Submitted:
22 February 2026
Posted:
25 February 2026
You are already at the latest version
Abstract
Keywords:
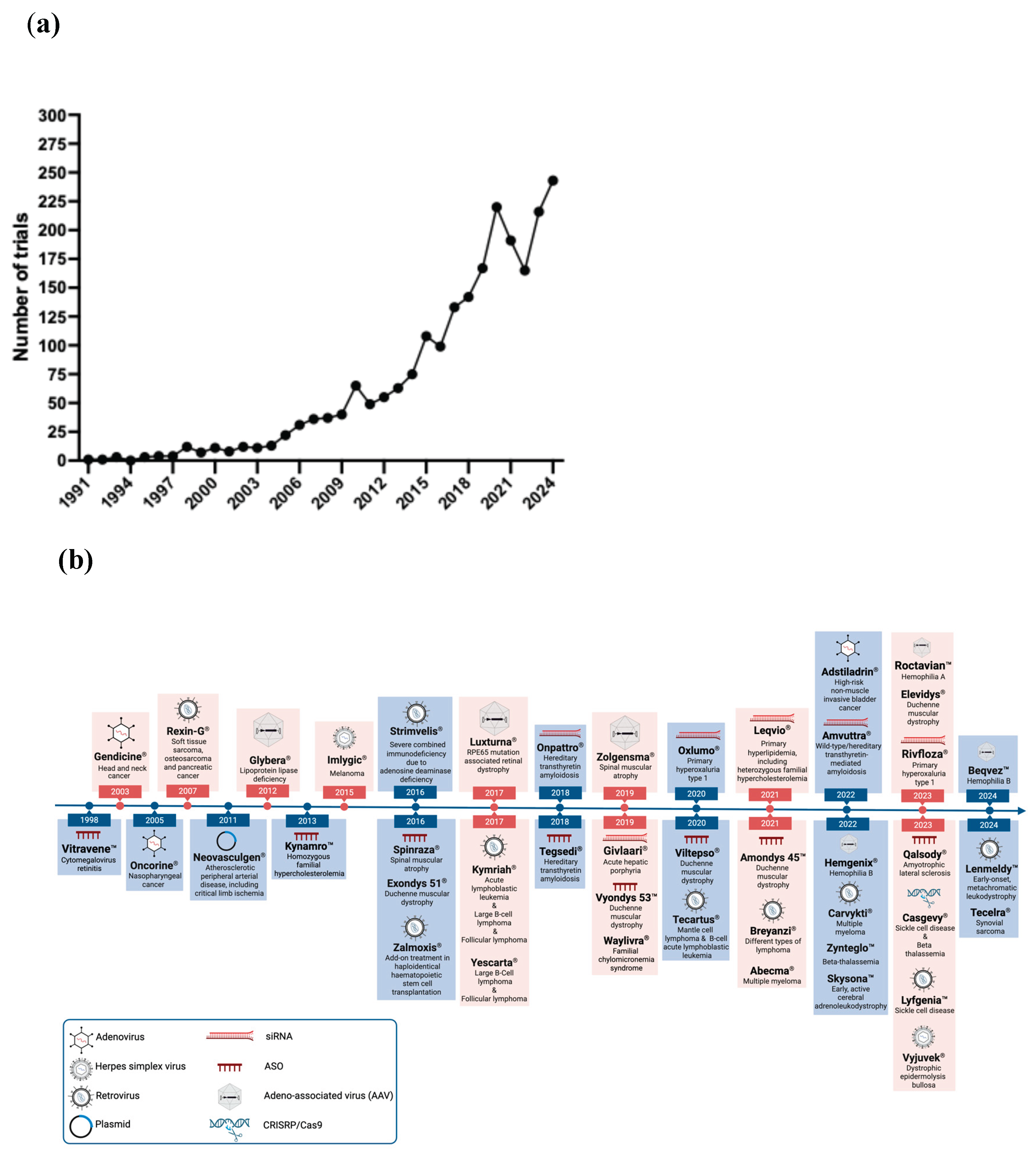
1. Introduction
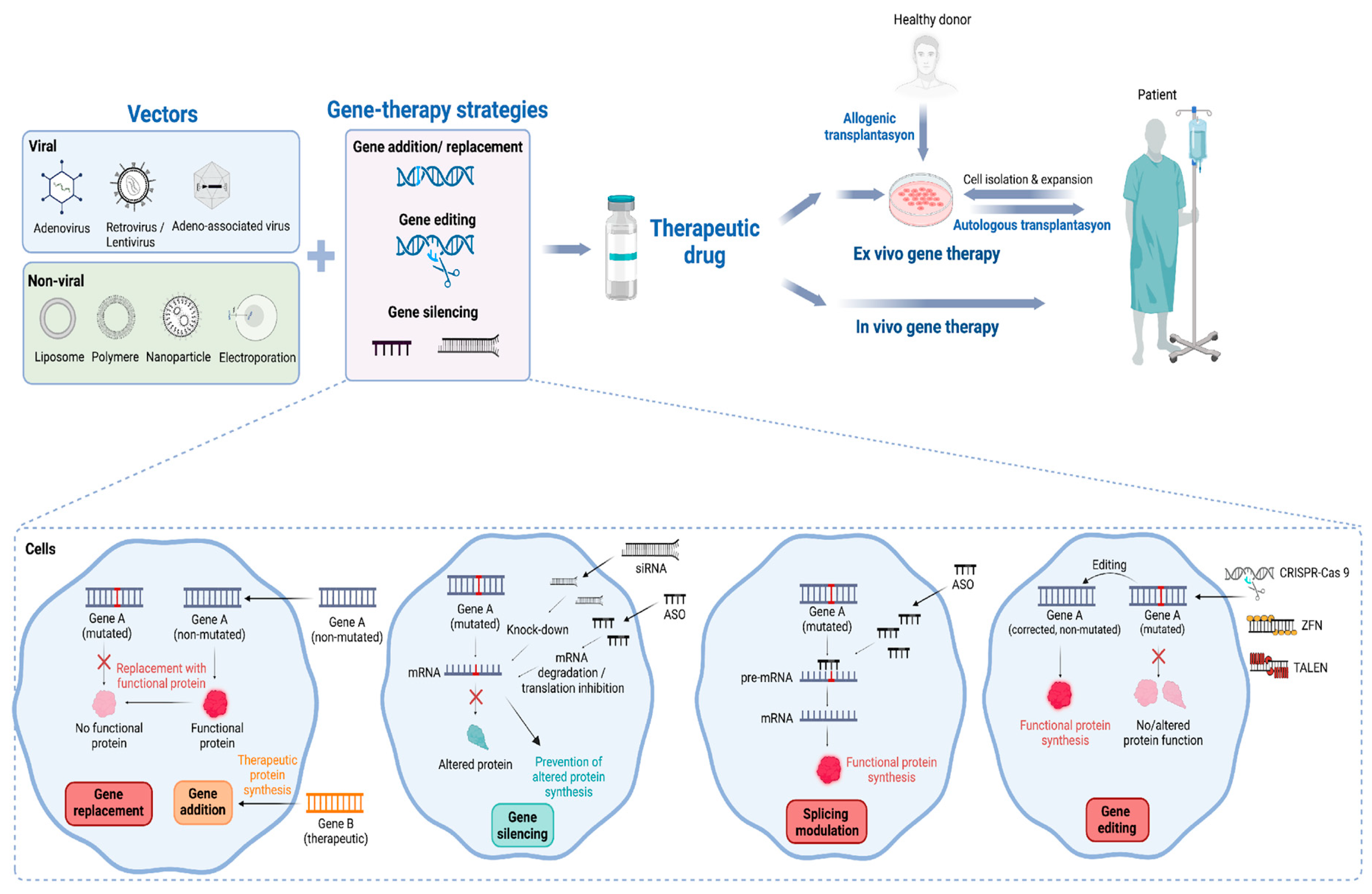
2. Delivery Methods, Vectors, and Strategies for Gene-Targeted Therapies
3. Gene-Targeted Therapy Approaches for Neurodegenerative Diseases
3.1. Gene Addition and Replacement
3.2. Gene Editing
3.3. Gene Silencing
3.4. Splice Modulating
4. Current Gene-Targeted Therapies in Neurodegenerative Disorders: SMA and ALS
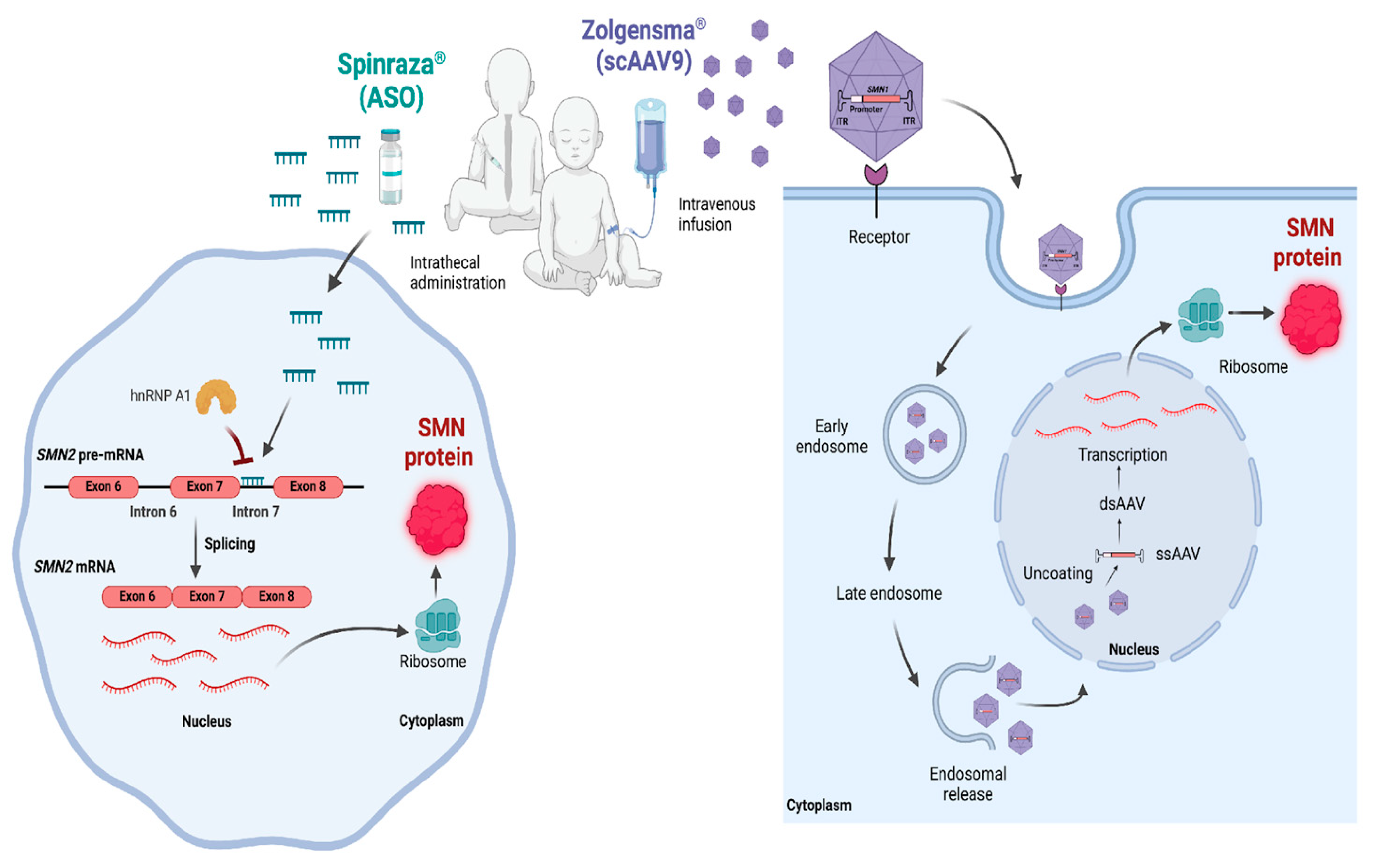
4.1. Spinal Muscular Atrophy (SMA)
4.2. ALS
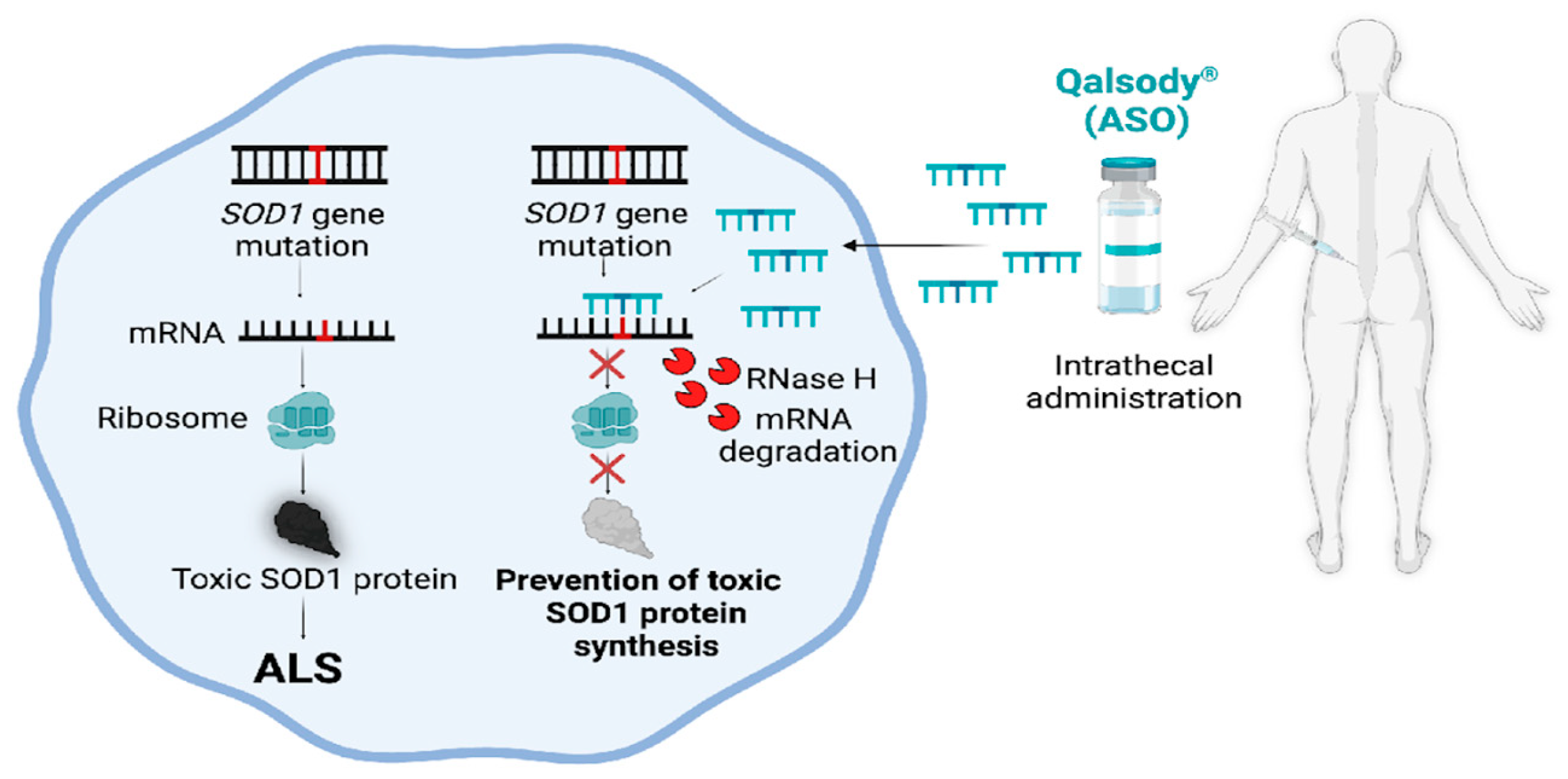
4.2.1. SOD1
4.2.2. FUS
4.2.3. C9orf72
4.2.4. TARDBP
5. Clinical Trial Readiness and Future Considerations
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AAV | Adeno-associated virus |
| AAV9 | Adeno-associated virus serotype 9 |
| Ad | Adenovirus |
| ALS | Amyotrophic lateral sclerosis |
| ALSFRS-R | Amyotrophic Lateral Sclerosis Functional Rating Scale–Revised |
| ASO | Antisense oligonucleotide |
| C9orf72 | Chromosome 9 open reading frame 72 |
| CAR-T | Chimeric antigen receptor T-cell |
| Cas9 | CRISPR-associated protein 9 |
| Cas13d | CRISPR-associated protein 13d |
| CNS | Central nervous system |
| CRISPR | Clustered Regularly Interspaced Short Palindromic Repeats |
| CSF | Cerebrospinal fluid |
| DNA | Deoxyribonucleic acid |
| dsDNA | Double-stranded DNA |
| EMA | European Medicines Agency |
| fALS | Familial amyotrophic lateral sclerosis |
| FDA | Food and Drug Administration |
| FTD | Frontotemporal dementia |
| FUS | Fused in sarcoma |
| gRNA | Guide RNA |
| HR | Homologous recombination |
| iPSC | Induced pluripotent stem cell |
| ITR | Inverted terminal repeat |
| LTR | Long terminal repeat |
| miRNA | MicroRNA |
| mRNA | Messenger ribonucleic acid |
| NDDs | Neurodegenerative diseases |
| NfL | Neurofilament light chain |
| NHEJ | Non-homologous end joining |
| RNA | Ribonucleic acid |
| RNAi | RNA interference |
| RNaseH1 | Ribonuclease H1 |
| sALS | Sporadic amyotrophic lateral sclerosis |
| scAAV | Self-complementary adeno-associated virus |
| sgRNA | Single guide RNA |
| siRNA | Small interfering RNA |
| SMA | Spinal muscular atrophy |
| SMN | Survival motor neuron |
| SMN1 | Survival motor neuron 1 |
| SMN2 | Survival motor neuron 2 |
| SOD1 | Superoxide dismutase 1 |
| STMN2 | Stathmin-2 |
| TALEN | Transcription activator-like effector nuclease |
| TARDBP | TAR DNA-binding protein |
| TDP-43 | TAR DNA-binding protein 43 |
| UTR | Untranslated region |
| ZFNs | Zinc finger nucleases |
References
- Arabi F, Mansouri V, Ahmadbeigi N. Gene therapy clinical trials, where do we go? An overview. Biomed Pharmacother. 2022;153:113324. [CrossRef]
- What is gene therapy? [Internet]. Silver Spring (MD): FDA. Available online: https://www.fda.gov/vaccines-blood-biologics/cellular-gene-therapy-products/what-gene-therapy (accessed on 27 October 2025).
- Razi Soofiyani S, Baradaran B, Lotfipour F, Kazemi T, Mohammadnejad L. Gene therapy, early promises, subsequent problems, and recent breakthroughs. Adv Pharm Bull. 2013;3(2):249-55. [CrossRef]
- Seow Y, Wood MJ. Biological gene delivery vehicles: beyond viral vectors. Mol Ther. 2009;17(5):767-77. [CrossRef]
- Alhakamy NA, Curiel DT, Berkland CJ. The era of gene therapy: From preclinical development to clinical application. Drug Discov Today. 2021;26(7):1602-19. [CrossRef]
- Shin MH, He Y, Marrogi E, Piperdi S, Ren L, Khanna C, et al. A RUNX2-Mediated Epigenetic Regulation of the Survival of p53 Defective Cancer Cells. PLoS Genet. 2016;12(2):e1005884. [CrossRef]
- Zuris JA, Thompson DB, Shu Y, Guilinger JP, Bessen JL, Hu JH, et al. Cationic lipid-mediated delivery of proteins enables efficient protein-based genome editing in vitro and in vivo. Nat Biotechnol. 2015;33(1):73-80. [CrossRef]
- Wang M, Zuris JA, Meng F, Rees H, Sun S, Deng P, et al. Efficient delivery of genome-editing proteins using bioreducible lipid nanoparticles. Proc Natl Acad Sci USA. 2016;113(11):2868-73. [CrossRef]
- Li C, Samulski RJ. Engineering adeno-associated virus vectors for gene therapy. Nat Rev Genet. 2020;21(4):255-72; [CrossRef]
- Bulcha JT, Wang Y, Ma H, Tai PWL, Gao G. Viral vector platforms within the gene therapy landscape. Signal Transduct Target Ther. 2021;6(1):53. [CrossRef]
- Alba R, Bosch A, Chillon M. Gutless adenovirus: last-generation adenovirus for gene therapy. Gene Ther. 2005;12 Suppl 1:S18-27. [CrossRef]
- Atchison RW, Casto BC, Hammon WM. Adenovirus-associated defective virus particles. Science. 1965;149(3685):754-6. [CrossRef]
- Wang JH, Gessler DJ, Zhan W, Gallagher TL, Gao G. Adeno-associated virus as a delivery vector for gene therapy of human diseases. Signal Transduct Target Ther. 2024;9(1):78. [CrossRef]
- Penaud-Budloo M, Le Guiner C, Nowrouzi A, Toromanoff A, Cherel Y, Chenuaud P, et al. Adeno-associated virus vector genomes persist as episomal chromatin in primate muscle. J Virol. 2008;82(16):7875-85. [CrossRef]
- Riyad JM, Weber T. Intracellular trafficking of adeno-associated virus (AAV) vectors: challenges and future directions. Gene Ther. 2021;28(12):683-96. [CrossRef]
- Ling Q, Herstine JA, Bradbury A, Gray SJ. AAV-based in vivo gene therapy for neurological disorders. Nat Rev Drug Discov. 2023;22(10):789-806. [CrossRef]
- Pupo A, Fernández A, Low SH, François A, Suárez-Amarán L, Samulski RJ. AAV vectors: The Rubik's cube of human gene therapy. Mol Ther. 2022;30(12):3515–3541. [CrossRef]
- Suarez-Amaran L, Song L, Tretiakova AP, Mikhail SA, Samulski RJ. AAV vector development, back to the future. Mol Ther. 2025;33(5):1903–1936. [CrossRef]
- van Haasteren J, Li J, Scheideler OJ, Murthy N, Schaffer DV. The delivery challenge: fulfilling the promise of therapeutic genome editing. Nat Biotechnol. 2020;38(7):845-55. [CrossRef]
- Ramamoorth M, Narvekar A. Non viral vectors in gene therapy- an overview. J Clin Diagn Res. 2015;9(1):GE01-6. [CrossRef]
- Picanco-Castro V, Pereira CG, Covas DT, Porto GS, Athanassiadou A, Figueiredo ML. Emerging patent landscape for non-viral vectors used for gene therapy. Nat Biotechnol. 2020;38(2):151-7. [CrossRef]
- Shahryari A, Burtscher I, Nazari Z, Lickert H. Engineering Gene Therapy: Advances and Barriers. Advanced Therapeutics. 2021;4(9). [CrossRef]
- Zhou H, He Y, Xiong W, Jing S, Duan X, Huang Z, et al. MSC based gene delivery methods and strategies improve the therapeutic efficacy of neurological diseases. Bioact Mater. 2023;23:409-37. [CrossRef]
- Zu H, Gao D. Non-viral Vectors in Gene Therapy: Recent Development, Challenges, and Prospects. AAPS J. 2021;23(4):78. [CrossRef]
- Giovannelli I, Higginbottom A, Kirby J, Azzouz M, Shaw PJ. Prospects for gene replacement therapies in amyotrophic lateral sclerosis. Nat Rev Neurol. 2023;19(1):39-52. [CrossRef]
- Goswami R, Subramanian G, Silayeva L, Newkirk I, Doctor D, Chawla K, et al. Gene Therapy Leaves a Vicious Cycle. Front Oncol. 2019;9(927):297. [CrossRef]
- Poletto E, Baldo G, Gomez-Ospina N. Genome Editing for Mucopolysaccharidoses. Int J Mol Sci. 2020;21(2):500. [CrossRef]
- Sun J, Roy S. Gene-based therapies for neurodegenerative diseases. Nat Neurosci. 2021;24(3):297-311. [CrossRef]
- Kim H, Kim JS. A guide to genome engineering with programmable nucleases. Nat Rev Genet. 2014;15(5):321-34. [CrossRef]
- Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. 2012;337(6096):816-21. [CrossRef]
- Mojica FJM D-VJ, García-Martínez Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J Mol Evol. 2005;60:174-82. [CrossRef]
- Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science. 2007;315(5819):1709-12. https://doi.org.10.1126/science.113814.
- Wang JY, Doudna JA. CRISPR technology: A decade of genome editing is only the beginning. Science. 2023;379(6629):eadd8643. [CrossRef]
- Jiang F, Doudna JA. CRISPR–Cas9 Structures and Mechanisms. Annual Review of Biophysics. 2017;46(Volume 46, 2017):505-29. [CrossRef]
- Chen PJ, Liu DR. Prime editing for precise and highly versatile genome manipulation. Nat Rev Genet. 2023;24(3):161-77. [CrossRef]
- Newby GA, Liu DR. In vivo somatic cell base editing and prime editing. Mol Ther. 2021;29(11):3107-24. [CrossRef]
- Casas-Mollano JA, Zinselmeier MH, Erickson SE, Smanski MJ. CRISPR-Cas Activators for Engineering Gene Expression in Higher Eukaryotes. CRISPR J. 2020;3(5):350-64. [CrossRef]
- Qi LS, Larson MH, Gilbert LA, Doudna JA, Weissman JS, Arkin AP, et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell. 2013;152(5):1173-83. [CrossRef]
- Komor AC, Kim YB, Packer MS, Zuris JA, Liu DR. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature. 2016;533(7603):420-4. [CrossRef]
- Kang Y, Chu C, Wang F, Niu Y. CRISPR/Cas9-mediated genome editing in nonhuman primates. Dis Model Mech. 2019;12:dmm039982. [CrossRef]
- Anzalone AV, Randolph PB, Davis JR, Sousa AA, Koblan LW, Levy JM, Chen PJ, Wilson C, Newby GA, Raguram A, Liu DR. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature. 2019;576(7785):149-157. [CrossRef]
- Chen PJ, Liu DR. Prime editing for precise and highly versatile genome manipulation. Nat Rev Genet. 2023;24(3):161–177. [CrossRef]
- Zhao Z, Shang P, Mohanraju P, Geijsen N. Prime editing: advances and therapeutic applications. Trends Biotechnol. 2023;41(8):1000-1012. [CrossRef]
- Schoch KM, Miller TM. Antisense Oligonucleotides: Translation from Mouse Models to Human Neurodegenerative Diseases. Neuron. 2017;94(6):1056-70. [CrossRef]
- Bennett CF, Kordasiewicz HB, Cleveland DW. Antisense Drugs Make Sense for Neurological Diseases. Annu Rev Pharmacol Toxicol. 2021;61:831-52. [CrossRef]
- Marrosu E, Ala P, Muntoni F, Zhou H. Gapmer Antisense Oligonucleotides Suppress the Mutant Allele of COL6A3 and Restore Functional Protein in Ullrich Muscular Dystrophy. Mol Ther Nucleic Acids. 2017;8:416-27. [CrossRef]
- Wu H, Lima WF, Zhang H, Fan A, Sun H, Crooke ST. Determination of the role of the human RNase H1 in the pharmacology of DNA-like antisense drugs. J Biol Chem. 2004;279(17):17181-9. [CrossRef]
- Lima WF, De Hoyos CL, Liang XH, Crooke ST. RNA cleavage products generated by antisense oligonucleotides and siRNAs are processed by the RNA surveillance machinery. Nucleic Acids Res. 2016;44(7):3351-63. [CrossRef]
- Lauffer MC, van Roon-Mom W, Aartsma-Rus A, Collaborative N. Possibilities and limitations of antisense oligonucleotide therapies for the treatment of monogenic disorders. Commun Med (Lond). 2024;4(1):6. [CrossRef]
- Amado DA, Davidson BL. Gene therapy for ALS: A review. Mol Ther. 2021;29(12):3345-58. [CrossRef]
- Brillante S, Volpe M, Indrieri A. Advances in MicroRNA Therapeutics: From Preclinical to Clinical Studies. Hum Gene Ther. 2024;35(17-18):628-648. [CrossRef]
- Borel F, Kay MA, Mueller C. Recombinant AAV as a platform for translating the therapeutic potential of RNA interference. Mol Ther. 2014;22(4):692–701. [CrossRef]
- Bennett CF. Therapeutic Antisense Oligonucleotides Are Coming of Age. Annu Rev Med. 2019;70:307-21. [CrossRef]
- Leung AK, Tam YY, Cullis PR. Lipid nanoparticles for short interfering RNA delivery. Adv Genet. 2014;88:71-110. [CrossRef]
- Dahlman JE, Kauffman KJ, Langer R, Anderson DG. Nanotechnology for in vivo targeted siRNA delivery. Adv Genet. 2014;88:37-69. [CrossRef]
- Geary RS, Norris D, Yu R, Bennett CF. Pharmacokinetics, biodistribution and cell uptake of antisense oligonucleotides. Adv Drug Deliv Rev. 2015;87:46-51. [CrossRef]
- Garreau M, Weidner J, Hamilton R, Kolosionek E, Toki N, Stavenhagen K, et al. Chemical modification patterns for microRNA therapeutic mimics: a structure-activity relationship (SAR) case-study on miR-200c. Nucleic Acids Research. 2024;52(6):2792-2807. [CrossRef]
- Wang P, Zhou Y, Richards AM. Effective tools for RNA-derived therapeutics: siRNA interference or miRNA mimicry. Theranostics. 2021 ;11(18):8771-8796. [CrossRef]
- Alterman JF, Godinho BMDC, Hassler MR, Ferguson CM, Echeverria D, Sapp E, et al. A divalent siRNA chemical scaffold for potent and sustained modulation of gene expression throughout the central nervous system. Nat Biotechnol. 2019;37(8):884-894. [CrossRef]
- Rinaldi C, Wood MJA. Antisense oligonucleotides: the next frontier for treatment of neurological disorders. Nat Rev Neurol. 2018;14(1):9-21. [CrossRef]
- Havens MA, Hastings ML. Splice-switching antisense oligonucleotides as therapeutic drugs. Nucleic Acids Res. 2016;44(14):6549-63. [CrossRef]
- Wirth B. Spinal Muscular Atrophy: In the Challenge Lies a Solution. Trends Neurosci. 2021;44(4):306-22. [CrossRef]
- Yeo CJJ, Tizzano EF, Darras BT. Challenges and opportunities in spinal muscular atrophy therapeutics. Lancet Neurol. 2024;23(2):205–218. [CrossRef]
- Shababi M, Habibi J, Yang HT, Vale SM, Sewell WA, Lorson CL. Cardiac defects contribute to the pathology of spinal muscular atrophy models. Hum Mol Genet. 2010;19(20):4059-71. [CrossRef]
- Allardyce H, Kuhn D, Hernandez-Gerez E, Hensel N, Huang YT, Faller K, et al. Renal pathology in a mouse model of severe Spinal Muscular Atrophy is associated with downregulation of Glial Cell-Line Derived Neurotrophic Factor (GDNF). Hum Mol Genet. 2020;29(14):2365-78. [CrossRef]
- Somers E, Lees RD, Hoban K, Sleigh JN, Zhou H, Muntoni F, et al. Vascular Defects and Spinal Cord Hypoxia in Spinal Muscular Atrophy. Ann Neurol. 2016;79(2):217-30. [CrossRef]
- Hensel N, Claus P. The Actin Cytoskeleton in SMA and ALS: How Does It Contribute to Motoneuron Degeneration? Neuroscientist. 2018;24(1):54-72. [CrossRef]
- Kaymaz AY, Bal SK, Bora G, Talim B, Ozon A, Alikasifoglu A, et al. Alterations in insulin-like growth factor system in spinal muscular atrophy. Muscle Nerve. 2022;66(5):631-8. [CrossRef]
- Bora G, Hensel N, Rademacher S, Koyunoğlu D, Sunguroğlu M, Aksu-Mengeş E, et al. Microtubule-associated protein 1B dysregulates microtubule dynamics and neuronal mitochondrial transport in spinal muscular atrophy. Hum Mol Genet. 2021;29(24):3935-44. [CrossRef]
- Chaytow H, Huang YT, Gillingwater TH, Faller KME. The role of survival motor neuron protein (SMN) in protein homeostasis. Cell Mol Life Sci. 2018;75(21):3877-94. [CrossRef]
- Tapken I, Schweitzer T, Paganin M, Schüning T, Detering NT, Sharma G, Niesert M, Saffari A, Kuhn D, Glynn A, Cieri F, Santonicola P, Cannet C, Gerstner F, Faller KME, Huang YT, Kothary R, Gillingwater TH, Di Schiavi E, Simon CM, Claus P. The systemic complexity of a monogenic disease: the molecular network of spinal muscular atrophy. Brain. 2025;148(2):580–596. [CrossRef]
- Özer PZ, Koyunoğlu D, Son ÇD, Erdem-Yurter H, Bora G. SMN loss dysregulates microtubule-associated proteins in spinal muscular atrophy model. Mol Cell Neurosci. 2022;120:103725. [CrossRef]
- Dayangaç-Erden D, Bora G, Ayhan P, Kocaefe C, Dalkara S, Yelekçi K, et al. Histone deacetylase inhibition activity and molecular docking of (e )-resveratrol: its therapeutic potential in spinal muscular atrophy. Chem Biol Drug Des. 2009;73(3):355-64. [CrossRef]
- Chaytow H, Faller KME, Huang YT, Gillingwater TH. Spinal muscular atrophy: From approved therapies to future therapeutic targets for personalized medicine. Cell Rep Med. 2021;2(7):100346. [CrossRef]
- Mercuri E, Pera MC, Scoto M, Finkel R, Muntoni F. Spinal muscular atrophy - insights and challenges in the treatment era. Nat Rev Neurol. 2020;16(12):706-15. [CrossRef]
- Jablonka S, Hennlein L, Sendtner M. Therapy development for spinal muscular atrophy: perspectives for muscular dystrophies and neurodegenerative disorders. Neurol Res Pract. 2022;4(1):2. [CrossRef]
- Singh NK, Singh NN, Androphy EJ, Singh RN. Splicing of a critical exon of human Survival Motor Neuron is regulated by a unique silencer element located in the last intron. Mol Cell Biol. 2006;26(4):1333-46. [CrossRef]
- Hua Y, Vickers TA, Okunola HL, Bennett CF, Krainer AR. Antisense masking of an hnRNP A1/A2 intronic splicing silencer corrects SMN2 splicing in transgenic mice. Am J Hum Genet. 2008;82(4):834-48. [CrossRef]
- Biogen. Spinraza® (nusinersen) [Internet]. Cambridge (MA): Biogen; c2024 Available online: https://wwwspinrazacom/ (accessed on 13 May 2025).
- Zolgensma®. Bannockburn (IL): Novartis; c2024. Available online: https://www.zolgensma.com/ (14 May 2025).
- Thomsen G, Burghes AHM, Hsieh C, Do J, Chu BTT, Perry S, et al. Biodistribution of onasemnogene abeparvovec DNA, mRNA and SMN protein in human tissue. Nat Med. 2021;27(10):1701-11. [CrossRef]
- Roche. FDA approves Roche’s Evrysdi tablet as first and only tablet for Spinal Muscular Atrophy (SMA). 2025 Feb 12. https://www.roche.com/media/releases/med-cor-2025-02-12.
- Ottesen EW, Singh NN, Luo D, Kaas B, Gillette BJ, Seo J, et al. Diverse targets of SMN2-directed splicing-modulating small molecule therapeutics for spinal muscular atrophy. Nucleic Acids Res. 2023;51(12):5948-80. [CrossRef]
- Wirth B, Karakaya M, Kye MJ, Mendoza-Ferreira N. Twenty-Five Years of Spinal Muscular Atrophy Research: From Phenotype to Genotype to Therapy, and What Comes Next. Annu Rev Genomics Hum Genet. 2020;21:231-61. [CrossRef]
- Singh RN, Ottesen EW, Singh NN. The First Orally Deliverable Small Molecule for the Treatment of Spinal Muscular Atrophy. Neurosci Insights. 2020;15:2633105520973985. [CrossRef]
- Chiriboga CA, Bruno C, Duong T, Fischer D, Mercuri E, Kirschner J, Kostera-Pruszczyk A, Jaber B, Gorni K, Kletzl H, Carruthers I, Martin C, Scalco RS, Fontoura P, Muntoni F, JEWELFISH Study Group. JEWELFISH: 24-month results from an open-label study in non-treatment-naïve patients with SMA receiving treatment with risdiplam. J Neurol. 2024;271(8):4871–4884. [CrossRef]
- Bekircan-Kurt CE, Subramanian S, Chagat S, Mackenzie SJ, Iammarino M, Reash N, Richardson C, Tsao CY, Noritz G, Gushue C, Kotha K, Paul G, Shell R, Alfano LN, Lowes LP, Connolly AM, Waldrop MA. Transitioning from nusinersen to risdiplam for spinal muscular atrophy in clinical practice: A single-center experience. Muscle Nerve. 2025;71(3):414–421. [CrossRef]
- Oechsel KF, Cartwright MS. Combination therapy with onasemnogene and risdiplam in spinal muscular atrophy type 1. Muscle Nerve. 2021;64(4):487–490. [CrossRef]
- Cure SMA. SMA drug pipeline [Internet]. Chicago (IL): Cure SMA. Available online: https://www.curesma.org/sma-drug-pipeline/ (accessed on 13 May 2025).
- Hardiman O, Al-Chalabi A, Chio A, Corr EM, Logroscino G, Robberecht W, et al. Amyotrophic lateral sclerosis. Nat Rev Dis Primers. 2017;3:17071. [CrossRef]
- Van Damme P, Robberecht W, Van Den Bosch L. Modelling amyotrophic lateral sclerosis: progress and possibilities. Dis Model Mech. 2017;10(5):537-49. [CrossRef]
- Shatunov A, Al-Chalabi A. The genetic architecture of ALS. Neurobiol Dis. 2021;147:105156. [CrossRef]
- Chiò A, Logroscino G, Traynor BJ, Collins J, Simeone JC, Goldstein LA, et al. Global epidemiology of amyotrophic lateral sclerosis: a systematic review of the published literature. Neuroepidemiology. 2013;41(2):118-30. [CrossRef]
- Xu L, Liu T, Liu L, Yao X, Chen L, Fan D, et al. Global variation in prevalence and incidence of amyotrophic lateral sclerosis: a systematic review and meta-analysis. J Neurol. 2020;267(4):944-53. [CrossRef]
- Logroscino G, Traynor BJ, Hardiman O, Chiò A, Mitchell D, Swingler RJ, et al. Incidence of amyotrophic lateral sclerosis in Europe. J Neurol Neurosurg Psychiatry. 2010;81(4):385-90. [CrossRef]
- Ruffo P, Traynor BJ, Conforti FL. Advancements in genetic research and RNA therapy strategies for amyotrophic lateral sclerosis (ALS): current progress and future prospects. J Neurol. 2025;272(3):233. [CrossRef]
- Feldman EL, Goutman SA, Petri S, Mazzini L, Savelieff MG, Shaw PJ, et al. Amyotrophic lateral sclerosis. Lancet. 2022;400(10360):1363-80. [CrossRef]
- Chia R, Chiò A, Traynor BJ. Novel genes associated with amyotrophic lateral sclerosis: diagnostic and clinical implications. Lancet Neurol. 2018;17(1):94-102. [CrossRef]
- Akcimen F, Lopez ER, Landers JE, Nath A, Chio A, Chia R, et al. Amyotrophic lateral sclerosis: translating genetic discoveries into therapies. Nat Rev Genet. 2023;24(9):642-58. [CrossRef]
- Renton AE, Chiò A, Traynor BJ. State of play in amyotrophic lateral sclerosis genetics. Nat Neurosci. 2014;17(1):17-23. [CrossRef]
- Müller K, Brenner D, Weydt P, Meyer T, Grehl T, Petri S, et al. Comprehensive analysis of the mutation spectrum in 301 German ALS families. J Neurol Neurosurg Psychiatry. 2018;89(8):817-27. [CrossRef]
- Sherman L, Dafni N, Lieman-Hurwitz J, Groner Y. Nucleotide sequence and expression of human chromosome 21-encoded superoxide dismutase mRNA. Proc Natl Acad Sci USA. 1983;80(18):5465-9. [CrossRef]
- Niwa J, Yamada S, Ishigaki S, Sone J, Takahashi M, Katsuno M, et al. Disulfide bond mediates aggregation, toxicity, and ubiquitylation of familial amyotrophic lateral sclerosis-linked mutant SOD1. J Biol Chem. 2007;282(38):28087-95. [CrossRef]
- Fridovich I. Biological effects of the superoxide radical. Arch Biochem Biophys. 1986;247(1):1-11. [CrossRef]
- Saccon RA, Bunton-Stasyshyn RK, Fisher EM, Fratta P. Is SOD1 loss of function involved in amyotrophic lateral sclerosis? Brain. 2013;136(Pt 8):2342-58. [CrossRef]
- Lim CKW, Gapinske M, Brooks AK, Woods WS, Powell JE, Zeballos CM, et al. Treatment of a Mouse Model of ALS by In Vivo Base Editing. Mol Ther. 2020;28(4):1177-89. [CrossRef]
- Duan W, Guo M, Yi L, Liu Y, Li Z, Ma Y, et al. The deletion of mutant SOD1 via CRISPR/Cas9/sgRNA prolongs survival in an amyotrophic lateral sclerosis mouse model. Gene Ther. 2020;27(3-4):157-69. [CrossRef]
- Smith RA, Miller TM, Yamanaka K, Monia BP, Condon TP, Hung G, et al. Antisense oligonucleotide therapy for neurodegenerative disease. J Clin Invest. 2006;116(8):2290-6. [CrossRef]
- Miller TM, Pestronk A, David W, Rothstein J, Simpson E, Appel SH, et al. An antisense oligonucleotide against SOD1 delivered intrathecally for patients with SOD1 familial amyotrophic lateral sclerosis: a phase 1, randomised, first-in-man study. Lancet Neurol. 2013;12(5):435-42. [CrossRef]
- McCampbell A, Cole T, Wegener AJ, Tomassy GS, Setnicka A, Farley BJ, et al. Antisense oligonucleotides extend survival and reverse decrement in muscle response in ALS models. J Clin Invest. 2018;128(8):3558-67. [CrossRef]
- Jin J, Zhong XB. ASO drug Qalsody (tofersen) targets amyotrophic lateral sclerosis. Trends Pharmacol Sci. 2023;44(12):1043-4. [CrossRef]
- Miller T, Cudkowicz M, Shaw PJ, Andersen PM, Atassi N, Bucelli RC, et al. Phase 1-2 Trial of Antisense Oligonucleotide Tofersen for SOD1 ALS. N Engl J Med. 2020;383(2):109-19. [CrossRef]
- Miller TM, Cudkowicz ME, Genge A, Shaw PJ, Sobue G, Bucelli RC, et al. Trial of Antisense Oligonucleotide Tofersen for SOD1 ALS. N Engl J Med. 2022;387(12):1099-110. [CrossRef]
- Blair HA. Tofersen: First Approval. Drugs. 2023;83:1039–1043. [CrossRef]
- Borel F, Gernoux G, Cardozo B, Metterville JP, Toro Cabrera GC, Song L, et al. Therapeutic rAAVrh10 Mediated SOD1 Silencing in Adult SOD1(G93A) Mice and Nonhuman Primates. Hum Gene Ther. 2016;27(1):19-31. [CrossRef]
- Arrowhead Pharmaceuticals. Poster for Arrowhead Pharmaceuticals [Internet]. Preclinical profile of ARO-SOD1, an siRNA therapy for SOD1-ALS. Available online: https://ir.arrowheadpharma.com/static-files/804aa94c-7323-4917-a04f-bd0812d2e2fb (accessed on 5 June 2025).
- Mueller C, Berry JD, McKenna-Yasek DM, Gernoux G, Owegi MA, Pothier LM, et al. SOD1 suppression with adeno-associated virus and microRNA in familial ALS. N Engl J Med. 2020;383(2):151–158. [CrossRef]
- Ye J, Jiang L, Wang L, Pan Y, Wang X, Qu H, et al. RAG-17, a Novel siRNA Therapy for SOD1-ALS: Safety and Preliminary Efficacy from a First-in-human Trial (S5.003). Neurology. 2025;104(7_Supplement_1). [CrossRef]
- Assoni AF, Foijer F, Zatz M. Amyotrophic Lateral Sclerosis, FUS and Protein Synthesis Defects. Stem Cell Rev Rep. 2023;19(3):625-38. [CrossRef]
- Tarlarini C, Lunetta C, Mosca L, Avemaria F, Riva N, Mantero V, et al. Novel FUS mutations identified through molecular screening in a large cohort of familial and sporadic amyotrophic lateral sclerosis. Eur J Neurol. 2015;22(11):1474-81. [CrossRef]
- Akiyama T, Warita H, Kato M, Nishiyama A, Izumi R, Ikeda C, et al. Genotype-phenotype relationships in familial amyotrophic lateral sclerosis with FUS/TLS mutations in Japan. Muscle Nerve. 2016;54(3):398-404. [CrossRef]
- Vance C, Rogelj B, Hortobágyi T, De Vos KJ, Nishimura AL, Sreedharan J, et al. Mutations in FUS, an RNA processing protein, cause familial amyotrophic lateral sclerosis type 6. Science. 2009;323(5918):1208-11. [CrossRef]
- Xiao X, Li M, Ye Z, He X, Wei J, Zha Y. FUS gene mutation in amyotrophic lateral sclerosis: a new case report and systematic review. Amyotroph Lateral Scler Frontotemporal Degener. 2024;25(1-2):1-15. [CrossRef]
- Reber S, Stettler J, Filosa G, Colombo M, Jutzi D, Lenzken SC, et al. Minor intron splicing is regulated by FUS and affected by ALS-associated FUS mutants. EMBO J. 2016;35(14):1504-21. [CrossRef]
- Korobeynikov VA, Lyashchenko AK, Blanco-Redondo B, Jafar-Nejad P, Shneider NA. Antisense oligonucleotide silencing of FUS expression as a therapeutic approach in amyotrophic lateral sclerosis. Nat Med. 2022;28(1):104-16. [CrossRef]
- Conte A, Lattante S, Zollino M, Marangi G, Luigetti M, Del Grande A, et al. P525L FUS mutation is consistently associated with a severe form of juvenile amyotrophic lateral sclerosis. Neuromuscul Disord. 2012;22(1):73-5. [CrossRef]
- Shneider NA, Harms MB, Korobeynikov VA, Rifai OM, Hoover BN, Harrington EA, et al. Antisense oligonucleotide jacifusen for FUS-ALS: an investigator-initiated, multicentre, open-label case series. Lancet. 2025;405(10494):2075-86. [CrossRef]
- Renton AE, Majounie E, Waite A, Simón-Sánchez J, Rollinson S, Gibbs JR, et al. A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron. 2011;72(2):257-68. [CrossRef]
- Smeyers J, Banchi EG, Latouche M. C9ORF72: What It Is, What It Does, and Why It Matters. Front Cell Neurosci. 2021;15:661447. [CrossRef]
- DeJesus-Hernandez M, Mackenzie IR, Boeve BF, Boxer AL, Baker M, Rutherford NJ, et al. Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron. 2011;72(2):245-56. [CrossRef]
- Majounie E, Renton AE, Mok K, Dopper EG, Waite A, Rollinson S, et al. Frequency of the C9orf72 hexanucleotide repeat expansion in patients with amyotrophic lateral sclerosis and frontotemporal dementia: a cross-sectional study. Lancet Neurol. 2012;11(4):323-30. [CrossRef]
- Van Mossevelde S, van der Zee J, Cruts M, Van Broeckhoven C. Relationship between C9orf72 repeat size and clinical phenotype. Curr Opin Genet Dev. 2017;44:117-24. [CrossRef]
- Iacoangeli A, Al Khleifat A, Jones AR, Sproviero W, Shatunov A, Opie-Martin S, et al. C9orf72 intermediate expansions of 24-30 repeats are associated with ALS. Acta Neuropathol Commun. 2019;7(1):115. [CrossRef]
- Sellier C, Corcia P, Vourc'h P, Dupuis L. C9ORF72 hexanucleotide repeat expansion: From ALS and FTD to a broader pathogenic role? Rev Neurol (Paris). 2024;180(5):417-428. [CrossRef]
- Braems E, Swinnen B, Van Den Bosch L. C9orf72 loss-of-function: a trivial, stand-alone or additive mechanism in C9 ALS/FTD? Acta Neuropathol. 2020;140(5):625-43. [CrossRef]
- Shi Y, Lin S, Staats KA, Li Y, Chang WH, Hung ST, et al. Haploinsufficiency leads to neurodegeneration in C9ORF72 ALS/FTD human induced motor neurons. Nat Med. 2018;24(3):313–325. Swinnen B, Robberecht W, Van Den Bosch L. RNA toxicity in non-coding repeat expansion disorders. Embo j. 2020;39(1):e101112. https://doi.org/10.15252/embj.2018101112. [CrossRef]
- Ash PE, Bieniek KF, Gendron TF, Caulfield T, Lin WL, Dejesus-Hernandez M, et al. Unconventional translation of C9ORF72 GGGGCC expansion generates insoluble polypeptides specific to c9FTD/ALS. Neuron. 2013;77(4):639-46. [CrossRef]
- Lagier-Tourenne C, Baughn M, Rigo F, Sun S, Liu P, Li HR, et al. Targeted degradation of sense and antisense C9orf72 RNA foci as therapy for ALS and frontotemporal degeneration. Proc Natl Acad Sci U S A. 2013;110(47):E4530-9. [CrossRef]
- Tran H, Moazami MP, Yang H, McKenna-Yasek D, Douthwright CL, Pinto C, et al. Suppression of mutant C9orf72 expression by a potent mixed backbone antisense oligonucleotide. Nat Med. 2022;28(1):117-24. [CrossRef]
- Jiang J, Zhu Q, Gendron TF, Saberi S, McAlonis-Downes M, Seelman A, et al. Gain of Toxicity from ALS/FTD-Linked Repeat Expansions in C9ORF72 Is Alleviated by Antisense Oligonucleotides Targeting GGGGCC-Containing RNAs. Neuron. 2016;90(3):535-50. [CrossRef]
- van den Berg LH, Rothstein JD, Shaw PJ, Babu S, Benatar M, Bucelli RC, et al. Safety, tolerability, and pharmacokinetics of antisense oligonucleotide BIIB078 in adults with C9orf72-associated amyotrophic lateral sclerosis: a phase 1, randomised, double blinded, placebo-controlled, multiple ascending dose study. Lancet Neurol. 2024: 23(9):901-912. [CrossRef]
- Meijboom KE, Brown RH. Approaches to Gene Modulation Therapy for ALS. Neurotherapeutics. 2022;19(4):1159-79. [CrossRef]
- Ratti A, Buratti E. Physiological functions and pathobiology of TDP-43 and FUS/TLS proteins. J Neurochem. 2016;138 Suppl 1:95-111. [CrossRef]
- Guo L, Shorter J. Biology and Pathobiology of TDP-43 and Emergent Therapeutic Strategies. Cold Spring Harb Perspect Med. 2017;7(9). [CrossRef]
- Giordana MT, Piccinini M, Grifoni S, De Marco G, Vercellino M, Magistrello M, et al. TDP-43 redistribution is an early event in sporadic amyotrophic lateral sclerosis. Brain Pathol. 2010;20(2):351-60. [CrossRef]
- Suk TR, Rousseaux MWC. The role of TDP-43 mislocalization in amyotrophic lateral sclerosis. Mol Neurodegener. 2020;15(1):45. [CrossRef]
- Kabashi E, Lin L, Tradewell ML, Dion PA, Bercier V, Bourgouin P, et al. Gain and loss of function of ALS-related mutations of TARDBP (TDP-43) cause motor deficits in vivo. Hum Mol Genet. 2010;19(4):671-83. [CrossRef]
- Babazadeh A, Rayner SL, Lee A, Chung RS. TDP-43 as a therapeutic target in neurodegenerative diseases: Focusing on motor neuron disease and frontotemporal dementia. Ageing Res Rev. 2023;92:102085. [CrossRef]
- Takeuchi T, Maeta K, Ding X, Oe Y, Takeda A, Inoue M, et al. Sustained therapeutic benefits by transient reduction of TDP-43 using ENA-modified antisense oligonucleotides in ALS/FTD mice. Mol Ther Nucleic Acids. 2023;31:353-66. [CrossRef]
- Becker LA, Huang B, Bieri G, Ma R, Knowles DA, Jafar-Nejad P, et al. Therapeutic reduction of ataxin-2 extends lifespan and reduces pathology in TDP-43 mice. Nature. 2017;544(7650):367-71. [CrossRef]
- Amado DA, Robbins AB, Smith AR, Whiteman KR, Chillon Bosch G, Chen Y, et al. AAV-based delivery of RNAi targeting Ataxin-2 improves survival, strength, and pathology in mouse models of rapidly and slowly progressive sporadic ALS. BioRxiv. 2024. [CrossRef]
- Biogen and Ionis Pharmaceuticals. Biogen and Ionis announce topline Phase 1/2 study results of investigational drug in amyotrophic lateral sclerosis. Biogen. Available online: https://investors.biogen.com/news-releases/news-release-details/biogen-and-ionis-announce-topline-phase-12-study-results (accessed on 16 May 2024 ).
- Melamed Z, Lopez-Erauskin J, Baughn MW, Zhang O, Drenner K, Sun Y, et al. Premature polyadenylation-mediated loss of stathmin-2 is a hallmark of TDP-43-dependent neurodegeneration. Nat Neurosci. 2019;22(2):180-90. [CrossRef]
- Klim JR, Williams LA, Limone F, Guerra San Juan I, Davis-Dusenbery BN, Mordes DA, et al. ALS-implicated protein TDP-43 sustains levels of STMN2, a mediator of motor neuron growth and repair. Nat Neurosci. 2019;22(2):167-79. [CrossRef]
- Shin JE, Geisler S, DiAntonio A. Dynamic regulation of SCG10 in regenerating axons after injury. Exp Neurol. 2014;252:1-11. [CrossRef]
- Baughn MW, Melamed Z, Lopez-Erauskin J, Beccari MS, Ling K, Zuberi A, et al. Mechanism of STMN2 cryptic splice-polyadenylation and its correction for TDP-43 proteinopathies. Science. 2023;379(6637):1140-9. [CrossRef]
- Genge A, Salmon K, Polzer J, Martinez C, Boggs B, Eon V, et al. QRL-201-01—A Multi-center, Randomized, Double-blind, Placebo-controlled Multiple Ascending Dose Study to Evaluate the Safety and Tolerability of QRL-201 in Amyotrophic Lateral Sclerosis (P6-11.010). Neurology. 2024;102(7_supplement_1). [CrossRef]
- Phares S, Phillip K, Trusheim M. Clinical development success rates for durable cell and gene therapies. Nat Rev Drug Discov. 2025;24(5):329-30. [CrossRef]
- Sumner CJ, Crawford TO. Early treatment is a lifeline for infants with SMA. Nat Med. 2022;28(7):1348-9. [CrossRef]
- Rothgangl T, Dennis MK, Lin PJC, Oka R, Witzigmann D, Villiger L, et al. In vivo adenine base editing of PCSK9 in macaques reduces LDL cholesterol levels. Nat Biotechnol. 2021;39(8):949-57. [CrossRef]
- Janssen P, Isa T, Lanciego J, Leech K, Logothetis N, Poo MM, et al. Visualizing advances in the future of primate neuroscience research. Curr Res Neurobiol. 2023;4:100064. [CrossRef]
- Westerling-Bui AD, Fast EM, Soare TW, Venkatachalan S, DeRan M, Fanelli AB, et al. Transplanted organoids empower human preclinical assessment of drug candidate for the clinic. Sci Adv. 2022;8(27):eabj5633. [CrossRef]
- Evangelisti C, Ramadan S, Orlacchio A, Panza E. Experimental Cell Models for Investigating Neurodegenerative Diseases. Int J Mol Sci. 2024;25(17). [CrossRef]
- Fernandopulle MS, Prestil R, Grunseich C, Wang C, Gan L, Ward ME. Transcription Factor-Mediated Differentiation of Human iPSCs into Neurons. Curr Protoc Cell Biol. 2018;79(1):e51. [CrossRef]
- Giacomelli E, Vahsen BF, Calder EL, Xu Y, Scaber J, Gray E, et al. Human stem cell models of neurodegeneration: From basic science of amyotrophic lateral sclerosis to clinical translation. Cell Stem Cell. 2022;29(1):11-35. [CrossRef]
- Answer ALS. Answer ALS [Internet]. New Orleans (LA): Answer ALS; Available online: https://www.answerals.org (accessed on 27 May 2025).
- Okano H, Morimoto S. iPSC-based disease modeling and drug discovery in cardinal neurodegenerative disorders. Cell Stem Cell. 2022;29(2):189-208. [CrossRef]
- Grass T, Dokuzluoglu Z, Buchner F, Rosignol I, Thomas J, Caldarelli A, et al. Isogenic patient-derived organoids reveal early neurodevelopmental defects in spinal muscular atrophy initiation. Cell Rep Med. 2024;5(8):101659. [CrossRef]
- Faravelli I, Rinchetti P, Tambalo M, Simutin I, Mapelli L, Mancinelli S, et al. Targeted Antisense Oligonucleotide Treatment Rescues Developmental Alterations in Spinal Muscular Atrophy Organoids. bioRxiv 2025. [CrossRef]
- Szebenyi K, Wenger LMD, Sun Y, Dunn AWE, Limegrover CA, Gibbons GM, et al. Human ALS/FTD brain organoid slice cultures display distinct early astrocyte and targetable neuronal pathology. Nat Neurosci. 2021;24(11):1542-54. [CrossRef]
- Gao C, Shi Q, Pan X, Chen J, Zhang Y, Lang J, et al. Neuromuscular organoids model spinal neuromuscular pathologies in C9orf72 amyotrophic lateral sclerosis. Cell Rep. 2024;43(3):113892. [CrossRef]
- van der Geest AT, Jakobs CE, Ljubikj T, Huffels CFM, Canizares Luna M, Vieira de Sa R, et al. Molecular pathology, developmental changes and synaptic dysfunction in (pre-) symptomatic human C9ORF72-ALS/FTD cerebral organoids. Acta Neuropathol Commun. 2024;12(1):152. [CrossRef]
- U.S. Food and Drug Administration. Rare diseases: natural history studies for drug development. Draft guidance for industry. FDA-2019-D-0481. Available online: https://www.fda.gov/regulatory-information/search-fda-guidance-documents/rare-diseases-natural-history-studies-drug-development (Accessed on 25 October 2025).
- Alqahtani A, Kokkinis A, Zizzi C, Dilek N, Fischbeck KH, Heatwole CR, et al. Patient-Reported Impact of Symptoms in Spinal and Bulbar Muscular Atrophy. Neurol Clin Pract. 2023;13(6):e200213. [CrossRef]
- Grunseich C, Patankar A, Amaya J, Watts JA, Li D, Ramirez P, et al. Clinical and Molecular Aspects of Senataxin Mutations in Amyotrophic Lateral Sclerosis 4. Ann Neurol. 2020;87(4):547-55. [CrossRef]
- FDA-NIH Biomarker Working Group, 2017, BEST (Biomarkers, EndpointS, and other Tools) Resource, Silver Spring, Maryland: Food and Drug Administration, and Bethesda, Maryland:National Institutes of Health Available online https://www.ncbi.nlm.nih.gov/books/NBK326791/pdf/Bookshelf_NBK326791.pdf (accessed on October 2025).
- Klickovic U, Zampedri L, Sinclair CDJ, Wastling SJ, Trimmel K, Howard RS, et al. Skeletal muscle MRI differentiates SBMA and ALS and correlates with disease severity. Neurology. 2019;93(9):e895-e907. [CrossRef]
- Tebbenkamp AT, Huggett SB, Lombardi V, Zampedri L, AlQahtani A, Kokkinis A, et al. Protein biomarker signature in patients with spinal and bulbar muscular atrophy. JCI Insight. 2024;9(13). [CrossRef]
- Xing X, Liu X, Li X, Li M, Wu X, Huang X, et al. Insights into spinal muscular atrophy from molecular biomarkers. Neural Regen Res. 2025;20(7):1849-63. [CrossRef]
- Ruffo P, Traynor BJ, Conforti FL. Advancements in genetic research and RNA therapy strategies for amyotrophic lateral sclerosis (ALS): current progress and future prospects. J Neurol. 2025;272(3):233. [CrossRef]
- Arbab M, Matuszek Z, Kray KM, Du A, Newby GA, Blatnik AJ, et al. Base editing rescue of spinal muscular atrophy in cells and in mice. Science. 2023;380(6642):eadg6518. [CrossRef]
- Alves CRR, Ha LL, Yaworski R, Sutton ER, Lazzarotto CR, Christie KA, et al. Optimization of base editors for the functional correction of SMN2 as a treatment for spinal muscular atrophy. Nat Biomed Eng. 2024;8(2):118-31. [CrossRef]
- Kempthorne L, Vaizoglu D, Cammack AJ, Carcolé M, Roberts MJ, Mikheenko A, et al. Dual-targeting CRISPR-CasRx reduces C9orf72 ALS/FTD sense and antisense repeat RNAs in vitro and in vivo. Nat Commun. 2025;16(1):459. [CrossRef]
- McCallister TX, Lim CKW, Singh M, Zhang S, Ahsan NS, Terpstra WM, et al. A high-fidelity CRISPR-Cas13 system improves abnormalities associated with C9ORF72-linked ALS/FTD. Nat Commun. 2025;16(1):460. [CrossRef]
| National Clinical Trial Identifier | Target | Interventions | Modality | Phases | Study Status |
|---|---|---|---|---|---|
| NCT03764488 NCT02623699 NCT03070119 NCT04856982 |
SOD1 | BIIB067 (Tofersen) | ASO | III | Approved for marketing |
| NCT05903690 NCT05903690 |
SOD1 | RAG-17 | RNAi | I | Completed Recruiting |
| NCT06100276 | SOD1 | AMT-162 (APB-102) |
AAV-RNAi (miRNA) | I/II | Active, not recruiting |
| NCT04768972 | FUS | ION363 (Jacifusen) | ASO | III | Active, not recruiting |
| NCT04632225 NCT05176093 NCT02039401 |
HGF addition | Engensis (VM202) | Non-viral plasmid gene delivery | II | Completed |
| NCT05633459 | STMN2 | QRL-201 | ASO | I | Active, not recruiting |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.





