Submitted:
13 February 2026
Posted:
27 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Plasmodium Parasite Lines
2.3. Cell Culture and In Vitro Infection of HeLa Cells
2.4. Stable Expression of GFP-Tagged SNARE Constructs in HeLa Cells
2.5. Generation of SNARE Knockout Cell Lines Using CRISPR/Cas9
2.6. Western Blotting
2.7. Sporozoite Isolation from Mosquito Salivary Glands
2.8. Live Cell Imaging
2.9. Immunofluorescence Analysis
2.10. Pre-Staining Expansion Microscopy (PS-ExM)
2.11. Liver Stage Parasite Size Measurement
2.12. Quantification of Lysosome (LE) and Late Endosome Association with Plasmodium PVM
2.13. Image Acquisition and Analysis
3. Results
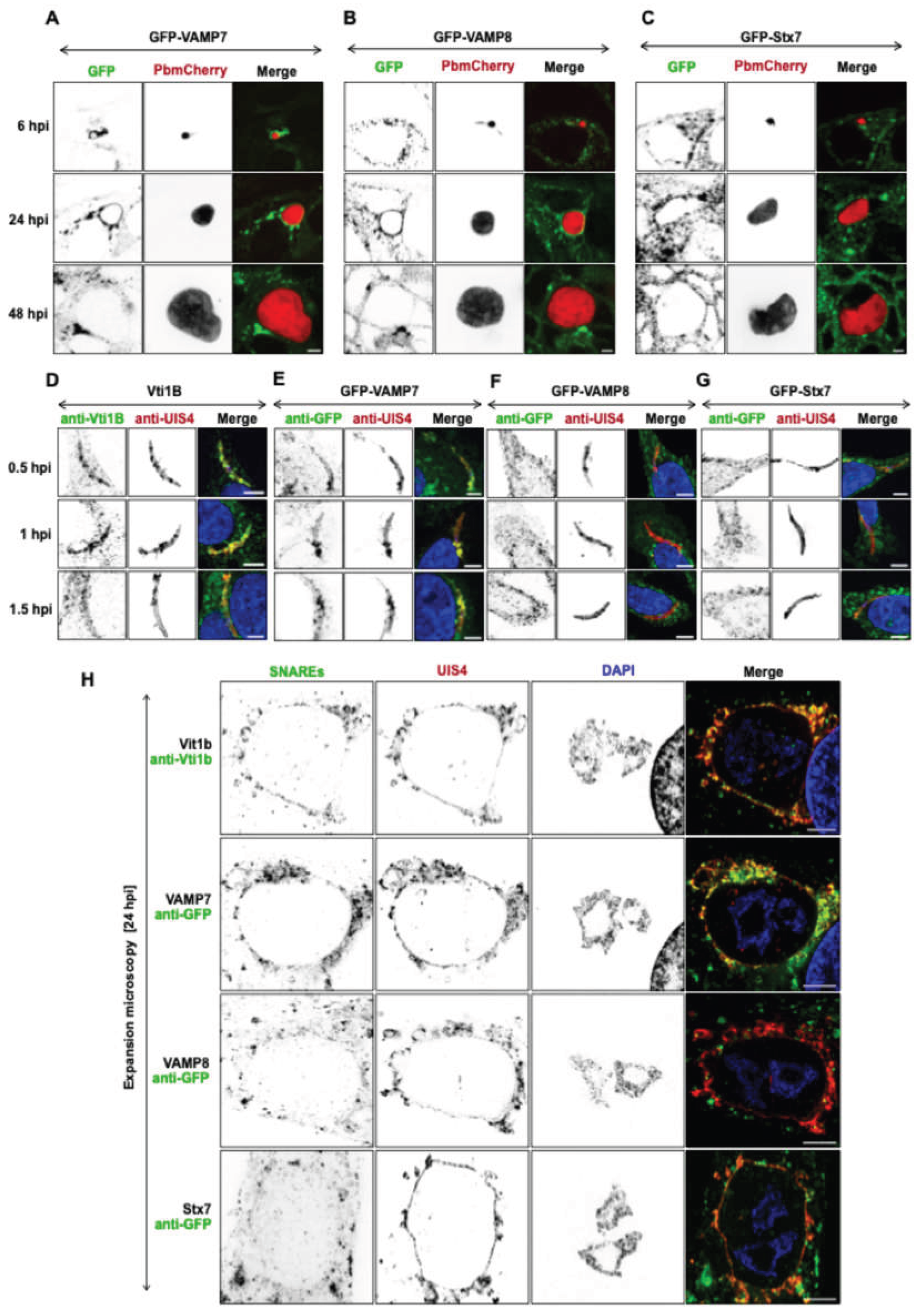
3.1. SNARE Proteins Are Recruited to the Plasmodium PVM during Liver stage Infection
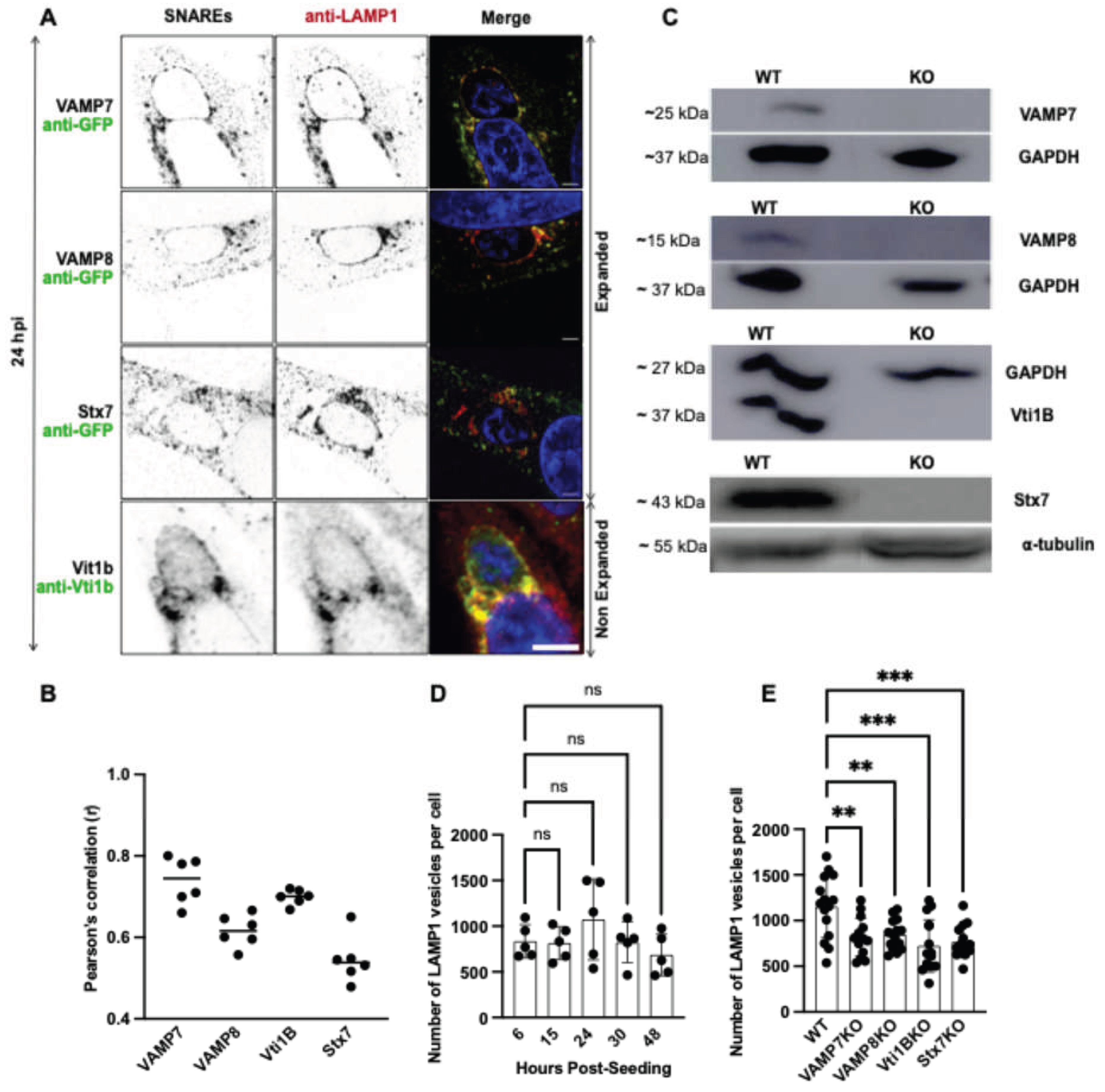
3.2. SNARE Proteins Colocalize with LAMP1-Positive Lysosomes and Regulate Lysosomal Homeostasis
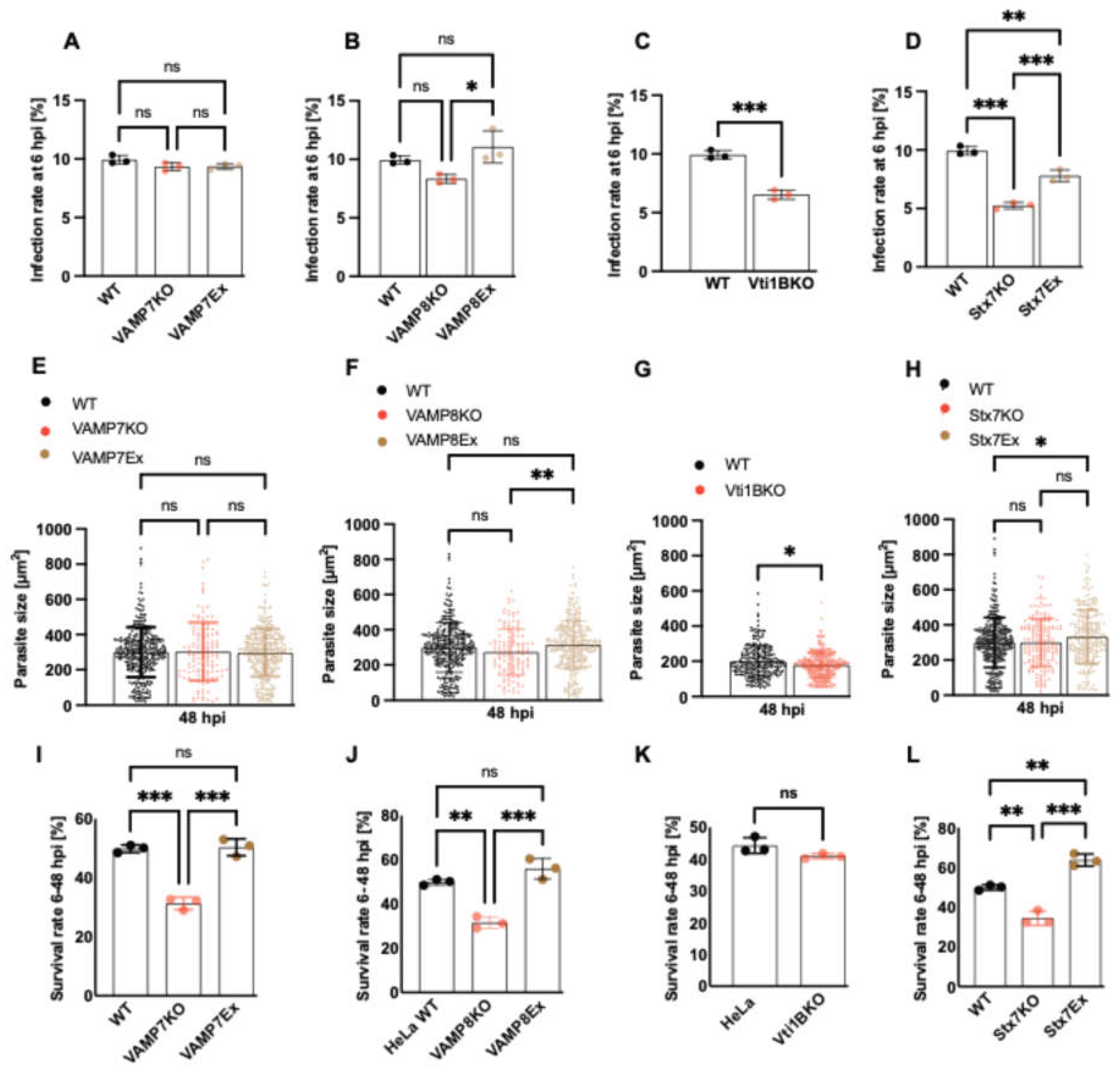
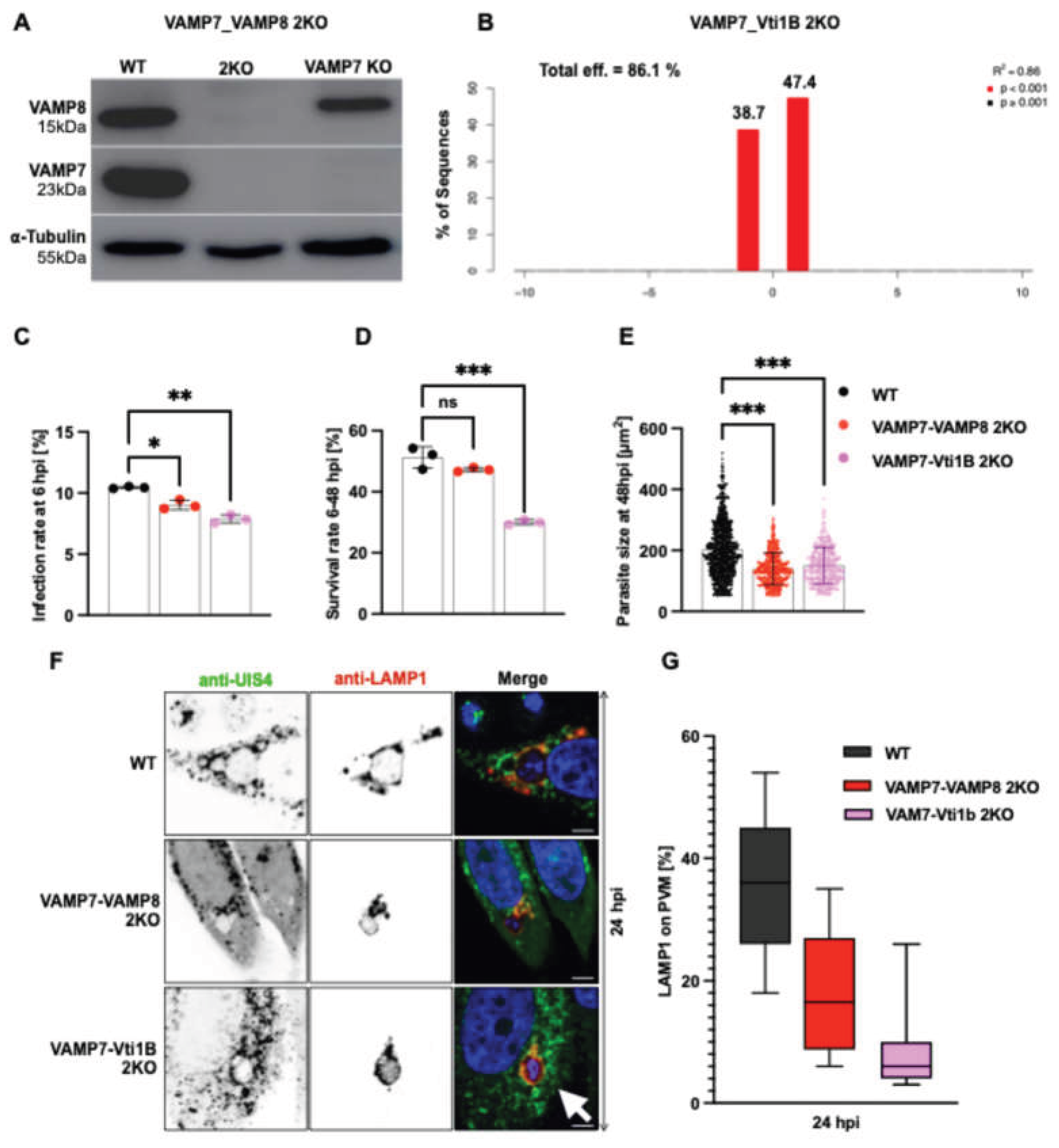
3.3. Effect of SNARE Protein Depletion on Parasite Infection and Survival
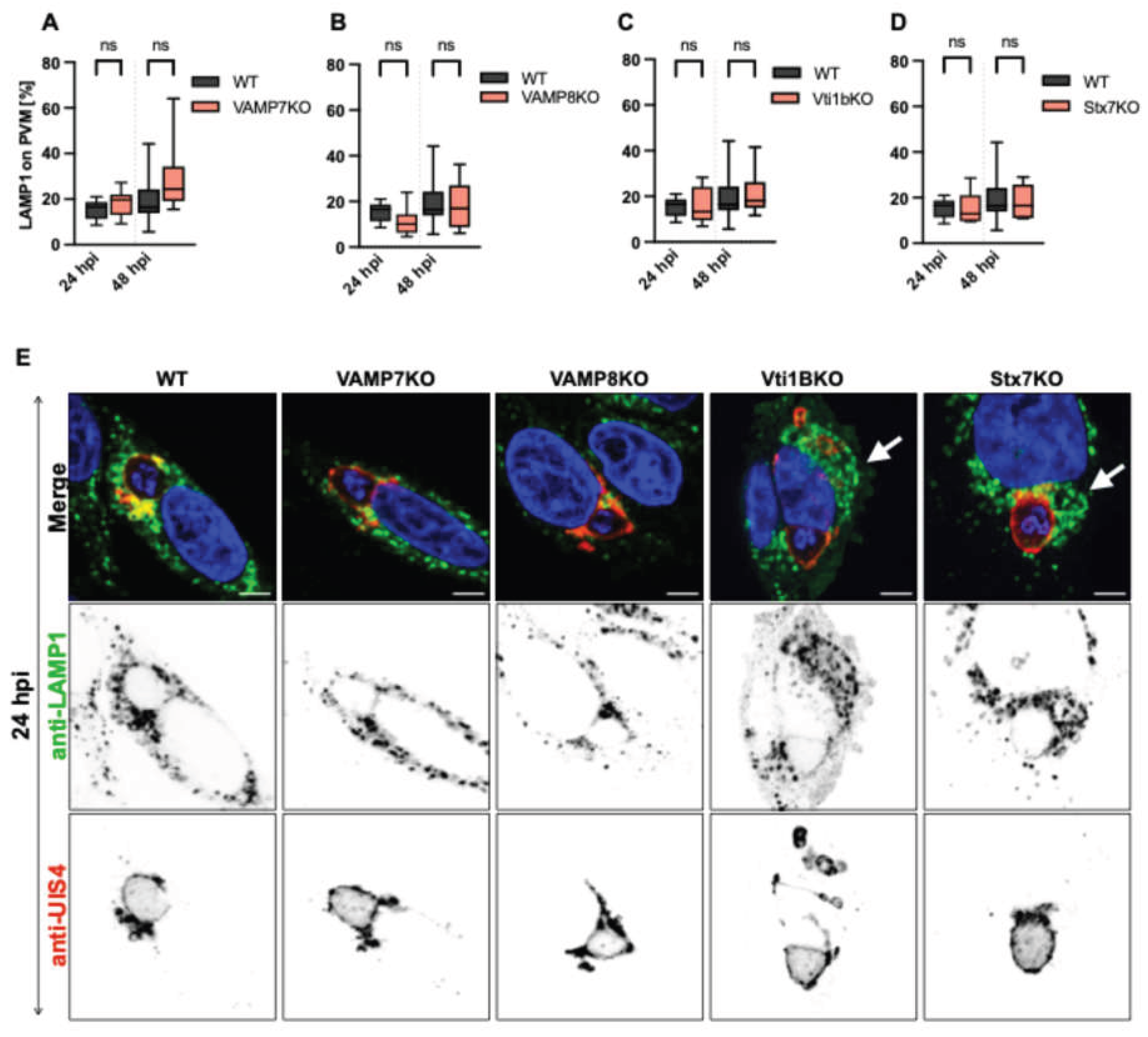
3.4. SNARE Protein Depletion Does Not Impair Lysosomal Recruitment to the PVM but Alters Vesicle Distribution
3.5. Double Knockout of SNARE Genes Impairs Parasite Development and Lysosome-PVM Fusion
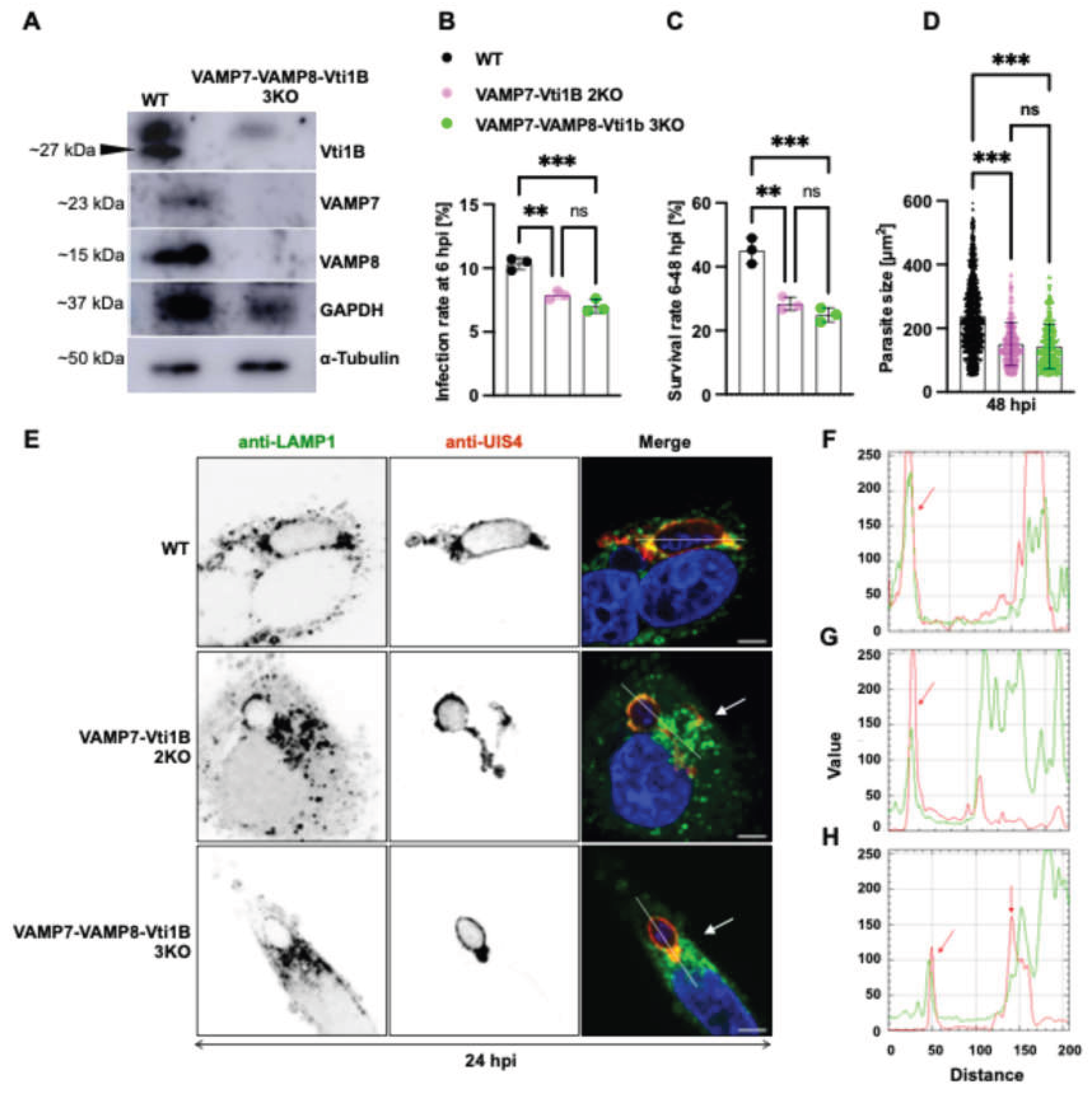
3.6. Triple SNARE Knockout Further Impairs Parasite Liver stage Development and Lysosome-PVM Interaction
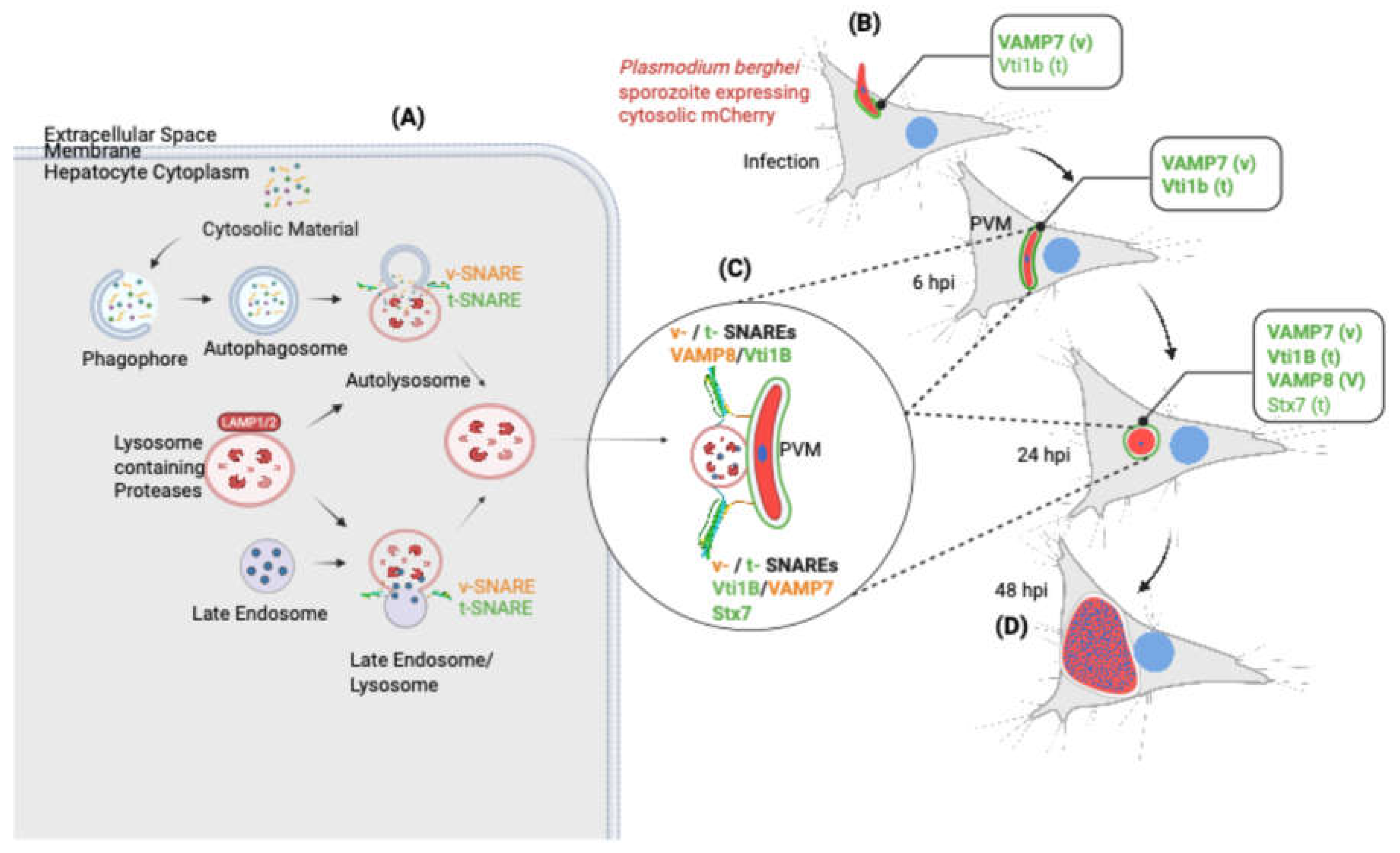
3.7. Proposed Model of SNARE-Mediated Host Lysosome–PVM Interactions During Plasmodium berghei Liver Stage Development
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| P. berghei | Plasmodium berghei |
| PV | Parasitophorous Vacuole |
| PVM | Parasitophorous Vacuole Membrane |
| KO | Knock Out |
| SNARE | Soluble N-ethylmaleimide-sensitive factor attachment protein receptors |
| LAMP1 | Lysosome-Associated Membrane Protein 1 |
| TIDE | Tracking of Indels by Decomposition |
| VAMP | Vesicle-Associated Membrane Protein |
| VAMP7 | Vesicle-Associated Membrane Protein 7 |
| VAMP8 | Vesicle-Associated Membrane Protein 8 |
| Vti1B | Vesicle Transport Through Interaction With T-SNAREs 1B |
| Stx | Syntaxin |
| Stx7 | Syntaxin 7 |
| WB | Western Blot |
| GAPDH | Glyceraldehyde-3-phosphate dehydrogenase |
References
- Venkatesan, P. The 2023 WHO World malaria report. Lancet Microbe 2024, 5(no. 3), e214. [Google Scholar] [CrossRef]
- Baer, K.; Klotz, C.; Kappe, S. H. I.; Schnieder, T.; Frevert, U. Release of hepatic Plasmodium yoelii merozoites into the pulmonary microvasculature. PLoS Pathog. 2007, 3(no. 11), 1651–1668. [Google Scholar] [CrossRef] [PubMed]
- Prudêncio, M.; Rodriguez, A.; Mota, M. M. The silent path to thousands of merozoites: the Plasmodium liver stage. Nature Reviews Microbiology 2006, 4(no. 11), 849–856. [Google Scholar] [CrossRef]
- Sturm, A.; et al. Manipulation of host hepatocytes by the malaria parasite for delivery into liver sinusoids. Science (1979). 2006, 313(no. 5791), 1287–1290. [Google Scholar] [CrossRef] [PubMed]
- Castelli, F. Waiting for the Malaria Vaccine: The Complex Epidemiological Transition Toward Malaria Elimination. Clin. Infect. Dis. 2020, 71(no. 2), 381–382. [Google Scholar] [CrossRef]
- De Niz, M.; et al. , Hijacking of the host cell Golgi by Plasmodium berghei liver stage parasites. J. Cell Sci. 2021, 134(no. 10), jcs252213. [Google Scholar] [CrossRef]
- Agop-Nersesian, C.; Niklaus, L.; Wacker, R.; Heussler, V. T. Host cell cytosolic immune response during Plasmodium liver stage development. FEMS Microbiol. Rev. 2018, 42(no. 3), 324. [Google Scholar] [CrossRef]
- Prado, M.; et al. , Long-term live imaging reveals cytosolic immune responses of host hepatocytes against Plasmodium infection and parasite escape mechanisms. Autophagy 2015, 11(no. 9), 1561–1579. [Google Scholar] [CrossRef] [PubMed]
- Burda, P. C.; Roelli, M. A.; Schaffner, M.; Khan, S. M.; Janse, C. J.; Heussler, V. T. A Plasmodium Phospholipase Is Involved in Disruption of the Liver Stage Parasitophorous Vacuole Membrane. PLoS Pathog. 2015, 11(no. 3), e1004760. [Google Scholar] [CrossRef]
- Labaied, M.; et al. , Plasmodium salvages cholesterol internalized by LDL and synthesized de novo in the liver. Cell. Microbiol. 2011, 13(no. 4), 569–586. [Google Scholar] [CrossRef]
- Lopes da Silva, M.; et al. , The host endocytic pathway is essential for Plasmodium berghei late liver stage development. Traffic 2012, 13(no. 10), 1351–1363. [Google Scholar] [CrossRef]
- Petersen, W.; et al. , Sequestration of cholesterol within the host late endocytic pathway restricts liver-stage Plasmodium development. Mol. Biol. Cell 2017, 28(no. 6), 726–735. [Google Scholar] [CrossRef]
- Thieleke-Matos, C.; et al. , Host cell autophagy contributes to Plasmodium liver development. Cell. Microbiol. 2016, 18(no. 3), 437–450. [Google Scholar] [CrossRef] [PubMed]
- Vijayan, K.; et al. , Plasmodium Secretion Induces Hepatocyte Lysosome Exocytosis and Promotes Parasite Entry. iScience 2019, 21, 603–611. [Google Scholar] [CrossRef] [PubMed]
- Niklaus, L.; Agop-Nersesian, C.; Schmuckli-Maurer, J.; Wacker, R.; Grünig, V.; Heussler, V. T. Deciphering host lysosome-mediated elimination of Plasmodium berghei liver stage parasites. Sci. Rep. 2019, 9(no. 1). [Google Scholar] [CrossRef] [PubMed]
- Gioseffi, A.; Edelmann, M. J.; Kima, P. E. Intravacuolar Pathogens Hijack Host Extracellular Vesicle Biogenesis to Secrete Virulence Factors. Front. Immunol. 2021, 12. [Google Scholar] [CrossRef]
- Romano, J. D.; et al. The parasite Toxoplasma sequesters diverse Rab host vesicles within an intravacuolar network. J. Cell Biol. 2017, 216(no. 12), 4235. [Google Scholar] [CrossRef]
- Wenbo, L.; Yewei, Y.; Hui, Z.; Zhongyu, L. Hijacking host cell vesicular transport: New insights into the nutrient acquisition mechanism of Chlamydia. Virulence 2024, 15(no. 1), 2351234. [Google Scholar] [CrossRef]
- Jahn, R.; Scheller, R. H. SNAREs — engines for membrane fusion. Nature Reviews Molecular Cell Biology 2006, 7(no. 9), 631–643. [Google Scholar] [CrossRef]
- Südhof, T. C.; Rothman, J. E. Membrane fusion: grappling with SNARE and SM proteins. Science 2009, 323(no. 5913), 474–477. [Google Scholar] [CrossRef]
- Parlati, F.; McNew, J. A.; Fukuda, R.; Miller, R.; Söllner, T. H.; Rothman, J. E. Topological restriction of SNARE-dependent membrane fusion. Nature 2000, 407(no. 6801), 194–198. [Google Scholar] [CrossRef] [PubMed]
- Antonin, W.; Fasshauer, D.; Becker, S.; Jahn, R.; Schneider, T. R. Crystal structure of the endosomal SNARE complex reveals common structural principles of all SNAREs. Nat. Struct. Biol. 2002, 9(no. 2), 107–111. [Google Scholar] [CrossRef]
- Rizo, J.; Rosenmund, C. Synaptic vesicle fusion. Nat. Struct. Mol. Biol. 2008, 15(no. 7), 665–674. [Google Scholar] [CrossRef]
- Dingjan, I.; Linders, P. T. A.; Verboogen, D. R. J.; Revelo, N. H.; Ter Beest, M.; Van Den Bogaart, G. Endosomal and phagosomal SNAREs. Physiol. Rev. 2018, 98(no. 3), 1465–1492. [Google Scholar] [CrossRef] [PubMed]
- Ayong, L.; Pagnotti, G.; Tobon, A. B.; Chakrabarti, D. Identification of Plasmodium falciparum family of SNAREs. Mol. Biochem. Parasitol. 2007, 152(no. 2), 113–122. [Google Scholar] [CrossRef]
- Antonin, W.; Holroyd, C.; Tikkanen, R.; Honing, S.; Jahn, R. The R-SNARE endobrevin/VAMP-8 mediates homotypic fusion of early endosomes and late endosomes. Mol. Biol. Cell 2000, 11(no. 10), 3289–3298. [Google Scholar] [CrossRef]
- Pryor, P. R.; et al. Combinatorial SNARE complexes with VAMP7 or VAMP8 define different late endocytic fusion events. EMBO Rep. 2004, 5(no. 6), 590–595. [Google Scholar] [CrossRef]
- Hassan, Z.; Kumar, N. D.; Reggiori, F.; Khan, G. How Viruses Hijack and Modify the Secretory Transport Pathway. Cells 2021, 10(no. 10), 2535. [Google Scholar] [CrossRef]
- Cueto, J.A.; et al. , Soluble N-ethylmaleimide-sensitive factor attachment protein receptors required during Trypanosoma cruzi parasitophorous vacuole development. Cell. Microbiol. 2017, 19(no. 6). [Google Scholar] [CrossRef]
- Abdallah, D.; Viscogliosi, E.; Certad, G. Cryptosporidium-host interactions: What’s new? Current Research in Parasitology & Vector-Borne Diseases 2025, 8, 100285. [Google Scholar] [CrossRef]
- Delevoye, C.; et al. , SNARE Protein Mimicry by an Intracellular Bacterium. PLoS Pathog. 2008, 4(no. 3), e1000022. [Google Scholar] [CrossRef]
- Fu, J.; Zhao, L.; Yang, J.; Chen, H.; Cao, S.; Jia, H. An unconventional SNARE complex mediates exocytosis at the plasma membrane and vesicular fusion at the apical annuli in Toxoplasma gondii. PLoS Pathog. 2023, 19(no. 3), e1011288. [Google Scholar] [CrossRef]
- Agop-Nersesian, C.; De Niz, M.; Niklaus, L.; Prado, M.; Eickel, N.; Heussler, V. T. Shedding of host autophagic proteins from the parasitophorous vacuolar membrane of Plasmodium berghei. Scientific Reports 2017 2017, 7(no. 1), 1–14. [Google Scholar] [CrossRef]
- Rosenbluh, J.; et al. , Genetic and Proteomic Interrogation of Lower Confidence Candidate Genes Reveals Signaling Networks in β-Catenin-Active Cancers. Cell Syst. 2016, 3(no. 3), 302–316.e4. [Google Scholar] [CrossRef]
- Bindschedler, A.; et al. LC3B labeling of the parasitophorous vacuole membrane of Plasmodium berghei liver stage parasites depends on the V-ATPase and ATG16L1. Mol. Microbiol. 2024, 121(no. 6), 1095–1111. [Google Scholar] [CrossRef]
- Sanjana, N. E.; Shalem, O.; Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 2014, 11(no. 8), 783–784. [Google Scholar] [CrossRef] [PubMed]
- Frischknecht, F.; et al. , Imaging movement of malaria parasites during transmission by Anopheles mosquitoes. Cell. Microbiol. 2004, 6(no. 7), 687–694. [Google Scholar] [CrossRef]
- Frischknecht, F.; Martin, B.; Thiery, I.; Bourgouin, C.; Menard, R. Using green fluorescent malaria parasites to screen for permissive vector mosquitoes. Malar. J. 2006, 5(no. 1), 1–8. [Google Scholar] [CrossRef] [PubMed]
- Rankin, K.E.; Graewe, S.; Heussler, V. T.; Stanway, R. R. Imaging liver-stage malaria parasites. Cell. Microbiol. 2010, 12(no. 5), 569–579. [Google Scholar] [CrossRef] [PubMed]
- Schmuckli-Maurer, J.; et al. , Plasmodium berghei liver stage parasites exploit host GABARAP proteins for TFEB activation. Communications Biology 2024, 7(no. 1), 1–13. [Google Scholar] [CrossRef]
- Van De Sand, C.; et al. , The liver stage of Plasmodium berghei inhibits host cell apoptosis. Mol. Microbiol. 2005, 58(no. 3), 731–742. [Google Scholar] [CrossRef]
- Atchou, Kodzo.; Berger, B. M.; Heussler, V.; Ochsenreiter, T. Pre-gelation staining expansion microscopy for visualisation of the Plasmodium liver stage. J. Cell Sci. 2023, 136(no. 22). [Google Scholar] [CrossRef]
- Pino, P.; et al. “A multistage antimalarial targets the plasmepsins IX and X essential for invasion and egress,”. Science (1979). 2017, 358(no. 6362), 522–528. [Google Scholar] [CrossRef]
- Itakura, E.; Kishi-Itakura, C.; Mizushima, N. The hairpin-type tail-anchored SNARE syntaxin 17 targets to autophagosomes for fusion with endosomes/lysosomes. Cell 2012, 151(no. 6), 1256–1269. [Google Scholar] [CrossRef]
- Moore, E. R.; Mead, D. J.; Dooley, C. A.; Sager, J.; Hackstadt, T. The trans-Golgi SNARE syntaxin 6 is recruited to the chlamydial inclusion membrane. Microbiology (Reading). 2011, 157 no. Pt 3, 830–838. [Google Scholar] [CrossRef]
- Martinez-Arca, S.; Alberts, P.; Zahraoui, A.; Louvard, D.; Galli, T. Role of tetanus neurotoxin insensitive vesicle-associated membrane protein (TI-VAMP) in vesicular transport mediating neurite outgrowth. J. Cell Biol. 2000, 149(no. 4), 889–899. [Google Scholar] [CrossRef]
- Verboogen, D. R. J.; Mancha, N. G.; ter Beest, M.; van den Bogaart, G. Fluorescence Lifetime Imaging Microscopy reveals rerouting of SNARE trafficking driving dendritic cell activation. Elife 2017, 6. [Google Scholar] [CrossRef]
- Mori, Y.; ichiro Takenaka, K.; Fukazawa, Y.; Takamori, S. The endosomal Q-SNARE, Syntaxin 7, defines a rapidly replenishing synaptic vesicle recycling pool in hippocampal neurons. Communications Biology 2021, 4(no. 1), 1–13. [Google Scholar] [CrossRef] [PubMed]
- Pols, M.S.; et al. , hVps41 and VAMP7 function in direct TGN to late endosome transport of lysosomal membrane proteins. Nature Communications 2013, 4(no. 1), 1–12. [Google Scholar] [CrossRef] [PubMed]
- Atlashkin, V.; Kreykenbohm, V.; Eskelinen, E.-L.; Wenzel, D.; Fayyazi, A.; Fischer von Mollard, G. Deletion of the SNARE vti1b in mice results in the loss of a single SNARE partner, syntaxin 8. Mol. Cell. Biol. 2003, 23(no. 15), 5198–5207. [Google Scholar] [CrossRef]
- Nyboer, B.; Heiss, K.; Mueller, A. K.; Ingmundson, A. The Plasmodium liver-stage parasitophorous vacuole: A front-line of communication between parasite and host. International Journal of Medical Microbiology 2018, 308(no. 1), 107–117. [Google Scholar] [CrossRef] [PubMed]
- Caldelari, R.; et al. , Transcriptome analysis of Plasmodium berghei during exo-erythrocytic development. Malar. J. 2019, 18(no. 1). [Google Scholar] [CrossRef] [PubMed]
- Parish, L. A.; Rayner, J. C. Plasmodium falciparum secretory pathway: Characterization of PfStx1, a plasma membrane Qa-SNARE. Mol. Biochem. Parasitol. 2009, 164(no. 2), 153–156. [Google Scholar] [CrossRef]
- Bisio, H.; et al. The zip code of vesicle trafficking in apicomplexa: Sec1/munc18 and snare proteins. mBio 2020, 11(no. 5), 1–21. [Google Scholar] [CrossRef]
- Wang, Y.; et al. Control of infection by LC3-associated phagocytosis, CASM, and detection of raised vacuolar pH by the V-ATPase-ATG16L1 axis. Sci. Adv. 2022, 8(no. 43). [Google Scholar] [CrossRef]
- Furuta, N.; Fujita, N.; Noda, T.; Yoshimori, T.; Amano, A. Combinational soluble N-ethylmaleimide-sensitive factor attachment protein receptor proteins VAMP8 and Vti1b mediate fusion of antimicrobial and canonical autophagosomes with lysosomes. Mol. Biol. Cell 2010, 21(no. 6), 1001–1010. [Google Scholar] [CrossRef]
- Marshall, M. R.; et al. VAMP8-dependent fusion of recycling endosomes with the plasma membrane facilitates T lymphocyte cytotoxicity. J. Cell Biol. 2015, 210(no. 1), 135–151. [Google Scholar] [CrossRef]
- Bui, D. C.; Jorgenson, L. M.; Ouellette, S. P.; Rucks, E. A. Eukaryotic SNARE VAMP3 Dynamically Interacts with Multiple Chlamydial Inclusion Membrane Proteins. Infect. Immun. 2021, 89(no. 2), e00409–20. [Google Scholar] [CrossRef]
- Shi, X.; Halder, P.; Yavuz, H.; Jahn, R.; Shuman, H. A. Direct targeting of membrane fusion by SNARE mimicry: Convergent evolution of Legionella effectors. Proc. Natl. Acad. Sci. U. S. A. 2016, 113(no. 31), 8807–8812. [Google Scholar] [CrossRef]
- Ghosh, S.; O’Connor, T. J. Beyond paralogs: The multiple layers of redundancy in bacterial pathogenesis. Front. Cell. Infect. Microbiol. 2017, 7, 467. [Google Scholar] [CrossRef] [PubMed]
- Wacker, R.; et al. , LC3-association with the parasitophorous vacuole membrane of Plasmodium berghei liver stages follows a noncanonical autophagy pathway. Cell. Microbiol. 2017, 19(no. 10). [Google Scholar] [CrossRef] [PubMed]
- Fischer, T. D.; Wang, C.; Padman, B. S.; Lazarou, M.; Youle, R. J. STING induces LC3B lipidation onto single-membrane vesicles via the V-ATPase and ATG16L1-WD40 domain. J. Cell Biol. 2020, 219(no. 12), e202009128. [Google Scholar] [CrossRef] [PubMed]
- Lahree, A.; et al. Active APPL1 sequestration by Plasmodium favors liver-stage development. Cell Rep. 2022, 39(no. 9). [Google Scholar] [CrossRef] [PubMed]
| SNARE Protein |
Subcellular localization [24] | Construct | (*) Construct origin |
(†) Association with PVM |
|---|---|---|---|---|
| SNAP29 | Cytosolic, PM, EE, RE | GFP-SNAP29 | Lab | - |
| Flag-SNAP29 | [44] | |||
| Stx3 | PM, Phagosomes | Antibody | Lab | + |
| Stx6 | PM, EE, RE, LE, Golgi | GFP-Stx6 | Lab | + |
| 3xFlag-Stx6 | [45] | |||
| Stx7 | PM, EE, RE, LE, Lysosomes | Stx7-GFP | Lab | ++ |
| Stx7-Myc | Gift by J. Brummel | |||
| Stx8 | EE, RE, LE, Lysosomes | GFP-Stx8 | Lab | + |
| Stx17 | Phagosomes, Autophagosomes | GFP-Stx17 | Lab | + |
| Flag-Stx17 | [44] | |||
| Stx11 | PM, EE, RE, LE, Lysosomes, TGN |
GFP-Stx11 | Lab | - |
| Vti1b | EE, LE, Lysosomes, Golgi | antibody | Lab | +++ |
| Flag-Vti1b | [44] | |||
| VAMP3 | EE, RE, Phagosomes | GFP-VAMP3 | Lab | - |
| VAMP7 | PM, EE, LE, Lysosomes, Autophagosomes | GFP-VAMP7 | Lab [46] |
+++ |
| VAMP8 | PM, EE, RE, LE, Lysosomes, TGN | VAMP8-GFP | Lab | +++ |
| VAMP8-mCherry | [47] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




