR Systems
- Authentication and Authorization: The system employs OAuth 2.0 protocols, as defined by SMART on FHIR, to securely authenticate users and authorize access to patient data. This ensures that sensitive medical information remains protected while maintaining seamless usability.
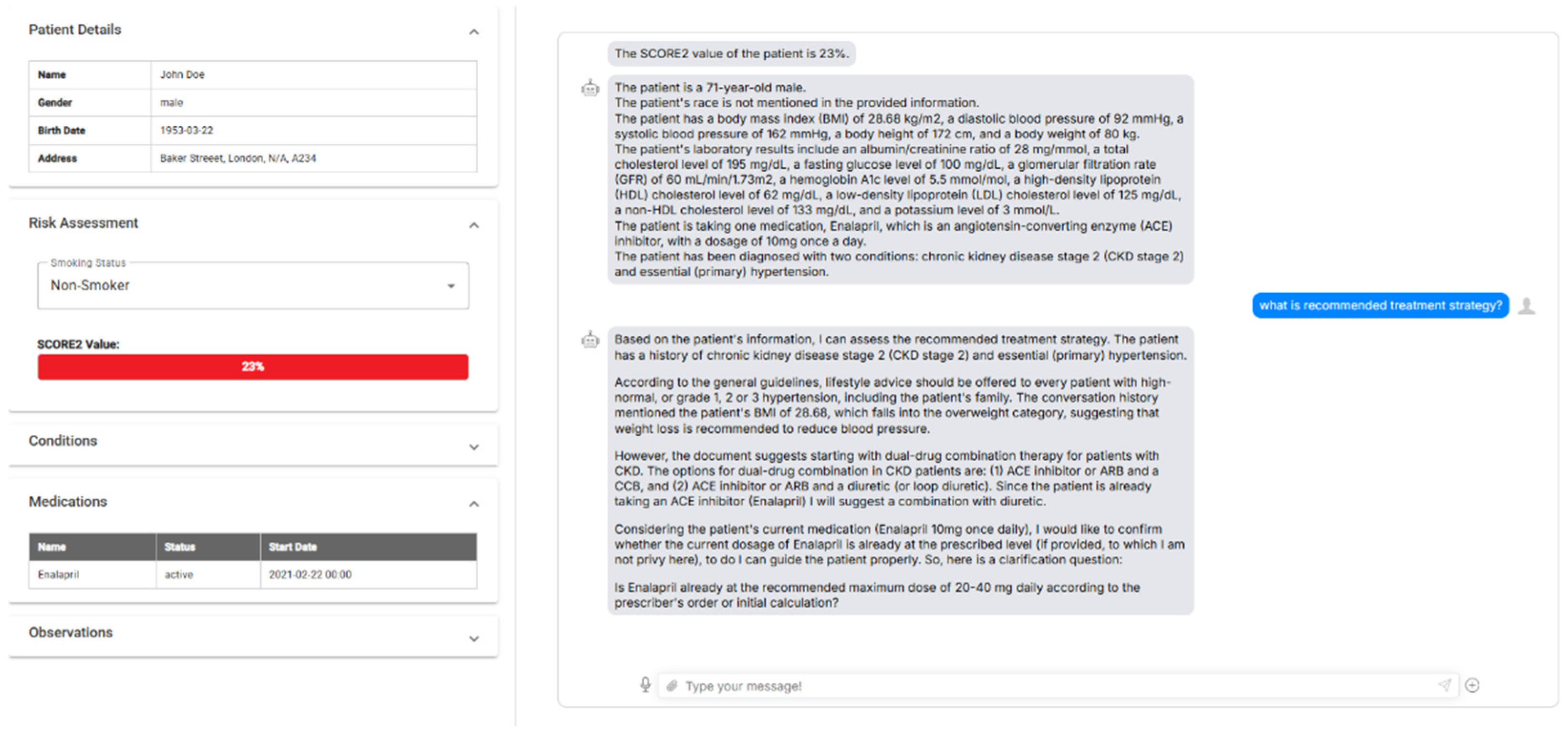
- Data Retrieval and Management: The integration enables real-time retrieval of patient information, such as demographics, medications, conditions, and observations, from any compliant FHIR server. All recent condition, medication, and observation resources are retrieved individually and then combined into a FHIR bundle for interpretation by the FHIR-RAG-MEDS system.
- Interoperable Query Processing: Once patient data is retrieved and summarized, it is sent to the FHIR-RAG-MEDS system for interpretation and recommendation generation as explained in the next section. The system’s query endpoints provide evidence-based clinical suggestions tailored to the patient’s specific medical context.
- Extensibility and Compatibility: The SMART on FHIR integration is designed to be extensible, allowing it to support additional use cases, such as integration with third-party applications, EHR systems, and telehealth platforms. By adhering to open standards, it ensures compatibility across diverse healthcare ecosystems.
2.2.3. Processing FHIR Bundles
2.2.4. RAG Execution
2.3. Evaluation Framework
- BERTScore can be the primary metric because, in healthcare, capturing the correct meaning of an answer is more important than exact word matches.
- ROUGE can be used to ensure that the model captures all necessary key terms from the guideline.
- METEOR can be a good middle-ground metric if flexibility is needed in language while still maintaining word overlap.
3. Results
3.1. Dementia Results Interpretation
- The BERTScore F1 value of 0.6372 for FHIR-RAG-MEDS demonstrates strong semantic alignment between the generated responses and the reference answers, indicating high semantic accuracy. This score clearly exceeds those of the other models, with Llama 3.1 8B achieving 0.5851, OpenBioLLM 0.5751, Meditron 0.5513, and BioMistral 0.5365. These results confirm that FHIR-RAG-MEDS produced responses that were semantically closer to the guideline-based reference answers.
- The ROUGE-L F1 score of 0.2543 achieved by FHIR-RAG-MEDS highlights its superior ability to preserve key clinical concepts and important guideline terminology. This score is substantially higher than those of the other models, which ranged from 0.1394 (Meditron) to 0.1531 (OpenBioLLM). This indicates that the proposed system more effectively incorporated relevant guideline content into its responses.
- The METEOR score of 0.3781 further demonstrates the strong performance of FHIR-RAG-MEDS in generating linguistically flexible yet accurate responses. This score significantly surpasses those of Llama 3.1 8B (0.2800), OpenBioLLM (0.2667), Meditron (0.2566), and BioMistral (0.1847). This suggests that FHIR-RAG-MEDS was more effective in maintaining the intended meaning while allowing natural variations in wording.
3.2. COPD Results Interpretation
- BERTScore F1: FHIR-RAG-MEDS achieved the highest score of 0.6311, indicating the strongest semantic alignment with the reference answers. The competing models showed slightly lower performance, with OpenBioLLM scoring 0.5809, Llama 3.1 8B 0.5780, Meditron 0.5623, and BioMistral 0.5624. These results confirm that FHIR-RAG-MEDS generated responses that were semantically closer to the guideline-based ground truth.
- ROUGE-L F1: FHIR-RAG-MEDS again achieved the best result with a score of 0.2585, demonstrating its superior ability to preserve important clinical terminology and key concepts from the COPD guideline. OpenBioLLM followed with 0.1800, while Meditron (0.1661), BioMistral (0.1587), and Llama 3.1 8B (0.1557) achieved lower scores. Although the gap is smaller than in the dementia guideline, FHIR-RAG-MEDS still clearly outperforms the other models.
- METEOR: FHIR-RAG-MEDS maintained its leading position with a score of 0.3642, reflecting its strong capability to generate responses that preserve meaning while allowing natural linguistic variation. Llama 3.1 8B achieved the second-highest score of 0.3075, followed by Meditron (0.2727), OpenBioLLM (0.2719), and BioMistral (0.2140). These results further confirm the advantage of the proposed system in producing flexible yet accurate responses.
3.3. Hypertension Results Interpretation
- BERTScore F1: FHIR-RAG-MEDS achieved the highest score of 0.6493, indicating excellent semantic alignment between the generated responses and the reference answers. This score substantially exceeds those of OpenBioLLM (0.5450), Llama 3.1 8B (0.5398), BioMistral (0.5258), and Meditron (0.5145). These results confirm that FHIR-RAG-MEDS produced responses that most closely preserved the intended clinical meaning.
- ROUGE-L F1: With a score of 0.2986, FHIR-RAG-MEDS clearly outperformed all competing models, demonstrating its superior ability to capture and retain key clinical terminology and important guideline content. The other models achieved considerably lower scores, including Meditron (0.1444), OpenBioLLM (0.1432), BioMistral (0.1367), and Llama 3.1 8B (0.1140). This large margin highlights the effectiveness of the proposed system in incorporating relevant guideline information into its responses.
- METEOR: FHIR-RAG-MEDS again achieved the best performance with a score of 0.4634, reflecting its strong capability to generate accurate responses while maintaining flexibility in language expression. The competing models achieved notably lower scores, including Llama 3.1 8B (0.2576), OpenBioLLM (0.2546), BioMistral (0.2121), and Meditron (0.1857). This further confirms the robustness of FHIR-RAG-MEDS in preserving meaning while allowing natural linguistic variation.
3.4. Sarcopenia Results Interpretation
- BERTScore F1: FHIR-RAG-MEDS achieved the highest score of 0.7367, indicating excellent semantic alignment between the generated responses and the reference answers. The closest competing models were OpenBioLLM (0.6142) and Meditron (0.6132), followed by BioMistral (0.5901) and Llama 3.1 8B (0.5538). These results demonstrate the superior capability of FHIR-RAG-MEDS in preserving the intended clinical meaning of the guideline content.
- ROUGE-L F1: With a score of 0.4648, FHIR-RAG-MEDS significantly outperformed all other models, more than doubling the performance of the closest competitor, OpenBioLLM (0.1968). The remaining models achieved lower scores, including Meditron (0.1835), BioMistral (0.1767), and Llama 3.1 8B (0.1230). This large performance gap highlights the effectiveness of FHIR-RAG-MEDS in accurately capturing and reproducing key clinical concepts and terminology from the sarcopenia guideline.
- METEOR: FHIR-RAG-MEDS achieved an outstanding score of 0.6401, demonstrating its strong ability to generate responses that maintain meaning while allowing natural variations in wording. The baseline models performed substantially worse, with OpenBioLLM scoring 0.3468, Meditron 0.3305, Llama 3.1 8B 0.2758, and BioMistral 0.2460. These results further confirm the robustness and linguistic flexibility of the proposed system.
3.5. Human Evaluation of FHIR-RAG-MEDS System
- COPD: The average physician score was 4.38 (±0.12), indicating strong agreement regarding the clinical relevance and accuracy of the responses.
- Dementia: The average score was 3.67 (±0.15), with variability reflecting the complexity of the scenarios and occasional gaps in contextual understanding.
- Hypertension: The system achieved the highest average score of 4.45 (±0.10), demonstrating both high consistency and reliability in this domain.
- Sarcopenia: The average score was 4.36 (±0.14), with qualitative feedback highlighting areas for improvement, particularly in providing actionable recommendations.
5. Discussions
5.1. Principal Findings
5.2. Limitations and Future Work
6. Conclusions
Supplementary Materials
Abbreviations
| AI | Artificial Intelligence |
| BLEU | Bilingual Evaluation Understudy |
| CDSS | Clinical Decision Support Systems |
| COPD | Chronic Obstructive Pulmonary Disease |
| CRG | Clinical Reference Group |
| EC | European Commission |
| EHR | Electronic Health Records |
| FHIR | Fast Healthcare Interoperability Resources |
| HL7 | Health Level 7 |
| LLM | Large Language Model |
| MCI | Mild Cognitive Impairment |
| MD | Mild Dementia |
| METEOR | Metric for Evaluation of Translation with Explicit Ordering |
| RAG | Retrieval-Augmented Generation |
| RAGAS | Retrieval Augmented Generation Assessment |
| RLHF | Reinforcement Learning with Human Feedback |
| ROUGE | Recall-Oriented Understudy for Gisting Evaluation |
References
- Lugtenberg, M.; Burgers, J.S.; Westert, G.P. Effects of Evidence-Based Clinical Practice Guidelines on Quality of Care: A Systematic Review. Qual. Saf. Health Care 2009, 18, 385–392. [Google Scholar] [CrossRef] [PubMed]
- Lichtner, G.; Spies, C.; Jurth, C.; Bienert, T.; Mueller, A.; Kumpf, O.; Piechotta, V.; Skoetz, N.; Nothacker, M.; Boeker, M.; et al. Automated Monitoring of Adherence to Evidenced-Based Clinical Guideline Recommendations: Design and Implementation Study. J. Med. Internet Res. 2023, 25, e41177. [Google Scholar] [CrossRef] [PubMed]
- Fischer, F.; Lange, K.; Klose, K.; Greiner, W.; Kraemer, A. Barriers and Strategies in Guideline Implementation—A Scoping Review. Healthcare 2016, 4, 36. [Google Scholar] [CrossRef]
- Riaño, D.; Peleg, M.; ten Teije, A. Ten Years of Knowledge Representation for Health Care (2009–2018): Topics, Trends, and Challenges. Artif. Intell. Med. 2019, 100, 101713. [Google Scholar] [CrossRef]
- Laleci Erturkmen, G.B.; Yuksel, M.; Sarigul, B.; Arvanitis, T.N.; Lindman, P.; Chen, R.; Zhao, L.; Sadou, E.; Bouaud, J.; Traore, L.; et al. A Collaborative Platform for Management of Chronic Diseases via Guideline-Driven Individualized Care Plans. Comput. Struct. Biotechnol. J. 2019, 17, 869–885. [Google Scholar] [CrossRef]
- García-Lorenzo, B.; Gorostiza, A.; González, N.; Larrañaga, I.; Mateo-Abad, M.; Ortega-Gil, A.; Bloemeke, J.; Groene, O.; Vergara, I.; Mar, J.; et al. Assessment of the Effectiveness, Socio-Economic Impact and Implementation of a Digital Solution for Patients with Advanced Chronic Diseases: The ADLIFE Study Protocol. Int. J. Environ. Res. Public Health 2023, 20, 3152. [Google Scholar] [CrossRef]
- Ulgu, M.M.; Laleci Erturkmen, G.B.; Yuksel, M.; Namli, T.; Postacı, Ş.; Gencturk, M.; Kabak, Y.; Sinaci, A.A.; Gonul, S.; Dogac, A.; et al. A Nationwide Chronic Disease Management Solution via Clinical Decision Support Services: Software Development and Real-Life Implementation Report. JMIR Med. Inform. 2024, 12, e49986. [Google Scholar] [CrossRef]
- Gencturk, M.; Laleci Erturkmen, G.B.; Akpinar, A.E.; Pournik, O.; Ahmad, B.; Arvanitis, T.N.; Schmidt-Barzynski, W.; Robbins, T.; Alcantud Corcoles, R.; Abizanda, P. Transforming Evidence-Based Clinical Guidelines into Implementable Clinical Decision Support Services: The CAREPATH Study for Multimorbidity Management. Front. Med. (Lausanne). 2024, 11. [Google Scholar] [CrossRef]
- Chen, Zeming, A.H.C. MEDITRON-70B: Scaling Medical Pretraining for Large Language Models. ArXiv 2023, abs/2311.16079. [Google Scholar]
- Labrak, Yanis, A.B.E.M.P.-A.G.M.R.R.D. BioMistral: A Collection of Open-Source Pretrained Large Language Models for Medical Domains. In Proceedings of the ACL 2024 - Proceedings of the 62st Annual Meeting of the Association for Computational Linguistics, 2024. [Google Scholar]
- Open Source Biomedical Large Language Model 2024.
- Lewis, Patrick, E.P.A.P.F.P.V.K.N.G.H.K.M.L.W.Y.T.R.S.R.D.K. Retrieval-Augmented Generation for Knowledge-Intensive NLP Tasks. ArXiv 2021. [Google Scholar] [CrossRef]
- Miao, J.; Thongprayoon, C.; Suppadungsuk, S.; Garcia Valencia, O.A.; Cheungpasitporn, W. Integrating Retrieval-Augmented Generation with Large Language Models in Nephrology: Advancing Practical Applications. Medicina (B Aires). 2024, 60, 445. [Google Scholar] [CrossRef]
- Tian, S.; Jin, Q.; Yeganova, L.; Lai, P.-T.; Zhu, Q.; Chen, X.; Yang, Y.; Chen, Q.; Kim, W.; Comeau, D.C.; et al. Opportunities and Challenges for ChatGPT and Large Language Models in Biomedicine and Health. Brief. Bioinform. 2023, 25. [Google Scholar] [CrossRef] [PubMed]
- Ji, Z.; Lee, N.; Frieske, R.; Yu, T.; Su, D.; Xu, Y.; Ishii, E.; Bang, Y.J.; Madotto, A.; Fung, P. Survey of Hallucination in Natural Language Generation. ACM Comput. Surv. 2023, 55, 1–38. [Google Scholar] [CrossRef]
- Gao, Yunfan, Y.X.X.G.K.J.J.P.Y.B.Y.D.J.S.M.W.H.W. Retrieval-Augmented Generation for Large Language Models: A Survey. ArXiv 2024. [Google Scholar] [CrossRef]
- Zhao, Penghao, H.Z.Q.Y.Z.W.Y.G.F.F.L.Y.W.Z.J.J.B.C. Retrieval-Augmented Generation for AI-Generated Content: A Survey. ArXiv 2024. [Google Scholar] [CrossRef]
- Wu, Junde, J.Z.Y.Q. Medical Graph RAG: Towards Safe Medical Large Language Model via Graph Retrieval-Augmented Generation. ArXiv 2024. [Google Scholar] [CrossRef]
- Zakka, C.; Shad, R.; Chaurasia, A.; Dalal, A.R.; Kim, J.L.; Moor, M.; Fong, R.; Phillips, C.; Alexander, K.; Ashley, E.; et al. Almanac — Retrieval-Augmented Language Models for Clinical Medicine. NEJM AI 2024, 1. [Google Scholar] [CrossRef]
- Ke, YuHe, L.J.K.E.H.R.A.N.L.A.T.H.S.C.R.S.J.Y.M.T.J.C.L.O.D.S.W.T. Development and Testing of Retrieval Augmented Generation in Large Language Models -- A Case Study Report. ArXiv 2024. [Google Scholar] [CrossRef]
- Al Nazi, Z.; Peng, W. Large Language Models in Healthcare and Medical Domain: A Review. Informatics 2024, 11, 57. [Google Scholar] [CrossRef]
- Li, Lingyao, J.Z.Z.G.W.H.L.F.H.Y.L.H.Y.Z.T.L.A.L.H.S.M. A Scoping Review of Using Large Language Models (LLMs) to Investigate Electronic Health Records (EHRs). ArXiv 2024. [Google Scholar] [CrossRef]
- Schmiedmayer, Paul. A.R.P.Z.V.R.A.Z.A.F.O.A. LLM on FHIR -- Demystifying Health Records. ArXiv 2024. [Google Scholar] [CrossRef]
- Li, Y.; Wang, H.; Yerebakan, H.Z.; Shinagawa, Y.; Luo, Y. FHIR-GPT Enhances Health Interoperability with Large Language Models. NEJM AI 2024, 1. [Google Scholar] [CrossRef]
- Lewis, M.; Thio, S.; Roberts, A.; Siju, C.; Mukit, W.; Kuruvilla, R.; Jiang, Z.J.; Möller-Grell, N.; Borakati, A.; Dobson, R.J.; et al. Grounding Large Language Models in Clinical Evidence: A Retrieval-Augmented Generation System for Querying UK NICE Clinical Guidelines; 2025. [Google Scholar]
- Tung, J.Y.M.; Le, Q.; Yao, J.; Huang, Y.; Lim, D.Y.Z.; Sng, G.G.R.; Lau, R.S.E.; Tan, Y.G.; Chen, K.; Tay, K.J.; et al. Performance of Retrieval-Augmented Generation Large Language Models in Guideline-Concordant Prostate-Specific Antigen Testing: Comparative Study With Junior Clinicians. J. Med. Internet Res. 2025, 27, e78393–e78393. [Google Scholar] [CrossRef] [PubMed]
- Alkhalaf, M.; Yu, P.; Yin, M.; Deng, C. Applying Generative AI with Retrieval Augmented Generation to Summarize and Extract Key Clinical Information from Electronic Health Records. J. Biomed. Inform. 2024, 156, 104662. [Google Scholar] [CrossRef]
- Kresevic, S.; Giuffrè, M.; Ajcevic, M.; Accardo, A.; Crocè, L.S.; Shung, D.L. Optimization of Hepatological Clinical Guidelines Interpretation by Large Language Models: A Retrieval Augmented Generation-Based Framework. NPJ Digit. Med. 2024, 7, 102. [Google Scholar] [CrossRef]
- Unlu, O.; Shin, J.; Mailly, C.J.; Oates, M.F.; Tucci, M.R.; Varugheese, M.; Wagholikar, K.; Wang, F.; Scirica, B.M.; Blood, A.J.; et al. Retrieval-Augmented Generation–Enabled GPT-4 for Clinical Trial Screening. NEJM AI 2024, 1. [Google Scholar] [CrossRef]
- Xiong, G.; Jin, Q.; Lu, Z.; Zhang, A. Benchmarking Retrieval-Augmented Generation for Medicine; 2024. [Google Scholar]
- FHIR-RAG-MED Interpretation System. Available online: https://github.com/srdc/fhir-rag-med-interpret (accessed on 24 February 2026).
- CAREPATH Project Website. Available online: https://cordis.europa.eu/project/id/945169 (accessed on 24 February 2026).
- Robbins, T.D.; Muthalagappan, D.; O’Connell, B.; Bhullar, J.; Hunt, L.-J.; Kyrou, I.; Arvanitis, T.N.; Keung, S.N.L.C.; Muir, H.; Pournik, O.; et al. Protocol for Creating a Single, Holistic and Digitally Implementable Consensus Clinical Guideline for Multiple Multi-Morbid Conditions. In Proceedings of the Proceedings of the 10th International Conference on Software Development and Technologies for Enhancing Accessibility and Fighting Info-exclusion, New York, NY, USA, August 31 2022; ACM; pp. 1–6. [Google Scholar]
- LangChain Open Source Library. 2026.
- Chroma Open Source AI Application Database 2026.
- Llama 3.1 8B LLM 2026.
- Lin, Chin-Yew. ROUGE: A Package for Automatic Evaluation of Summaries. In Proceedings of the In Proceedings of the Workshop on Text Summarization Branches Out (WAS 2004), 2004. [Google Scholar]
- Banerjee, Satanjeev, A.L. METEOR: An Automatic Metric for MT Evaluation with Improved Correlation with Human Judgments. In Proceedings of the Proceedings of the ACL Workshop on Intrinsic and Extrinsic Evaluation Measures for Machine Translation and/or Summarization, 2005. [Google Scholar]
- Zhang, Tianyi, V.K.F.W.K.Q.W.Y.A. BERTScore: Evaluating Text Generation with BERT. In Proceedings of the International Conference on Learning Representations, 2020. [Google Scholar]
- Papineni, K.; Roukos, S.; Ward, T.; Zhu, W.-J. BLEU. In Proceedings of the Proceedings of the 40th Annual Meeting on Association for Computational Linguistics - ACL ’02, 2001; Association for Computational Linguistics: Morristown, NJ, USA; p. 311. [Google Scholar]
- Yan, Z. Evaluating the Effectiveness of LLM-Evaluators (Aka LLM-as-Judge). Available online: https://eugeneyan.com/writing/llm-evaluators/ (accessed on 24 February 2026).
- Kim, Seungone, J.S.S.L.B.Y.L.J.S.S.W.G.N.M.L.K.L.M.S. Prometheus 2: An Open Source Language Model Specialized in Evaluating Other Language Models. ArXiv 2024. [Google Scholar] [CrossRef]
- RAGAS Evaluation Framework 2026.
- Mahadevaiah, G.; RV, P.; Bermejo, I.; Jaffray, D.; Dekker, A.; Wee, L. Artificial Intelligence-based Clinical Decision Support in Modern Medical Physics: Selection, Acceptance, Commissioning, and Quality Assurance. Med. Phys. 2020, 47. [Google Scholar] [CrossRef]
- Golden, G.; Popescu, C.; Israel, S.; Perlman, K.; Armstrong, C.; Fratila, R.; Tanguay-Sela, M.; Benrimoh, D. Applying Artificial Intelligence to Clinical Decision Support in Mental Health: What Have We Learned? Health Policy Technol. 2024, 13, 100844. [Google Scholar] [CrossRef]
- Chaudhari, S.; Aggarwal, P.; Murahari, V.; Rajpurohit, T.; Kalyan, A.; Narasimhan, K.; Deshpande, A.; Castro da Silva, B. RLHF Deciphered: A Critical Analysis of Reinforcement Learning from Human Feedback for LLMs. ACM Comput. Surv. 2026, 58, 1–37. [Google Scholar] [CrossRef]
| ###Task Description: An instruction (might include an Input inside it), a response to evaluate, a reference answer that gets a score of 5, and a score rubric representing a evaluation criteria are given. 1. Write detailed feedback that assesses the quality of the response strictly based on the given score rubric, not evaluating in general. 2. After writing a feedback, write a score that is an integer between 1 and 5. You should refer to the score rubric. 3. The output format should look as follows: “Feedback: {{write a feedback for criteria}} [RESULT] {{an integer number between 1 and 5}}” 4. Please do not generate any other opening, closing, and explanations. Be sure to include [RESULT] in your output. ###The instruction to evaluate: {instruction} ###Response to evaluate: {response} ###Reference Answer (Score 5): {reference_answer} ###Score Rubrics: [Is the response correct, accurate, and factual based on the reference answer?] Score 1: The response is completely incorrect, inaccurate, and/or not factual. Score 2: The response is mostly incorrect, inaccurate, and/or not factual. Score 3: The response is somewhat correct, accurate, and/or factual. Score 4: The response is mostly correct, accurate, and factual. Score 5: The response is completely correct, accurate, and factual. ###Feedback: |
| Type | Metric | Description |
| Retriever Metrics |
Context Precision | In simple terms how relevant is context retrieved to the question asked. |
| Context Recall | Is the retriever able to retrieve all the relevant context pertaining to ground truth? | |
| Response Generation Metrics |
Answer Relevancy | How relevant is the generated answer to the question. |
| Faithfulness | Factual consistency of generated answers with the given context. | |
| Comprehensive Metrics | Answer Correctness | Answer correctness encompasses two critical aspects: semantic similarity between the generated answer and the ground truth, as well as factual similarity. |
| Answer Similarity | The semantic resemblance between the generated answer and the ground truth. |
| LLM NAME |
PROMETHEUS 2 AVERAGE SCORE |
BERTSCORE F1 | ROUGE-L F1 | METEOR |
| BIOMISTRAL | 3.5000 | 0.5365 | 0.1418 | 0.1847 |
| LLAMA 3.1 8B | 3.1667 | 0.5851 | 0.1526 | 0.2800 |
| MEDITRON | 2.6667 | 0.5513 | 0.1394 | 0.2566 |
| OPENBIOLLM | 3.0000 | 0.5751 | 0.1531 | 0.2667 |
| FHIR-RAG-MEDS | 4.0000 | 0.6372 | 0.2543 | 0.3781 |
| METRIC | DEMENTIA | COPD | Hypertension | Sarcopenia |
| ANSWER CORRECTNESS | 0.780 | 0.882 | 0.952 | 0.878 |
| ANSWER RELEVANCY | 0.806 | 0.821 | 0.872 | 0.846 |
| ANSWER SIMILARITY | 0.942 | 0.929 | 0.908 | 0.939 |
| CONTEXT PRECISION | 0.903 | 0.962 | 0.958 | 1.000 |
| CONTEXT RECALL | 0.776 | 0.897 | 0.905 | 0.935 |
| FAITHFULNESS | 0.859 | 0.767 | 0.789 | 0.785 |
| LLM NAME | AVERAGE SCORE | BERTSCORE F1 | ROUGE-L F1 | METEOR |
| BioMistral | 3.8462 | 0.5624 | 0.1587 | 0.2140 |
| Llama 3.1 8B | 4.0769 | 0.5780 | 0.1557 | 0.3075 |
| Meditron | 3.2308 | 0.5623 | 0.1661 | 0.2727 |
| OpenBioLLM | 4.1538 | 0.5809 | 0.1800 | 0.2719 |
| FHIR-RAG-MEDS | 4.3846 | 0.6311 | 0.2585 | 0.3642 |
| LLM NAME | AVERAGE SCORE | BERTSCORE F1 | ROUGE-L F1 | METEOR |
| BioMistral | 3.5135 | 0.5258 | 0.1367 | 0.2121 |
| Llama 3.1 8B | 3.3514 | 0.5398 | 0.1140 | 0.2576 |
| Meditron | 2.9189 | 0.5145 | 0.1444 | 0.1857 |
| OpenBioLLM | 3.0811 | 0.5450 | 0.1432 | 0.2546 |
| FHIR-RAG-MEDS | 4.4474 | 0.6493 | 0.2986 | 0.4634 |
| LLM NAME | AVERAGE SCORE | BERTSCORE F1 | ROUGE-L F1 | METEOR |
| BioMistral | 3.5455 | 0.5901 | 0.1767 | 0.2460 |
| Llama 3.1 8B | 3.9091 | 0.5538 | 0.1230 | 0.2758 |
| Meditron | 3.6364 | 0.6132 | 0.1835 | 0.3305 |
| OpenBioLLM | 3.0909 | 0.6142 | 0.1968 | 0.3468 |
| FHIR-RAG-MEDS | 4.3636 | 0.7367 | 0.4648 | 0.6401 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





