1. Introduction
Lactic acid bacteria (LAB) are widely used in food fermentations and biotechnological applications, where the efficiency of biomass production and substrate utilization directly affects product yield and process optimization [
1,
2,
3].
These bacteria produce organic acids (mainly lactic acid) through the catabolism of sugars present in the substrate, leading to a decrease in culture pH [
3,
4]. Biomass accumulation critically depends on culture pH, as it influences cellular metabolism, intracellular homeostasis, enzyme activity, and nutrient uptake [
4,
5,
6,
7]. Previous studies have shown that biomass production reaches a maximum within an optimal pH range (e.g., near neutral or mildly acidic conditions) for many LAB strains. In contrast, excessively low pH values significantly reduce biomass accumulation and may even lead to cell death, as acid stress inhibits metabolism and cell division [
8,
9,
10] and can impair LAB growth more severely than nutrient depletion [
4,
11].
Acid accumulation increases the proportion of undissociated acid species, which can diffuse back into the cell, dissipate the proton motive force, and drain cellular energy as bacteria expel excess protons—ultimately inhibiting growth and reducing biomass yield [
12,
13,
14,
15]. Thus, in LAB fermentations, biomass formation is directly linked to the availability of growth-limiting nutrients, particularly nitrogen sources, and is strongly modulated by culture acidification resulting from lactic acid production [
4,
16,
17].
LAB are also capable of producing bacteriocins—antibacterial peptides often considered pH-dependent primary metabolites—which are highly sensitive to the time course of culture pH during fermentation [
18]. Therefore, changes in culture pH can modulate bacteriocin synthesis [
19,
20], likely through regulation of biosynthetic genes [
21,
22], modulation of the enzymatic activity responsible for post-translational processing of prebacteriocins into their active forms [
15,
23], and adsorption of bacteriocins to the cell wall of the producer strain [
24].
Furthermore, total sugar and nitrogen consumption are governed by biomass production, as these nutrients serve as energy sources to support growth and cellular maintenance, are incorporated into new biomass, and are utilized for metabolite production [
17,
25,
26].
Mathematical modeling of LAB fermentations is an essential tool for understanding, predicting, and optimizing microbial growth, metabolite production, and substrate utilization in complex culture systems [
25]. Among the available empirical approaches, logistic-type equations have been widely and successfully applied to describe batch biomass growth, lactic acid formation, antimicrobial metabolite production, substrate consumption, and pH evolution during LAB fermentations due to their simplicity and good fitting performance across different media and strains [
27,
28,
29,
30]. These models assume self-limiting behavior and capture the sigmoidal dynamics typically observed in batch cultures [
27,
28].
However, a major limitation of these equations is that they describe culture variables solely as functions of time, implicitly assuming that microbial growth, product formation, and nutrient consumption in batch cultures are purely autocatalytic processes. Moreover, such models do not explicitly account for the underlying biological and biochemical mechanisms driving growth, metabolite production, and nutrient utilization in LAB fermentations.
Consequently, treating kinetic parameters—such as growth, production, and consumption rates—as independent constants may limit the predictive power and physiological relevance of these models.
By linking the kinetic parameters of the logistic equations through biologically meaningful relationships—accounting for pH effects, nutrient limitation, and growth–production coupling—the proposed approach provides a more precise, comprehensive, and integrated description of LAB fermentation dynamics while retaining the mathematical simplicity of logistic models. Such models not only enhance our understanding of microbial physiology but also provide practical benefits for the design and control of fermentation processes. This approach may improve the predictive accuracy of growth models and support the optimization of culture conditions, ultimately contributing to more efficient fermentation processes and a deeper understanding of microbial physiology.
To address the limitations of logistic-type equations in describing the kinetics of LAB fermentation, the present study focuses on developing an improved modeling framework based on coupled logistic models that incorporate the relationships among biomass growth, lactic acid production, total antibacterial activity, pH evolution, and nutrient consumption (total sugars and nitrogen).
3. Results and Discussion
The efficacy of the coupled logistic models (16)–(21) in describing the time courses of six culture variables—namely biomass growth (X), product formation (LA and TAA), culture pH, and nutrient consumption (TS and TN)—was demonstrated using experimental data from three lactic acid bacteria strains (CECT 539, CECT 4043, and NRRL B–5627) cultivated in different culture media [
31,
32,
33,
34,
35,
36].
To further validate the accuracy of the proposed models in describing the six culture variables, the results obtained from the coupled models (16)–(21) were compared with those generated by the corresponding unmodified logistic models (1), (2), (3), (6), (7), and (8).
In addition, the models were validated using data from batch cultures of
Pediococcus acidilactici LB42
–923 (a pediocin AcH producer),
Lactococcus lactis subsp.
lactis ATCC 11454 (a nisin producer),
Leuconostoc carnosum Lm1 (a leuconocin Lcm1 producer), and
Lactobacillus sakei LB 706 (a sakacin A producer), grown in TGE broth at uncontrolled pH [
23].
As only experimental data on growth, bacteriocin production, and pH time courses were available in this case [
23], only models (16), (18), and (19) were validated, and the corresponding results were compared with those obtained using the unmodified models (1), (3), and (6), respectively.
Considering that lactic acid is primarily a growth-associated product, as it is predominantly synthesized by LAB strains during the active growth phase [
30,
42], model (19) was modified by substituting the lactic acid time-course data, LA(t), and its corresponding constants (
KLA and
aLA) with those of biomass growth (X(t),
KX, and
aX), as indicated below:
Where bX–pH represents the rate of pH decline as influenced by biomass growth.
3.1. Modeling Batch Growth, Product Formation, and Nutrient Consumption by Lactococcus lactis CECT 539 in Concentrated Whey and MRS Broth
3.1.1. Uncoupled Models
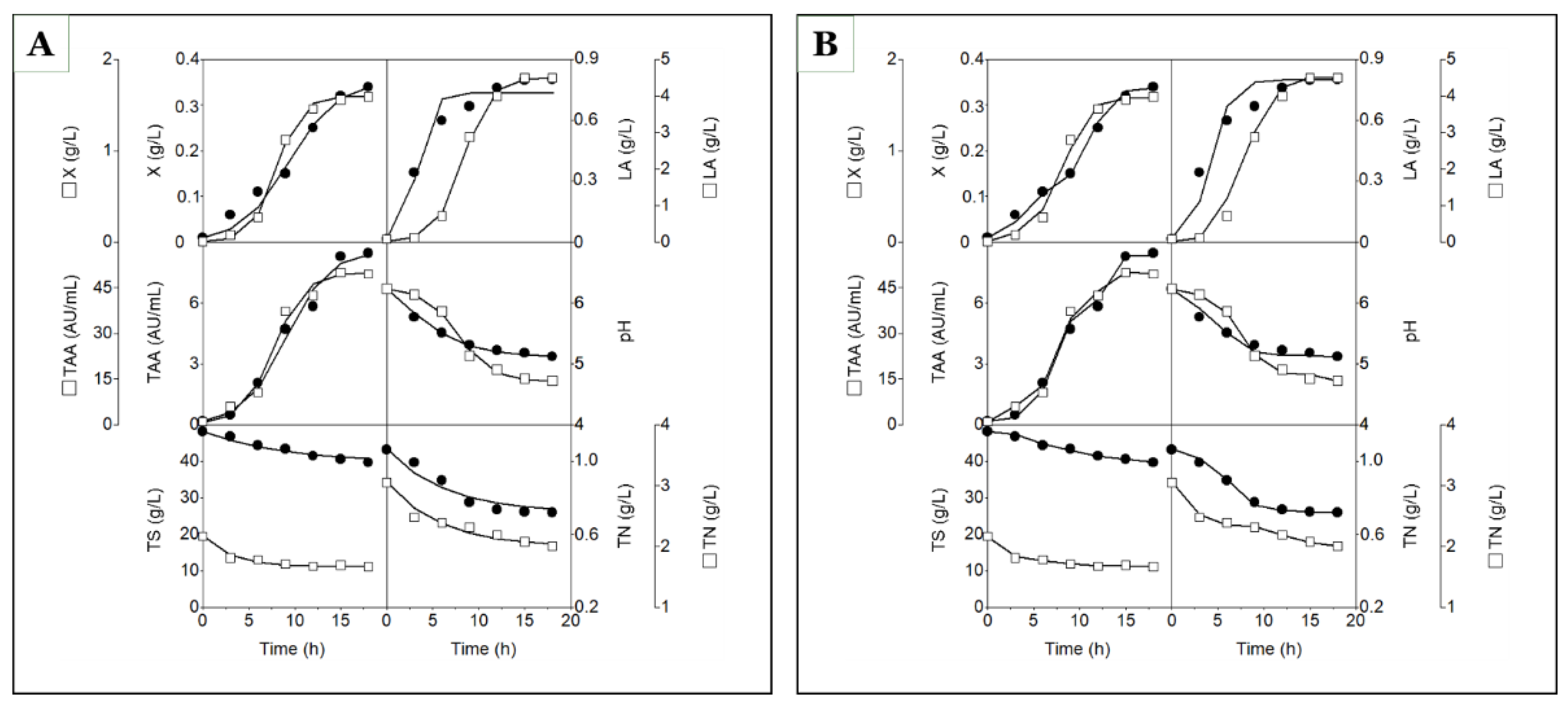
Figure 2A shows the time courses of experimental biomass growth, lactic acid production, total antibacterial activity synthesis, culture pH, and consumption of total sugars and nitrogen by strain CECT 539 in concentrated whey (CW) and MRS broth, together with the predictions of models (1), (2), (3), (6), (7), and (8).
For the CW culture (
Figure 2A and
Table 2), models (1), (3), and (6) provided an adequate description of the experimental data for biomass growth (
R2X = 0.9846,
SSD–
X = 2.29 × 10
–3), total antibacterial activity production (
R²
TAA = 0.9838,
SSD–
TAA = 1.18), and culture pH (
R²
pH = 0.9952,
SSD–
pH = 6.12 × 10
–3). However, models (2), (7), and (8) did not satisfactorily describe the time courses of lactic acid (
R²
LA = 0.9500,
SSD–
LA = 2.58 × 10
–2), total sugars (
R²
TS = 0.9638,
SSD–
TS = 3.34), and total nitrogen (
R²
TN = 0.9512,
SSD–
TN = 8.27 × 10
–3), respectively (
Figure 2A and
Table 2).
The failure of model (2) to adequately describe the kinetics of lactic acid synthesis in CW medium can be attributed to the fact that lactic acid production did not follow the typical logistic-type profile observed in MRS broth (see the upper right panel of
Figure 2A). Even when the constraint requiring that the model-predicted values at time zero (
Yp₀) match the corresponding experimental values (
Yexp₀) was removed, the
R²
LA value increased to 0.9793 and the
SSD–
LA decreased to 1.18 × 10⁻². However, the predicted initial lactic acid concentration increased to 0.082 g/L, which is considerably higher than the experimental value (0.016 g/L).
The relatively low
R2 obtained for models (7) and (8) can be explained by the fact that the time courses of total sugars (TS) and total nitrogen (TN) did not strictly follow a C-shaped profile during CW fermentation (see the lower left and lower right panels of
Figure 2A).
With regard to the culture in MRS broth, the modeling results were slightly better than those obtained in CW medium (
Figure 2A), with higher
R2 values (0.9979, 0.9981 and 0.9809) obtained for models (1), (2), and (7), respectively (
Table 2). However, the predictions of models (3), (6), and (8) for the kinetics of total antibacterial activity (TAA) synthesis, culture pH, and total nitrogen (TN) were similar in both culture media, yielding comparable
R2 values.
The differences observed in the predictions of the models (1), (2), (3), (6), (7), and (8) between fermentations conducted in the two culture media can be attributed solely to the degree of agreement between the experimental time-course data for the corresponding culture variables and the mathematical structure of each model.
3.1.2. Coupled Models
Uncoupled models assume that each variable evolves independently with its own logistic time constant. In LAB fermentations, the growth, the synthesis of lactic acid and total antibacterial activity, pH drop and consumption of nutrients (TS and TN) are not independent processes evolving only with time, because, fermentation is a closed biochemical network, not six independent time
–driven phenomena [
27,
29,
30,
31,
32]. Then, developing of coupled models could reflect this biological interdependence.
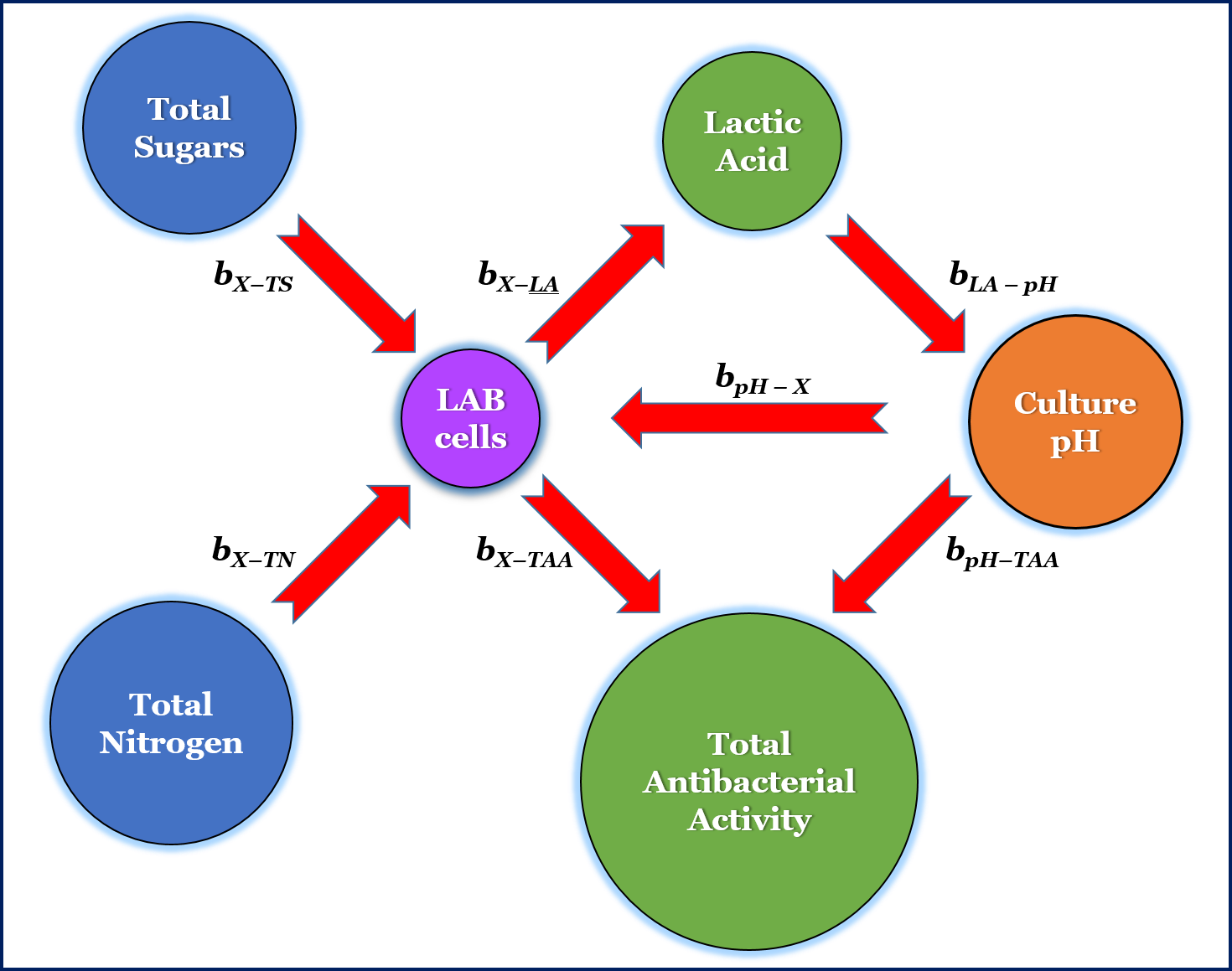
In this study, the following simple biologically linked through cause–effect relationships between the six variables were assumed: i) pH affects cell growth kinetics, ii) lactic acid production decreases pH, iii) primary metabolite production (LA) depends on growth, iv) antibacterial activity depends on both biomass and pH evolution and, v) biomass formation drives sugar and nitrogen consumption.
The coupled models (16)–(21) were fitted simultaneously to the experimental data for the six culture variables (X, LA, TAA, pH, TS, and TN), incorporating biologically meaningful cause–effect relationships among them. The statistical performance of these models in concentrated whey (CW) and MRS broth is summarized in
Table 3, and the corresponding fits are presented in
Figure 2B.
In CW medium, the coupled formulation significantly improved the overall description of the fermentation process compared with the uncoupled approach (
Table 2 and
Table 3). The coupled biomass model provided an excellent fit (
R²
X = 0.9962;
SSD–X = 6.25 × 10⁻⁴), outperforming the uncoupled model (
R²
X = 0.9846;
SSD–X = 2.29 × 10⁻³). The approximately one-order-of-magnitude reduction in
SSD–X indicates that incorporating the pH-dependent correction of the growth parameter (
bpH–X = 0.29 h⁻¹) enhanced the representation of growth dynamics. This confirms that biomass evolution in CW is strongly influenced by culture acidification.
In contrast, the coupled model did not substantially improve the description of lactic acid production (
R²
LA = 0.9492;
SSD–LA = 3.99 × 10⁻²) compared with the uncoupled model (
R²
LA = 0.9500;
SSD–LA = 2.58 × 10⁻²). The lower goodness-of-fit reflects the deviation of lactic acid production in CW from a strictly logistic profile (
Figure 2A and 2B). Although coupling with biomass (
bX–LA = 0.51 h⁻¹) ensures biological consistency, it does not fully capture the irregular kinetic pattern observed experimentally. This suggests that additional metabolic constraints or substrate limitations may influence acid synthesis in CW.
A marked improvement was observed for TAA production (R²TAA = 0.9943; SSD–TAA = 0.47) compared with the uncoupled model (R²TAA = 0.9838; SSD–TAA = 1.18). The simultaneous dependence on biomass (bX–TAA = 0.23 h⁻¹) and pH (bpH–TAA = 0.12 h⁻¹) enhanced the predictive capacity of the model. The lower SSD indicates that antibacterial activity is better described when both growth and acidification effects are considered, supporting its classification as a growth-associated metabolite influenced by environmental pH.
Although the fit remained acceptable (
R²
pH = 0.9750), the coupled pH model showed a lower
R² and higher
SSD–pH (4.13 × 10⁻²) than the uncoupled formulation (
R²
pH = 0.9952;
SSD–pH = 6.12 × 10⁻³). This slight deterioration is likely due to the imposed dependence of pH on lactic acid production (
bLA–pH = 0.83 h⁻¹), which did not follow a strictly logistic time profile (
Figure 2A and 2B). Nevertheless, the coupled model ensures mechanistic coherence between acid production and pH decline.
A substantial improvement was observed for sugar consumption. The R²TS increased from 0.9638 to 0.9926, while SSD–TS decreased from 3.34 to 0.56. By directly linking total sugar (TS) consumption to biomass formation (bX–TS = 0.40 h⁻¹), the coupled model more accurately represented substrate depletion, confirming that sugar utilization is primarily growth-driven rather than purely time-dependent.
An even more pronounced improvement was observed for nitrogen consumption. The R²TN increased from 0.9512 to 0.9954, and SSD–TN decreased by nearly one order of magnitude (from 8.27 × 10⁻³ to 8.88 × 10⁻⁴). This indicates that total nitrogen (TN) depletion is strongly correlated with biomass production (bX–TN = 0.44 h⁻¹), and that the coupling strategy effectively captures this relationship.
Overall, in CW medium, the coupled models improved the description of biomass, TAA, TS, and TN, maintained a similar fit for lactic acid, and slightly reduced the statistical performance for pH. Nevertheless, the coupled system provides a more coherent and biologically consistent representation of the fermentation process (
Table 2 and
Table 3).
In MRS broth, where the uncoupled models already provided excellent fits, the improvements introduced by coupling were less pronounced, although still relevant.
For biomass, R²X decreased slightly from 0.9979 to 0.9938, and the SSD–X increased modestly. Nevertheless, the fit remained excellent, and the growth–pH interaction (bpH–X = 0.51 h⁻¹) confirms that acidification influences growth even in a nutritionally rich medium.
For lactic acid (LA) synthesis, the coupled model slightly reduced
R²
LA (0.9915 vs. 0.9981), although the fit remained very good. Because LA production in MRS follows a well-defined logistic profile, the uncoupled model was already sufficient (
Figure 2A); thus, coupling mainly improved mechanistic interpretation rather than statistical performance.
In the case of the TAA model, a slight improvement was observed (
R²
TAA = 0.9958 vs. 0.9856), accompanied by a marked reduction in
SSD–TAA (0.12 vs. 42.07). The dual dependence on biomass and pH significantly enhanced model accuracy, highlighting the synergistic regulation of antibacterial compound synthesis (
Figure 2A and 2B).
The coupled pH model slightly improved the fit (
R²
pH = 0.9969 vs. 0.9949; lower
SSD–pH), indicating that linking pH decline to LA production (
bLA–pH = 0.68 h⁻¹) is appropriate in MRS broth (
Table 2 and
Table 3).
For total sugars (TS) and total nitrogen (TN), the coupled models showed clear improvements. For TS, R²TS increased from 0.9809 to 0.9966, while SSD–TS decreased from 1.28 to 0.18. For TN, R²TN increased from 0.9563 to 0.9957; with a substantial reduction in SSD–TN. These results further confirm that nutrient depletion is tightly coupled to biomass formation and that the coupled models provide a more accurate description of substrate utilization kinetics.
Overall, across both media, the coupled models improved or maintained high R² values for most variables and significantly reduced SSD for biomass, TAA, TS, and TN. However, they slightly reduced statistical flexibility for LA (in CW) and pH (in CW) due to the imposed biological constraint (initial experimental LA value = predicted LA value).
The most notable improvements were observed for nutrient consumption and antibacterial activity, variables that strongly depend on biomass and environmental conditions.
To further demonstrate the effectiveness of the coupled models, kinetic data from fermentations conducted in media containing different initial concentrations of total nitrogen (TN) or total sugars (TS) were used for model fitting. Under these conditions, the time-course profiles of TS and TN differed substantially [
31,
32,
33,
34,
35,
36], allowing a more rigorous evaluation of the ability of the coupled models to describe the kinetics of the six culture variables.
Accordingly, both the uncoupled and coupled models were applied to describe the evolution of the six culture variables in DW and MPW media supplemented with different concentrations of glutamic acid, as well as in DW medium supplemented with varying concentrations of glucose.
3.2. Modeling of Batch Growth, Product Formation, Culture pH Drop, and Nutrient Consumption by Strain CECT 539 in DW and MPW Media Supplemented with Different Concentrations of Glutamic Acid
3.2.1. Uncoupled Models
The results of fitting the uncoupled models (1), (2), (3), (6), (7), and (8) to the fermentation kinetics of
L. lactis CECT 539 in both unsupplemented and glutamic acid–supplemented DW and MPW media are presented in
Figure 3A and 3B and
Table 4 and
Table 5, respectively.
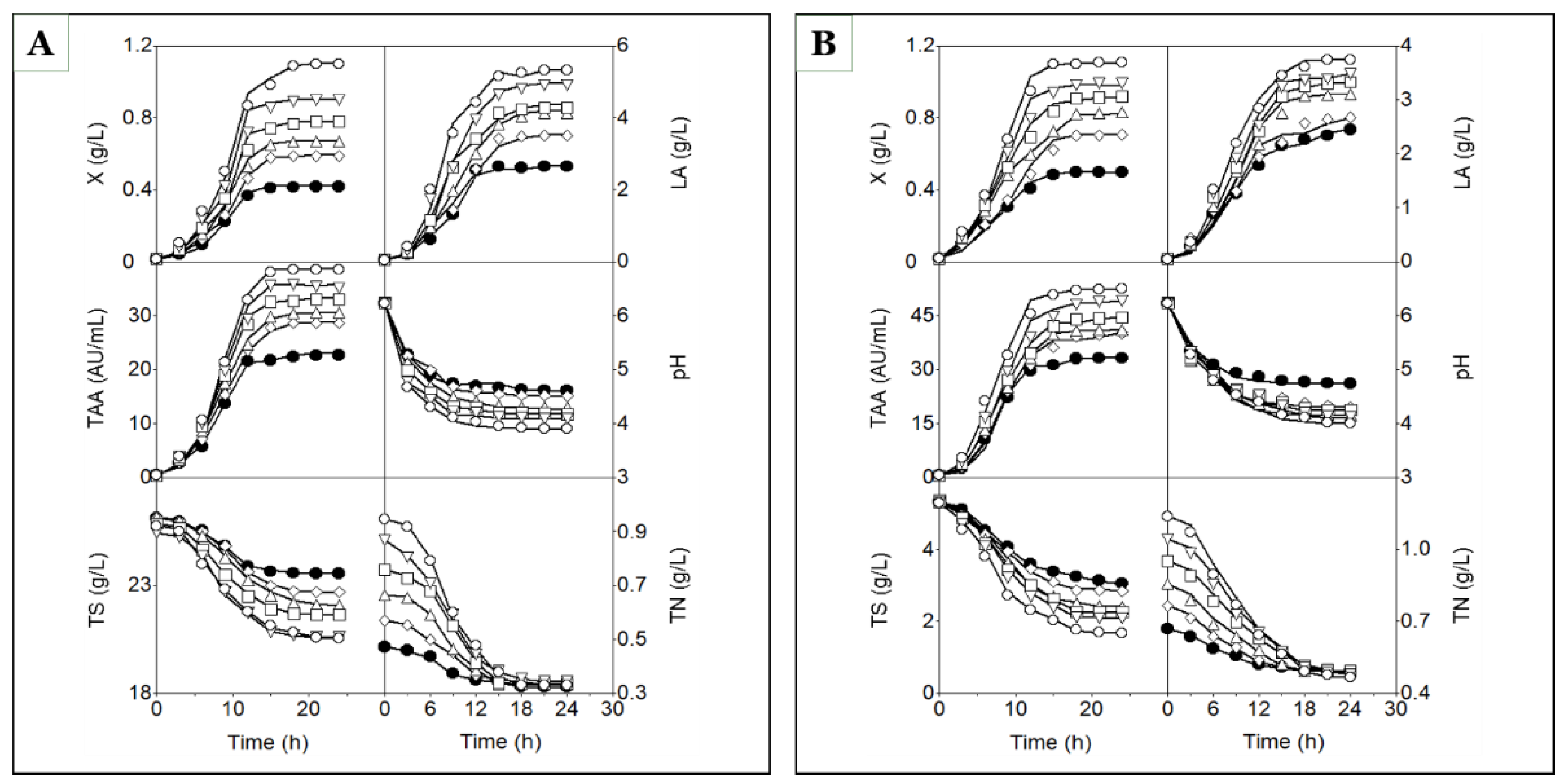
In both media, biomass growth and the production of lactic acid and total antibacterial activity followed well-defined sigmoidal (S-shaped) logistic profiles, whereas culture pH exhibited a C-shaped profile (
Figure 3A and 3B). The good agreement between the experimental time courses and the model predictions for these four variables (X, LA, TAA, and pH) supported the fitting of models (1), (2), (3), and (6), respectively (
Table 4 and
Table 5).
However, supplementation of DW and MPW media with glutamic acid resulted in more pronounced inverted S-shaped profiles for total sugars (TS) and total nitrogen (TN) compared with the corresponding unsupplemented media (
Figure 3A and 3B). This behavior affected the fitting of the uncoupled TS and TN models in DW cultures, which showed the poorest fits, with
R² values ranging from 0.9066 to 0.9545 for TS and from 0.9041 to 0.9479 for TN (
Table 4).
In MPW cultures, the R
2 values obtained for the TS model (0.9614–0.9765) and the TN model (0.9645–0.9846) (
Table 5) were higher than those observed in DW media (
Table 4). This difference was likely due to the wider variation ranges of TS and TN in DW, DW–GA1, DW–GA2, DW–GA3, DW–GA4, and DW–GA5 media (
Figure 3A) compared with the corresponding MPW media (
Figure 3B).
In DW, DW–GA1, DW–GA2, DW–GA3, DW–GA4, and DW–GA5 media, TS concentrations ranged from 26.10 to 23.56, 26.16 to 22.70, 26.19 to 22.10, 25.87 to 21.64, 25.44 to 20.60, and 25.76 to 20.55 g/L, respectively. The corresponding TN concentrations ranged from 0.473 to 0.323, 0.570 to 0.334, 0.664 to 0.332, 0.758 to 0.335, 0.867 to 0.346, and 0.945 to 0.332 g/L (
Figure 3A). In contrast, in MPW media, TS concentrations ranged from 5.32 to 3.05, 5.32 to 2.84, 5.32 to 2.41, 5.36 to 2.26, 5.31 to 2.09, and 5.31 to 1.68 g/L, respectively, while TN concentrations ranged from 0.668 to 0.492, 0.765 to 0.492, 0.857 to 0.490, 0.951 to 0.493, 1.043 to 0.481, and 1.137 to 0.468 g/L (
Figure 3B).
3.2.2. Coupled Models
The results of fitting the coupled models (16)
–(21) to the fermentation kinetics of
L. lactis CECT 539 in both unsupplemented and glutamic acid–supplemented DW and MPW media are presented in
Figure 4A and 4B and
Table 6 and
Table 7, respectively.
A comparison between the uncoupled (
Table 4 and
Table 5) and coupled models (
Table 6 and
Table 7) clearly shows that coupling the models substantially improved the overall goodness of fit, particularly for substrate (TS) and nitrogen (TN) consumption, while maintaining similar predictive performance for biomass (X), lactic acid (LA), total antibacterial activity (TAA), and pH.
For biomass in DW media,
R²
X values (0.9945–0.9985) were above 0.9900 for all cultures with the uncoupled model (
Table 4). When the coupled model was applied (
Table 6), slight decreases in
R² were observed specifically in DW–GA2 (from 0.9985 to 0.9955), DW–GA3 (from 0.9966 to 0.9899), and DW–GA4 (from 0.9966 to 0.9872). In DW and DW–GA5, the differences were minimal (DW increased from 0.9945 to 0.9984, whereas DW–GA5 changed slightly from 0.9945 to 0.9941) (
Table 4 and
Table 6).
In MPW media,
R² values for X were generally comparable between approaches (
Table 5 and
Table 7). A clear improvement with the coupled model was observed in MPW–GA2 (from 0.9797 to 0.9968). In the remaining MPW cultures, differences were small: slight decreases occurred in MPW (0.9900 to 0.9920, actually a small improvement), MPW–GA1 (0.9925 to 0.9919), MPW–GA3 (0.9861 to 0.9913, improvement), MPW–GA4 (0.9892 to 0.9886), and MPW–GA5 (0.9938 to 0.9940, slight improvement). Overall, the changes were minor and confirm that biomass was already well described by the uncoupled logistic model.
For LA and TAA, the coupled models produced similar or slightly higher
R² values compared with the uncoupled models, especially in DW cultures (
Table 4,
Table 5,
Table 6 and
Table 7). The improvement was more noticeable for TAA in DW medium, where
R²
TAA increased to values up to 0.9991 (
Table 4 and
Table 6), indicating that linking TAA production to biomass growth and pH enhanced the model’s descriptive capacity.
The most remarkable improvement was observed for TS and TN. In DW cultures, the coupled models increased
R² dramatically to values between 0.9982–0.9993 for TS and 0.9973–1.0000 for TN (
Table 4 and
Table 6). A similar trend was observed in MPW media, where
R² values for TS increased from 0.9614–0.9765 (uncoupled) to 0.9938–0.9981 (coupled), and for TN from 0.9645–0.9846 to approximately 0.9942–0.9968 (
Table 5 and
Table 7). These results indicate that substrate and nitrogen consumption were more accurately described when directly linked to biomass formation (
Figure 4A and 4B).
The
SSD values confirm the trends observed for
R². For X, LA, TAA, and pH,
SSD values were of the same order of magnitude in both approaches, with slight improvements in some cases under the coupled formulation (
Table 4 and
Table 6).
However, for TS and TN in DW medium, the reduction in
SSD was substantial. For example,
SSD–TS decreased from values between 1.38–4.12 (uncoupled) to 1.86×10⁻²–7.63×10⁻² (coupled), representing a reduction of approximately two orders of magnitude. An even more pronounced decrease was observed for TN, where
SSD values dropped from 10⁻³–10⁻² to 10⁻⁴–10⁻⁵ (
Table 4 and
Table 6).
Similar, though less dramatic, reductions were observed in MPW cultures (
Table 5 and
Table 7). This marked decrease demonstrates the superior predictive accuracy of the coupled models for nutrient consumption.
The estimated constants for biomass (KX and aX) followed similar increasing trends with glutamic acid concentration in both modeling strategies, indicating that coupling did not distort the biological interpretation of growth stimulation by glutamic acid. Likewise, KLA and KTAA increased consistently with supplementation in both approaches, confirming the positive effect of glutamic acid on metabolite production.
In the coupled models, the relatively stable values of the coupling coefficients (bX–LA, bX–TAA, bLA–pH, bX–TS), which link product formation, pH decrease, and nutrient consumption to biomass growth and/or lactic acid production, across different media and supplementation levels suggest consistent physiological relationships among growth, acidification, and substrate utilization. In particular, the strong coupling between biomass and TS/TN consumption (bX–TS and bX–TN) explains the substantial improvement in the fit obtained for these variables.
These results emphasize that the coupled modeling approach provides a more robust and integrated framework for describing the kinetics of growth, product formation, and nutrient consumption in batch cultures of L. lactis CECT 539, particularly when interactions among variables play a significant role.
3.3. Comparison between Uncoupled and Coupled Models in Fermentations of Glucose-Supplemented DW Media with Strain CECT 539
In the previous set of cultures, the fermentation media were supplemented with different concentrations of glutamic acid to generate distinct TN time-course profiles (
Figure 3 and
Figure 4) and to evaluate the ability of the coupled models to describe these variations.
In the present analysis, the focus was on modeling the time courses of the six culture variables in glucose-supplemented DW media (
Figure 5A and 5B), where different TS consumption profiles were observed [
36]. To assess the performance of the coupled models under these conditions, a comparison between the fitting results obtained with the uncoupled (
Table 8) and coupled models (
Table 9) was carried out.
The comparison reveals clear differences in model performance, particularly for substrate (TS) and nitrogen (TN) consumption (
Figure 5A and 5B).
For biomass, both approaches provided very good fits (
R² generally > 0.9700) (
Table 8 and
Table 9). However, in contrast to the glutamic acid experiments, the uncoupled model performed slightly better in some cultures. For example, in DW,
R²
X decreased from 0.9984 (uncoupled) to 0.9824 (coupled), and
SSD–X increased from 8.10×10⁻⁴ to 1.27×10⁻². A similar trend was observed in DW–G20.
In the case of lactic acid production, both approaches yielded high
R² values (>0.9800 in most cases). The coupled model improved the fit in several cultures, notably in DW (
R²
LA increased from 0.9926 to 0.9970, with
SSD–LA reduced by half) and in DW–G20 (
R²
LA increased from 0.9821 to 0.9972, with
SSD–LA decreasing from 4.19×10⁻² to 5.27×10⁻³). In other cases (e.g., DW–G5), the uncoupled model showed slightly lower
SSD–LA values. Overall, coupling LA production to biomass formation (
bX–LA) improved consistency to some extent but did not substantially alter the predictive capacity, as the uncoupled model already provided an adequate fit (
Table 8 and
Table 9).
For TAA, both models performed well (
R² ≈ 0.9900). The coupled model improved
R²
TAA values in most glucose-supplemented cultures (e.g., DW–G15: 0.9931 to 0.9964; DW–G20: 0.9868 to 0.9983), although
SSD–
TAA values were not always lower (e.g., DW–G5). The inclusion of coupling terms (
bX–TAA and
bpH–TAA) enabled a more mechanistic representation of TAA formation, particularly at higher glucose concentrations, where growth and acidification effects are more pronounced (
Figure 5).
For pH, both approaches showed good agreement with the experimental data (R²pH between 0.9600 and 0.9900). Differences between models were relatively small. In some cases (DW–G10 and DW–G15), the coupled model slightly improved R²pH and reduced SSD–pH, whereas in others (DW and DW–G20) the uncoupled model performed marginally better. Thus, coupling pH decrease to lactic acid production (bLA–pH) provided a biologically consistent formulation but did not systematically enhance the statistical indicators.
The most significant differences were observed for TS. In the uncoupled model, R²TS values were highly variable and, in some cases, low, particularly in DW (R²TS = 0.8194, SSD–TS = 36.16). Even in glucose-supplemented media, R²TS values ranged from 0.9425 to 0.9758. In contrast, the coupled model dramatically improved the fit for all cultures, with R²TS values between 0.9972 and 0.9995 and a substantial reduction in SSD–TS (e.g., from 36.16 to 9.46×10⁻² in DW). This improvement confirms that TS consumption is strongly growth-associated and is more accurately described when directly linked to biomass formation (bX–TS).
A similar pattern was observed for TN. The uncoupled model yielded moderate
R²
TN values (0.8766–0.9209), indicating limited descriptive capacity (
Table 8). The coupled model increased
R²
TN to 0.9955–0.9989 and reduced
SSD–TN by approximately one to two orders of magnitude (e.g., from 6.21×10⁻³ to 1.87×10⁻⁴ in DW). This marked improvement demonstrates that nitrogen consumption is closely associated with biomass growth and cannot be adequately represented as an independent process.
Overall, in glucose-supplemented DW cultures, the uncoupled models were generally sufficient to describe biomass growth, lactic acid production, TAA formation, and pH evolution. However, they were less accurate in capturing the dynamics of substrate (TS) and nitrogen (TN) consumption, particularly under conditions where glucose supplementation altered TS profiles.
The coupled models provided a more integrated and physiologically consistent description of the fermentation process. Although improvements for X, LA, TAA, and pH were moderate and not systematic, the enhancement in the fitting of TS and TN was substantial, as reflected by the marked increase in R² values and the pronounced reduction in SSD.
Therefore, as observed in the glutamic acid experiments, the coupled modeling approach is particularly advantageous for describing nutrient consumption kinetics when strong interactions exist among growth, substrate utilization, and metabolite production.
3.4. Performance of Uncoupled and Coupled Models in Describing Fermentations in CW, DW, and MRS Media with Lactobacillus casei CECT 4043 and Pediococcus acidilactici NRRL B–5627
Experimental data from batch cultures of
Lact. casei CECT 4043 [
34,
35] and
Ped. acidilactici NRRL B–5627 [
31,
32] grown in concentrated whey (CW), diluted whey (DW), and MRS broth were used to further evaluate the ability of the coupled models to describe the fermentation kinetics of additional LAB strains.
The time courses for strain CECT 4043 are presented in
Figure 6A and 6B, while the estimated parameters and the corresponding
R² and
SSD values are summarized in
Table 10 (uncoupled models) and
Table 11 (coupled models).
For strain CECT 4043, the coupled models consistently yielded higher R² values and lower SSD values than the uncoupled models for the six culture variables (X, LA, TAA, pH, TS, and TN). Although the uncoupled biomass model already provided high R²X values (0.9676–0.9909), the coupled model further improved the fit in CW and MRS media (R²X up to 0.9961), with a marked reduction in SSD–X (e.g., from 1.05 × 10⁻³ to 2.20 × 10⁻⁴ in CW).
For lactic acid production, both approaches performed well. However, the coupled models slightly increased R²LA in CW and DW and substantially reduced SSD–LA in DW. In MRS medium, although R²LA remained high (>0.9900), SSD–LA increased in the coupled model, suggesting that under this specific condition the additional coupling did not significantly improve the dispersion of residuals despite maintaining a strong overall correlation.
In the case of total antibacterial activity (TAA), improvement with the coupled formulation was particularly evident in CW and MRS media, where R²TAA increased to ≥0.9972 and SSD–TAA decreased considerably (e.g., from 1.58 to 0.44 in MRS).
Regarding culture pH, both models described pH evolution accurately (R²pH ≈ 0.9900). Differences in SSD–pH were small, indicating that pH dynamics were already well captured by the uncoupled formulation, although slight improvements were observed in CW and MRS media with the coupled approach.
The culture variables TS and TN showed the most pronounced improvement when modeled using the coupled models (
Figure 6A and 6B;
Table 10 and
Table 11). In MRS medium, the
R²
TS value increased dramatically (from 0.8823 to 0.9973), and
SSD–TS decreased from 29.48 to 1.32. Similarly, TN fitting improved across all media, with
R²
TN values approaching or exceeding 0.9900 and substantial reductions in
SSD–TN.
A similar trend was observed for
Ped. acidilactici NRRL B–5627 in the three culture media (
Figure 7A and 7B;
Table 10 and
Table 11). For biomass production,
R²
X values were already very high for both modeling approaches (>0.9800); however, the coupled models generally reduced
SSD–X, indicating improved precision even when correlation was strong.
For lactic acid production, the coupled approach significantly improved the fit in CW and MRS media (R²LA up to 0.9944 and 0.9936, respectively), accompanied by marked reductions in SSD–LA (e.g., from 1.25 × 10⁻² to 9.51 × 10⁻⁴ in CW).
Substantial improvements were also observed for TAA synthesis in CW and MRS media, with R²TAA values increasing to ≥0.9960 and SSD–TAA decreasing considerably (e.g., from 4417.129 to 1182.13 in MRS). Although SSD–TAA values remained relatively high due to the magnitude of TAA measurements, the relative reduction reflects improved predictive accuracy.
Both pH models provided excellent fits (
R² ≥ 0.9600), and improvements with coupling were moderate, reflecting the already robust performance of the uncoupled models (
Table 10 and
Table 11).
Modeling of TS showed one of the most striking improvements in MRS medium, where
R²
TS increased from 0.6789 to 0.9867 and
SSD–TS decreased from 23.89 to 0.80. Similar improvements were observed in CW and DW (
Table 10 and
Table 11). In addition, the coupled models substantially improved TN prediction, with
R²
TN values reaching up to 0.9997 in CW and SSD
–TN decreasing by several orders of magnitude.
Overall, while the uncoupled models already described biomass growth and pH evolution reasonably well, coupling the kinetic expressions to biologically meaningful interactions (e.g., X–LA, X–TS, X–TN, and LA–pH relationships) significantly enhanced the mechanistic consistency and predictive robustness of the models.
Therefore, the coupled modeling strategy provides a more reliable and biologically realistic framework for describing LAB fermentations in CW, DW, and MRS media.
3.5. Performance of Uncoupled and Coupled Models in Describing Fermentations in TGE Broth with Pediococcus acidilactici LB42–923, Lactococcus lactis subsp. lactis ATCC 11454, Leuconostoc carnosum Lm1, and Lactobacillus sakei LB 706
The performance of the uncoupled (models 1, 3, and 6) and coupled (models 16, 18, and 22) formulations was evaluated using batch fermentation data in TGE broth for strains LB42–923, ATCC 11454, Lm1, and LB 706 (
Table 12 and
Table 13;
Figure 8). In this case, the comparison was based on the coefficients of determination (
R²) and the sums of squared differences (
SSD) for biomass (X), bacteriocin production (B or TAA), and pH.
For all four strains, the uncoupled models already provided very good fits, with
R²
X values ranging from 0.9786 to 0.9939 (
Table 12). However, the coupled models systematically improved biomass prediction. Specifically,
R²
X increased from 0.9920 to 0.9964 for strain LB42–923, from 0.9786 to 0.9911 for strain ATCC 11454, from 0.9845 to 0.9975 for strain Lm1, and from 0.9939 to 0.9996 for strain LB 706. Correspondingly,
SSD–
X values decreased from 0.16 to 8.36 × 10⁻² (LB42–923), from 3.25 × 10⁻² to 1.25 × 10⁻² (ATCC 11454), from 6.56 × 10⁻² to 1.23 × 10⁻² (Lm1), and markedly from 3.28 × 10⁻² to 2.77 × 10⁻³ (LB 706).
Greater differences between the modeling approaches were observed for bacteriocin production. For strain LB42–923, R²TAA increased from 0.9774 to 0.9894, while SSD–TAA decreased dramatically from 34.35 to 0.18. For strain ATCC 11454, R²TAA increased from 0.9886 to 0.9999, with SSD–TAA decreasing from 4.89 to 4.59 × 10⁻², indicating an almost perfect fit with the coupled model. In the case of strain Lm1, R²TAA improved from 0.9902 to 0.9974, and SSD–TAA decreased from 13.21 to 3.55. However, for strain LB 706, R²TAA decreased slightly (from 0.9948 to 0.9821) and SSD–TAA increased (from 4.86 to 12.62) with the coupled formulation. This suggests that, for this strain, bacteriocin production may not be strongly influenced by the additional coupling terms, or that the uncoupled model already described the kinetics adequately.
The pH profiles were described with high accuracy by the uncoupled models, with
R²
pH values ranging from 0.9945 to 0.9985 (
Table 12). The coupled models produced only minor changes: slight increases in
R²
pH for strains LB42–923 and ATCC 11454, and small decreases for strains Lm1 and LB 706 (table 13). Similarly,
SSD–pH values exhibited only modest variations, without a consistent trend toward improvement.
These results indicate that pH dynamics were adequately captured by both modeling approaches and that explicitly coupling pH to biomass did not substantially enhance predictive performance in most cases.
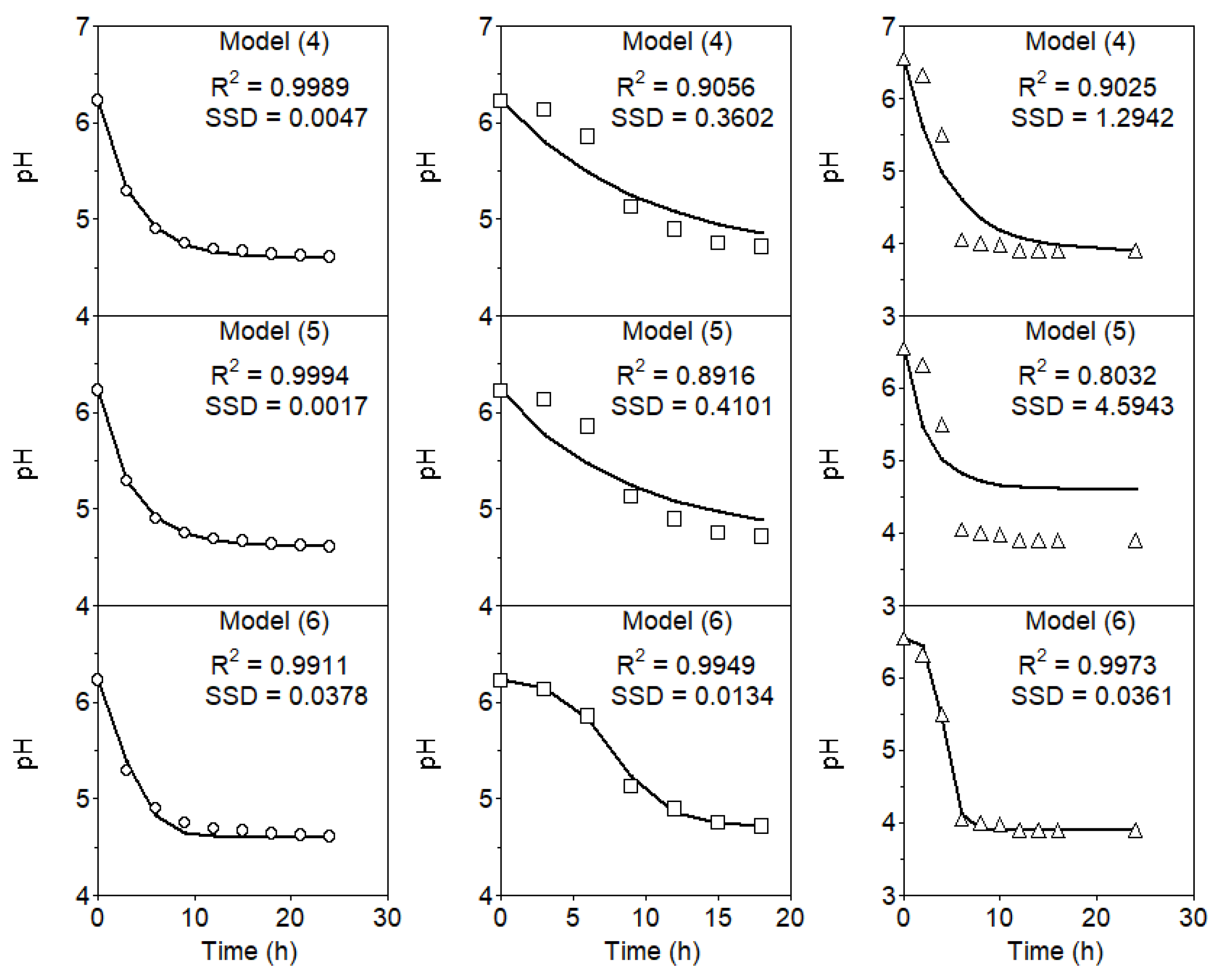
Figure 1.
Time courses of pH in cultures of
L. lactis CECT 539 grown in diluted whey (open circles) and MRS broth (open squares), and
L. lactis subsp.
lactis ATCC 11454 grown in TGE broth (open triangles) [
23]. SSD: sum of squared differences.
Figure 1.
Time courses of pH in cultures of
L. lactis CECT 539 grown in diluted whey (open circles) and MRS broth (open squares), and
L. lactis subsp.
lactis ATCC 11454 grown in TGE broth (open triangles) [
23]. SSD: sum of squared differences.
Figure 2.
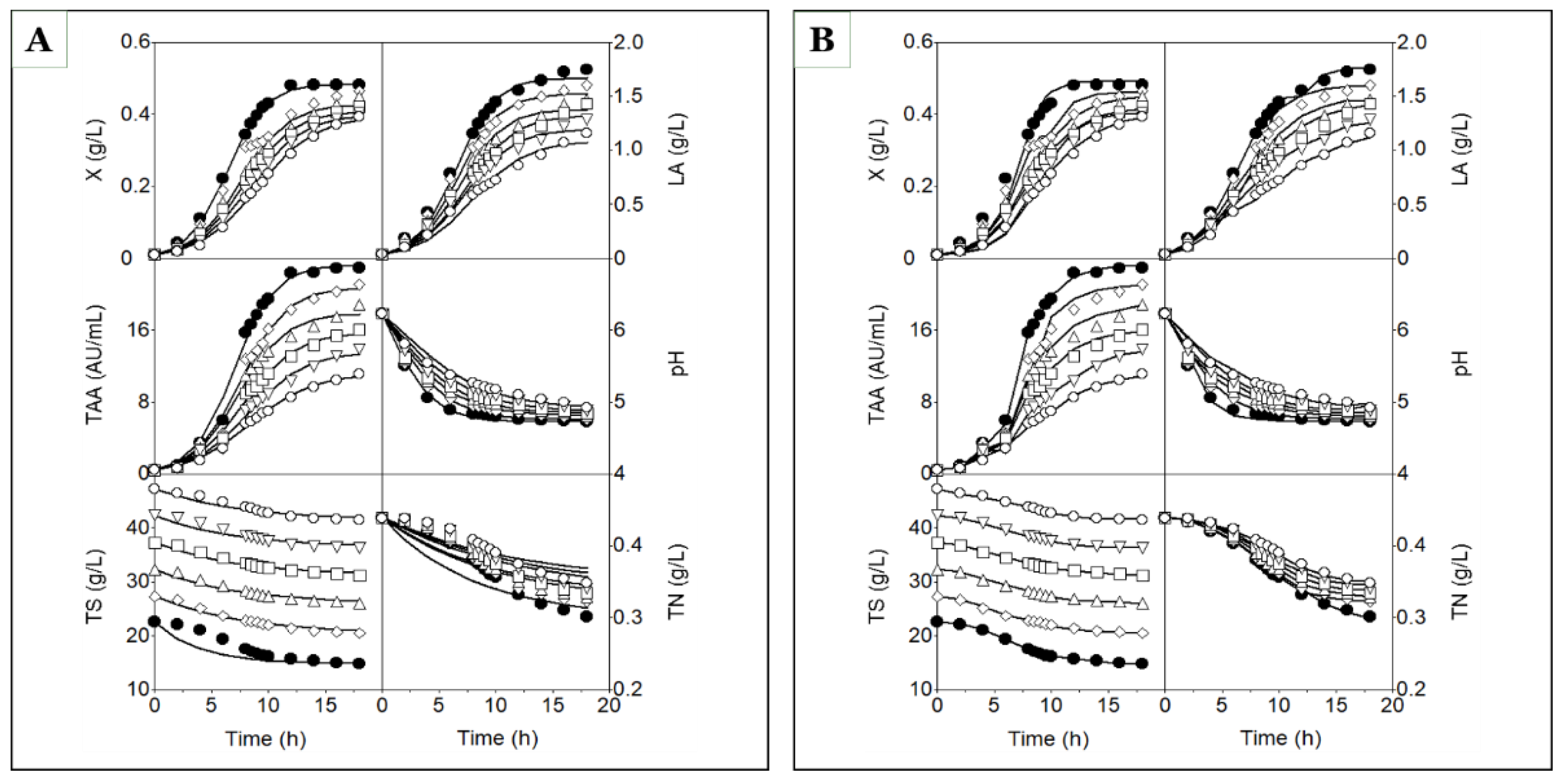
Time courses of biomass (X), lactic acid (LA), total antibacterial activity (TAA), pH, total sugars (TS), and total nitrogen (TN) in batch cultures of strain CECT 539 grown in concentrated whey (closed circles) and MRS broth (open squares). The lines fitted to the experimental data (symbols) for X, LA, TAA, pH, TS, and TN were generated using models (1), (2), (3), (6), (7), and (8), respectively (A), or models (16), (17), (18), (19), (20), and (21), respectively (B).
Figure 2.
Time courses of biomass (X), lactic acid (LA), total antibacterial activity (TAA), pH, total sugars (TS), and total nitrogen (TN) in batch cultures of strain CECT 539 grown in concentrated whey (closed circles) and MRS broth (open squares). The lines fitted to the experimental data (symbols) for X, LA, TAA, pH, TS, and TN were generated using models (1), (2), (3), (6), (7), and (8), respectively (A), or models (16), (17), (18), (19), (20), and (21), respectively (B).
Figure 3.
Time courses of biomass (X), lactic acid (LA), total antibacterial activity (TAA), pH, total sugars (TS), and total nitrogen (TN) in batch cultures of strain CECT 539 grown in DW medium (A) and MPW medium (B), either unsupplemented (closed circles) or supplemented with glutamic acid at concentrations of 1 (open diamonds), 2 (open triangles), 3 (open squares), 4 (open inverted triangles), and 5 (open circles) g/L. Lines fitted to the experimental data (symbols) for X, LA, TAA, pH, TS, and TN were generated using models (1), (2), (3), (6), (7), and (8), respectively (A and B).
Figure 3.
Time courses of biomass (X), lactic acid (LA), total antibacterial activity (TAA), pH, total sugars (TS), and total nitrogen (TN) in batch cultures of strain CECT 539 grown in DW medium (A) and MPW medium (B), either unsupplemented (closed circles) or supplemented with glutamic acid at concentrations of 1 (open diamonds), 2 (open triangles), 3 (open squares), 4 (open inverted triangles), and 5 (open circles) g/L. Lines fitted to the experimental data (symbols) for X, LA, TAA, pH, TS, and TN were generated using models (1), (2), (3), (6), (7), and (8), respectively (A and B).
Figure 4.
Time courses of biomass (X), lactic acid (LA), total antibacterial activity (TAA), pH, total sugars (TS), and total nitrogen (TN) in batch cultures of strain CECT 539 grown in DW medium (A) and MPW medium (B), either unsupplemented (closed circles) or supplemented with glutamic acid at concentrations of 1 (open diamonds), 2 (open triangles), 3 (open squares), 4 (open inverted triangles), and 5 (open circles) g/L. Lines fitted to the experimental data (symbols) for X, LA, TAA, pH, TS, and TN were generated using models (16), (17), (18), (19), (20), and (21), respectively.
Figure 4.
Time courses of biomass (X), lactic acid (LA), total antibacterial activity (TAA), pH, total sugars (TS), and total nitrogen (TN) in batch cultures of strain CECT 539 grown in DW medium (A) and MPW medium (B), either unsupplemented (closed circles) or supplemented with glutamic acid at concentrations of 1 (open diamonds), 2 (open triangles), 3 (open squares), 4 (open inverted triangles), and 5 (open circles) g/L. Lines fitted to the experimental data (symbols) for X, LA, TAA, pH, TS, and TN were generated using models (16), (17), (18), (19), (20), and (21), respectively.
Figure 5.
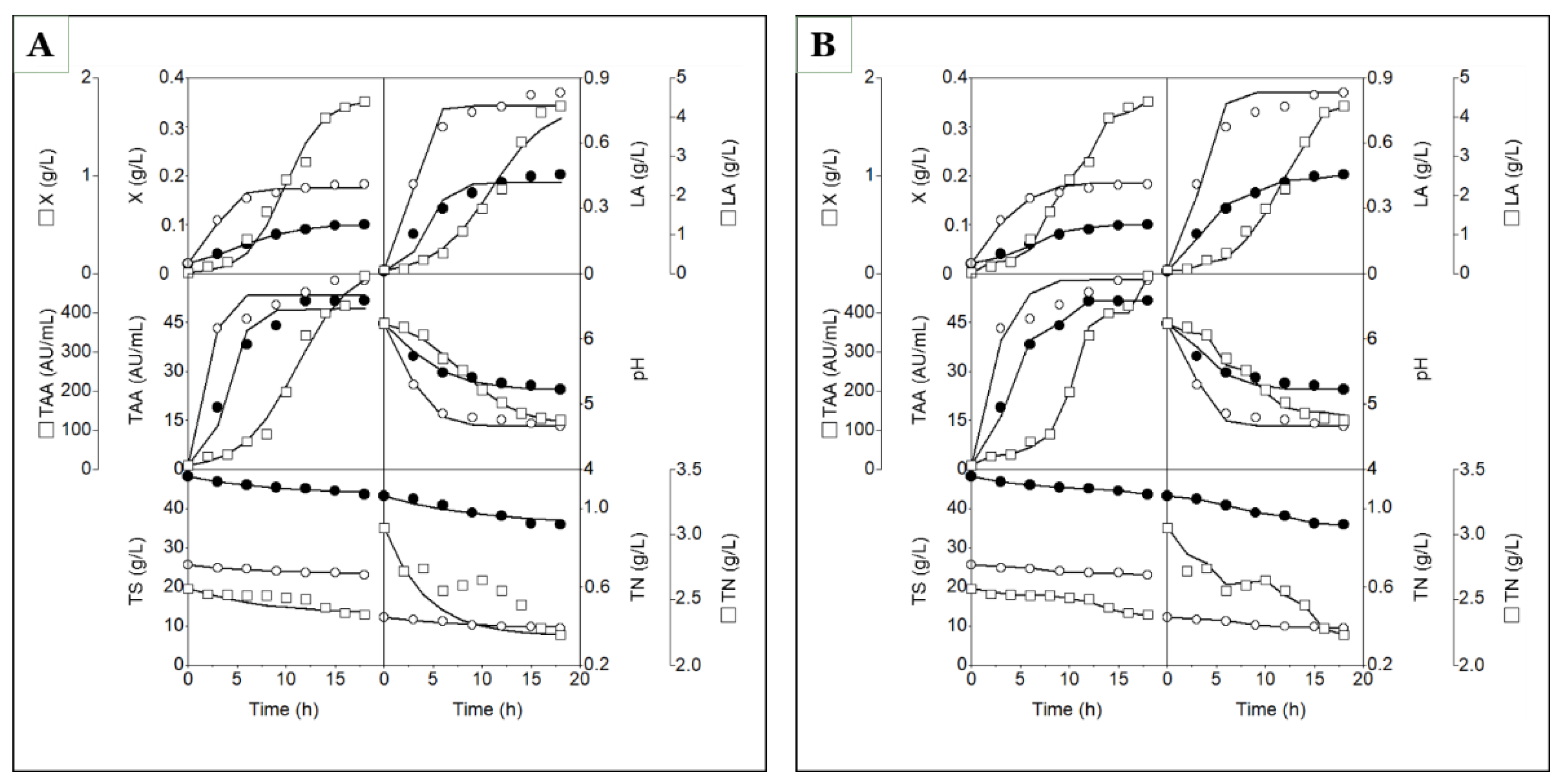
Time courses of biomass (X), lactic acid (LA), total antibacterial activity (TAA), pH, total sugars (TS), and total nitrogen (TN) in batch cultures of strain CECT 539 grown in DW medium, either unsupplemented (closed circles) or supplemented with glucose at concentrations of 5 (open diamonds), 10 (open triangles), 15 (open squares), 20 (open inverted triangles), and 25 (open circles) g/L. The lines fitted to the experimental data (symbols) for X, LA, TAA, pH, TS, and TN were gener-ated using models (1), (2), (3), (6), (7), and (8), respectively (A), or models (16), (17), (18), (19), (20), and (21), respectively (B).
Figure 5.
Time courses of biomass (X), lactic acid (LA), total antibacterial activity (TAA), pH, total sugars (TS), and total nitrogen (TN) in batch cultures of strain CECT 539 grown in DW medium, either unsupplemented (closed circles) or supplemented with glucose at concentrations of 5 (open diamonds), 10 (open triangles), 15 (open squares), 20 (open inverted triangles), and 25 (open circles) g/L. The lines fitted to the experimental data (symbols) for X, LA, TAA, pH, TS, and TN were gener-ated using models (1), (2), (3), (6), (7), and (8), respectively (A), or models (16), (17), (18), (19), (20), and (21), respectively (B).
Figure 6.
Time courses of biomass (X), lactic acid (LA), total antibacterial activity (TAA), pH, total sugars (TS), and total nitrogen (TN) in batch cultures of strain CECT 4043 grown in concentrated whey (closed circles), diluted whey (open circles), and MRS broth (open squares) media. The lines fitted to the experimental data (symbols) for X, LA, TAA, pH, TS, and TN were generated using models (1), (2), (3), (6), (7), and (8), respectively (A), or models (16), (17), (18), (19), (20), and (21), respectively (B).
Figure 6.
Time courses of biomass (X), lactic acid (LA), total antibacterial activity (TAA), pH, total sugars (TS), and total nitrogen (TN) in batch cultures of strain CECT 4043 grown in concentrated whey (closed circles), diluted whey (open circles), and MRS broth (open squares) media. The lines fitted to the experimental data (symbols) for X, LA, TAA, pH, TS, and TN were generated using models (1), (2), (3), (6), (7), and (8), respectively (A), or models (16), (17), (18), (19), (20), and (21), respectively (B).
Figure 7.
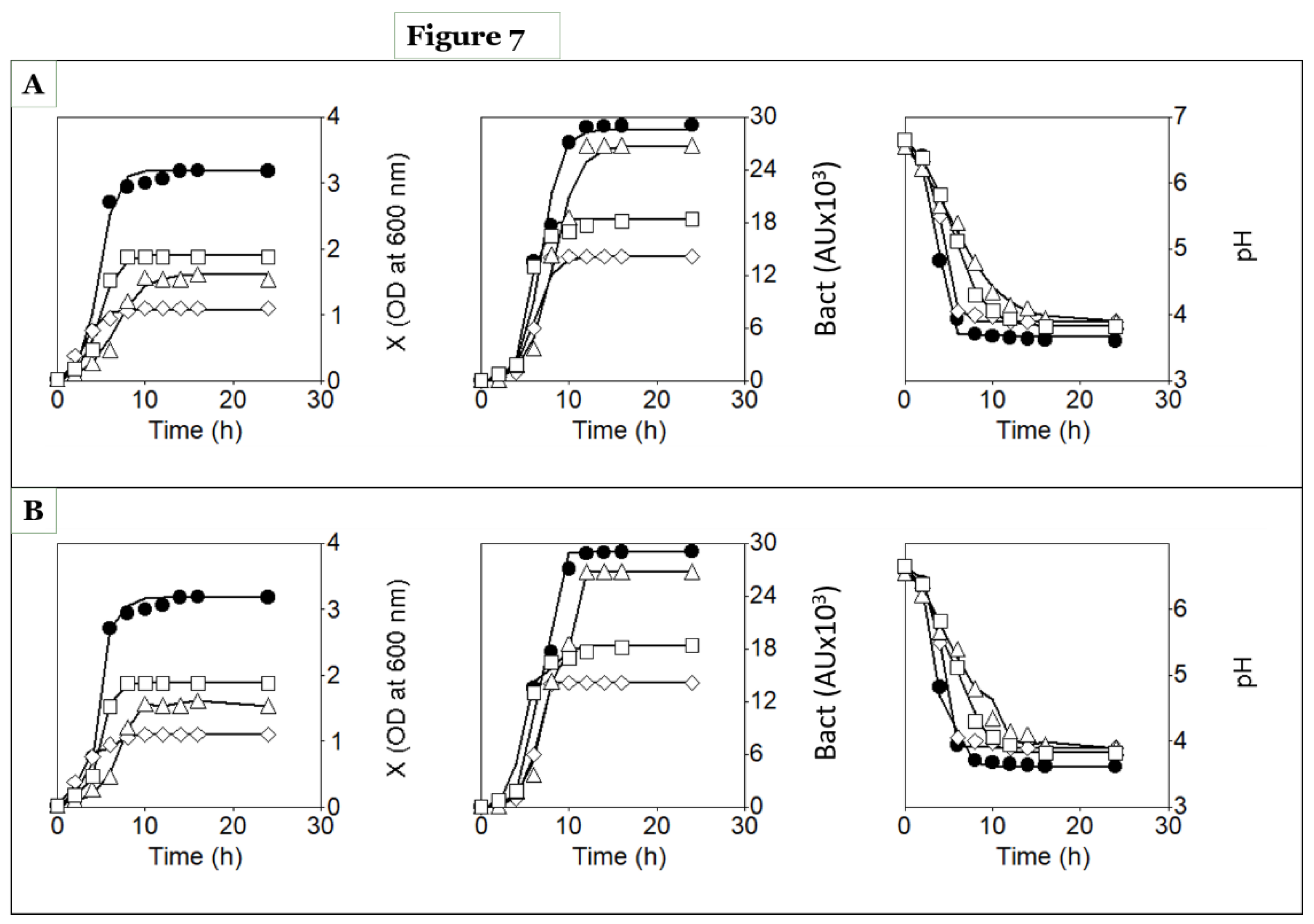
Time courses of biomass (X), lactic acid (LA), total antibacterial activity (TAA), pH, total sugars (TS), and total nitrogen (TN) in batch cultures of strain NRRL B-5627 grown in concentrated whey (closed circles), diluted whey (open circles), and MRS broth (open squares) media. The lines fitted to the experimental data (symbols) for X, LA, TAA, pH, TS, and TN were generated using models (1), (2), (3), (6), (7), and (8), respectively (A), or models (16), (17), (18), (19), (20), and (21), respectively (B).
Figure 7.
Time courses of biomass (X), lactic acid (LA), total antibacterial activity (TAA), pH, total sugars (TS), and total nitrogen (TN) in batch cultures of strain NRRL B-5627 grown in concentrated whey (closed circles), diluted whey (open circles), and MRS broth (open squares) media. The lines fitted to the experimental data (symbols) for X, LA, TAA, pH, TS, and TN were generated using models (1), (2), (3), (6), (7), and (8), respectively (A), or models (16), (17), (18), (19), (20), and (21), respectively (B).
Figure 8.
Time courses of biomass (X), bacteriocin (Bact), and pH in batch cultures of
Pediococcus acidilactici LB42–923 (closed circles),
Lactococcus lactis subsp.
lactis ATCC 11454 (open diamonds),
Leuconostoc carnosum Lm1 (open triangles), and
Lactobacillus sakei LB 706 (open squares) grown in TGE broth at uncontrolled pH [
23]. Lines fitted to the experimental data (symbols) for X, B, and pH were generated using models (1), (3), and (6), respectively (A), or models (16), (18), and (22), respectively (B).
Figure 8.
Time courses of biomass (X), bacteriocin (Bact), and pH in batch cultures of
Pediococcus acidilactici LB42–923 (closed circles),
Lactococcus lactis subsp.
lactis ATCC 11454 (open diamonds),
Leuconostoc carnosum Lm1 (open triangles), and
Lactobacillus sakei LB 706 (open squares) grown in TGE broth at uncontrolled pH [
23]. Lines fitted to the experimental data (symbols) for X, B, and pH were generated using models (1), (3), and (6), respectively (A), or models (16), (18), and (22), respectively (B).
Table 1.
Mean concentrations of total sugars and total nitrogen in the culture media used for batch cultures of
L. lactis CECT 539 [
31,
32,
33,
36],
Lact. casei CECT 4043 [
34,
35],
Ped. acidilactici NRRL B-5627 [
31,
32],
Ped. acidilactici LB42
–923,
L. lactis subsp.
lactis ATCC 11454,
Leuconostoc carnosum Lm1, and
Lact. sakei LB 706 [
23].
Table 1.
Mean concentrations of total sugars and total nitrogen in the culture media used for batch cultures of
L. lactis CECT 539 [
31,
32,
33,
36],
Lact. casei CECT 4043 [
34,
35],
Ped. acidilactici NRRL B-5627 [
31,
32],
Ped. acidilactici LB42
–923,
L. lactis subsp.
lactis ATCC 11454,
Leuconostoc carnosum Lm1, and
Lact. sakei LB 706 [
23].
| Growing Strain |
Culture medium* |
Total sugars (g/L) |
Total nitrogen (g/L) |
| CECT 539, CECT 4043, NRRL B-5627 |
CW |
48.11 ± 0.44 |
1.064 ± 0.018 |
| MRS |
19.59 ± 0.36 |
3.055 ± 0.005 |
| DW |
20.54 ± 0.51 |
0.453 ± 0.014 |
| CECT 539 |
DW–GA1 |
22.16 ± 0.61 |
0.570 ± 0.030 |
| DW–GA2 |
22.19 ± 0.32 |
0.664 ± 0.011 |
| DW–GA3 |
22.87 ± 0.17 |
0.758 ± 0.002 |
| DW–GA4 |
22.44 ± 0.01 |
0.867 ± 0.014 |
| DW–GA5 |
22.76 ± 0.02 |
0.945 ± 0.007 |
| DW–G5 |
27.30 ± 0.03 |
0.439 ± 0.011 |
| DW–G10 |
32.30 ± 0.01 |
0.439 ± 0.012 |
| DW–G15 |
37.30 ± 0.05 |
0.438 ± 0.012 |
| DW–G20 |
42.30 ± 0.06 |
0.438 ± 0.011 |
| DW–G25 |
47.32 ± 0.09 |
0.436 ± 0.009 |
| MPW |
5.33 ± 0.21 |
0.654 ± 0.009 |
| MPW–GA1 |
5.32 ± 0.01 |
0.765 ± 0.020 |
| MPW–GA2 |
5.32 ± 0.01 |
0.857 ± 0.012 |
| MPW–GA3 |
5.32 ± 0.01 |
0.951 ± 0.010 |
| MPW–GA4 |
5.31 ± 0.00 |
1.043 ± 0.022 |
| MPW–GA5 |
5.31 ± 0.00 |
1.137 ± 0.011 |
| LB42–923, ATCC 11454, Lm1, LB 706 |
TGE |
10 ± 0.01 |
2.337 ± 0.140** |
Table 2.
Estimated values for the constants of models (1), (2), (3), (6), (7), and (8), together with the coefficient of determination (R2) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of L. lactis CECT 539 in concentrated whey (CW) medium and MRS broth.
Table 2.
Estimated values for the constants of models (1), (2), (3), (6), (7), and (8), together with the coefficient of determination (R2) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of L. lactis CECT 539 in concentrated whey (CW) medium and MRS broth.
| |
Culture medium |
| Parameter |
CW medium |
MRS broth |
|
| KX |
0.35 |
1.60 |
|
|
aX
|
34.19 |
336.42 |
|
| bX |
0.38 |
0.73 |
|
|
R2X
|
0.9846 |
0.9979 |
|
| SSD–X |
2.29×10-3
|
9.20×10-3
|
|
| KLA |
0.74 |
4.50 |
|
| aLA |
44.80 |
261.64 |
|
| bLA |
1.16 |
0.67 |
|
|
R2LA
|
0.9500 |
0.9981 |
|
| SSD–LA |
2.58×10-2
|
5.05×10-2
|
|
| KTAA |
8.54 |
50.10 |
|
| aTAA |
49.55 |
85.08 |
|
| bTAA |
0.43 |
0.58 |
|
|
R2TAA
|
0.9838 |
0.9856 |
|
| SSD–TAA |
1.18 |
42.07 |
|
| pHi |
6.23 |
6.23 |
|
| pHf |
5.12 |
4.72 |
|
| apH |
0.61 |
1.41×10-2
|
|
| bpH |
0.30 |
0.55 |
|
|
R2pH
|
0.9952 |
0.9949 |
|
| SSD–pH |
6.12×10-3
|
1.34×10-2
|
|
| TSi |
48.16 |
19.63 |
|
| TSf |
39.64 |
11.11 |
|
| bTS |
0.11 |
0.33 |
|
|
R2TS
|
0.9638 |
0.9809 |
|
| SSD–TS |
3.34 |
1.28 |
|
| TNi |
1.07 |
3.06 |
|
| TNf |
0.72 |
2.00 |
|
| bTN |
0.16 |
0.17 |
|
|
R2TN
|
0.9512 |
0.9563 |
|
| SSD–TN |
8.27×10-3
|
3.60×10-2
|
|
Table 3.
Estimated values for the constants of models (16)–(21), together with the coefficient of determination (R2) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of L. lactis CECT 539 in concentrated whey (CW) medium and MRS broth.
Table 3.
Estimated values for the constants of models (16)–(21), together with the coefficient of determination (R2) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of L. lactis CECT 539 in concentrated whey (CW) medium and MRS broth.
| |
Culture medium |
| Parameter |
CW medium |
MRS broth |
|
| KX |
0.34 |
1.60 |
|
|
aX
|
33.02 |
159.29 |
|
| bpH-X |
0.29 |
0.51 |
|
|
R2X
|
0.9962 |
0.9938 |
|
| SSD–X |
6.25×10-4
|
2.07×10-2
|
|
| KLA |
0.80 |
4.50 |
|
| aLA |
49.07 |
261.64 |
|
| bX-LA |
0.51 |
0.67 |
|
|
R2LA
|
0.9492 |
0.9915 |
|
| SSD–LA |
3.99×10-2
|
0.30 |
|
| KTAA |
8.45 |
50.10 |
|
| aTAA |
49.55 |
50.56 |
|
| bX-TAA |
0.23 |
0.25 |
|
| bpH-TAA |
0.12 |
0.31 |
|
|
R2TAA
|
0.9943 |
0.9958 |
|
| SSD–TAA |
0.47 |
0.12 |
|
| pHi |
6.23 |
6.23 |
|
| pHf |
5.12 |
4.72 |
|
| apH |
0.58 |
2.20×10-2
|
|
| bLA-pH |
0.83 |
0.68 |
|
|
R2pH
|
0.9750 |
0.9969 |
|
| SSD–pH |
4.13×10-2
|
8.43×10-3
|
|
| TSi |
48.16 |
19.42 |
|
| TSf |
39.64 |
11.11 |
|
| bX-TS |
0.40 |
0.64 |
|
|
R2TS
|
0.9926 |
0.9966 |
|
| SSD–TS |
0.56 |
0.18 |
|
| TNi |
1.07 |
3.06 |
|
| TNf |
0.72 |
2.00 |
|
| bX-TN |
0.44 |
0.63 |
|
|
R2TN
|
0.9954 |
0.9957 |
|
| SSD–TN |
8.88×10-4
|
3.57×10-3
|
|
Table 4.
Estimated values for the constants of models (1), (2), (3), (6), (7), and (8), together with the coefficient of determination (R2) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of L. lactis CECT 539 in DW medium supplemented with different glutamic acid concentrations.
Table 4.
Estimated values for the constants of models (1), (2), (3), (6), (7), and (8), together with the coefficient of determination (R2) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of L. lactis CECT 539 in DW medium supplemented with different glutamic acid concentrations.
| |
Culture medium |
| Parameter |
DW |
DW–GA1 |
DW–GA2 |
DW–GA3 |
DW–GA4 |
DW–GA5 |
| KX |
0.43 |
0.61 |
0.68 |
0.79 |
0.91 |
1.09 |
|
aX
|
28.03 |
40.04 |
44.30 |
52.62 |
59.50 |
71.88 |
| bX |
0.39 |
0.40 |
0.43 |
0.43 |
0.45 |
0.47 |
|
R2X
|
0.9945 |
0.9957 |
0.9985 |
0.9966 |
0.9966 |
0.9945 |
| SSD–X |
1.460×10-3
|
2.135×10-3
|
9.76×10-4
|
3.16×10-3
|
4.68×10-3
|
1.20×10-2
|
| KLA |
2.72 |
3.56 |
4.12 |
4.22 |
4.80 |
5.15 |
| aLA |
51.25 |
67.50 |
81.40 |
80.12 |
94.95 |
102.08 |
| bLA |
0.46 |
0.44 |
0.47 |
0.53 |
0.54 |
0.63 |
|
R2LA
|
0.9876 |
0.9926 |
0.9960 |
0.9948 |
0.9832 |
0.9863 |
| SSD–LA |
0.13 |
0.14 |
0.12 |
0.14 |
0.63 |
0.56 |
| KTAA |
22.82 |
28.79 |
30.42 |
32.96 |
35.57 |
38.60 |
| aTAA |
50.28 |
63.70 |
66.61 |
73.07 |
78.04 |
84.77 |
| bTAA |
0.49 |
0.48 |
0.50 |
0.52 |
0.52 |
0.53 |
|
R2TAA
|
0.9945 |
0.9988 |
0.9961 |
0.9971 |
0.9951 |
0.9969 |
| SSD–TAA |
3.94 |
2.15 |
6.83 |
6.67 |
11.88 |
10.02 |
| pHi |
6.23 |
6.23 |
6.23 |
6.23 |
6.23 |
6.23 |
| pHf |
4.60 |
4.50 |
4.29 |
4.18 |
4.09 |
3.90 |
| apH |
0.39 |
0.47 |
0.55 |
0.55 |
0.66 |
0.81 |
| bpH |
0.53 |
0.41 |
0.46 |
0.46 |
0.58 |
0.53 |
|
R2pH
|
0.9911 |
0.9796 |
0.9901 |
0.9880 |
0.9887 |
0.9871 |
| SSD–pH |
3.78×10-2
|
6.72×10-2
|
4.87×10-2
|
5.42×10-2
|
6.89×10-2
|
9.23×10-2
|
| TSi |
26.10 |
26.16 |
26.19 |
26.00 |
25.44 |
25.76 |
| TSf |
23.54 |
22.69 |
22.06 |
21.65 |
20.59 |
20.54 |
| bTS |
0.11 |
0.09 |
0.10 |
0.10 |
0.10 |
0.10 |
|
R2TS
|
0.9066 |
0.9066 |
0.9298 |
0.9460 |
0.9251 |
0.9545 |
| SSD–TS |
1.38 |
2.63 |
2.78 |
2.87 |
4.12 |
2.82 |
| TNi |
0.47 |
0.57 |
0.67 |
0.76 |
0.87 |
0.95 |
| TNf |
0.32 |
0.33 |
0.33 |
0.33 |
0.34 |
0.33 |
| bTN |
0.11 |
0.11 |
0.11 |
0.11 |
0.11 |
0.10 |
|
R2TN
|
0.9345 |
0.9305 |
0.9100 |
0.9041 |
0.9479 |
0.9332 |
| SSD–TN |
3.04×10-3
|
8.42×10-3
|
2.36×10-2
|
3.91×10-2
|
3.12×10-2
|
5.93×10-2
|
Table 5.
Estimated values for the constants of models (1), (2), (3), (6), (7), and (8), together with the coefficient of determination (R2) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of L. lactis CECT 539 in media prepared with mussel processing wastes supplemented with different glutamic acid concentrations.
Table 5.
Estimated values for the constants of models (1), (2), (3), (6), (7), and (8), together with the coefficient of determination (R2) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of L. lactis CECT 539 in media prepared with mussel processing wastes supplemented with different glutamic acid concentrations.
| |
|
Culture medium |
| Parameter |
MPW |
MPW–GA1 |
MPW–GA2 |
MPW–GA3 |
MPW–GA4 |
MPW–GA5 |
| KX |
0.50 |
0.71 |
0.80 |
0.90 |
0.97 |
1.10 |
|
aX
|
26.40 |
38.22 |
43.76 |
48.82 |
53.00 |
60.32 |
| bX |
0.43 |
0.39 |
0.45 |
0.47 |
0.51 |
0.52 |
|
R2X
|
0.9900 |
0.9925 |
0.9797 |
0.9861 |
0.9892 |
0.9938 |
| SSD–X |
3.98×10-3
|
6.73×10-3
|
2.10×10-2
|
1.91×10-2
|
1.76×10-2
|
1.52×10-2
|
| KLA |
2.31 |
2.56 |
3.02 |
3.23 |
3.38 |
3.60 |
| aLA |
47.22 |
52.42 |
61.95 |
66.38 |
69.37 |
73.94 |
| bLA |
0.46 |
0.45 |
0.47 |
0.49 |
0.50 |
0.53 |
|
R2LA
|
0.9809 |
0.9742 |
0.9798 |
0.9785 |
0.9762 |
0.9805 |
| SSD–LA |
0.16 |
0.30 |
0.30 |
0.37 |
0.44 |
0.40 |
| KTAA |
32.90 |
38.80 |
40.62 |
43.30 |
47.35 |
51.26 |
| aTAA |
49.53 |
58.59 |
61.38 |
65.50 |
71.71 |
77.72 |
| bTAA |
0.52 |
0.50 |
0.51 |
0.52 |
0.54 |
0.59 |
|
R2TAA
|
0.9990 |
0.9921 |
0.9937 |
0.9870 |
0.9879 |
0.9872 |
| SSD–TAA |
1.426 |
17.566 |
16.604 |
37.417 |
42.851 |
53.497 |
| pHi |
6.23 |
6.23 |
6.23 |
6.17 |
6.23 |
6.23 |
| pHf |
4.74 |
4.31 |
4.16 |
4.25 |
4.11 |
4.00 |
| apH |
0.83 |
0.64 |
0.64 |
0.88 |
0.53 |
0.92 |
| bpH |
0.41 |
0.33 |
0.33 |
0.30 |
0.28 |
0.24 |
|
R2pH
|
0.9830 |
0.9755 |
0.9800 |
0.9762 |
0.9772 |
0.9868 |
| SSD–pH |
5.33×10-2
|
0.11 |
9.50×10-2
|
9.79×10-2
|
0.13 |
7.46×10-2
|
| TSi |
5.32 |
5.32 |
5.32 |
5.36 |
5.31 |
5.31 |
| TSf |
3.05 |
2.84 |
2.41 |
2.26 |
2.00 |
1.67 |
| bTS |
0.10 |
0.10 |
0.11 |
0.12 |
0.11 |
0.13 |
|
R2TS
|
0.9662 |
0.9682 |
0.9614 |
0.9693 |
0.9719 |
0.9765 |
| SSD–TS |
0.37 |
0.50 |
0.69 |
0.61 |
0.55 |
0.56 |
| TNi |
0.67 |
0.77 |
0.86 |
0.95 |
1.04 |
1.14 |
| TNf |
0.49 |
0.49 |
0.49 |
0.49 |
0.48 |
0.47 |
| bTN |
0.11 |
0.11 |
0.12 |
0.10 |
0.10 |
0.11 |
|
R2TN
|
0.9846 |
0.9846 |
0.9767 |
0.9701 |
0.9649 |
0.9645 |
| SSD–TN |
1.70×10-3
|
4.21×10-3
|
5.84×10-3
|
1.31×10-2
|
2.38×10-2
|
3.46×10-2
|
Table 6.
Estimated values for the constants of models (16), (17), (18), (19), (20), and (21), together with the coefficient of determination (R2) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of L. lactis CECT 539 in diluted whey medium supplemented with different glutamic acid concentrations.
Table 6.
Estimated values for the constants of models (16), (17), (18), (19), (20), and (21), together with the coefficient of determination (R2) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of L. lactis CECT 539 in diluted whey medium supplemented with different glutamic acid concentrations.
| |
Culture medium |
| Parameter |
DW |
DW–GA1 |
DW–GA2 |
DW–GA3 |
DW–GA4 |
DW–GA5 |
| KX |
0.42 |
0.60 |
0.67 |
0.78 |
0.90 |
1.11 |
|
aX
|
27.59 |
39.51 |
46.31 |
51.81 |
60.13 |
73.89 |
| bpH-X |
0.28 |
0.25 |
0.24 |
0.29 |
0.31 |
0.27 |
|
R2X
|
0.9984 |
0.9975 |
0.9955 |
0.9899 |
0.9872 |
0.9941 |
| SSD–X |
7.91×10-4
|
1.32×10-3
|
4.23×10-3
|
1.09×10-2
|
1.98×10-2
|
1.69×10-2
|
| KLA |
2.66 |
3.54 |
4.20 |
4.38 |
4.97 |
5.34 |
| aLA |
50.26 |
67.04 |
79.74 |
83.32 |
94.59 |
101.78 |
| bX-LA |
0.38 |
0.40 |
0.45 |
0.43 |
0.49 |
0.47 |
|
R2LA
|
0.9972 |
0.9949 |
0.9946 |
0.9963 |
0.9873 |
0.9918 |
| SSD–LA |
3.97×10-2
|
9.83×10-2
|
0.13 |
0.13 |
0.52 |
0.435 |
| KTAA |
23.19 |
28.69 |
30.61 |
33.61 |
35.89 |
38.89 |
| aTAA |
51.11 |
63.47 |
67.78 |
74.53 |
79.65 |
86.39 |
| bX-TAA |
0.25 |
0.20 |
0.34 |
0.28 |
0.31 |
0.30 |
| bpH-TAA |
0.10 |
0.13 |
5.73×10-2
|
0.11 |
0.10 |
0.10 |
|
R2TAA
|
0.9991 |
0.9989 |
0.9986 |
0.9979 |
0.9949 |
0.9959 |
| SSD–TAA |
0.68 |
1.37 |
2.18 |
5.21 |
12.88 |
15.01 |
| pHi |
6.23 |
6.23 |
6.23 |
6.23 |
6.23 |
6.23 |
| pHf |
4.60 |
4.50 |
4.26 |
4.17 |
4.08 |
3.90 |
| apH |
4.31 |
4.31 |
4.31 |
4.31 |
4.31 |
4.31 |
| bLA-pH |
0.48 |
0.49 |
0.49 |
0.55 |
0.61 |
0.68 |
|
R2pH
|
0.9949 |
0.9976 |
0.9968 |
0.9993 |
0.9978 |
0.9976 |
| SSD–pH |
1.16×10-2
|
7.02×10-3
|
1.32×10-2
|
3.70×10-3
|
1.17×10-2
|
1.64×10-2
|
| TSi |
26.16 |
26.16 |
26.20 |
25.88 |
25.45 |
25.78 |
| TSf |
23.55 |
22.68 |
22.01 |
21.63 |
20.59 |
20.54 |
| bX-TS |
0.41 |
0.42 |
0.41 |
0.41 |
0.47 |
0.47 |
|
R2TS
|
0.9982 |
0.9988 |
0.9986 |
0.9992 |
0.9993 |
0.9988 |
| SSD–TS |
1.86×10-2
|
2.29×10-2
|
7.63×10-2
|
4.94×10-2
|
2.63×10-2
|
6.19×10-2
|
| TNi |
0.47 |
0.57 |
0.67 |
0.76 |
0.87 |
0.95 |
| TNf |
0.32 |
0.33 |
0.33 |
0.33 |
0.35 |
0.33 |
| bX-TN |
0.43 |
0.42 |
0.43 |
0.43 |
0.50 |
0.50 |
|
R2TN
|
0.9973 |
0.9985 |
1.0000 |
0.9997 |
0.9981 |
0.9987 |
| SSD–TN |
1.00×10-4
|
1.21×10-4
|
2.89×10-5
|
1.55×10-4
|
8.66×10-4
|
9.60×10-4
|
Table 7.
Estimated values for the constants of models (16), (17), (18), (19), (20), and (21), together with the coefficient of determination (R2) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of L. lactis CECT 539 in media prepared with mussel processing wastes supplemented with different glutamic acid concentrations.
Table 7.
Estimated values for the constants of models (16), (17), (18), (19), (20), and (21), together with the coefficient of determination (R2) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of L. lactis CECT 539 in media prepared with mussel processing wastes supplemented with different glutamic acid concentrations.
| |
Culture medium |
| Parameter |
MPW |
MPW–GA1 |
MPW–GA2 |
MPW–GA3 |
MPW–GA4 |
MPW–GA5 |
| KX |
0.50 |
0.71 |
0.83 |
0.92 |
1.00 |
1.11 |
|
aX
|
26.72 |
38.26 |
45.42 |
50.20 |
54.39 |
60.78 |
| bpH-X |
0.37 |
0.21 |
0.21 |
0.20 |
0.18 |
0.18 |
|
R2X
|
0.9920 |
0.9919 |
0.9968 |
0.9913 |
0.9886 |
0.9940 |
| SSD–X |
3.33×10-3
|
7.19×10-3
|
3.38×10-3
|
1.16×10-2
|
1.74×10-2
|
1.31×10-2
|
| KLA |
2.46 |
2.67 |
3.11 |
3.32 |
3.48 |
3.75 |
| aLA |
50.22 |
54.74 |
63.80 |
63.80 |
71.61 |
77.13 |
| bX-LA |
0.63 |
0.40 |
0.44 |
0.44 |
0.48 |
0.49 |
|
R2LA
|
0.9884 |
0.9780 |
0.9820 |
0.9856 |
0.9878 |
0.9942 |
| SSD–LA |
0.10 |
0.21 |
0.312 |
0.33 |
0.28 |
0.16 |
| KTAA |
33.32 |
40.19 |
41.19 |
44.60 |
49.16 |
52.56 |
| aTAA |
50.17 |
60.72 |
62.25 |
67.49 |
74.49 |
79.71 |
| bX-TAA |
0.63 |
0.27 |
0.21 |
0.00 |
0.24 |
0.08 |
| bpH-TAA |
2.55×10-3
|
6.85×10-2
|
0.10 |
0.19 |
8.87×10-2
|
0.15 |
|
R2TAA
|
0.9951 |
0.9923 |
0.9954 |
0.9947 |
0.9898 |
0.9939 |
| SSD–TAA |
10.13 |
20.07 |
18.59 |
16.86 |
43.64 |
28.68 |
| pHi |
6.23 |
6.23 |
6.23 |
6.23 |
6.23 |
6.23 |
| pHf |
4.74 |
4.30 |
4.14 |
4.23 |
4.10 |
3.98 |
| apH |
4.31 |
4.31 |
4.31 |
4.31 |
4.31 |
4.31 |
| bLA-pH |
0.82 |
0.55 |
0.55 |
0.55 |
0.54 |
0.56 |
|
R2pH
|
0.9924 |
0.9760 |
0.9917 |
0.9887 |
0.9868 |
0.9890 |
| SSD–pH |
2.11×10-2
|
0.11 |
4.38×10-2
|
5.32×10-2
|
7.54×10-2
|
7.23×10-2
|
| TSi |
5.32 |
5.32 |
5.32 |
5.32 |
5.31 |
5.31 |
| TSf |
3.05 |
2.84 |
2.41 |
2.26 |
2.09 |
1.67 |
| bX-TS |
0.64 |
0.41 |
0.41 |
0.41 |
0.41 |
0.46 |
|
R2TS
|
0.9981 |
0.9969 |
0.9963 |
0.9947 |
0.9938 |
0.9964 |
| SSD–TS |
1.51×10-2
|
2.98×10-2
|
7.97×10-2
|
0.11 |
0.21 |
0.12 |
| TNi |
0.67 |
0.77 |
0.86 |
0.95 |
1.05 |
1.14 |
| TNf |
0.49 |
0.49 |
0.49 |
0.49 |
0.48 |
0.47 |
| bX-TN |
0.69 |
0.43 |
0.43 |
0.43 |
0.43 |
0.43 |
|
R2TN
|
0.9961 |
0.9958 |
0.9942 |
0.9966 |
0.9968 |
0.9963 |
| SSD–TN |
1.92×10-4
|
4.82×10-4
|
1.53×10-3
|
1.76×10-3
|
3.37×10-3
|
4.88×10-3
|
Table 8.
Estimated values for the constants of models (1), (2), (3), (6), (7), and (8), together with the coefficient of determination (R2) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of L. lactis CECT 539 in diluted whey medium supplemented with different glucose concentrations.
Table 8.
Estimated values for the constants of models (1), (2), (3), (6), (7), and (8), together with the coefficient of determination (R2) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of L. lactis CECT 539 in diluted whey medium supplemented with different glucose concentrations.
| |
|
Culture medium |
| Parameter |
DW |
DW–G5 |
DW–G10 |
DW–G15 |
DW–G20 |
DW–G25 |
| KX |
0.48 |
0.48 |
0.43 |
0.41 |
0.40 |
0.39 |
|
aX
|
47.42 |
47.42 |
41.84 |
39.99 |
38.98 |
38.35 |
| bX |
0.60 |
0.60 |
0.49 |
0.47 |
0.44 |
0.40 |
|
R2X
|
0.9984 |
0.9727 |
0.9917 |
0.9962 |
0.9985 |
0.9973 |
| SSD–X |
8.10×10-4
|
4.06×10-2
|
2.61×10-3
|
9.99×10-4
|
3.22×10-4
|
5.70×10-4
|
| KLA |
1.67 |
1.52 |
1.38 |
1.33 |
1.20 |
1.08 |
| aLA |
46.98 |
42.71 |
38.63 |
37.06 |
33.26 |
30.10 |
| bLA |
0.59 |
0.56 |
0.53 |
0.48 |
0.48 |
0.43 |
|
R2LA
|
0.9926 |
0.9874 |
0.9865 |
0.9784 |
0.9821 |
0.9808 |
| SSD–LA |
4.28×10-2
|
5.70×10-2
|
4.41×10-2
|
6.21×10-2
|
4.19×10-2
|
3.31×10-2
|
| KTAA |
23.27 |
20.71 |
17.87 |
15.69 |
13.58 |
11.11 |
| aTAA |
50.76 |
44.16 |
37.98 |
33.22 |
28.61 |
23.24 |
| bTAA |
0.56 |
0.50 |
0.49 |
0.45 |
0.40 |
0.37 |
|
R2TAA
|
0.9922 |
0.9901 |
0.9883 |
0.9931 |
0.9868 |
0.9948 |
| SSD–TAA |
7.70 |
6.63 |
5.90 |
2.39 |
3.21 |
0.86 |
| pHi |
6.23 |
6.23 |
6.23 |
6.23 |
6.20 |
6.23 |
| pHf |
4.73 |
4.76 |
4.81 |
4.84 |
4.87 |
4.92 |
| apH |
0.93 |
0.79 |
0.92 |
0.87 |
0.68 |
0.67 |
| bpH |
0.50 |
0.40 |
0.34 |
0.32 |
0.29 |
0.26 |
|
R2pH
|
0.9951 |
0.9798 |
0.9747 |
0.9668 |
0.9699 |
0.9822 |
| SSD–pH |
2.09×10-2
|
7.21×10-2
|
7.67×10-2
|
9.25×10-2
|
7.29×10-2
|
4.65×10-2
|
| TSi |
22.63 |
27.46 |
32.36 |
37.36 |
42.32 |
47.32 |
| TSf |
14.84 |
20.50 |
26.00 |
31.23 |
36.29 |
41.47 |
| bTS |
0.26 |
0.14 |
0.14 |
0.14 |
0.14 |
0.14 |
|
R2TS
|
0.8194 |
0.9758 |
0.9691 |
0.9683 |
0.9503 |
0.9425 |
| SSD–TS |
36.16 |
2.32 |
2.52 |
2.40 |
4.04 |
3.77 |
| TNi |
0.44 |
0.44 |
0.44 |
0.44 |
0.44 |
0.44 |
| TNf |
0.30 |
0.32 |
0.33 |
0.33 |
0.34 |
0.35 |
| bTN |
0.12 |
0.10 |
0.10 |
0.08 |
0.08 |
0.08 |
|
R2TN
|
0.8766 |
0.9192 |
0.9180 |
0.9209 |
0.9069 |
0.8793 |
| SSD–TN |
6.21×10-3
|
2.62×10-3
|
2.32×10-3
|
2.46×10-3
|
2.16×10-3
|
2.30×10-3
|
Table 9.
Estimated values for the constants of models (16), (17), (18), (19), (20), and (21), together with the coefficient of determination (R2) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of L. lactis CECT 539 in diluted whey medium supplemented with different glucose concentrations.
Table 9.
Estimated values for the constants of models (16), (17), (18), (19), (20), and (21), together with the coefficient of determination (R2) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of L. lactis CECT 539 in diluted whey medium supplemented with different glucose concentrations.
| |
|
Culture medium |
| Parameter |
DW |
DW–G5 |
DW–G10 |
DW–G15 |
DW–G20 |
DW–G25 |
| KX |
0.49 |
0.47 |
0.45 |
0.42 |
0.40 |
0.39 |
|
aX
|
48.43 |
45.54 |
44.04 |
41.01 |
39.40 |
38.30 |
| bpH-X |
0.58 |
0.37 |
0.31 |
0.27 |
0.36 |
0.23 |
|
R2X
|
0.9824 |
0.9847 |
0.9873 |
0.9938 |
0.9893 |
0.9960 |
| SSD–X |
1.27×10-2
|
8.05×10-3
|
6.42×10-3
|
2.49×10-3
|
4.66×10-3
|
1.37×10-3
|
| KLA |
1.76 |
1.61 |
1.49 |
1.43 |
1.28 |
1.16 |
| aLA |
49.57 |
45.05 |
41.61 |
41.61 |
35.69 |
32.25 |
| bX-LA |
0.60 |
0.53 |
0.48 |
0.48 |
0.43 |
0.40 |
|
R2LA
|
0.9970 |
0.9862 |
0.9894 |
0.9922 |
0.9972 |
0.9921 |
| SSD–LA |
2.17×10-2
|
7.89×10-2
|
5.02×10-2
|
4.86×10-2
|
5.27×10-3
|
1.35×10-2
|
| KTAA |
23.12 |
21.12 |
18.82 |
16.07 |
13.86 |
11.16 |
| aTAA |
50.42 |
45.05 |
40.04 |
34.13 |
29.22 |
23.33 |
| bX-TAA |
0.59 |
0.51 |
0.47 |
0.47 |
0.43 |
0.40 |
| bpH-TAA |
1.89×10-3
|
2.00×10-3
|
2.46×10-3
|
2.00×10-3
|
2.00×10-3
|
2.00×10-3
|
|
R2TAA
|
0.9993 |
0.9890 |
0.9941 |
0.9964 |
0.9983 |
0.9970 |
| SSD–TAA |
1.07 |
10.61 |
5.25 |
2.11 |
0.45 |
0.50 |
| pHi |
6.23 |
6.23 |
6.23 |
6.23 |
6.23 |
6.23 |
| pHf |
4.73 |
4.76 |
4.80 |
4.84 |
4.87 |
4.92 |
| apH |
0.14 |
0.48 |
0.68 |
1.00 |
0.19 |
0.84 |
| bLA-pH |
0.73 |
0.62 |
0.55 |
0.53 |
0.50 |
0.44 |
|
R2pH
|
0.9855 |
0.9795 |
0.9852 |
0.9804 |
0.9648 |
0.9839 |
| SSD–pH |
6.03×10-2
|
7.10×10-2
|
4.63×10-2
|
5.32×10-2
|
9.05×10-2
|
3.61×10-2
|
| TSi |
22.63 |
27.46 |
32.36 |
37.36 |
42.32 |
47.32 |
| TSf |
14.83 |
20.50 |
26.00 |
31.23 |
36.29 |
41.47 |
| bX-TS |
0.60 |
0.53 |
0.48 |
0.46 |
0.46 |
0.41 |
|
R2TS
|
0.9995 |
0.9972 |
0.9978 |
0.9989 |
0.9989 |
0.9976 |
| SSD–TS |
9.46×10-2
|
0.36 |
0.23 |
8.80×10-2
|
0.12 |
0.16 |
| TNi |
0.44 |
0.44 |
0.44 |
0.44 |
0.44 |
0.44 |
| TNf |
0.30 |
0.32 |
0.33 |
0.33 |
0.34 |
0.35 |
| bX-TN |
0.54 |
0.54 |
0.47 |
0.47 |
0.47 |
0.45 |
|
R2TN
|
0.9963 |
0.9975 |
0.9989 |
0.9976 |
0.9955 |
0.9960 |
| SSD–TN |
1.87×10-4
|
1.01×10-4
|
4.10×10-5
|
3.97×10-5
|
8.38×10-5
|
9.86×10-5
|
Table 10.
Estimated values for the constants of models (1), (2), (3), (6), (7), and (8), together with the coefficient of determination (R²) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of Lact. casei CECT 4043 and Pediococcus acidilactici NRRL B–562 in concentrated whey (CW), diluted whey (DW), and MRS broth.
Table 10.
Estimated values for the constants of models (1), (2), (3), (6), (7), and (8), together with the coefficient of determination (R²) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of Lact. casei CECT 4043 and Pediococcus acidilactici NRRL B–562 in concentrated whey (CW), diluted whey (DW), and MRS broth.
| |
|
Culture medium/Strain |
| Parameter |
CW/CECT 4043 |
DW/CECT 4043 |
MRS/CECT 4043 |
CW/NRRL B–562 |
DW/NRRL B–562 |
MRS/NRRL B–562 |
| KX |
0.18 |
0.33 |
2.22 |
0.10 |
0.17 |
1.77 |
|
aX
|
21.76 |
15.69 |
221.37 |
4.08 |
7.69 |
176.40 |
| bX |
0.26 |
0.45 |
0.49 |
0.30 |
0.81 |
0.52 |
|
R2X
|
0.9676 |
0.9909 |
0.9758 |
0.9990 |
0.9836 |
0.9854 |
| SSD–X |
1.05×10-3
|
1.02×10-3
|
0.20 |
6.40×10-6
|
3.43×10-4
|
8.60×10-2
|
| KLA |
1.33 |
2.17 |
22.56 |
0.42 |
0.77 |
4.28 |
| aLA |
44.29 |
47.71 |
50.63 |
45.05 |
45.38 |
50.08 |
| bLA |
0.33 |
0.63 |
0.30 |
0.88 |
1.29 |
0.36 |
|
R2LA
|
0.9597 |
0.9861 |
0.9982 |
0.9346 |
0.9734 |
0.9853 |
| SSD–LA |
8.26×10-2
|
7.51×10-2
|
0.87 |
1.25×10-2
|
1.42×10-2
|
0.48 |
| KTAA |
1.84 |
3.00 |
11.98 |
49.03 |
53.25 |
526.10 |
| aTAA |
43.60 |
42.81 |
49.02 |
46.88 |
45.07 |
53.40 |
| bTAA |
0.34 |
0.73 |
0.38 |
0.95 |
1.75 |
0.36 |
|
R2TAA
|
0.9708 |
0.9884 |
0.9905 |
0.9648 |
0.9563 |
0.9859 |
| SSD–TAA |
0.10 |
0.15 |
1.58 |
88.98 |
103.37 |
4417.13 |
| pHi |
7.02 |
7.01 |
6.42 |
6.23 |
6.23 |
6.23 |
| pHf |
5.38 |
4.34 |
4.39 |
5.22 |
4.66 |
4.68 |
| apH |
0.27 |
0.52 |
0.27 |
0.99 |
0.37 |
6.66×10-2
|
| bpH |
0.31 |
0.26 |
0.24 |
0.31 |
0.60 |
0.34 |
|
R2pH
|
0.9947 |
0.9935 |
0.9941 |
0.9881 |
0.9950 |
0.9968 |
| SSD–pH |
1.44×10-2
|
4.43×10-2
|
3.59×10-2
|
1.26×10-2
|
2.39×10-2
|
9.77×10-3
|
| TSi |
48.18 |
22.56 |
20.32 |
48.18 |
25.58 |
19.63 |
| TSf |
44.43 |
17.17 |
6.33 |
43.60 |
23.00 |
12.85 |
| bTS |
8.36×10-2
|
0.16 |
8.57×10-2
|
0.11 |
0.11 |
0.13 |
|
R2TS
|
0.9515 |
0.9731 |
0.8823 |
0.9661 |
0.9684 |
0.6789 |
| SSD–TS |
1.23 |
0.92 |
29.48 |
0.48 |
0.16 |
23.89 |
| TNi |
1.19 |
0.48 |
3.33 |
1.06 |
0.44 |
3.06 |
| TNf |
1.00 |
0.34 |
2.68 |
0.92 |
0.39 |
2.22 |
| bTN |
0.05 |
0.15 |
0.08 |
0.10 |
0.12 |
0.24 |
|
R2TN
|
0.9790 |
0.9467 |
0.9336 |
0.9365 |
0.9491 |
0.7400 |
| SSD–TN |
1.35×10-3
|
1.20×10-3
|
4.66×10-2
|
1.93×10-3
|
1.76×10-4
|
0.40 |
Table 11.
Estimated parameter values for models (16), (17), (18), (19), (20), and (21), together with the coefficient of determination (R²) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of Lact. casei CECT 4043 and Pediococcus acidilactici NRRL B–562 in concentrated whey (CW), diluted whey (DW), and MRS broth.
Table 11.
Estimated parameter values for models (16), (17), (18), (19), (20), and (21), together with the coefficient of determination (R²) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of Lact. casei CECT 4043 and Pediococcus acidilactici NRRL B–562 in concentrated whey (CW), diluted whey (DW), and MRS broth.
| |
|
Culture medium/Strain |
| Parameter |
CW/CECT 4043 |
DW/CECT 4043 |
MRS/CECT 4043 |
CW/NRRL B–562 |
DW/NRRL B–562 |
MRS/NRRL B–562 |
| KX |
0.21 |
0.34 |
2.22 |
0.10 |
0.18 |
1.76 |
|
aX
|
25.52 |
15.90 |
221.30 |
4.03 |
8.19 |
175.01 |
| bpH-X |
0.22 |
0.97 |
0.22 |
0.32 |
0.49 |
0.41 |
|
R2X
|
0.9951 |
0.9825 |
0.9961 |
0.9883 |
0.9916 |
0.9955 |
| SSD–X |
2.20×10-4
|
2.89×10-3
|
2.39×10-2
|
1.05×10-4
|
3.07×10-4
|
1.96×10-2
|
| KLA |
1.47 |
2.23 |
21.89 |
0.45 |
0.83 |
4.28 |
| aLA |
49.06 |
47.71 |
49.10 |
49.07 |
49.06 |
50.08 |
| bX-LA |
0.25 |
0.46 |
0.45 |
0.31 |
0.77 |
0.51 |
|
R2LA
|
0.9643 |
0.9901 |
0.9910 |
0.9944 |
0.9746 |
0.9936 |
| SSD–LA |
0.10 |
5.85×10-2
|
7.21 |
9.51×10-4
|
2.66×10-2
|
0.18 |
| KTAA |
2.08 |
3.14 |
11.98 |
51.70 |
57.93 |
493.32 |
| aTAA |
23.30 |
43.47 |
49.05 |
49.49 |
49.11 |
50.01 |
| bX-TAA |
2.07×10-3
|
0.44 |
1.62×10-2
|
0.28 |
0.73 |
0.13 |
| bpH-TAA |
0.21 |
1.25×10-3
|
0.21 |
3.12×10-2
|
2.30×10-2
|
0.31 |
|
R2TAA
|
0.9972 |
0.9840 |
0.9979 |
0.9964 |
0.9609 |
0.9963 |
| SSD–TAA |
1.0×10-2
|
0.18 |
0.44 |
9.95 |
143.32 |
1182.13 |
| pHi |
7.00 |
7.00 |
6.42 |
6.23 |
6.23 |
6.23 |
| pHf |
5.37 |
4.34 |
4.39 |
5.22 |
4.66 |
4.75 |
| apH |
0.71 |
0.86 |
0.36 |
0.59 |
0.71 |
3.83×10-2
|
| bLA-pH |
0.31 |
0.57 |
0.31 |
0.80 |
1.16 |
0.36 |
|
R2pH
|
0.9910 |
0.9863 |
0.9975 |
0.9659 |
0.9902 |
0.9889 |
| SSD–pH |
2.85×10-2
|
8.64×10-2
|
1.13×10-2
|
4.17×10-2
|
4.12×10-2
|
4.10×10-2
|
| TSi |
48.18 |
22.56 |
20.32 |
48.18 |
25.58 |
19.63 |
| TSf |
44.43 |
17.17 |
6.33 |
43.60 |
23.00 |
12.85 |
| bX-TS |
0.24 |
0.41 |
0.46 |
0.30 |
0.48 |
0.60 |
|
R2TS
|
0.9833 |
0.9582 |
0.9973 |
0.9988 |
0.9828 |
0.9867 |
| SSD–TS |
0.24 |
0.84 |
1.32 |
1.76×10-2
|
0.14 |
0.80 |
| TNi |
1.19 |
0.48 |
3.33 |
1.06 |
0.44 |
3.06 |
| TNf |
1.00 |
0.34 |
2.68 |
0.92 |
0.39 |
2.22 |
| bX-TN |
0.23 |
0.45 |
0.49 |
0.30 |
0.48 |
0.54 |
|
R2TN
|
0.9886 |
0.9922 |
0.9952 |
0.9997 |
0.9918 |
0.9693 |
| SSD–TN |
5.00×10-4
|
1.67×10-4
|
2.60×10-3
|
7.14×10-6
|
4.50×10-5
|
2.27×10-2
|
Table 12.
Estimated parameter values for models (1), (3), and (6), together with the coefficient of determination (R²) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of Pediococcus acidilactici LB42–923, Lactococcus lactis subsp. lactis ATCC 11454, Leuconostoc carnosum Lm1, and Lactobacillus sakei LB 706 in concentrated TGE broth.
Table 12.
Estimated parameter values for models (1), (3), and (6), together with the coefficient of determination (R²) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of Pediococcus acidilactici LB42–923, Lactococcus lactis subsp. lactis ATCC 11454, Leuconostoc carnosum Lm1, and Lactobacillus sakei LB 706 in concentrated TGE broth.
| |
Strain |
| Parameter |
LB42–923 |
ATCC 11454 |
Lm1 |
LB 706 |
|
| KX |
3.19 |
1.08 |
1.61 |
1.91 |
|
|
aX
|
155.53 |
34.86 |
58.70 |
100.37 |
|
| bX |
1.07 |
1.15 |
0.62 |
0.96 |
|
|
R2X
|
0.9920 |
0.9786 |
0.9845 |
0.9939 |
|
| SSD–X |
0.16 |
3.25×10-2
|
6.56×10-2
|
3.28×10-2
|
|
| KTAA |
28.58 |
14.12 |
26.79 |
18.36 |
|
| aTAA |
548.61 |
306.07 |
285.56 |
1363.75 |
|
| bTAA |
0.93 |
0.94 |
0.69 |
1.33 |
|
|
R2TAA
|
0.9774 |
0.9886 |
0.9902 |
0.9948 |
|
| SSD–TAA |
34.35 |
4.89 |
13.21 |
4.86 |
|
| pHi |
6.65 |
6.55 |
6.55 |
6.65 |
|
| pHf |
3.61 |
3.90 |
3.90 |
3.82 |
|
| apH |
5.15 |
2.98×10-3
|
0.17 |
5.06×10-2
|
|
| bpH |
1.27 |
1.37 |
0.34 |
0.55 |
|
|
R2pH
|
0.9945 |
0.9973 |
0.9948 |
0.9985 |
|
| SSD–pH |
8.22×10-2
|
3.61×10-2
|
4.59×10-2
|
1.57×10-2
|
|
Table 13.
Estimated parameter values for models (16), (18), and (22), together with the coefficient of determination (R²) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of Pediococcus acidilactici LB42–923, Lactococcus lactis subsp. lactis ATCC 11454, Leuconostoc carnosum Lm1, and Lactobacillus sakei LB 706 in concentrated TGE broth.
Table 13.
Estimated parameter values for models (16), (18), and (22), together with the coefficient of determination (R²) and the sum of squared differences (SSD) between predicted and experimental values, for the batch growth of Pediococcus acidilactici LB42–923, Lactococcus lactis subsp. lactis ATCC 11454, Leuconostoc carnosum Lm1, and Lactobacillus sakei LB 706 in concentrated TGE broth.
| |
Strain |
| Parameter |
LB42–923 |
ATCC 11454 |
Lm1 |
LB 706 |
|
| KX |
3.19 |
1.11 |
1.62 |
1.89 |
|
|
aX
|
158.53 |
35.92 |
59.00 |
99.65 |
|
| bpH-X |
0.94 |
1.18 |
0.37 |
0.56 |
|
|
R2X
|
0.9964 |
0.9911 |
0.9975 |
0.9996 |
|
| SSD–X |
8.36×10-2
|
1.25×10-2
|
1.23×10-2
|
2.77×10-3
|
|
| KTAA |
29.07 |
14.10 |
26.80 |
18.37 |
|
| aTAA |
558.06 |
541.42 |
558.06 |
6121.68 |
|
| bX-TAA |
0.37 |
0.38 |
0.65 |
0.94 |
|
| bpH-TAA |
0.57 |
0.73 |
5.70×10-2
|
5.70×10-2
|
|
|
R2TAA
|
0.9894 |
0.9999 |
0.9974 |
0.9821 |
|
| SSD–TAA |
0.18 |
4.59×10-2
|
3.55 |
12.62 |
|
| pHi |
6.65 |
6.55 |
6.55 |
6.65 |
|
| pHf |
3.61 |
3.90 |
3.90 |
3.82 |
|
| apH |
3.00×10-2
|
8.48×10-3
|
0.12 |
4.86×10-2
|
|
| bX-pH |
1.02 |
1.09 |
0.76 |
1.11 |
|
|
R2pH
|
0.9947 |
0.9978 |
0.9899 |
0.9947 |
|
| SSD–pH |
7.10×10-2
|
4.26×10-2
|
0.11 |
7.11×10-2
|
|