Submitted:
25 February 2026
Posted:
25 February 2026
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
2. Osmotic Stress in Halophilic Bacteria
3. Thermal Stress in Thermophilic Bacteria
4. pH Extremes: Acidophiles and Alkaliphiles
5. Comparative Synthesis of Stress Networks Across Extremophiles
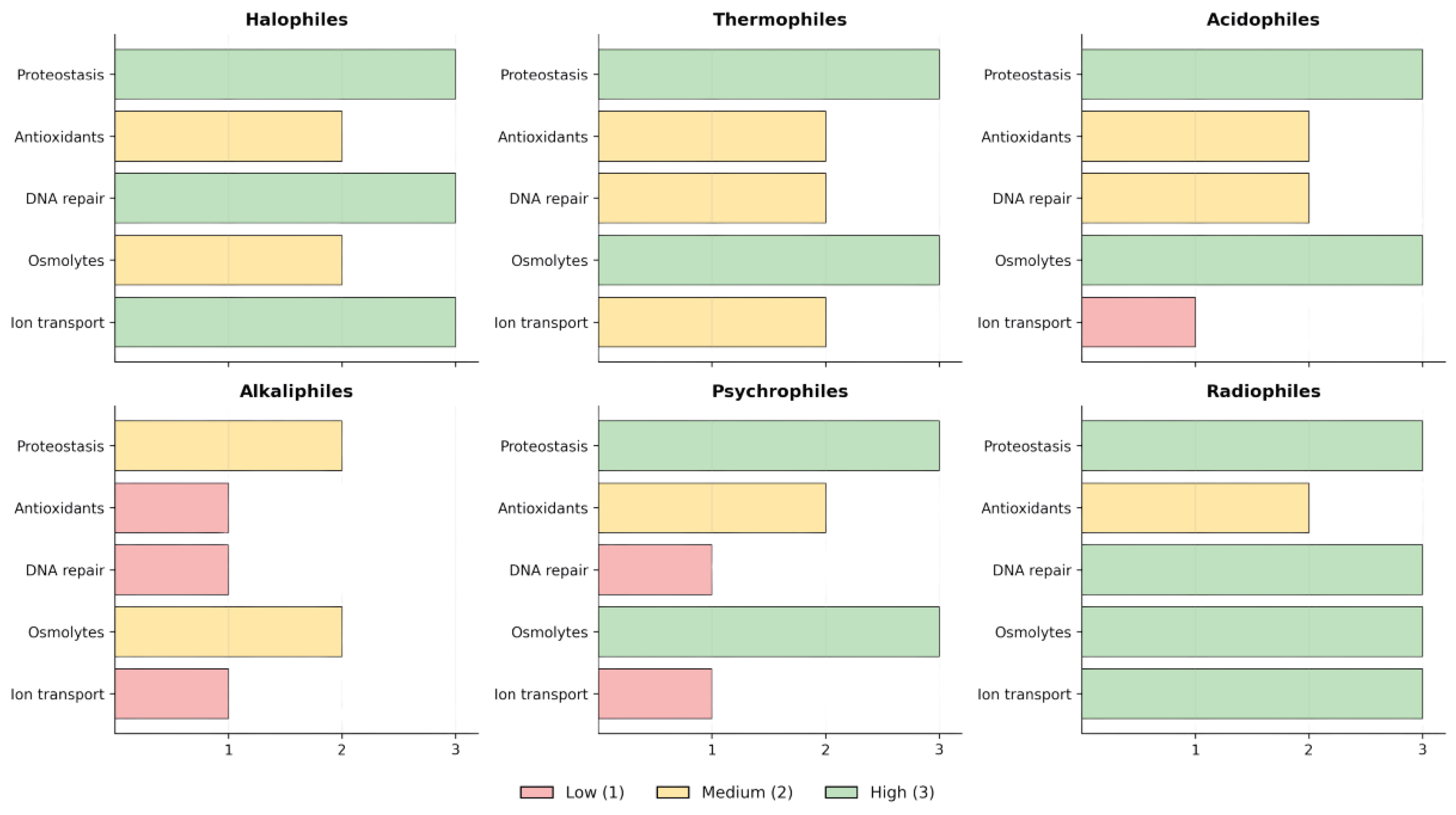
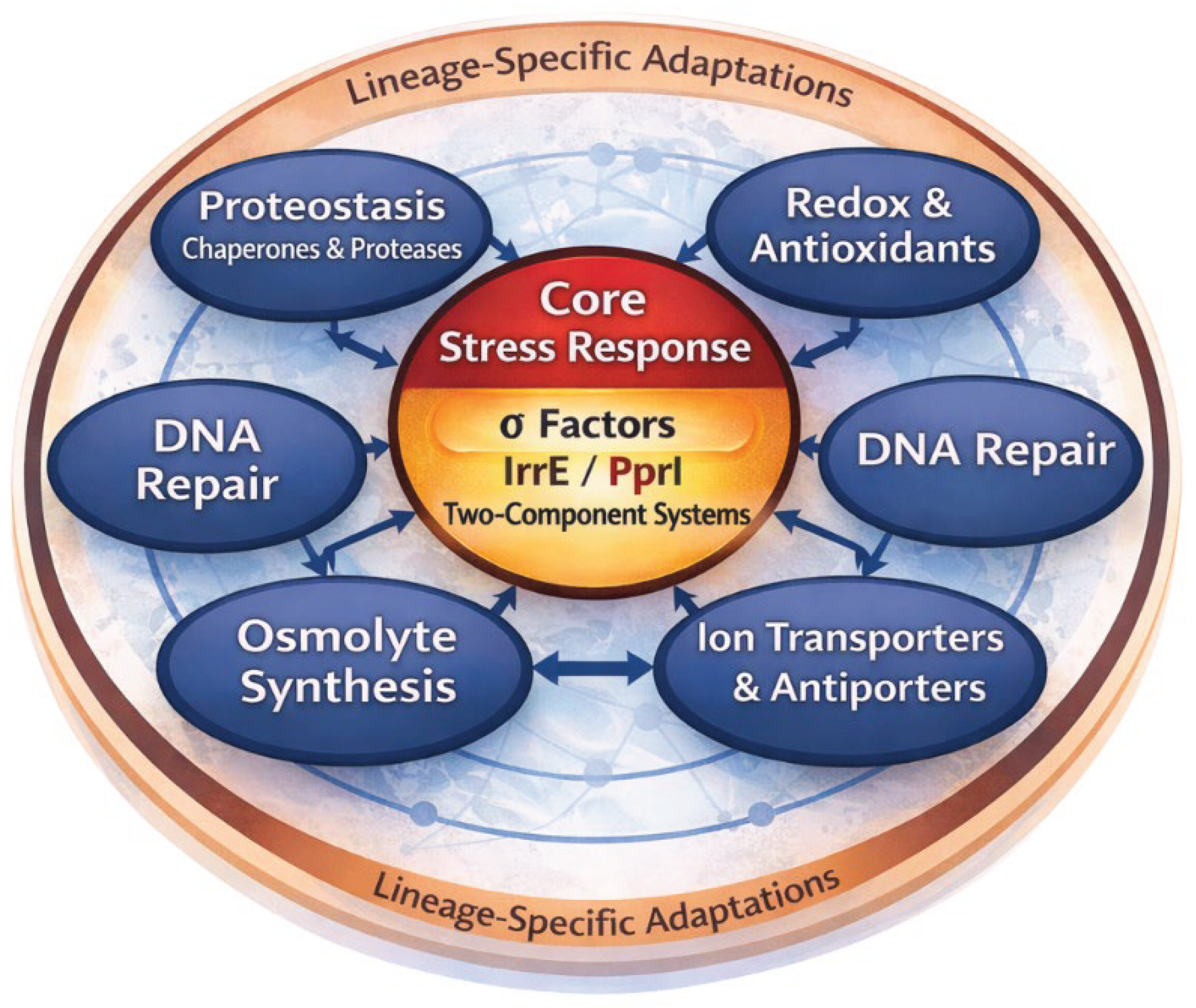
6. Oxidative Stress as a Cross-Cutting Axis
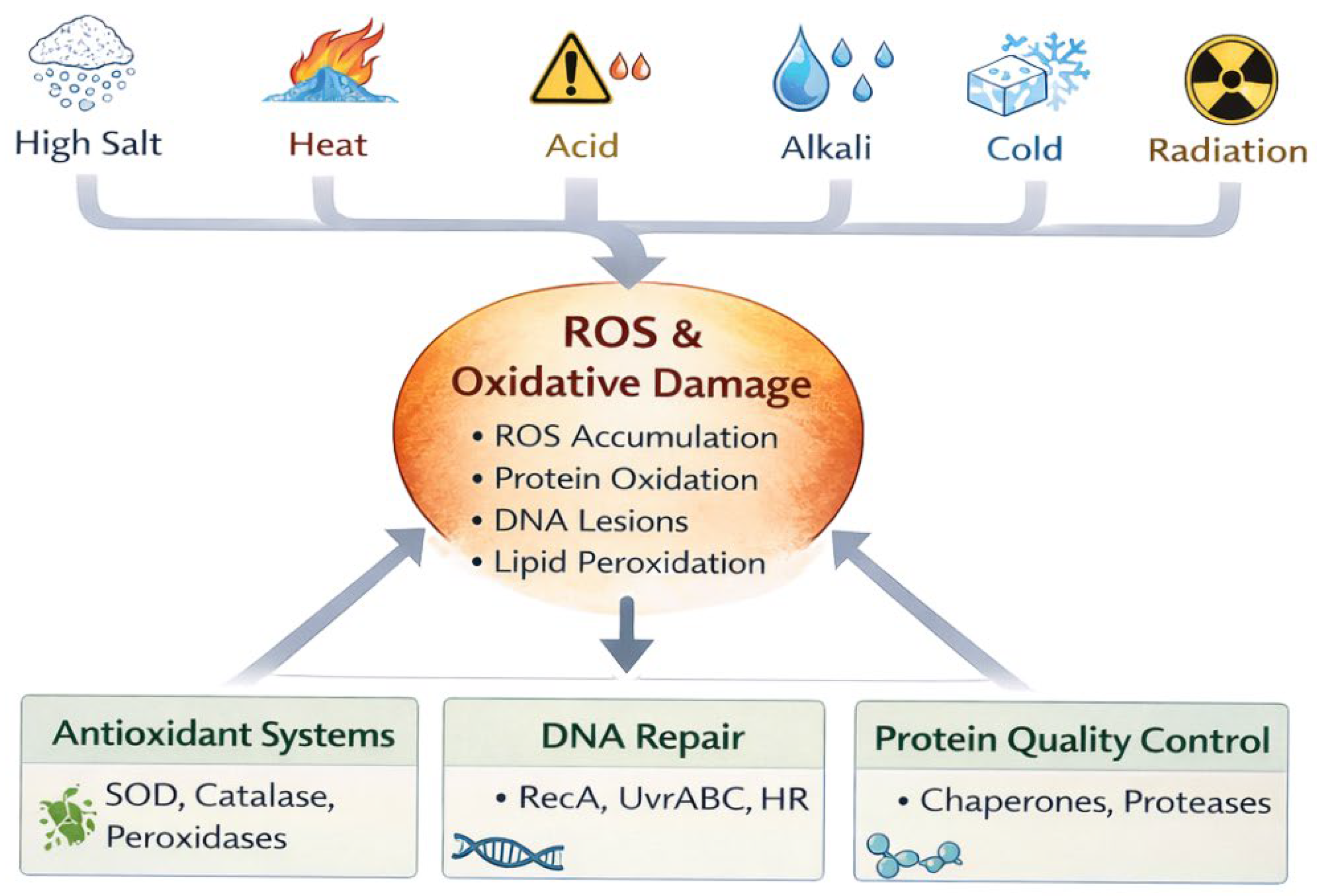
7. Mechanistic Insights from Psychrophiles and Radiophiles
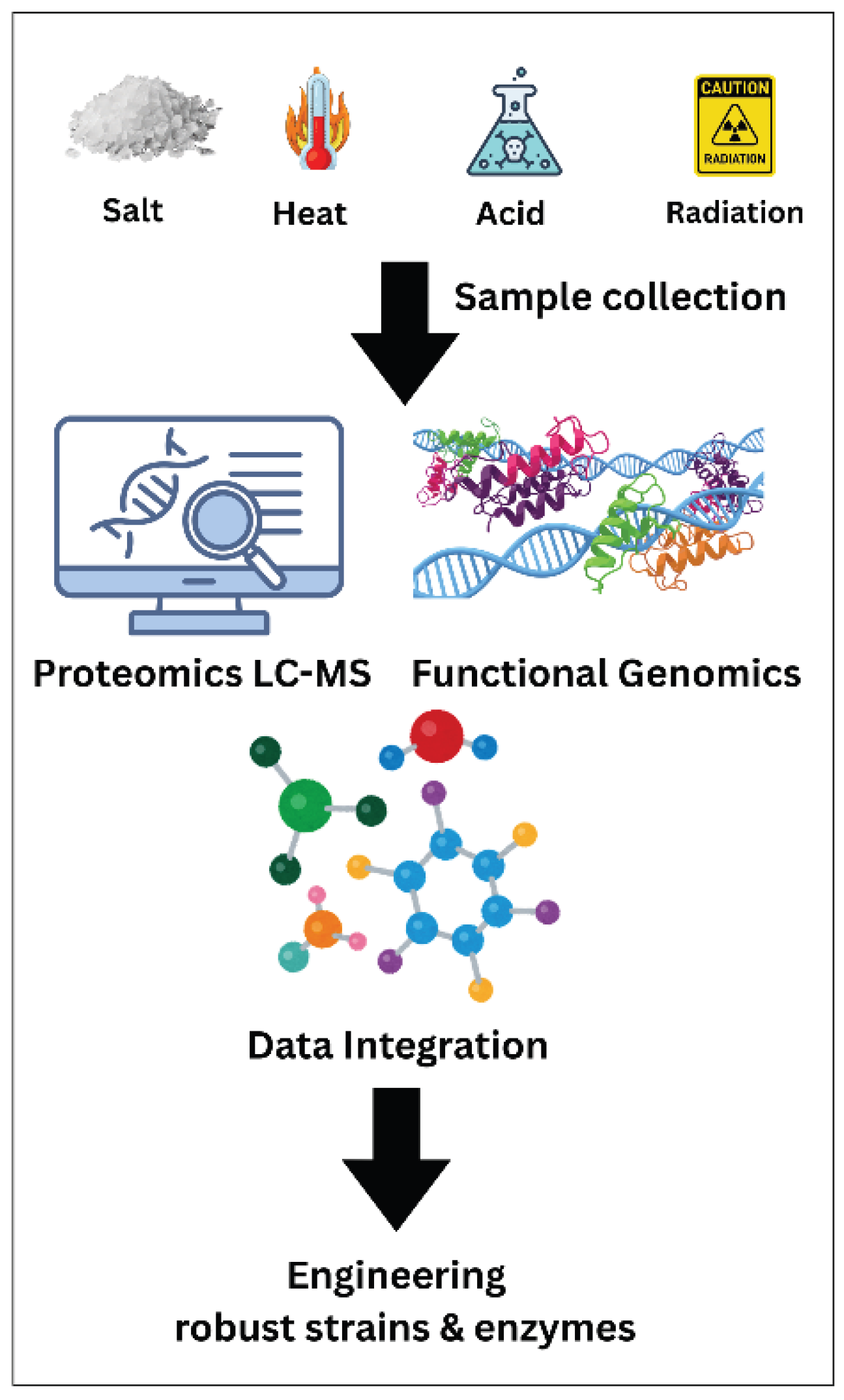
Radiophile Adaptations (Radiation-Resistant Extremophiles)
8. Conclusions and Future Perspectives
Author Contributions
Conflicts of Interest
References
- Rubiano-Labrador, C.; Bland, C.; Miotello, G.; Armengaud, J.; Baena, S. Salt Stress Induced Changes in the Exoproteome of the Halotolerant Bacterium Tistlia consotensis Deciphered by Proteogenomics. PloS one 2015, 10, e0135065. [Google Scholar] [CrossRef]
- Remonsellez, F.; Castro-Severyn, J.; Pardo-Esté, C.; Aguilar, P.; Fortt, J.; Salinas, C.; Barahona, S.; León, J.; Fuentes, B.; Areche, C.; Hernández, K. L.; Aguayo, D.; Saavedra, C. P. Characterization and Salt Response in Recurrent Halotolerant Exiguobacterium sp. SH31 Isolated From Sediments of Salar de Huasco, Chilean Altiplano. Frontiers in microbiology 2018, 9, 2228. [Google Scholar] [CrossRef]
- Purohit, H. V. Nucleoid-associated proteins: molecular mechanisms in microbial adaptation. World journal of microbiology & biotechnology 2025, 41, 277. [Google Scholar] [CrossRef]
- Valdez-Nuñez, L. F.; Kappler, A.; Ayala-Muñoz, D.; Chávez, I. J.; Mansor, M. Acidophilic sulphate-reducing bacteria: Diversity, ecophysiology, and applications. Environmental microbiology reports 2024, 16, e70019. [Google Scholar] [CrossRef]
- Matarredona, L.; Zafrilla, B.; Camacho, M.; Bonete, M. J.; Esclapez, J. Understanding the tolerance of halophilic archaea to stress landscapes. Environmental microbiology reports 2024, 16, e70039. [Google Scholar] [CrossRef]
- Altinisik Kaya, F. E.; Avci, F. G.; Sayar, N. A.; Kazan, D.; Sayar, A. A.; Sariyar Akbulut, B. What Are the Multi-Omics Mechanisms for Adaptation by Microorganisms to High Alkalinity? A Transcriptomic and Proteomic Study of a Bacillus Strain with Industrial Potential. Omics: a journal of integrative biology 2018, 22, 717–732. [Google Scholar] [CrossRef] [PubMed]
- Atakav, Y.; Pinar, O.; Kazan, D. Investigation of the Physiology of the Obligate Alkaliphilic Bacillus marmarensis GMBE 72T Considering Its Alkaline Adaptation Mechanism for Poly(3-hydroxybutyrate) Synthesis. Microorganisms 2021, 9, 462. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Wang, F.; Chen, H.; Yang, Z.; Ning, Y.; Chang, C.; Yang, D. New Insights into Radio-Resistance Mechanism Revealed by (Phospho)Proteome Analysis of Deinococcus Radiodurans after Heavy Ion Irradiation. International journal of molecular sciences 2023, 24, 14817. [Google Scholar] [CrossRef]
- Han, J. M.; Mwiti, G.; Yeom, S. J.; Lim, J.; Kim, W. S.; Lim, S.; Lim, S. T.; Byun, E. B. Radiation-Resistant Bacteria Deinococcus radiodurans-Derived Extracellular Vesicles as Potential Radioprotectors. Advanced healthcare materials 2025, 14, e2403192. [Google Scholar] [CrossRef]
- Fongaro, G.; Maia, G. A.; Rogovski, P.; Cadamuro, R. D.; Lopes, J. C.; Moreira, R. S.; Camargo, A. F.; Scapini, T.; Stefanski, F. S.; Bonatto, C.; Marques Souza, D. S.; Stoco, P. H.; Duarte, R. T. D.; Cabral da Cruz, A. C.; Wagner, G.; Treichel, H. Extremophile Microbial Communities and Enzymes for Bioenergetic Application Based on Multi-Omics Tools. Current genomics 2020, 21, 240–252. [Google Scholar] [CrossRef] [PubMed]
- Nie, T.; Wang, L.; Liu, Y.; Fu, S.; Wang, J.; Cui, K.; Wang, L. A Halophilic Bacterium for Bioremediation of Saline-Alkali Land: The Triadic and Synergetic Response Mechanism of Oceanobacillus picturae DY09 to Salt Stress. Microorganisms 2025, 13, 1474. [Google Scholar] [CrossRef]
- Kung, C.; Martinac, B.; Sukharev, S. Mechanosensitive channels in microbes. Annual review of microbiology 2010, 64, 313–329. [Google Scholar] [CrossRef] [PubMed]
- Belin, B. J.; Busset, N.; Giraud, E.; Molinaro, A.; Silipo, A.; Newman, D. K. Hopanoid lipids: from membranes to plant-bacteria interactions. Nature reviews. Microbiology 2018, 16, 304–315. [Google Scholar] [CrossRef]
- Yin, L.; Xue, Y.; Ma, Y. Global Microarray Analysis of Alkaliphilic Halotolerant Bacterium Bacillus sp. N16-5 Salt Stress Adaptation. PloS one 2015, 10, e0128649. [Google Scholar] [CrossRef]
- Vandrich, J.; Pfeiffer, F.; Alfaro-Espinoza, G.; Kunte, H. J. Contribution of mechanosensitive channels to osmoadaptation and ectoine excretion in Halomonas elongata. Extremophiles: life under extreme conditions 2020, 24, 421–432. [Google Scholar] [CrossRef]
- Oren, A. Microbial life at high salt concentrations: phylogenetic and metabolic diversity. Saline systems 2008, 4, 2. [Google Scholar] [CrossRef]
- Enuh, B. M.; Aytar Çelik, P. Genome Analysis of Halomonas elongata Strain 153B and Insights Into Polyhydroxyalkanoate Synthesis and Adaptive Mechanisms to High Saline Environments. Current microbiology 2022, 80, 18. [Google Scholar] [CrossRef]
- Dildar, T.; Cui, W.; Ikhwanuddin, M.; amp; Ma, H. Aquatic Organisms in Response to Salinity Stress: Ecological Impacts, Adaptive Mechanisms, and Resilience Strategies. Biology 2025, 14, 667. [Google Scholar] [CrossRef]
- Chan, Y. F.; Chen, C. Y.; Lu, C. Y.; Tu, Y. C.; Tandon, K.; Shikina, S.; Tang, S. L. A first insight into the heat-induced changes in proteomic profiles of the coral symbiotic bacterium Endozoicomonas montiporae. Frontiers in Marine Science 2022, 9, 808132. [Google Scholar] [CrossRef]
- Sato, Y.; Okano, K.; Honda, K. Effects of small heat shock proteins from thermotolerant bacteria on the stress resistance of Escherichia coli to temperature, pH, and hyperosmolarity. Extremophiles: life under extreme conditions 2024, 28, 12. [Google Scholar] [CrossRef] [PubMed]
- Izquierdo-Fiallo, K.; Muñoz-Villagrán, C.; Orellana, O.; Sjoberg, R.; Levicán, G. Comparative genomics of the proteostasis network in extreme acidophiles. PloS one 2023, 18, e0291164. [Google Scholar] [CrossRef] [PubMed]
- Zhao, N.; Jiao, L.; Xu, J.; Zhang, J.; Qi, Y.; Qiu, M.; Wei, X.; Fan, M. Integrated transcriptomic and proteomic analysis reveals the response mechanisms of Alicyclobacillus acidoterrestris to heat stress. Food research international (Ottawa, Ont.) 2022, 151, 110859. [Google Scholar] [CrossRef]
- Shih, T. W.; Pan, T. M. Stress responses of thermophilic Geobacillus sp. NTU 03 caused by heat and heat-induced stress. Microbiological research 2011, 166, 346–359. [Google Scholar] [CrossRef] [PubMed]
- Ranawat, P.; amp; Rawat, S. Stress response physiology of thermophiles. Archives of microbiology 2017, 199, 391–414. [Google Scholar] [CrossRef]
- Belnap, C. P.; Pan, C.; VerBerkmoes, N. C.; Power, M. E.; Samatova, N. F.; Carver, R. L.; Hettich, R. L.; Banfield, J. F. Cultivation and quantitative proteomic analyses of acidophilic microbial communities. The ISME journal 2010, 4, 520–530. [Google Scholar] [CrossRef] [PubMed]
- Cortez, D.; Neira, G.; González, C.; Vergara, E.; Holmes, D. S. A Large-Scale Genome-Based Survey of Acidophilic Bacteria Suggests That Genome Streamlining Is an Adaption for Life at Low pH. Frontiers in microbiology 2022, 13, 803241. [Google Scholar] [CrossRef]
- Somayaji, A.; Dhanjal, C. R.; Lingamsetty, R.; Vinayagam, R.; Selvaraj, R.; Varadavenkatesan, T.; Govarthanan, M. An insight into the mechanisms of homeostasis in extremophiles. Microbiological research 2022, 263, 127115. [Google Scholar] [CrossRef]
- Huanca-Juarez, J.; Nascimento-Silva, E. A.; Silva, N. H.; Silva-Rocha, R.; Guazzaroni, M. E. Identification and functional analysis of novel protein-encoding sequences related to stress-resistance. Frontiers in microbiology 2023, 14, 1268315. [Google Scholar] [CrossRef]
- Marzban, G.; Tesei, D. The Extremophiles: Adaptation Mechanisms and Biotechnological Applications. Biology 2025, 14, 412. [Google Scholar] [CrossRef]
- Purohit, H. V.; Chakraborty, J. Metagenomic approaches for studying ubiquitous yet diverse nucleoid associated proteins in microbial communities: challenges and advances. World journal of microbiology & biotechnology 2025, 41, 383. [Google Scholar] [CrossRef]
- Baraúna, R. A.; Freitas, D. Y.; Pinheiro, J. C.; Folador, A. R.; Silva, A. A Proteomic Perspective on the Bacterial Adaptation to Cold: Integrating OMICs Data of the Psychrotrophic Bacterium Exiguobacterium antarcticum B7. Proteomes 2017, 5, 9. [Google Scholar] [CrossRef] [PubMed]
- Boas Lichty, K. E.; Gregory, G. J.; Boyd, E. F. NhaR, LeuO, and H-NS are part of an expanded regulatory network for ectoine biosynthesis expression. Applied and Environmental Microbiology 2023, 89, e00479-23. [Google Scholar] [CrossRef]
- García-Descalzo, L.; García-López, E.; Cid, C. Comparative proteomic analysis of psychrophilic vs. mesophilic bacterial species reveals different strategies to achieve temperature adaptation. Frontiers in Microbiology 2022, 13, 841359. [Google Scholar] [CrossRef]
- Baker-Austin, C.; Dopson, M. Life in acid: pH homeostasis in acidophiles. Trends in microbiology 2007, 15, 165–171. [Google Scholar] [CrossRef]
- Padan, E.; Bibi, E.; Ito, M.; Krulwich, T. A. Alkaline pH homeostasis in bacteria: new insights. Biochimica et biophysica acta (BBA)-biomembranes 2005, 1717, 67–88. [Google Scholar] [CrossRef]
- Kumar, S.; Suyal, D. C.; Yadav, A.; Shouche, Y.; Goel, R. Psychrophilic Pseudomonas helmanticensis proteome under simulated cold stress. Cell Stress and Chaperones 2020, 25, 1025–1032. [Google Scholar] [CrossRef]
- Qi, H. Z.; Wang, W. Z.; He, J. Y.; Ma, Y.; Xiao, F. Z.; He, S. Y. Antioxidative system of Deinococcus radiodurans. Research in microbiology 2020, 171, 45–54. [Google Scholar] [CrossRef]
- Jomova, K.; Alomar, S. Y.; Alwasel, S. H.; Nepovimova, E.; Kuca, K.; Valko, M. Several lines of antioxidant defense against oxidative stress: antioxidant enzymes, nanomaterials with multiple enzyme-mimicking activities, and low-molecular-weight antioxidants. Archives of toxicology 2024, 98, 1323–1367. [Google Scholar] [CrossRef]
- Martínez-Espinosa, R. M. Halocins and C50 Carotenoids from Haloarchaea: Potential Natural Tools against Cancer. Marine drugs 2024, 22, 448. [Google Scholar] [CrossRef] [PubMed]
- M, S.; R. P., N; Chakraborty, A.; Rajendrasozhan, S. Proteomic profiling of Deinococcus radiodurans with response to thioredoxin reductase inhibitor and ionizing radiation treatment. Journal of proteomics 2022, 267, 104697. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Li, N.; Zhang, Y. The radioresistant and survival mechanisms of Deinococcus radiodurans. Radiation Medicine and Protection 2023, 4, 70–79. [Google Scholar] [CrossRef]
- Orban, K.; Finkel, S. E. Dps Is a Universally Conserved Dual-Action DNA-Binding and Ferritin Protein. Journal of bacteriology 2022, 204, e0003622. [Google Scholar] [CrossRef]
- Vauclare, P.; Wulffelé, J.; Lacroix, F.; Servant, P.; Confalonieri, F.; Kleman, J. P.; Bourgeois, D.; Timmins, J. Stress-induced nucleoid remodeling in Deinococcus radiodurans is associated with major changes in Heat Unstable (HU) protein dynamics. Nucleic acids research 2024, 52, 6406–6423. [Google Scholar] [CrossRef]
- Ge, P.; Rashid, F. M.; Dame, R. T. The role of nucleoid-associated proteins in mediating responses to environmental changes. Current opinion in microbiology 2025, 87, 102628. [Google Scholar] [CrossRef] [PubMed]
- Purohit, HV; Kanojia, H; Pandya, V; Nalla, Y; Raval, KY; Kapadiya, KM; Kamdar, JH. Soil as a host to the biotic community. In Soil Microbiome of the cold habitats: trends and applications; Gupta, P, Shahnawaz, M, Eds.; CRC, 2023; pp. pp 17–30. [Google Scholar] [CrossRef]
- Mogk, A.; Tomoyasu, T.; Goloubinoff, P.; Rüdiger, S.; Röder, D.; Langen, H.; Bukau, B. Identification of thermolabile Escherichia coli proteins: prevention and reversion of aggregation by DnaK and ClpB. The EMBO journal 1999. [Google Scholar] [CrossRef] [PubMed]
- Figaj, D. The role of heat shock protein (Hsp) chaperones in environmental stress adaptation and virulence of plant pathogenic bacteria. International Journal of Molecular Sciences 2025, 26, 528. [Google Scholar] [CrossRef]
- Deng, M.; Xu, X.; Yang, Z.; Lu, K. CSP15 attenuates chlorpyrifos toxicity through sequestration and bioactivation suppression in Nilaparvata lugens. Insect Biochemistry and Molecular Biology 2025, 104453. [Google Scholar] [CrossRef]
- Lu, H.; Hua, Y. PprI: the key protein in response to DNA damage in Deinococcus. Frontiers in Cell and Developmental Biology 2021, 8, 609714. [Google Scholar] [CrossRef]
- Fatima, K.; Naqvi, F.; amp; Younas, H. A review: Molecular chaperone-mediated folding,unfolding and disaggregation of expressed recombinant proteins. Cell Biochemistry and Biophysics 2021, 79, 153–174. [Google Scholar] [CrossRef] [PubMed]
- Weinisch, L.; Kühner, S.; Roth, R.; Grimm, M.; Roth, T.; Netz, D. J.; Filker, S. Identification of osmoadaptive strategies in the halophile, heterotrophic ciliate Schmidingerothrix salinarum. PLoS Biology 2018, 16, e2003892. [Google Scholar] [CrossRef]
- Silva, R.; Gonçalves, T.; Morone, J.; Moreira, G. A.; Morais, J.; Hentschke, G. S.; Lopes, G.; amp. Pigments profile and antioxidant potential of extremophile cyanobacteria isolated from the Mexican Volcanic Lake Chichonal. Algal Research 2024, 81, 103578. [Google Scholar] [CrossRef]
- Kanekar, P.P.; Kanekar, S.P. Radiophilic, Radioresistant, and Radiotolerant Microorganisms. In Diversity and Biotechnology of Extremophilic Microorganisms from India. Microorganisms for Sustainability; Springer: Singapore, 2022. [Google Scholar] [CrossRef]
- Yusof, N. A.; Hashim, N. H. F.; Bharudin, I. Cold Adaptation Strategies and the Potential of Psychrophilic Enzymes from the Antarctic Yeast, Glaciozyma antarctica PI12. Journal of fungi (Basel, Switzerland) 2021, 7, 528. [Google Scholar] [CrossRef] [PubMed]
- Son, Y.; Min, J.; Shin, Y.; Park, W. Morphological and physiological adaptations of psychrophilic Pseudarthrobacter psychrotolerans YJ56 under temperature stress. Scientific reports 2023, 13, 14970. [Google Scholar] [CrossRef]
- Basu, B. The radiophiles of Deinococcaceae family: Resourceful microbes for innovative biotechnological applications. Current research in microbial sciences 2022, 3, 100153. [Google Scholar] [CrossRef]
- Minton, K. W. DNA repair in the extremely radioresistant bacterium Deinococcus radiodurans. Molecular microbiology 1994, 13, 9–15. [Google Scholar] [CrossRef] [PubMed]
| Extremophile type | Typical conditions | Representative species | Primary stressors / notes | References |
| Halophiles | High salt (≈2–5 M NaCl) | Halobacterium salinarum, Halomonas spp. | Ionic/osmotic stress; reliance on compatible solutes and specialized Na⁺ transport. | [32] |
| Thermophiles / Hyperthermophiles | 60–120 °C | Thermus thermophilus, Pyrococcus spp. | Protein thermostability, membrane adaptations, heat-shock systems and ATP-dependent proteases. | [33] |
| Acidophiles | pH ~0–3 | Acidithiobacillus ferrooxidans, Ferroplasma | Proton toxicity, metal stress; low-permeability membranes and proton export systems. | [34] |
| Alkaliphiles | pH ≈9–12 | Bacillus halodurans, Halomonas alkaliphila | Proton scarcity → Na⁺-based bioenergetics and multi-subunit antiporters (Mrp/Mnh). | [35] |
| Psychrophiles | ~−5 to 10 °C | Psychromonas, Antarctic Pseudomonas spp. | Cold-adapted enzymes, RNA chaperones (Csps), membrane fluidity and cryoprotectants. | [36] |
| Radiophiles / Desiccation-resistant | High ionizing radiation / desiccation | Deinococcus radiodurans | Exceptional DNA-repair systems, PprI/DdrO regulatory circuit, strong antioxidant defenses. | [37] |
| Extremophile group | Proteomics signature (selected) | Functional genomics / genetic evidence | Takeaway (essential network components) | References |
| Halophiles | Strong induction of ectoine pathway enzymes; Na⁺ transporters | Knockouts or regulatory mutants in ectoine pathways or antiporters alter salt tolerance and ectoine production. | Compatible-solute synthesis + Na⁺ antiporters tightly coupled to proteostasis and metabolism. | [32] |
| Thermophiles | High levels of HSPs (DnaK/GroEL) and ATP-dependent proteases | Deletion or depletion of groESL, clpB, or lon reduces thermotolerance. | Proteostasis network (folding + targeted proteolysis) is essential for high-T survival. | [46,47] |
| Acidophiles | Upregulated proton exporters and low-permeability membrane components | Mutational studies show loss of pH tolerance with impaired proton export / membrane integrity genes. | Proton homeostasis + iron handling + ROS defenses shape acid tolerance. | [34] |
| Alkaliphiles | Elevated Na⁺-coupled ATP synthase subunits and Mrp/Mnh antiporters | Genetic disruption of multi-subunit antiporters compromises growth at high pH. | Na⁺-bioenergetics + antiport complexes are core to alkaline adaptation. | [35] |
| Psychrophiles | Upregulation of Csps and flexible metabolic enzymes | csp mutants show impaired cold growth; proteomics shows increased chaperones & desaturases. | RNA chaperones + membrane fluidity adjustments are central cold strategies. | [36,48] |
| Radiophiles | Enriched DNA-repair proteins & antioxidants | PprI / DdrO and RecA systems are essential; mutants show radiation sensitivity. | Robust DNA repair + antioxidant networks underpin extreme radio/desiccation resistance. | [49] |
| Protein class / module | Functional role | Extremophiles with strong proteomic induction | Typical examples (proteins) | References |
| Molecular chaperones | Prevent protein misfolding & refold damaged proteins | Thermophiles, halophiles, psychrophiles | DnaK (Hsp70), GroEL/GroES, small HSPs. | [33,50] |
| ATP-dependent proteases / proteolysis | Remove irreversibly damaged proteins; proteostasis | Thermophiles, radiophiles | Lon, ClpP/ClpX, FtsH. | [46] |
| Compatible solute synthesis & uptake | Osmotic balance / protein stabilisation | Halophiles, some psychrophiles | Ectoine biosynthetic enzymes, glycine-betaine transporters. | [32] |
| Ion transporters / antiporters | pH & ionic homeostasis | Halophiles, alkaliphiles, acidophiles | Na⁺/H⁺ antiporters (Mrp/Mnh), K⁺ transporters. | [51] |
| DNA-repair & genome maintenance | Repair double-strand breaks, base damage | Radiophiles, thermophiles | RecA, UvrABC, PprA / PprI regulatory elements. | [49] |
| Antioxidants & redox enzymes | Detoxify ROS; maintain redox balance | Radiophiles, acidophiles | Superoxide dismutase (SOD), catalase, peroxiredoxins. | [37] |
| Membrane remodeling enzymes | Preserve membrane integrity/fluidity | Thermophiles, psychrophiles, alkaliphiles | Fatty-acid desaturases, cardiolipin synthases, hopanoid biosynthesis. | [33] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




